Abstract

The sinonasal cavity is the gateway to the respiratory tract, playing an essential role in respiratory health. 1 The nasal microbiota is complex yet stable in health; disruption or dysbiosis of the microbiota is often associated with disease, including chronic rhinosinusitis.2,3 Techniques to study microbiota have evolved from growing bacteria on agar to DNA sequencing techniques (microbiome research) that target short hypervariable regions in the bacterial 16S rRNA gene allowing culture independent bacterial identification. This leap in technology significantly improved our ability to study sinonasal microbiome, especially considering that some species are difficult to culture. However, short read sequencing technology do not provide reliable species-level taxonomic resolution. 4
The recently developed third-generation sequencing technology, [eg, PacBio and Oxford Nanopore Technologies (ONT)], enable cost-effective long-read sequencing of the full-length bacterial 16S rRNA gene (~1500 base pairs), providing species-level identification. 5 To highlight the advantages of ONT technology in sinonasal microbiome research, we present a pilot study to determine the compositional differences in the bacterial nasal microbiome between participants with primary ciliary dyskinesia (PCD), subjects with chronic rhinosinusitis without nasal polyps (CRSsNP), and healthy controls.
Participants diagnosed with PCD, CRSsNP, and healthy controls were consented and enrolled. Nasal epithelial lining fluid was collected on leukosorb strips from each participant under endoscopic visualization of the middle meatus. Strips were stored at −80°C and subsequently subjected to DNA extraction. Full-length 16S rDNA library preparation and sequencing were performed following the manufacturer protocol (ONT). Details on patient recruitment, bioethics, methods, and analysis can be found in the Supplemental Material.
Samples from 53 individuals, including 15 with CRSsNP, 20 with PCD, and 18 healthy controls were analyzed. Bacterial composition was assessed by permutational multivariate analysis of variance and was found to be significantly different between PCD and CRSsNP (P = .009), and between PCD and healthy controls (P = .003). No significant difference was observed between CRSsNP and healthy controls (Figure 1A).
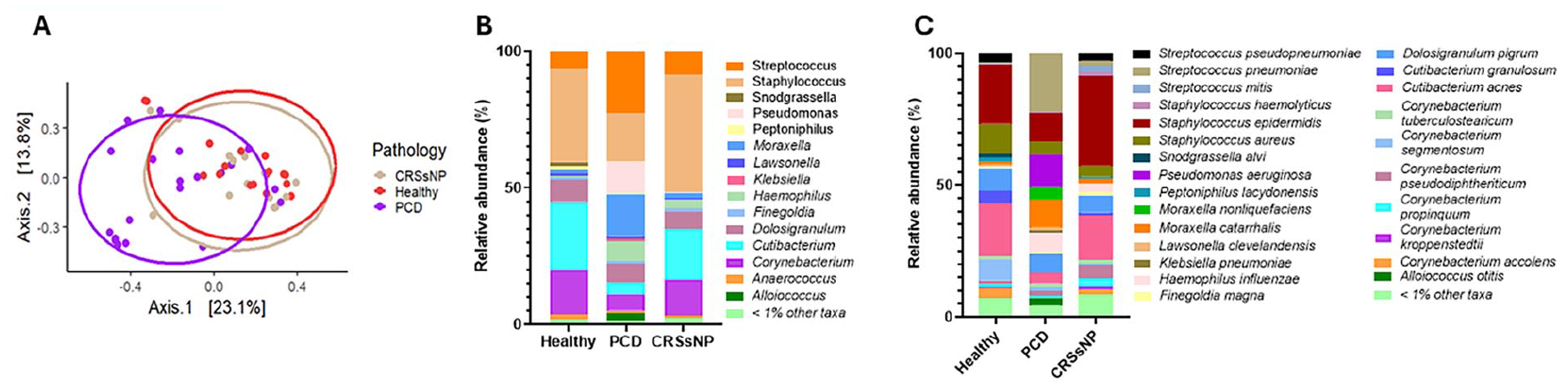
(A) Principal coordinates analysis using Bray–Curis distances and pairwise PERMANOVA statistical test, PCD versus CRSsNP P = .009, PCD versus Healthy P = .003, CRSsNP versus Healthy P = 1.00. (B) Relative abundance plot at genus taxonomic level showing taxa composition in Healthy, PCD, and CRSsNP. (C) Species-level relative abundance plot showing taxa composition in Healthy, PCD, and CRSsNP. PERMANOVA, permutational multivariate analysis of variance; PCD, primary ciliary dyskinesia; CRSsNP, chronic rhinosinusitis without nasal polyps.
Relative abundance analysis at genus level revealed an enrichment of Streptococcus, Pseudomonas, Haemophilus, and Moraxella in PCD compared to CRSsNP and healthy controls. In addition, a decrease in Staphylococcus, Cutibacterium, and Corynebacterium was observed in PCD compared to the other groups. In CRSsNP, we observed an enrichment of Staphylococcus compared to PCD and healthy controls (Figure 1B). At a species level, we found that the enrichment of Streptococcus, Pseudomonas, Haemophilus, and Moraxella in PCD could be attributed to single pathogenic species for each genus, including Streptococcus pneumoniae, Pseudomonas aeruginosa, Haemophilus influenzae, and Moraxella catarrhalis. In addition, the decrease of genera Staphylococcus and Cutibacterium in PCD can be attributable to the decrease in the species Staphylococcus epidermidis and Cutibacterium acnes. The decrease in the genus Corynebacterium in PCD is likely due to the decrease in multiple Corynebacterium species (Figure 1C).
In PCD, we observed profound nasal microbiome dysbiosis marked by an increase in Streptococcus pneumoniae, Pseudomonas aeruginosa, Haemophilus influenzae, and Moraxella catarrhalis, which are all considered key pathogens responsible for pulmonary infections in PCD. 6 Based on our data, we cannot determine if the sinonasal cavity or the lower respiratory tract is the site of primary infection. In CRSsNP, dysbiosis was more subtle, but reflected a marked increase in Staphylococcus epidermidis compared to the PCD and healthy groups. Staphylococcus epidermidis is commonly linked to chronic rhinosinusitis with and without nasal polyps. 7
Using third-generation sequencing technology, we characterized the nasal microbiome in healthy and diseased subjects to genus and species taxonomic levels (Figure 1B and C). To date, the majority of sinonasal microbiome studies have relied on second-generation short-read sequencing technologies, targeting 16S variable regions, 8 which yield genus-level data similar to that depicted in Figure 1B. However, a single bacterial genus, like Staphylococcus can include both commensal (eg, Staphylococcus epidermidis), and pathogenic (eg, Staphylococcus aureus) species, highlighting the importance of species-level identification for interpreting changes in the microbiome during health and disease.
Species or even strain-level identification are critical for the development of targeted therapies and the design of probiotics. In conclusion, the nasal microbiome composition between PCD compared to CRSsNP and healthy controls was significantly different. We illustrated that species-level identification is critical for further studies. For this reason, we propose a shift from genus level only microbiome analysis and reporting, to species-level analysis to direct future research of probiotics and biomarkers for diagnosis and disease prognosis.
Supplemental Material
sj-docx-1-ear-10.1177_01455613251341711 – Supplemental material for Advances in Sinonasal Microbiome Analysis
Supplemental material, sj-docx-1-ear-10.1177_01455613251341711 for Advances in Sinonasal Microbiome Analysis by Cristian Roca, Yasine Mirmozaffari, W. Jared Martin, Nicole Capps, Kelli M. Sullivan, Brian D. Thorp, Thomas W. Ferkol, Stephanie D. Davis, Adam Kimple and Matthew C. Wolfgang in Ear, Nose & Throat Journal
Footnotes
Acknowledgements
We thank all participants for their voluntarily collaboration with this study.
Ethical Considerations
Sample collections and de-identified data usage for this study received ethical approval from the University of North Carolina at Chapel Hill institutional review board (approval numbers: 20-1170 and 17-2677).
Consent to Participate
All participants (or a legal representative) provided written consent and/or assent to participate in this study.
Author Contributions
B.D.T., T.W.F., S.D.D., and A.K. contributed to the investigation design, sample acquisition, final approval of manuscript, and funding acquisition. Y.M., W.J.M., N.C., and K.M.S. contributed to the study methodology, sample acquisition, drafting of manuscript, and approval of final versions. C.R. and M.C.W. contributed to the study design, data analysis, write and revision of the manuscript, and approval of the final version.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study received funding from the Genetic Disorders of Mucociliary Clearance Consortium (U54HL096458), which is part of the National Center for Advancing Translational Sciences (NCATS) Rare Diseases Clinical Research Network (RDCRN). RDCRN is an initiative of the Office of Rare Diseases Research (ORDR) funded through a collaboration between NCATS and the National Heart, Lung, and Blood Institute (NHLBI).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data published are part of an ongoing study that will be deposited into a public repository once the entire project is completed. However, the de-identified data from this preliminary assessment is immediately available upon request.
Supplemental Material
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
