Abstract
Background
Nasopharyngeal carcinoma (NPC) is a malignancy with a 2 per 100 000 incidence rate in the world. Overall survival (OS) of patients in stage I–II disease is around 80%, whereas OS of patients in stage III-IVB disease drops to 60%, implying the importance of diagnosis to reduce NPC mortality. However, more than 70% patients of NPC were diagnosed at advanced stages (stage III and IV) in clinics, and it definitely contributes to little substantial improvement in the 5-year survival rates although NPC is sensitive to radio-and chemotherapy. Hence, development of novel biomarkers and targetable genes in NPC is eagerly awaited.
Methods
We had analyzed the dataset GSE12452 and found hundreds of genes trans-activated in NPC. Among them, this study focused on PARP-1 binding protein (PARPBP) whose overexpression was also validated in GSE13597 and GSE53819 datasets.
Results
Knockdown of PARPBP significantly reduced cell viability in NPC and also identified hundreds of differentially expressed genes including 377 downregulated and 518 upregulated genes in HONE-1 cells with stably knockdown PARPBP. Furthermore, PARPBP might promote cell migration and invasion in NPC through positive regulation of ubiquitin-conjugating enzyme 2C (UBE2C).
Conclusion
The results demonstrate the aberrant expression of PARPBP in NPC, and imply its importance in nasopharyngeal carcinogenesis which further opens up the possibility of PARPBP as a novel diagnostic biomarker for NPC therapy.
Introduction
Nasopharyngeal carcinoma (NPC) is a rare malignancy worldwide with a 2 per 100 000 incidence rate.1,2 NPC is remarkably prevalent in Southern China and Southeastern Asia. The major etiological factors of NPC are considered Epstein-Barr virus (EBV) infection, genetic components, and environmental influences.3,4 Consumption of certain foods including salted fish and preserved foods is reported to be associated with high risks of NPC in Southeastern Asia. It usually has no specific symptoms at early stage, whereas some common signs and symptoms are usually associated with distant metastasis. Currently, radiation therapy for stage I and II, and radiation combined chemoradiotherapy for stages IIB–IVB diseases is the standard therapy in NPC. Overall survival (OS) of patients in stage I–II disease is around 80%, whereas OS of patients in advanced stage (stage III–IVB) disease drops to 60%, implying the importance of diagnosis to reduce NPC mortality. 5 Hence, stage at diagnosis is the most crucial prognostic factors in NPC. However, more than 70% patients of NPC were diagnosed at advanced stages (stage III and IV) in clinics, 6 and it definitely contributes to little substantial improvement in the 5 year survival rates although NPC is sensitive to radio-and chemotherapy. 7 Hence, development of novel biomarkers and targetable genes in NPC is eagerly awaited.
PARP-1 binding protein (PARPBP) is believed to have important roles in the maintenance of genomic stability.8,9 Deletion of PARPBP sensitizes pancreatic ductal adenocarcinoma (PDAC) cells to DNA-damaging agents by suppressing PARP-1 activity through direct interaction. 10 PARPBP also interacts with PCNA and RAD51 and suppresses the formation of RAD51-DNA homologous recombination (HR) structures. 11 Aberrant PARPBP expression is implicated in numerous cancer types.10,12 However, there is no report regarding to PARPBP in NPC. 13 Ubiquitin-conjugating enzyme E2C (UBE2C) is highly increased in a variety of cancers and associated with poor clinical outcomes in human cancer.14,15 UBE2C promotes cell migration and invasion in cancers.16,17
In our study, we find elevated PARPBP expression in NPC tissue samples by reanalyzing microarray datasets and silencing PARPBP attenuates NPC cell viability. Through RNA-seq analysis, a set of PARPBP-regulated genes are identified including UBE2C. The expression level of PARPBP is positively correlated with UBE2C in NPC, and knockdown of PARPBP leads to a marked decrease in mesenchymal markers including N-cadherin and Vimentin partly through reducing UBE2C protein expression. Taken together, our results state that PARPBP is associated with cell migration and invasion partly through modulating UBE2C in NPC and argue the possibility of PARPBP as a novel therapeutic target against NPC.
Material and Methods
Microarray Data Collection and Analysis
Microarray data were obtained from The Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo) to identify differentially expressed genes (DEGs) between nasopharyngeal carcinoma and normal controls. GSE12452 (included 31 NPCs and 10 normal controls),18,19 GSE13597 (included 25 NPCs and 3 normal controls), 20 and GSE53819 (included 18 NPCs and 18 normal controls) 21 were used to examine PARPBP expression levels.
Cell line and cell culture
The human nasopharyngeal carcinoma cell lines C666-1, HONE-1, and 5-8F cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, China) supplemented with 10% fetal bovine serum (ScienCell, USA) and 1% antibiotic/antimycotic solution (Gibco, Shanghai, China) at 37°C in a humidified incubator with 5% CO2.
Small Interfering RNA (siRNA) Transfection and RNA Extraction
PARPBP-specific siRNA (#1:5′-GAAUAGAUUGUACGGCAAA-3′, #2:5′- CAUCAUCAUGGAACGUCUA-3′)and siEGFP (5′-GCAGCACGACUUCUUCAAG−3′) as a control siRNA were transfected into NPC cell lines with Lipofectamine® 2000 (Thermo Fisher Scientific, Inc, China), respectively. Total RNA was isolated from the cells by using Takara MiniBEST Universal
RNA Extraction kit (Takara, Dalian, China), and reverse transcribed to cDNA with PrimeScriptTM 1st strand cDNA Synthesis kit (Takara, Dalian, China) according to the manufacturer’s instruction. The primer sequences were as follows:
ACTB forward: 5′-TTGGCTTGACTCAGGATTTA-3’
ACTB reverse: 5′-ATGCTATCACCTCCCCTGTG-3’
PARPBP forward: 5′-CTCAGCTGGGAAAGCTACAGAT-3’
PARPBP reverse: 5′-CATGCCAGGTAGTTCTTCCATC-3’.
Each PCR regime involved initial denaturation at 94°C for 2 min followed by 23 cycles (for ACTB), 28 cycles (for PARPBP) at 94°C for 30 sec, 55°C for 30 sec, and 72°C for 1 min.
Lentivirus Packaging and Transduction
Short hairpin RNAs targeting PARPBP (shPARPBP) and negative control (shNC) were ligated into the LV-3 (pGLVH1/GFP+Puro) vector. The target sequences for PARPBP were 5′-GAAUAGAUUGUACGGCAAA-3′, and 5′-TTCTCCGAACGTGTCACGT-3′ as a negative control. The 293T cells were co-transfected with the lentiviral vectors LV-3-PARPBP (or the control lentiviral vectors) and the packaging vectors psPAX2 and pMD2G using Lipofectamine® 2000. Supernatant was collected at 48h after transfection, filtered, and stored at -80 degree. HONE-1 cells with 40–70% confluency were incubated with virus and 8 µg/ml polybrene for 24 h followed by subsequent selection with 2 µg/ml puromycin for 1-2 weeks. Knockdown efficiency was examined by semi-quantitative reverse transcription PCR.
Cell Viability Assay and Colony Formation Assay
Cells (5000 cells/well) seeded in 96-well plates were transfected with specific siRNA, respectively, and cell viability was measured at 96 hours using Cell Counting Kit-8 (Dojindo, Shanghai, China). Cells (5000 cells/well) transfected with specific siRNA for 4 days and cultured for additional 6 days were fixed and stained.
Cell Migration and Invasion Assay
The 24-well Transwell insert chamber (Corning costar, USA) and BioCoatTM Matrigel Invasion Chamber (Corning, USA) were used according to the manufacturer’s instruction. Cells (1*105 cells/insert for migration and 1*105 cells/insert for invasion) were seeded in the insert and incubated for 16–20 hours. The migrating or invading cells were fixed and stained.
Western Blot
Cells were lysed in RIPA buffer with complete protease inhibitor cocktail (Roche Applied Science, China). Anti-β-actin (A5441, Sigma-Aldrich, USA), anti-UBE2C (A18075, Abclonal, China), anti-N-Cadherin (#13116, Cell signaling, USA), and anti-Vimentin (A2584, Abclonal, China) were used.
RNA-Seq Analysis
RNA-seq analysis was performed as previously described. 22 Gene expression and differential transcription between shNC and shPARPBP treated cells were evaluated using Cuffdiff. Gene Ontology (GO) analysis was performed in the standard enrichment computation method. KEGG pathway analysis was performed by KOBAS, and P<0.05 was set as the cut-off criterion. 23
Statistical Analysis
Statistical analyses were performed using SPSS version 20.0 (IBM Corp., Armonk, NY, USA). Comparisons between two groups were analyzed using an independent two-sample t-test (two-tailed); comparisons among multiple groups were analyzed by one-way analysis of variance followed by the least significant difference post hoc test. Results were presented as mean ± standard deviation of three independent experiments. P < .05 was defined as significant.
Results
The mRNA level of PARPBP is enhanced in NPC. To identify differentially expressed genes (DEGs) between tumor and normal tissue in NPC, we reanalyzed the microarray expression dataset GSE12452 and found a set of DEGs. Of these, PARPBP was highly upregulated in NPC samples as compared to normal tissues (fold change of 4.08 with a P value of 6.49E-06, Figure 1A). Concordantly, the elevated PARPBP was also found in NPC microarray expression dataset GSE13597 and GSE53819 (fold change of 2.08 with a P value of 0.03 in GSE13597; fold change of 3.48 with a P value of 9.68E-06 in GSE53819, Figure 1A), implying the importance of PARPBP in NPC development and progression. In addition, through ROC analysis, we observed that the expression of PARPBP was likely to discriminate between NPC tumor tissues and normal tissues (AUC=0.9710, P < .0001 in GSE12452; AUC=0.9067, P < .05 in GSE13597; AUC=0.9352, P<0.0001 in GSE12452, Figure 1B). Collectively, these data implied that PARPBP was upregulated in NPC and might have crucial roles in the carcinogenesis of NPC. The expression of PARPBP is elevated in NPC tissues. (A) PARPBP mRNA expression is evaluated in NPC tissues by analyzing GSE12452, GSE13597, and GSE53819. (B) ROC curves and AUC values for NPC by analyzing GEO datasets.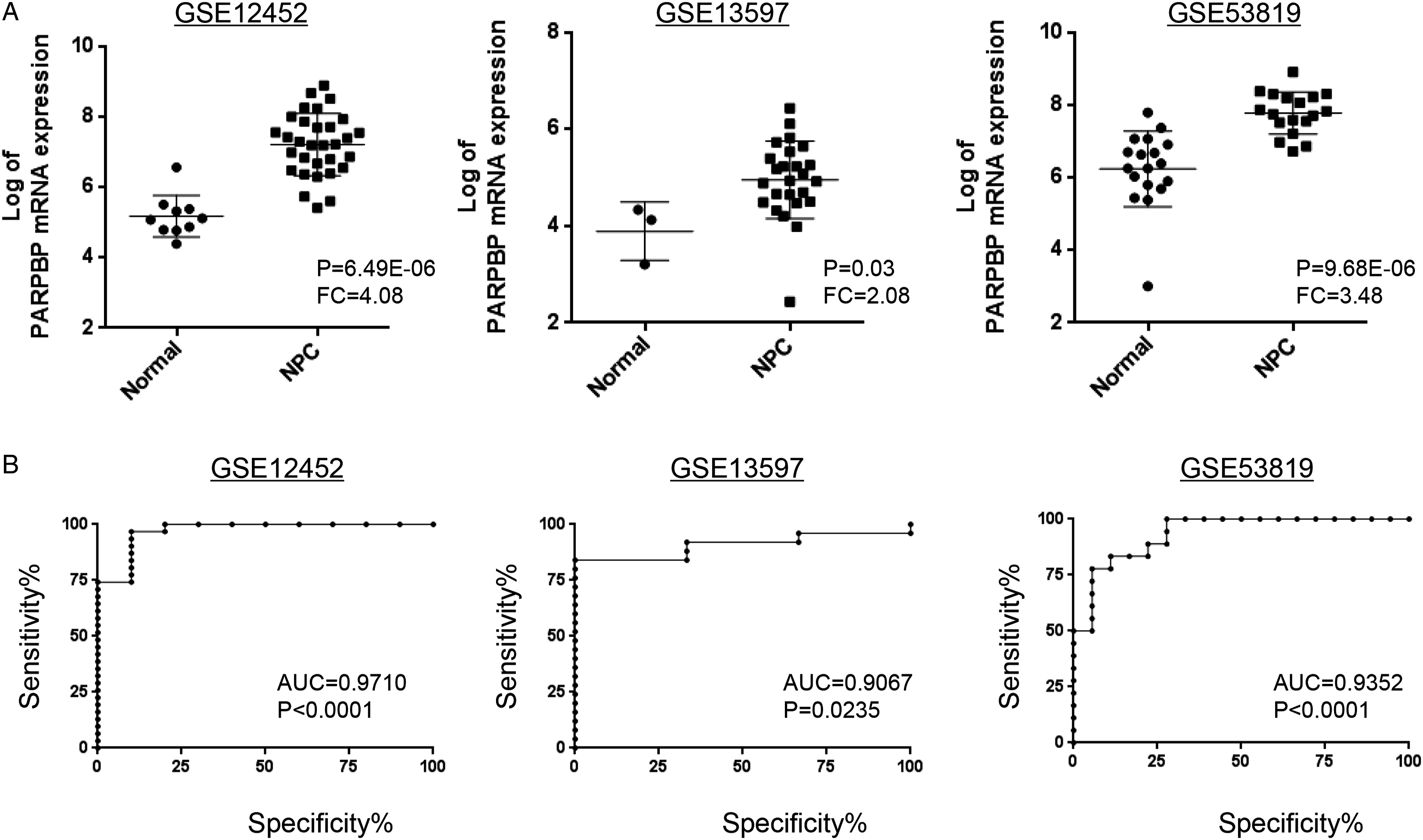
Depletion of PARPBP suppresses NPC cell proliferation. The expression of PARPBP in HONE-1, C666-1, and 5-8F NPC cell lines was examined and the results showed a higher expression of PARPBP in HONE-1 and C666-1 compared with 5-8F (Figure 2A). Thus, HONE-1 and C666-1 NPC cell lines were used for further study. To investigate the biological significance of PARPBP in NPC, HONE-1 and C666-1 cells were transfected with two independent siRNA specifically targeting PARPBP (si#1, si#2), respectively. Knockdown efficiency was examined by semi-quantitative reverse transcription-PCR. The expression of PARPBP in cells treated with si#1 or si#2 for 48 hours was significantly reduced compared with siEGFP control (Figure 2B). Meanwhile, we observed a marked decrease in cancer cell viability in PARPBP-deleted HONE-1 and C666-1 cells (Figure 2C). Colony formation assay was conducted to assess the functions of PARPBP on NPC cell proliferation. It revealed that deletion of PARPBP in HONE-1 and C666-1 cells led to drastic reduction in the number of viable cells and formed smaller colonies as compared to control group (Figure 2D). Thereafter, PARPBP stably knockdown cells were constructed using shRNA in HONE-1 cells and knockdown effect of PARPBP was confirmed by semi-quantitative reverse transcription-PCR (Figure 2E). Cell proliferation activity of negative control shRNA (shNC) and shPARPBP was measured. As a result, shPARPBP cells exhibited reduced cell growth rate than shNC cells (Figure 2F), and consistently, colony formation assay indicated that shPARPBP cells showed smaller and less colonies than shNC cells (Figure 2G). These findings indicated that PARPBP might have important roles in the growth of NPC cells. PARPBP depletion results in a marked reduction in NPC cell viability. (A) The expression of PARPBP is examined in NPC cells (HONE-1, C666-1, and 5-8F). (B) Semi-quantitative RT-PCR confirmed the knockdown of PARPBP in NPC cells (HONE-1 and C666-1) transfected with PARPBP-specific siRNA (si#1, si#2). (C) PARPBP knockdown NPC cells exhibits a drastic reduction in cell viability. (D) Colony formation assays with NPC cells (HONE-1 and C666-1) transfected with PARPBP-specific siRNA (si#1, si#2) or control siEGFP for 10 days. (E) The efficiency of knockdown by lentiviral delivery of shRNA is assayed by semi-quantitative RT-PCR. (F) The proliferation of HONE-1 cells stably expressing shNC or shPARPBP is measured. (G) Colony formation assays with HONE-1 cells stably expressing shNC or shPARPBP. Statistical results of colony formation results are presented. Mean ± SD are shown. *P < .05.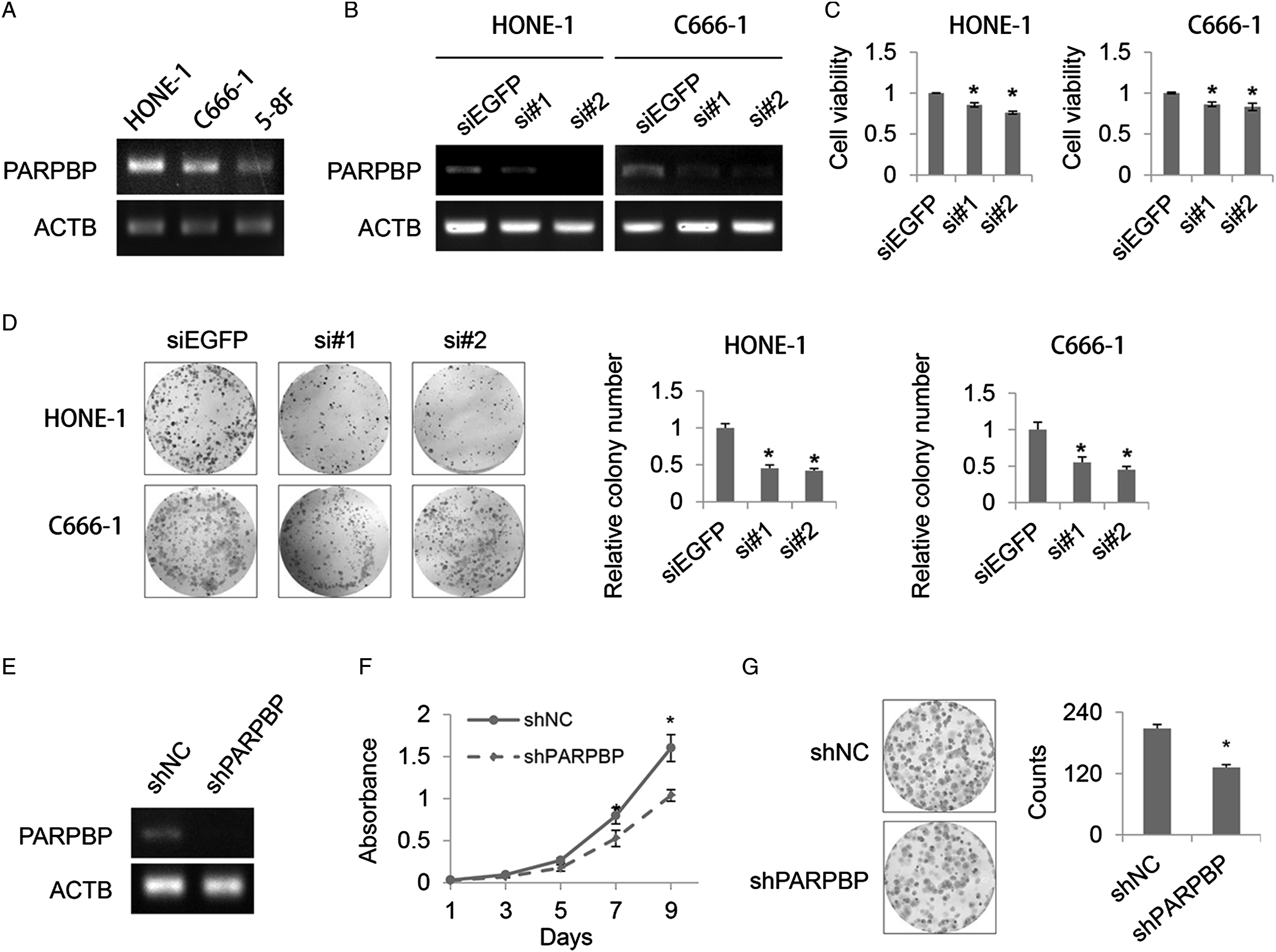
Identification of downstream genes of PARPBP. To understand the involvement of PARPBP in NPC, we conducted RNA sequencing (RNA-seq) gene expression analysis to identify genes and pathways associated with PARPBP in NPC cells on a genome-wide scale. HONE-1 cells stably expressing shRNA silencing PARPBP or control shNC were used to perform RNA-seq analysis. Totally, hundreds of DEGs including 377 downregulated genes and 518 upregulated genes in HONE-1 cells with stably expressing shPARPBP were found according to RNA-seq with the threshold of P < .05 and a fold-change≥2 as compared with the control (Figure 3A and B). Of those, the top 10 upregulated and downregulated genes were listed in Figure 3C and 3D. The DEGs matched for GO annotation were assigned to three main categories, including molecular function, cellular component, and biological process, and the subgroups identified with a significant adjusted P value less than 0.05 were ranked by gene numbers (Figure 3E). Protein binding (GO:0005515), insulin-like growth factor binding (GO:0005520), and MAP kinase tyrosine/serine/threonine phosphatase activity (GO:0017017) were the major enriched subgroups of molecular function. As for the cellular component, cytoplasm (GO:0005737), plasma membrane (GO:0005886), and cytosol (GO:0005829) were the most enriched subgroups. Positive regulation of transcription from RNA polymerase II promoter (GO:0045944), positive regulation of endothelial cell proliferation (GO:0001938), positive regulation of smooth muscle cell proliferation (GO:0048661), and mRNA splice site selection (GO:0006376) were significantly enriched in biological process category (Figure 3E). Next, we used scripts in house to enrich significant DEGs in KEGG pathways. Through enrichment analysis, the most significantly enriched pathways were identified as MAPK signaling pathway (ko04010), IL-17 signaling pathway (ko04657), and salmonella infection (ko05132) (Figure 3F). In addition, these significantly enriched pathways were associated with environmental information processing, organismal systems, human diseases, metabolism, and cellular processes. Taken together, these findings indicated the intriguing possibility of the involvement of PARPBP in NPC. Downstream genes of PARPBP are identified through RNA-seq. (A). Heatmap depicting 895 DEGs identified by genome-wide RNA sequencing (RNA-seq). (B) The volcano plot of all DEGs based on an adjusted P<0.05 and log FC≥ 1. The top 10 downregulated (C) and upregulated (D) genes upon knockdown of PARPBP in HONE-1 cells are listed. (E)Analysis of Gene ontology (GO) term enrichment of the main category of genes after PARPBP depletion. (F) KEGG pathway enrichment bubble chart of DEGs.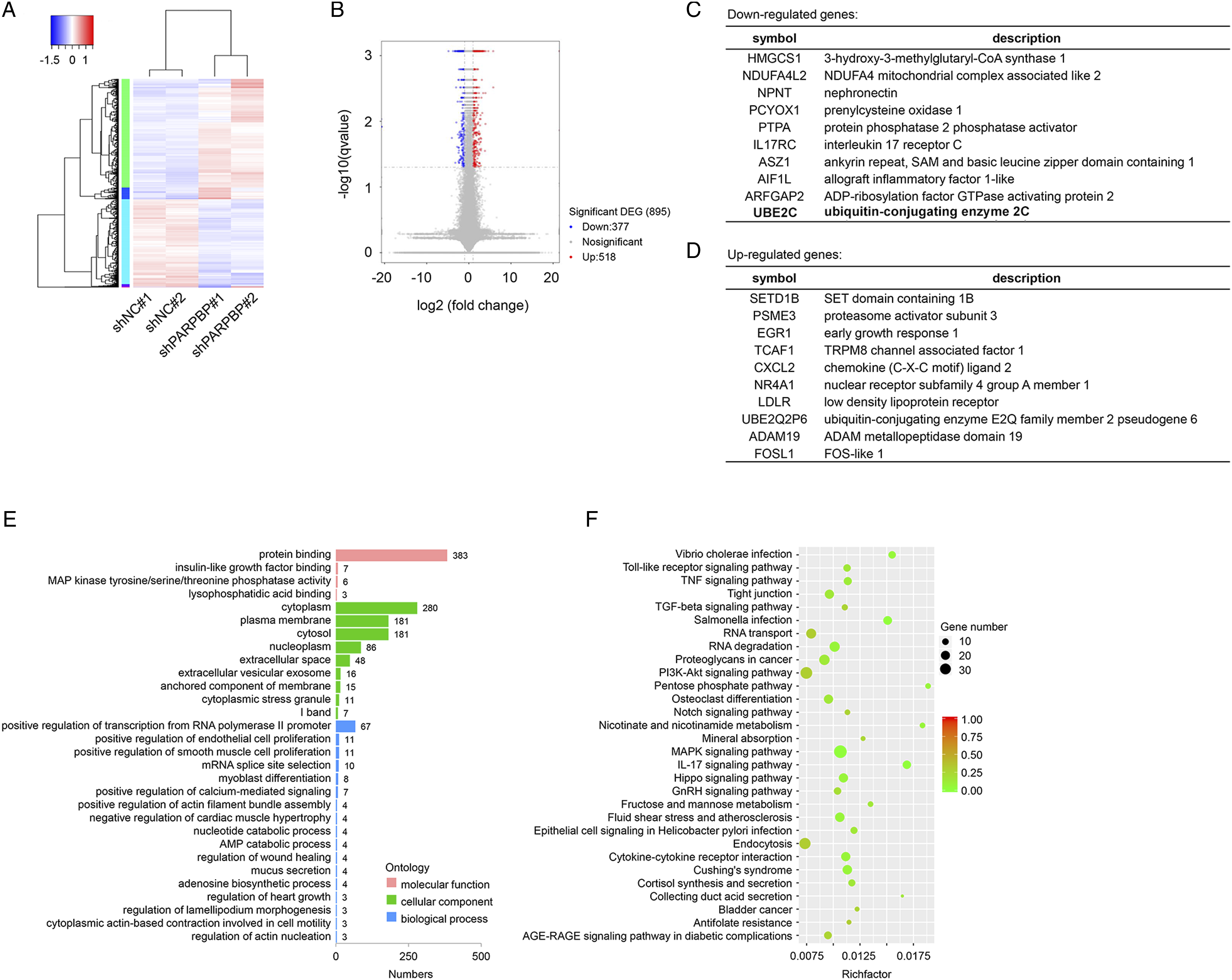
PARPBP promotes cell invasion and migration in NPC partly through regulating UBE2C. One of the most downregulated genes upon knockdown of PARPBP was UBE2C which possessed oncogenic roles in various cancers.18,24 Through analyzing GSE12452 dataset, we first found that UBE2C was positively correlated with PARPBP mRNA expression in NPC samples (Figure 4A, r=0.7517 with P < .0001) and also upregulated in NPC (Figure 4B). In addition, the protein level of UBE2C was notably decreased upon knockdown of PARPBP (Figure 4C). Previous studies report that UBE2C is closely involved in EMT process.16,17,25 Therefore, the levels of mesenchymal markers including N-Cadherin and Vimentin were examined. Notably, both N-Cadherin and Vimentin exhibited a decrease in PARPBP stably knockdown HONE-1 cells as compared to control shNC (Figure 4C). Next, we found that silencing PARPBP impeded cell migratory and invasive potential in HONE-1 NPC cells (Figure 4D). These findings illustrated that PARPBP would be involved in cell migration and invasion of NPC cells partly through modulating UBE2C expression. PARPBP regulates UBE2C expression and facilitates the migration and invasion of NPC cells. (A) Correlation between PARPBP and UBE2C in mRNA levels. There is a highly positive correlation between PARPBP and UBE2C within GSE12452 dataset (n = 31). R represents Spearman’s rho correlation coefficient. (B) UBE2C mRNA level is evaluated in dataset GSE12452 (left panel) and GSE13597 (right panel). (C) The levels of UBE2C as well as mesenchymal markers are decreased upon silencing PARPBP. The expression of UBE2C, N-cadherin, and Vimentin is confirmed by western blot. The β-actin is loaded as control. (D) Migration and invasion assays are conducted in shNC- and shPARPBP-stable HONE-1 cells and representative images are presented. Mean ± SD are shown. *P < .05.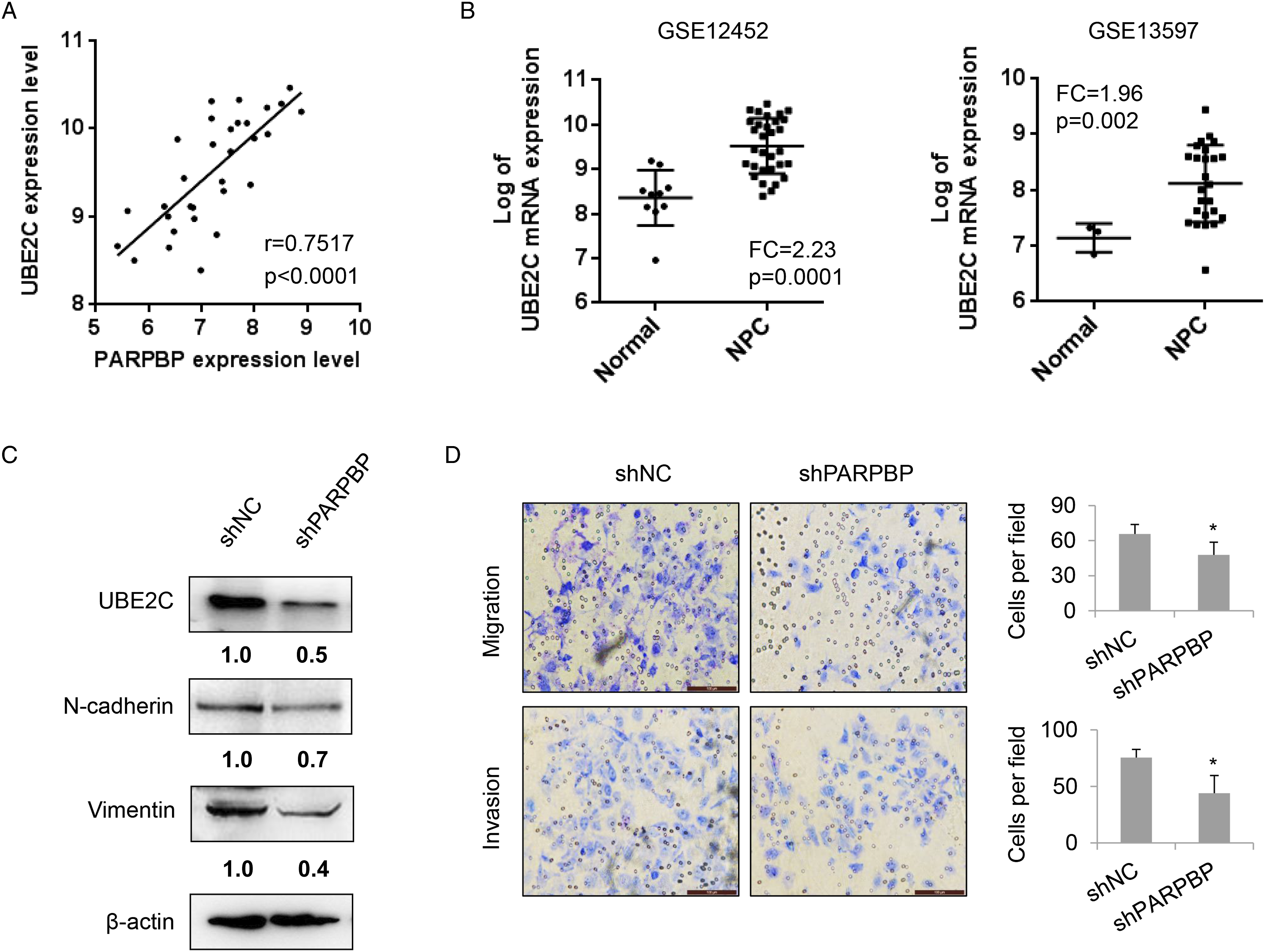
Discussion
Nasopharyngeal carcinoma (NPC) is one of the most prevalently diagnosed head and neck cancer in Southeast Asia and South China. At present, overall 5-year survival for NPC is 50–70%, and surgery, radiotherapy, and concurrent cisplatin or 5-fluorouracil-based chemoradiotherapy are the main treatment option. The mechanism of pathogenesis in NPC remains largely unknown and over 70% of NPC patients have locally advanced disease at the initial diagnosis and prone to disease recurrence and poor survival. 26 Therefore, it is urgent to identify genes critical to the development and progression of NPC.
In this study, through analyzing gene expression data from GEO series GSE12452, GSE13597, and GSE53819, we found a gene PARPBP which is commonly upregulated in NPC tissues. Deletion of PARPBP in NPC cells led to a marked reduction in cancer cell viability and proliferation, implicating its important functions in nasopharyngeal carcinogenesis. To our knowledge, this is the first report argued the involvement of PARPBP in NPC. In addition, enhanced expression of PARPBP is also reported in pancreatic cancer, gastric cancer, and leukemia.10,12,13,27 Importantly, the expression of PARPBP is hardly detectable in other normal adult organs except the testis, implying PARPBP to be a potent targetable molecule in multiple cancers including NPC. 10
PARPBP is encoded by a novel gene C12orf48 (chromosome 12 open reading frame 48) and contains an UvrD helicase-related domain. Although its molecular function is not well-characterized, previous studies imply that PARPBP might play important roles in genomic stability. AROM, a mouse homologue of PARPBP, is reported to have high binding affinity to single-stranded DNA and poly(A) homopolymers indicating a role in mRNA maturation/metabolism. 28 Previously, we reported PARPBP could bind to PARP-1, an essential protein in DNA repair pathway, and enhance its activity. 10 Through interacting with PCNA, PARPBP interferes the formation of RAD51-DNA homologous recombination (HR) structures.11,12 PARPBP deletion also suppresses the UV sensitivity of RAD18-silenced cells, indicating PARPBP as a negative regulator of HR, 8 and enhances replication stress and DNA-damage accumulation. These studies illustrate PARPBP is associated with the process of DNA repair pathway that is usually abnormal in therapy-resistant cancer. Indeed, elevated PARPBP is detected in some therapy-resistant malignancies, including relapsed small-cell lung cancer, cholangiocarcinoma, and castration-resistant prostate cancer.29,30 Currently, little is known about the regulatory mechanism of PARPBP in NPC. Thus, we perform RNA-seq analysis with PARPBP stably knockdown NPC cells to understand its associated genes and pathways in NPC. Totally, 377 down-regulated genes and 518 upregulated genes after PARPBP deletion were identified. These DEGs were employed to KEGG pathway. Accordingly, we revealed 42 PARPBP-associated pathways with significant enrichment, including MAPK signaling pathway (ko04010), IL-17 signaling pathway (ko04657), and salmonella infection (ko05132). Numerous well-known genes involved in MAPK signaling pathway such as MECOM, ERBB3, EFNA1, and VEGFA were downregulated, whereas NR4A1, FOS, JUN, and MET were upregulated upon PARPBP deletion. Further studies are required to explore the underlying mechanism.
Of these downregulated genes, we first revealed that an ubiquitin-conjugating enzyme UBE2C was regulated by PARPBP, and moreover, it is positively correlated with PARPBP in NPC tissue samples. Importantly, deletion of PARPBP decreased the levels of N-Cadherin and Vimentin, and in turn, impeded cell migratory and invasive potential of NPC cells partly through decreasing the expression of UBE2C. However, the molecular mechanisms by which PARPBP regulates UBE2C are still unknown and required to be explored.
Our findings illustrated that PARPBP possessed oncogenic roles in NPC, especially it was closely involved in cancer cell proliferation, migration, and invasion which provided some evidence in support of the hypothesis that PARPBP might be a targetable molecule for the treatment of NPC.
Footnotes
Authors’ contributions
Conceptualization: Li Ji
Data curation: Lianhua Piao
Formal analysis: Li Ji
Funding acquisition: Jian Wu
Investigation: Lianhua Piao
Methodology: Lianhua Piao
Project administration: Jian Wu and Qingquan Hua
Resources:Jian Wu
Software: Xiaofeng Gu
Supervision: Jian Wu,Qingquan Hua
Validation: Li Ji
Visualization: Feng Xiao
Writing—original draft: Lianhua Piao and Li Ji
Writing—review and editing: Li Ji
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was supported by The National Natural Science Foundation of China (grant no.81903661).
Ethical Approval
This study was approved by ethics committee of The Second People’s Hospital of Changzhou affiliated to Nanjing Medical University.
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
