Abstract
Objectives:
To analyze factors regarding patient characteristics, sampling techniques, and coronavirus disease 2019 (COVID-19) specific manifestations that may cause false-negative reverse-transcription polymerase chain reaction (RT-PCR).
Materials and Methods:
A cross-sectional study was conducted. For the diagnosis of COVID-19, patients with RT-PCR test positive in the first nasopharyngeal and oropharyngeal swabs were accepted as true positive, and patients with negative 3 consecutive swab results were considered true negative. Those who had a negative initial swab were considered false negatives if they subsequently tested positive on the second or third swab. Demographic data of the patients, the onset of the disease, presence of nasal septal deviation, presence of epistaxis, the clinician (otolaryngologist/other physicians [OP]) who collected the samples, and medical treatments for laryngopharyngeal reflux, allergic respiratory diseases, allergic rhinitis, which include proton pump inhibitors and nasal steroids (NS), were documented. The analysis of dependent variables was performed with the chi-square test. Binary logistic regression was performed for significant variables.
Results:
A total of 399 patients were included in the study, and 357 (89.5%) patients were detected as positive after 2 or 3 consecutive RT-PCR tests. The presence of ageusia, anosmia, and collecting the samples within 7 days following the onset of symptoms were determined as significant factors for positive RT-PCR results (
Conclusions:
Patient- and/or sample-related factors can affect RT-PCR results of possible COVID-19 cases. The presence of these factors can easily be determined in cases with high clinical suspicion and negative RT-PCR results. The presence of ageusia, anosmia, early sampling (<7days), and appropriate collection of swabs decrease false-negative RT-PCR results.
Introduction
SARS-CoV-2 is the coronavirus responsible for the novel coronavirus disease 2019 (COVID-19) pandemic. 1 The exact diagnosis is made by detecting the genetic material of the virus by reverse-transcription polymerase chain reaction (RT-PCR) in nasopharyngeal (NP) and pharyngeal swabs. The sensitivity of the RT-PCR test is closely related to the sample.2,3 The collection time of swabs, collection technique, and patient characteristics can affect the results of RT-PCR tests. 3 Therefore, clinical findings combined with the patient’s medical history—when supported by computed tomography findings—were found to be more sensitive when compared to RT-PCR. 2
The mucosal physiology of the nasopharynx can be affected by diseases such as laryngopharyngeal reflux (LPR), allergic/inflammatory respiratory diseases, local or systemic medications, and tobacco use. Laryngopharyngeal reflux is a chronic disease that can cause chronic pathological changes in the pharynx and nasopharynx.4,5 The proton pump inhibitors (PPIs) are the main treatment options for LPR 6 . Similarly, allergic rhinitis (AR) is a common condition in the population that originates from the nasal mucosa, and nasal steroids (NS) are often preferred in treatment.7,8 Recent studies documented that the immune response and microbial flora of allergic mucosa differ from that of healthy individuals. 9 It was also reported that local budesonide treatment (a local steroid agent used in AR and asthma treatment) inhibits rhinovirus replication and stimulates cytokine production in primary cultures of human tracheal epithelial cells. 10 Anatomical pathologies of the nasal cavities such as nasal septum deviation (NSD) can also diminish proper sampling by blocking access to the nasopharynx. Therefore, RT-PCR tests of NP swabs must comply with some preconditions in order to be accepted as reliable.
The main objective of the current study is to analyze the possible effect of the above-mentioned factors on outcomes of RT-PCR tests of probable cases with COVID-19. We also sought to evaluate if the presence of any of these factors correlates with false-negative RT-PCR tests.
Materials and Methods
The study was conducted in a secondary referral state hospital between March 2020 and July 2020. Verbal consent was obtained from all participating patients. The study was carried out in accordance with the 1964 Helsinki Declaration and subsequent amendments. The research protocol was approved by the provincial health directorate.
Consecutive patients with medical history, symptoms, clinical and laboratory findings that were compatible with COVID-19 were evaluated for the study. Only patients with a definite diagnosis of COVID-19 by 1 positive test of 3 consecutive RT-PCR tests were included. Data regarding swab-related variables were collected during the first sampling.
The demographic characteristics of the patients, medical data including symptoms, otolaryngological and systemic examination findings, comorbidities and chronic diseases, and medical treatments were recorded. The query was detailed for the presence of LPR, AR, and regular medications including NS and PPI. Cotton-tipped plastic swabs were used to collect both oropharyngeal and nasopharyngeal (OP/NP) samples. The OP/NP swabs were collected with 1 applicator: First, an OP sample was collected, then the same applicator was inserted into 1 nostril for the NP sample. Results of RT-PCR (Bosphore Novel Coronavirus 2019-nCoV Detection Kit) were determined as positive or negative according to the manufacturer protocol. Clinicians who collected the samples were grouped as otolaryngologists (ORL) and other physicians. The presence of epistaxis during NP sampling was recorded. Patients were grouped into 2 groups according to the sampling time following the onset of the symptoms (≤7 days or > 7 days).
Three consecutive RT-PCR tests were applied as an institutional protocol to provide treatment and isolation in cases with a high degree of COVID-19 infection suspected by clinical and laboratory findings. So, for the diagnosis of COVID-19, patients with RT-PCR test positive in the initial NP/OP swabs were accepted as true positive, and patients with negative 3 consecutive swab results were considered true negative. Those who had a negative initial swab were considered false negatives if they subsequently tested positive on the second or third swab. The second and third samples were collected 24 hours after the previous negative test. Patients who had 3 consecutive negative tests were excluded from the study. Besides, immunocompromised patients, patients with bleeding diathesis, anticoagulant drug use, and patients with a history of hyposmia and hypogeusia were excluded. During the nasal examination, patients with a total nasal obstruction due to severe NSD, turbinate hypertrophy, or nasal polyposis; patients with telangiectatic or crusted nasal mucosa; and patients younger than 18 years old were also eliminated.
SPSS 22.0 (IBM Corp) was used for statistical analysis. The frequency of symptoms and categorical variables were calculated. The analysis of categorical variables was performed with the chi-square test, which was followed by binary logistic regression analysis for significant variables.
Results
A total of 399 consecutive patients were evaluated for the study. Forty-two (10.5%) patients were excluded due to 3 consecutive negative tests. Three hundred fifty-seven patients were included in the final analysis. One hundred ninety-nine (55.7%) patients were females and 158 (44.3%) were males. The mean age was 52.1 ± 21.6 years. The mean duration between the onset of the symptoms and the admission was 7.9 ± 3.7 days. The most common symptoms regarding the head and neck region were anosmia (n = 157, 43.7%) and ageusia (n = 149, 41.5%). The RT-PCR samples were collected after an average of 7.9 ± 3.7 days from the onset of the symptoms. The first RT-PCR test was positive in 183 (51.3%) patients. The presence of ageusia and anosmia was detected as a significant indicator of positive RT-PCR results (
Distribution of Complaints According to PCR Test Results and
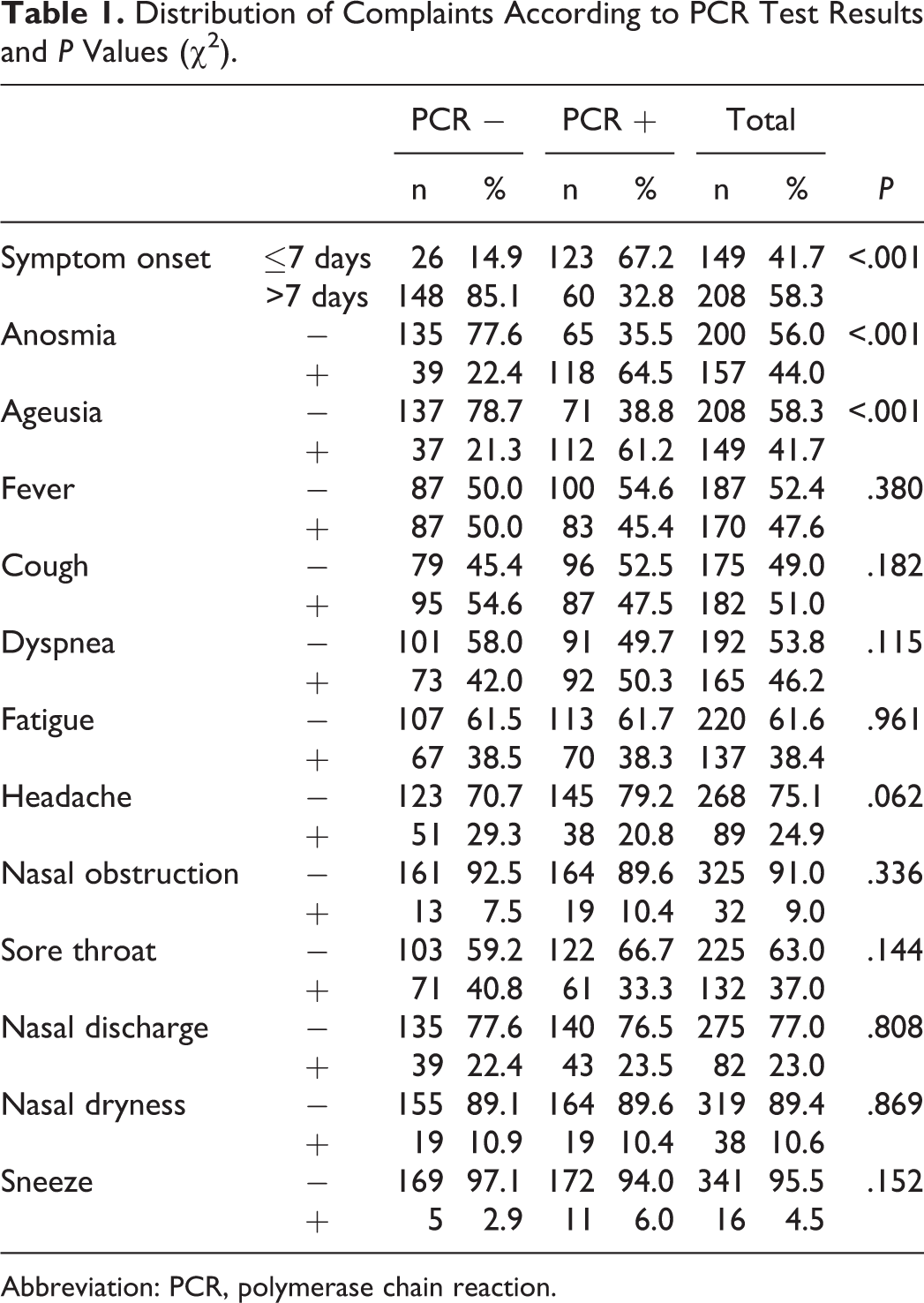
Abbreviation: PCR, polymerase chain reaction.
The distribution of patient- and swab-related characteristics of RT-PCR positive and negative groups is presented in Table 2. The profession of the clinician (other physicians), NS use, and presence of epistaxis during sampling were detected as significant factors for negative RT-PCR results (
Distribution of Patient- and Sample-Related Variables According to PCR Test Results and
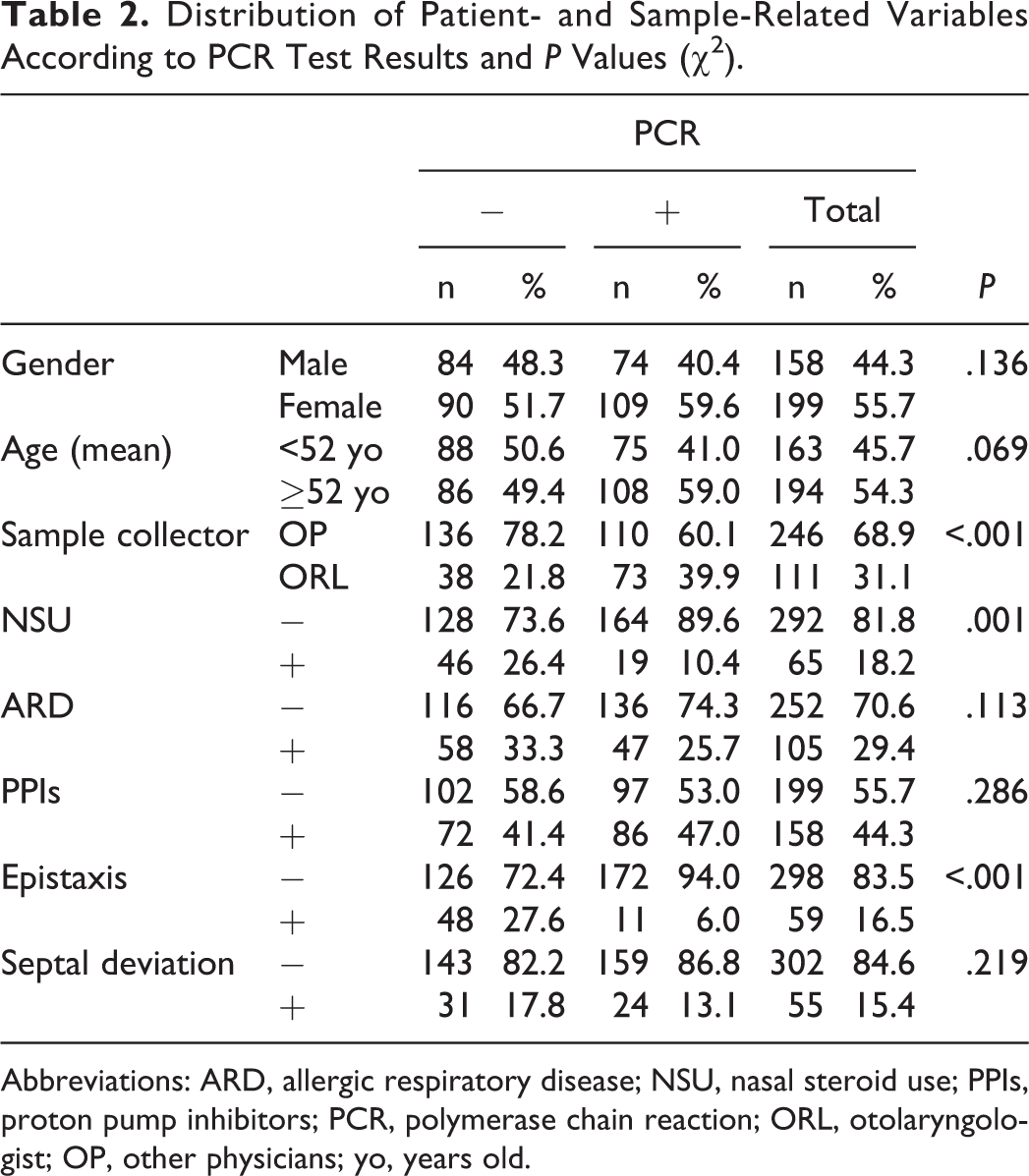
Abbreviations: ARD, allergic respiratory disease; NSU, nasal steroid use; PPIs, proton pump inhibitors; PCR, polymerase chain reaction; ORL, otolaryngologist; OP, other physicians; yo, years old.
Logistic Regression Analysis of Significant Variables for PCR Results in Univariate Analysis.a
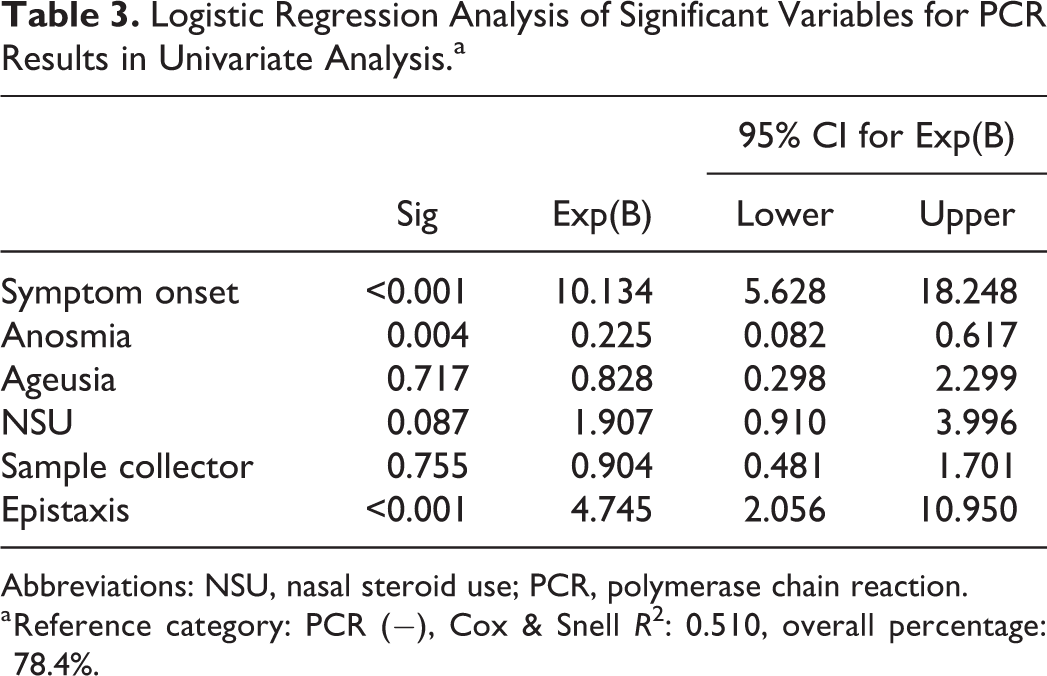
Abbreviations: NSU, nasal steroid use; PCR, polymerase chain reaction.
a Reference category: PCR (−), Cox & Snell
Discussion
False-negative RT-PCR tests cause problems in the diagnosis and early isolation of a patient with COVID-19 and can also mislead the clinician during the follow-up period. 11 The definite diagnosis of COVID-19 is made by RT-PCR tests, which detect SARS-CoV-2 viral RNA in pharyngeal or NP swabs or sputum, urine, and stool samples. 12 The virus dissemination was found to be high during the first week of the symptoms and to decline toward the end of the first week. 12 In addition, the incubation period of the virus is between 3 and 14 days.13,14 Therefore, the exact time of virus transmission to the patient, the onset time of symptoms, and the sampling time affect the results of the RT-PCR tests. Wölfel et al 12 investigated various specimens from patients who had close contact with patients diagnosed with COVID-19. They detected viral RNA in all of the OP or NP swabs collected from 9 patients in the first to the fifth day after the onset of symptoms. The authors reported that the initial symptoms were flu-like symptoms (66.6%; n = 6) and odor disorder (44%; n = 4). Wang et al 15 reported that the SARS-CoV-2 RNA was detected only in 32% of pharyngeal swabs (126/398) and in 63% of NP swabs (5/8). However, this report does not mention the collection time of the samples or the onset of symptoms. The current opinion in the literature is that the viral load in the upper respiratory tract is maximum in the first week after the onset of symptoms. Therefore, OP/NP swabs have the highest positivity rate in this period.16,17 Similar to the literature, we observed that the RT-PCR positivity rate was significantly higher (67.2% vs 32.8%) in patients with onset of symptoms less than 1 week. Thus, repeated RT-PCR tests should be recommended for patients who have symptoms for more than 1 week.
Pharyngeal and NP samplings are easy and practical procedures. However, anatomical pathologies such as NSD or turbinate hypertrophy may complicate the procedure. If the clinician collecting the samples lacks anatomical knowledge of the nasal cavity, the swabs may not be pushed forward properly through the nasal cavity to the nasopharynx and appropriate samples cannot be collected. Furthermore, the procedure can become painful and complications such as epistaxis may occur. 18 In the current study, the rate of ORL specialists as sample collectors was found to be significantly higher in the RT-PCR positive group (39.9% vs 21.8%). The NSD rate was higher in the RT-PCR negative group, but the difference was not statistically significant (17.8% vs 13.1%). The presence of epistaxis during sampling can be accepted as an indicator of insufficient/inappropriate sampling. We observed that the frequency of epistaxis was significantly higher in the RT-PCR negative group (27.6% vs 6.0%). Besides, 54 (91.6%) samples with epistaxis were collected by clinicians except for ORL, supporting previous findings. The occurrence of epistaxis during OP/NP sampling can increase the anxiety of the patient and the clinician, decrease patient compliance, and can also cause early termination of the procedure.
Nasal steroids are frequently preferred topical agents for the treatment of AR.7,19 They increase the recruitment of inflammatory cells to the airway mucosa, selectively suppress local cytokine expression, inhibit the release of mediators, and support the normal structure of the mucosa. 20 Ogimi et al 21 demonstrated a significant relationship between prolonged shedding of human coronavirus (HCoV) and the high-dose use of steroids in cases with hematopoietic cell transplants. Waltl et al 22 reported that betamethasone could decrease the epithelial damage caused by rhinovirus infection. Kim et al 23 showed the in vitro antiviral activity of budesonide, which inhibits the human rhinovirus replication by activation of autophagy. The authors recommended budesonide as a therapeutic option. Yamaya et al 10 demonstrated that budesonide reduced rhinovirus count, replication, and concentrations of cytokines of primary cultures of human tracheal epithelial cells. In our study, there was a significant difference in rates of patients with NS use between RT-PCR (−) and (+) groups (26.4% vs 10.4%), which may be explained by the antiviral effects of steroids indicated by the literature. The different types of NS and dosages, different durations of medications, and also the absence of in vivo and histopathological evidence are important limitations of the current study. Also, turbinate hypertrophy is another indication of NS use, and excluding patients with severe turbinate hypertrophy may have an impact on results by reducing the number of patients in the NS group. Yet, few studies investigate the effects of NS on HCoV infections in the literature, whereas no SARS-CoV-2 related investigations are reported. It is not recommended to stop the use of NS in patients diagnosed with COVID-19, but the effect of NS on viral replication is not clear. 24
Hyposmia and hypogeusia were reported as significant indicators for positive RT-PCR results. 25 We also observed that the RT-PCR positivity rates were significantly higher in cases with hyposmia and hypogeusia (OR = 6.2 and 5.8, respectively). Questioning the presence of these complaints while collecting OP/NP samples can provide accurate predictions regarding RT-PCR test results. The absence of an objective assessment of hyposmia or hypogeusia is also a limitation of our study.
Mucosal changes caused by LPR are well described in the literature. 4 Sella et al 26 revealed that LPR had a pathological effect on the nasal mucosa of patients with chronic rhinosinusitis. However, the effects of LPR on viral infections of the respiratory tract are not clear. In this study, the frequency of PPI use in the RT-PCR (+) group was found to be slightly higher, but the difference is not significant. The effects of LPR on the OP or NP mucosal barrier are already depicted, but histopathological studies with large sample groups can clarify the effect of LPR on RT-PCR results.
In this study, significant factors for false-negative RT-PCR results of cases with COVID-19 are presented. The presence of these patient- and/or sample-related factors should be questioned in cases with high suspicion of COVID-19 infection but negative RT-PCR results. Larger study groups are required to predict the prevalence and effect of these factors more accurately. Furthermore, the possible effects of NS on viral replication should be demonstrated in vivo.
Conclusions
Detecting viral RNA in the NP/OP swabs is the gold standard method for the definitive diagnosis of COVID-19. However, the presence of some factors, which are documented in this study, can affect RT-PCR results. The presence of these factors can easily be determined from the anamnesis and should be kept in mind in cases with high clinical suspicion and negative RT-PCR results. If possible, a consultation from an ORL may be requested for cases with difficult sampling. A short-term education and training of clinicians by an ORL can also be recommended.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
