Abstract
Nasal polyposis is a disease characterized with chronic inflammation of the nasal mucosa. Toll-like receptors (TLRs) are defined as essential receptors of the innate immune system and may play in the development of nasal polyposis. A total of 71 patients with nasal polyposis and 74 healthy controls were included in this study. Three single-nucleotide polymorphisms (SNPs); TLR2 (2258 A>G), TLR4 (896 A>G), and TLR4 (1196 C>T) were analyzed in all patients. The degree of pair-wise linkage disequilibrium and the genotype and haplotype analyses were conducted using regression in this logistic model and the Multifactor Dimensionality Reduction (MDR) software package was used to construct all possible interactions among different genotype variants belonging to the TLR gene. There was significant difference in genotype and allele frequencies of the TLR4 (1196 C>T) polymorphism between the nasal polyposis and control groups (0.017). Also, it was observed that the probability of nasal polyposis was 62.7% in the presence of TLR4 (1196 C>T) polymorphism with asthma (P = .007). As a conclusion, this study showed that TLR4 and TLR2 polymorphisms were predisposing factors for nasal polyposis. Further functional studies investigating the consequences of loss of TLR function are needed.
Introduction
Nasal polyposis is characterized with chronic inflammation of the nasal mucosa and pathophysiological mechanism of the disease is not completely clear. Nasal polyposis is characterized with typical symptoms such as nasal obstruction, nasal discharge, and hyposmia, which significantly affect daily life of patients. 1 Despite aggressive medical treatments and endoscopic sinus surgery, disease control cannot be achieved in many patients and has high recurrence rates. 2
The epithelial immune system plays an important role in the removal of molecules like dangerous bacteria, fungi, and viruses from the environment. Toll-like receptors (TLRs) are defined as essential receptors of the innate immune system. Activation of TLR-dependent signaling pathways results in the expression of cytokines and chemokines, and in this way immune response is generated. 3 Toll-like receptor 2, TLR-3, and TLR-4 have been implicated in the initiation of nasal mucosal epithelial immunological response. 4 Toll-like receptor 2 has the most ligand diversity and recognizes gram-positive and gram-negative bacteria with a wide variety. 5 Toll-like receptor 4 pathway is crucial for gram-negative bacterial infections in the airway and TLR-4 polymorphism has been associated with gram-negative bacterial infections in intensive care units. 6 The immunity of the nasal mucosa can play an important role in the development of nasal polyposis by developing an inappropriate inflammatory response.
Three single-nucleotide polymorphisms (SNPs); TLR2 (2258 A>G) (corresponding to an Arg753Gln substitution mutation; SNP database [dbSNP] accession number rs5743708), TLR4 (896 A>G) (corresponding to an Asp299Gly substitution mutation; dbSNP accession number rs4986790), and TLR4 (1196 C>T) (corresponding to a Thr399Ile substitution mutation; dbSNP accession number rs4986791) are associated with receptor unresponsiveness. 7 However, the association of these polymorphisms with nasal polyposis has not been fully elucidated. The aim of this study is to investigate whether there is an association between presence of TLR2 (2258 A>G), TLR4 (896 A>G), and TLR4 (1196 C>T) polymorphisms and nasal polyposis development.
Patients and Methods
Study Group
Seventy-one nasal polyposis patients and 74 preoperative septoplasty patients as control group were included in the study in the period of March 2017 to June 2017. This study was approved by Yıldırım Beyazıt University Yenimahalle Training and Research Hospital Institutional Ethics Committee (approval number: 2017/02). The informed consent was obtained from all participants.
Nasal polyposis was diagnosed according to EPOS 2012 criteria by endoscopic nasal examination and paranasal computerized tomography. 1 Control group was consisting of 74 preoperative septoplasty patients who underwent paranasal computed tomography (CT) because of preoperative evaluation. Patients who were included in the control group were examined for nasal mucosal disease and patients who have normal nasal mucosa were included in the control group. Patients who have chronic rhinosinusitis or history of rhinosinusitis were excluded from the control group. Exclusion criteria of this study were personal and family history of immune deficiency, diseases with known immune-related etiology, purulent nasal infection, cranio-facial abnormalities, and congenital defects.
Systemic and local drugs were recorded, and patients were ordered to discontinue both systemic and local drugs 2 weeks before endoscopic examination, CT scan and blood sampling. Patients were recommended to use nasal irrigation with isotonic saline at low pressure and large volume for these 2 weeks. After 2 weeks, CT scan and blood samples were obtained after being assured that there is no infection by endoscopic examination. The paranasal sinus CT scans in nasal polyposis group were obtained in the noninfected state of each patient. Polyp sizes were quantified endoscopically according to the Lildholdt classification in the noninfected state of each patient; 1: polyps only in the middle meatus; 2: polyps that reached the upper surface of the inferior turbinate; and 3: polyps that completely obstructed the nasal cavity.
All patients were evaluated for probable asthma by pulmonology department. Patients were questioned about past medical history, underwent detailed respiratory examination by pulmonologist, and asthma diagnosis was confirmed with pulmonary function tests. Patients whose asthma diagnosis was confirmed by pulmonology department were accepted as asthmatic. Patients having aspirin reaction history were evaluated by pulmonologist and underwent aspirin provocation test; patients with positive aspirin provocation test were diagnosed as having aspirin intolerance.
Total serum immunoglobulin E (IgE) concentration was determined with the nephelometric assays method. Skin prick tests were performed according to the European Academy of Allergy and Clinical Immunology using 18 allergens (ALK Abello, Madrid, Spain) in all participants. Skin prick tests were considered positive if at least one allergen elicited a wheal reaction >3 mm in diameter after subtraction of the diameter of the wheal produced by the negative control. The patient was considered atopic if she/he had at least one positive skin prick test result.
Detection of Genetic Variations of the Toll-Like Receptors
Blood obtained in EDTA containing tubes from each participant. A standard kit (NucleoSpin blood DNA, Macherey-Nagel GmbH & Co. Kg, Düren, Germany) was used according to the manufacturer’s protocols for genomic DNA extraction from peripheral blood samples. Detection of the polymorphisms was performed by using allele-specific polymerase chain reaction (PCR), followed by restriction fragment length polymorphism (PCR-RFLP) analysis. All primers used for PCR-RFLP are summarized in Table 1. Primers for the detection of the TLR4-D299G SNP and TLR4-T399I SNP were forward 3′ and cut point 5′. The nucleotides in parentheses were modified and changed to the underlined ones.
The Primers Used for PCR-RFLP.
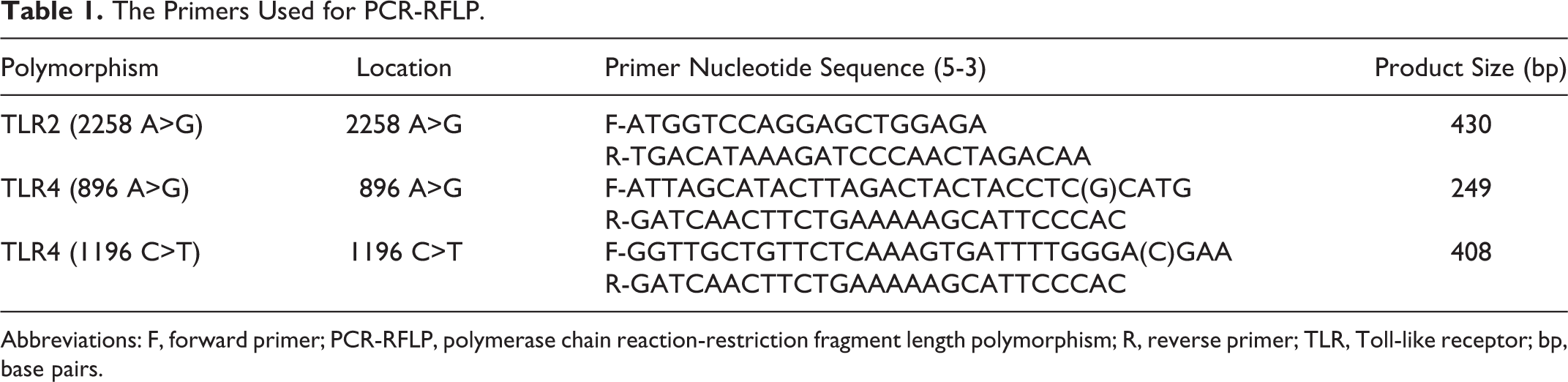
Abbreviations: F, forward primer; PCR-RFLP, polymerase chain reaction-restriction fragment length polymorphism; R, reverse primer; TLR, Toll-like receptor; bp, base pairs.
For detection of the TLR2-R753Q polymorphism, the protocol was designed on the basis of the fact that the polymorphism results in the creation of a DNA sequence recognized by the restriction enzyme SfcI (New England BioLabs, Ipswich, Massachusetts). 8 The PCR conditions were 2 minutes at 94°C, followed by 38 cycles of 94°C for 30 seconds, 64°C for 30 seconds, and 72°C for 45 seconds with 5 minutes at 72°C after the last cycle. The presence of undigested PCR products (430 bp) was indicative of wild-type samples, whereas the presence of the polymorphism resulted in the digestion of the PCR products to 307-bp and 123-bp fragments.
Molecular detection of the TLR4-D299G and TLR4-T399I SNPs was performed by PCR-RFLP. 9 In brief, the forward primers were modified at the 5′ end in 2 reactions, creating restriction enzyme recognition sites (NcoI for the TLR4-D299G polymorphism and HinfI for the TLR4-T399I polymorphism), so that if a polymorphism was present, PCR-RFLP analysis would create digestion fragments visible on agarose gels. 9 The conditions of both PCRs were 5 minutes at 94°C, followed by 38 cycles of 94°C for 30 seconds, 61°C for 30 seconds, and 72°C for 60 seconds, and 5 minutes at 72°C after the last cycle. For the TLR4-D299G SNP, a 249-bp fragment was amplified by PCR and subjected to NcoI digestion (Invitrogen, Carlsbad, California) overnight at 37°C. The presence of undigested PCR products was indicative of wild-type samples, whereas the presence of the polymorphism resulted in the digestion of the PCR products to 226-bp and 23-bp fragments. For the TLR4-T399I SNP, a 408-bp fragment was amplified by PCR and subjected to HinfI digestion (FastDigest; Thermo Fisher Scientific, Waltham, Massachusetts) for 4 hours at 37°C. The presence of undigested PCR products was indicative of wild-type samples, whereas the presence of the polymorphism resulted in the digestion of the PCR products to 378-bp and 29-bp fragments.
For all PCRs described here, a total of 2 µL (20-50 ng/µL) of DNA was amplified in a 25-µL reaction mixture using, 2.5 µL of 10X buffer (Complete; Bioron Inc, Ludwigshafen, Germany), 0.5 µL of 10 mM deoxynucleoside triphosphate, 0.5 µL of 20 pmol of each primer, 2.5 µL of 25 mM MgCl2, 0.2 µL (1.0 U) of Taq polymerase (Invitrogen, Carlsbad, California) and 16 µL of dH2O in a buffer supplied by the manufacturer. All PCR and digestion procedures were carried out in a PCR thermal cycler (T100 Thermal Cyler; Bio-Rad, Hercules, California), and the PCR and digestion products were analyzed in 2% tris-borate-EDTA agarose gels. For confirmation of the PCR-RFLP analysis results, randomly chosen PCR products positive and negative for the TLR polymorphisms were purified by use of a kit (NucleoFast 96 PCR; Macherey-Nagel GmbH & Co. Kg) and were directly sequenced using an ABI PRISM 3130 Genetic Analyzer (Applied Biosystems, Foster City, California) and a sequencing kit (BigDye Terminator V3.1 Cycle Sequencing; Applied Biosystems).
Statistical Analysis
Statistical analyses were performed using SPSS 16.0 (SPSS Inc, Chicago, Illinois). Student t test was used to compare age. The χ2 tests were used to assess the differences for sex distribution, the alleles, and genotypes. The frequency of each genotype was evaluated using SNPStats software (http://bioinfo.iconcologia.net/index.php?module=Snpstats) for concordance with Hardy-Weinberg equilibrium. 10 The degree of pair-wise linkage disequilibrium and the genotype and haplotype analyses were conducted using regression in this logistic model and expressed as the odds ratios and 95% confidence interval (95% CI). 11 The association between SNPs and a risk of nasal polyposis was estimated by calculating the ORs and 95% CIs. The Multifactor Dimensionality Reduction (MDR) software package (version 1.0.0, available at www.epistasis.org) was used to construct all possible interactions among different genotype variants belonging to the TLR gene. 12 Multifactor Dimensionality Reduction is a novel computational tool described by Hahn et al, and it is a nonparametric method (ie, no parameters are estimated). It can detect SNP X SNP and SNP X phenotype interactions. 13 P < .05 was considered statistically significant.
Results
Characteristics of Patients
A total of 71 patients with nasal polyposis and 74 healthy controls were included in this study. The demographic characteristics of nasal polyposis group and control group are shown in Table 2. The mean age of the nasal polyposis group was 40.28 (10.72) years and the mean age of the control group was 39.48 (10.40) years. There was no statistically significant difference between groups in terms of mean age of patients (P = .651). Of the nasal polyposis group, 37 (52.1%) were male and 34 (47.8%) were female. Of the control group, 44 (59.4%) were male and 30 (40.5%) were female. There was no difference in male/female ratio between the groups (P = .373). Asthma and aspirin intolerance were higher in nasal polyposis group than in the control group (P = .002, P = .016, respectively). There was no difference between groups in terms of atopy (P = .215).
Characteristics of the Study Population.a
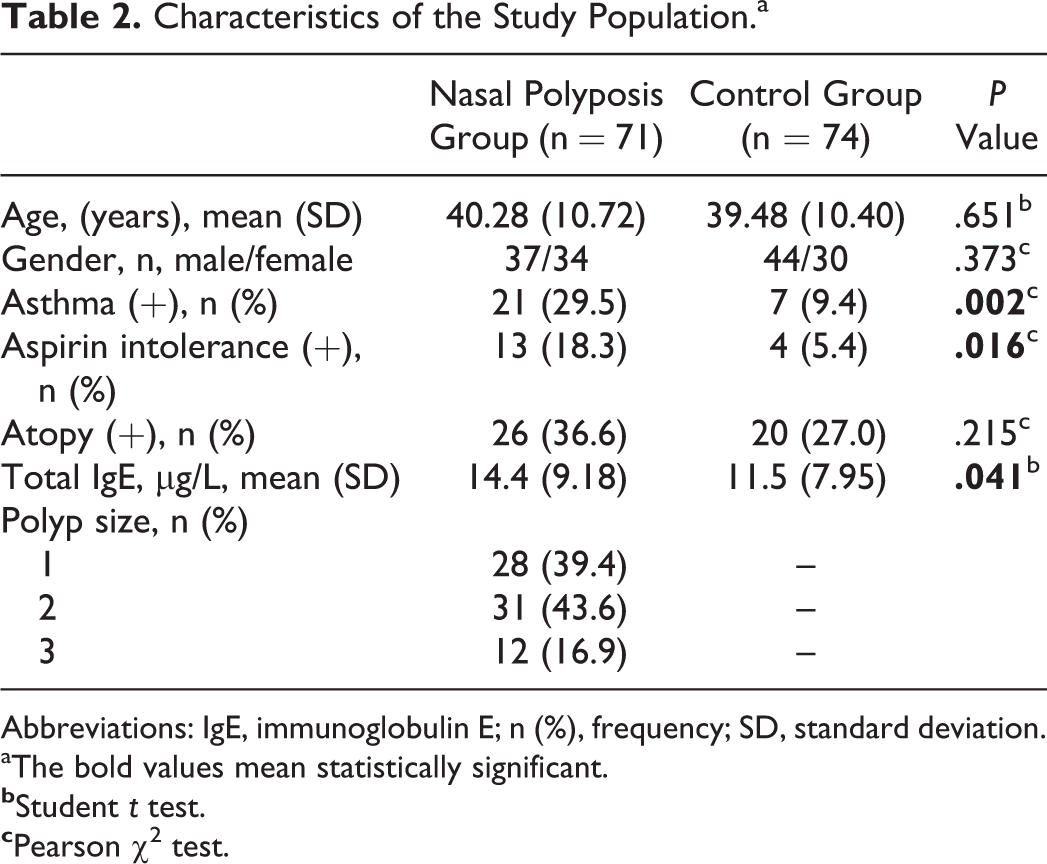
Abbreviations: IgE, immunoglobulin E; n (%), frequency; SD, standard deviation.
aThe bold values mean statistically significant.
Hardy-Weinberg Equilibrium
In the nasal polyposis and control groups, the genotypes in TLR2 (2258 A>G), TLR4 (896 A>G), and TLR4 (1196 C>T) polymorphisms preserved the H-W equilibrium (All P > .005; Table 3). 14 –17
The TLR Gene Polymorphisms.
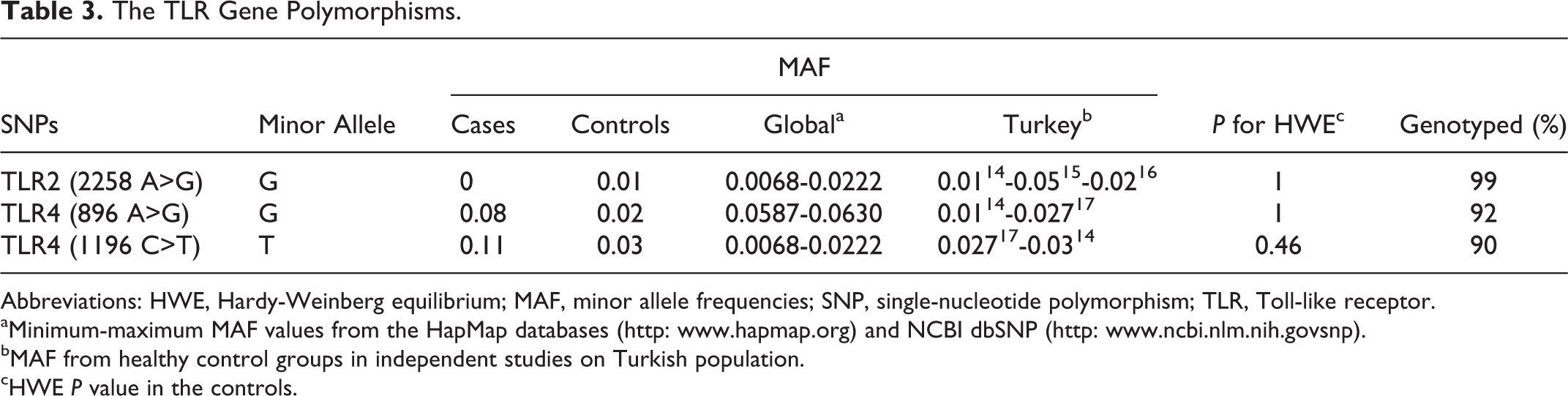
Abbreviations: HWE, Hardy-Weinberg equilibrium; MAF, minor allele frequencies; SNP, single-nucleotide polymorphism; TLR, Toll-like receptor.
aMinimum-maximum MAF values from the HapMap databases (http: www.hapmap.org) and NCBI dbSNP (http: www.ncbi.nlm.nih.govsnp).
bMAF from healthy control groups in independent studies on Turkish population.
cHWE P value in the controls.
Association of the TLR Genotypes With Nasal Polyposis
Genotype analysis revealed that TLR4 (1196 C>T) genotype was more frequent in nasal polyposis group than control group in the codominant, dominant, overdominant, and log-additive models (P = .033, P = .009, P = .036, P = .017, respectively). In other words, T-dominant all-inheritance models for TLR4 (1196 C>T; except for recessive TLR4 [1196 C>T]) were associated with a significantly increased risk of nasal polyposis (Table 4). When the minor allele frequency of the TLR4 (896 A>G) and TLR4 (1196 C>T) was considered, significant differences were observed between patients with nasal polyposis and the controls (P = .013, P = .017, respectively; Table 4).
Frequencies of TLR SNP Genotypes and Alleles.a
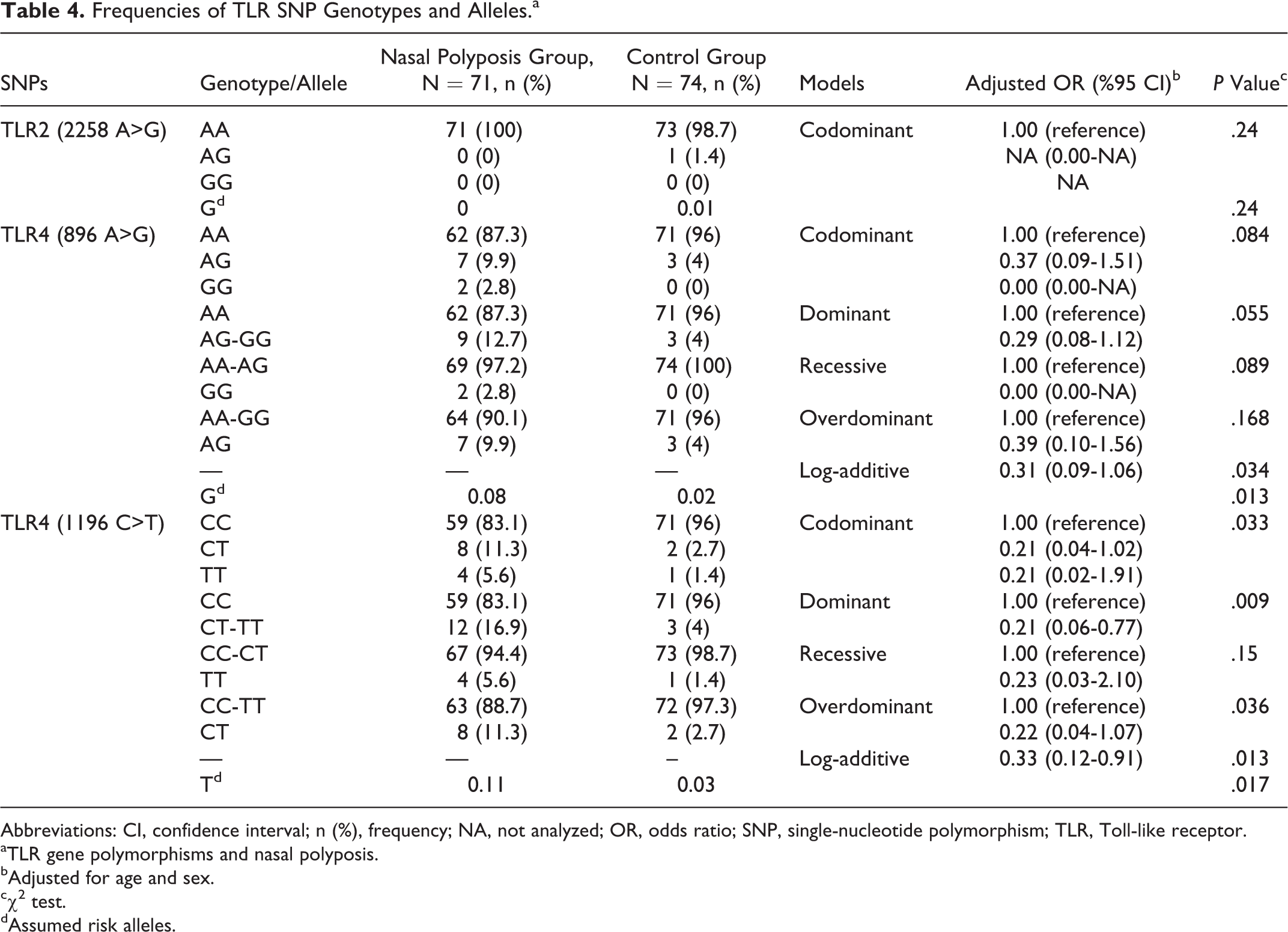
Abbreviations: CI, confidence interval; n (%), frequency; NA, not analyzed; OR, odds ratio; SNP, single-nucleotide polymorphism; TLR, Toll-like receptor.
aTLR gene polymorphisms and nasal polyposis.
bAdjusted for age and sex.
cχ2 test.
dAssumed risk alleles.
Association of the TLR Genotypes With Nasal Polyposis-Related Phenotypes
The frequencies of the TLR4 (1196 C>T) CT and TT genotypes were higher in asthmatic patients compared with those without asthma (OR = 0.22, 95% CI = 0.06-0.81, P = .017). There was no significant association between aspirin intolerance, atopy, total IgE levels, polyp size and TLR2 (2258 A>G), TLR4 (896 A>G) and TLR4 (1196 C>T) genotypes (all P > .05; data not shown).
Multifactor Dimensionality Reduction Analyses
Multifactor Dimensionality Reduction software was used to analyze the interaction of the SNPs and phenotypes that might affect nasal polyposis (Table 5). It was found that TLR4 (1196 C>T) was the best single-focus predictive model and this model is able to correctly predict nasal polyposis with 57.24% accuracy. The TLR4 (1196 C>T) CT and TT genotypes were significantly associated with nasal polyposis and had a 4-fold increased risk for nasal polyposis (testing balanced accuracy = 0.5642, cross-validation consistency (CVC) = 10/10, OR = 4.8103, CI = 1.2969-17.8657, P = 0.01; Figure 1).
Multifactor Dimensionality Reduction Analysis of the Common TLR Polymorphisms.a
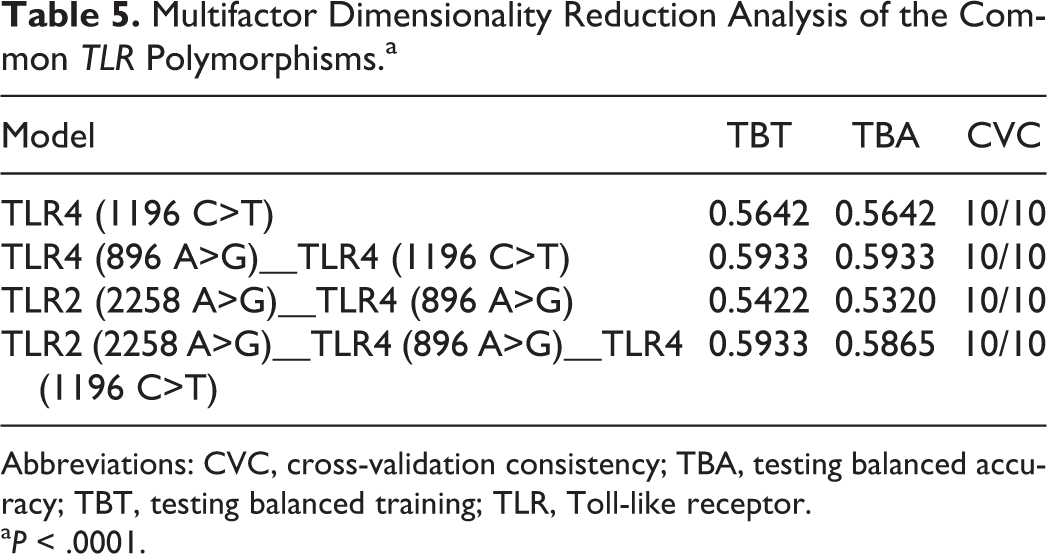
Abbreviations: CVC, cross-validation consistency; TBA, testing balanced accuracy; TBT, testing balanced training; TLR, Toll-like receptor.
a P < .0001.
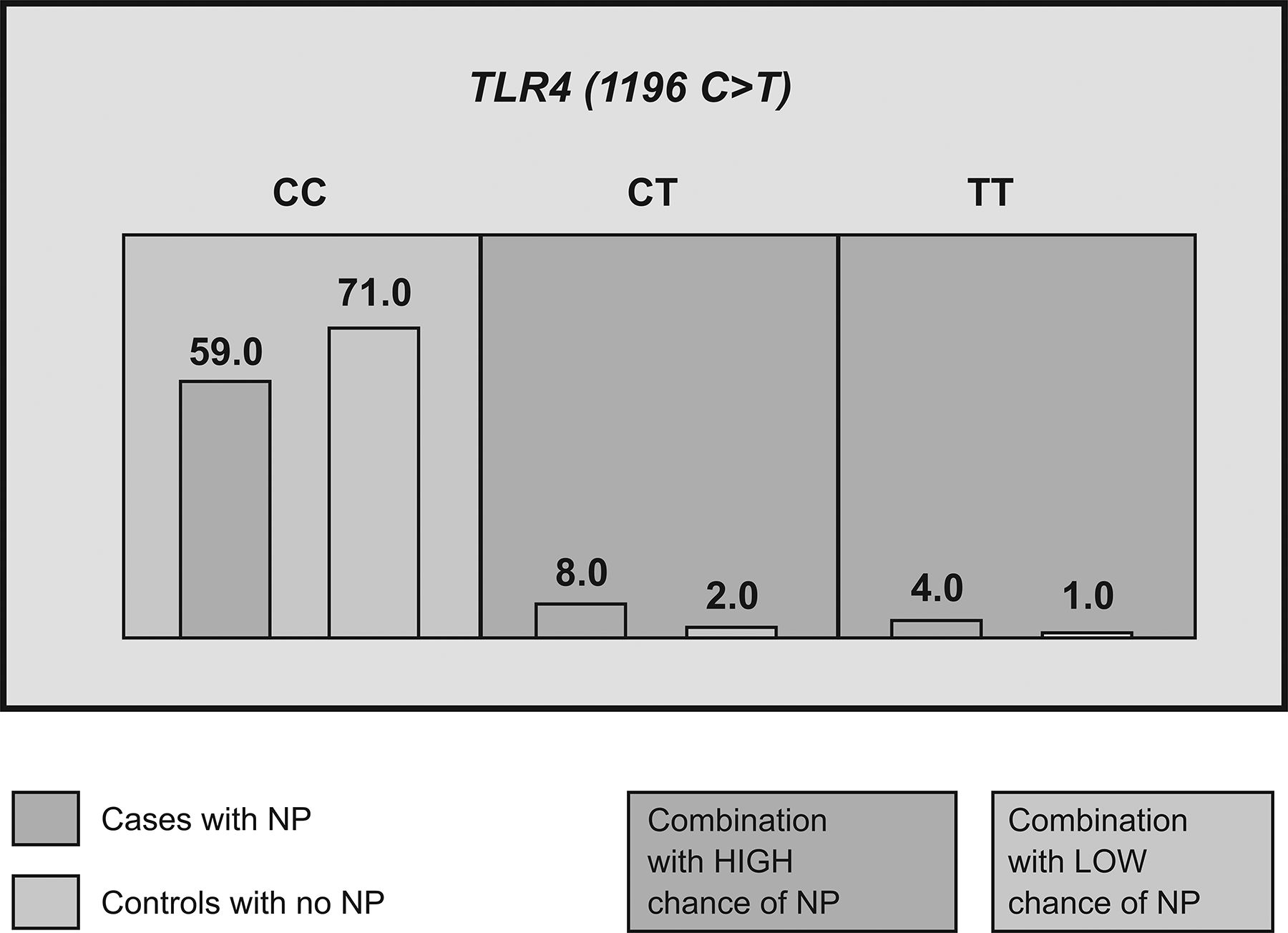
Distribution of the TLR4 (1196 C>T) genotype in the cases with nasal polyposis (NP) and control groups on MDR. Single-locus model demonstrating the effect of the TLR4 (1196 C>T) which is able to correctly predict nasal polyposis with 57.24% accuracy. The TLR4 (1196 C>T) CT and TT genotypes had a 4-fold increased risk of nasal polyposis. For each genotype, the number of cases is displayed in the histogram on the left in each cell while the number of controls is displayed in the histogram on the right. Darker shade indicates the high-risk group (TBA = 0.5642, CVC = 10/10, OR = 4.8103, CI = 1.2969-17.8657, P = .01). CVC indicates cross-validation consistency; CI, confidence interval; MDR, Multifactor Dimensionality Reduction; TBA, testing balanced accuracy; TLR, Toll-like receptor; OR, odds ratio.
According to the TLR4 (896 A>G) and (1196 C>T) genotype combinations in the 2-locus predictive models, patients carrying the AA+CT, AA+TT, AG+CC, AG+CT, and GG+CC diplotpes showed a testing balanced accuracy (TBA) of 59.3% and a cross validation consistency of 10/10 for prediction of risk of nasal polyposis. Toll-like receptor 4 (896 A>G) + (1196 C>T) AA+CC, AA+CT, AA+TT, AG+CC, AG+CT, and GG+CC diplotpes had increased risk of nasal polyposis 0.8-fold, 3-fold, 4-fold, 1.6-fold, ∞-fold, and ∞-fold, respectively (Figure 2). It was found that the pattern of high and low risk for the TLR4 (896 A>G) differs depending on the value of the TLR4 (1196 C>T) (TBA = 0.593, CVC = 10/10, OR = 4.14, CI = 0.183-93.687, P = .0048). In the 3 locus model, it was found that the strongest interaction is between the TLR4 (896 A>G) and TLR4 (1196 C>T) polymorphisms (TBA = 0.586, CVC = 10/10, OR = 3.4, CI = 0.1794-68.1879, P = .0048).
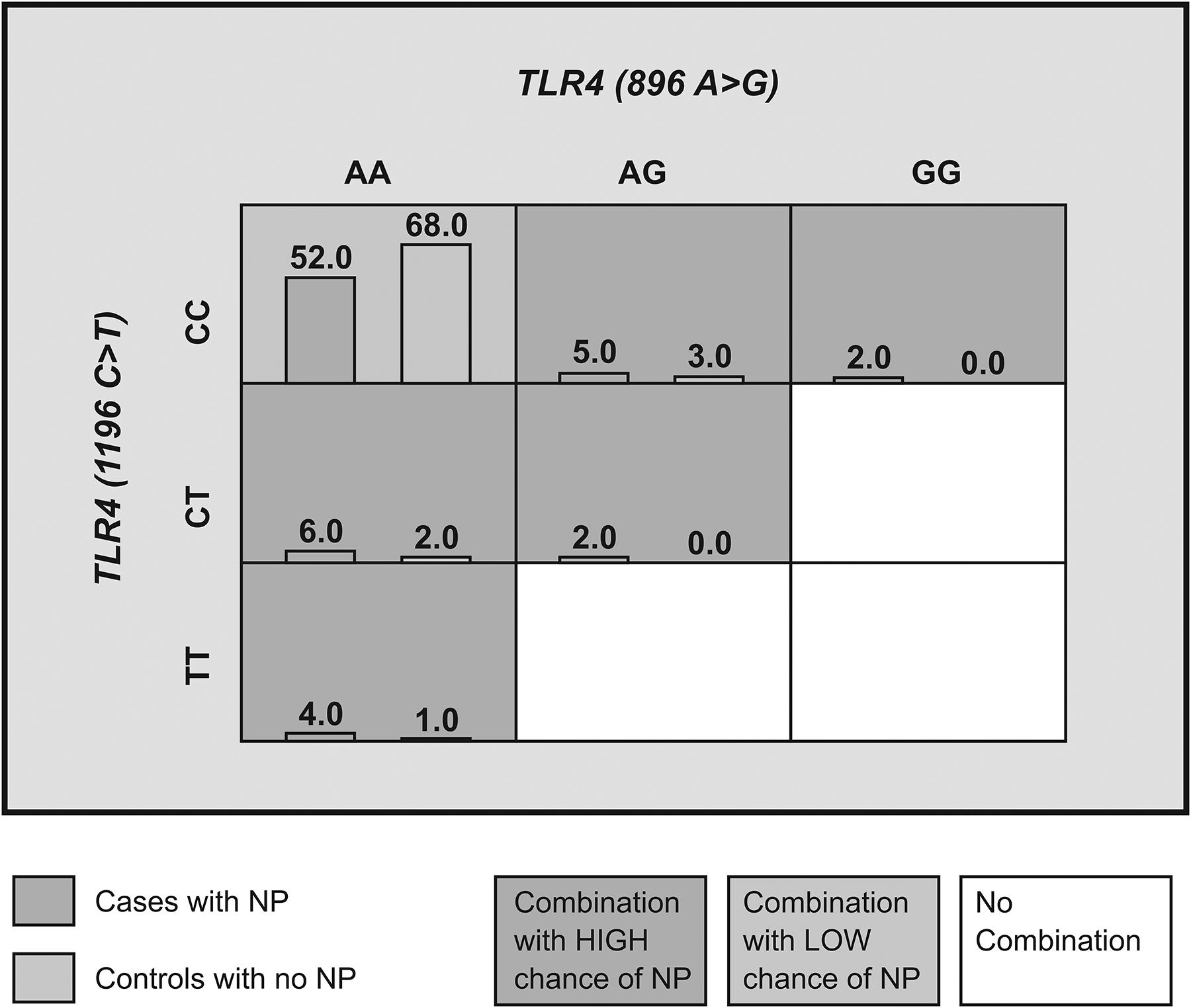
Distribution of TLR4 (896 A>G) and (1196 C>T) genotype combinations in the study and the control groups in application of MDR. The 2-locus model demonstrating the effect of TLR4 (896 A>G) and (1196 C>T), which correctly predicted NP with 60.0% accuracy. TLR4 AA+CC, AA+CT, AA+TT, AG+CC, AG+CT, and GG+CC diplotpes increased risk of NP 0.8-fold, 3-fold, 4-fold, 1.6-fold, ∞-fold, and ∞-fold, respectively. For each diplotpe, the number of cases is displayed in the histogram on the left in each cell while the number of controls is displayed by the histogram on the right. Darker shade indicates those combinations that were classified as high-risk in the original analysis while lighter shade indicates those combinations that were classified as low-risk in the original analysis. Note that the pattern of high and low risk for the TLR4 (896 A>G) differs depending on the value of the TLR4 (1196 C>T) (TBA = 0.593, CVC = 10/10, OR = 4.14, CI = 0.183-93.687, P = .0048). CI indicates confidence interval; CVC, cross-validation consistency; MDR, Multifactor Dimensionality Reduction; NP, nasal polyposis; OR, odds ratio; TBA, testing balanced accuracy.
In addition, the individuals with the TLR4 (1196 C>T) genotype CT and TT as well as asthma were in the high-risk group. To examine potential genotype X phenotype interactions, in the 2-factors model MDR analysis demonstrating effect of the asthma and TLR4 (1196 C>T) genotype, it was determined that this model is able to correctly predict nasal polyposis (NP) with 62.7%. The asthma + CC, asthma + CT, and asthma + TT had a 2-fold, ∞-fold, and ∞-fold increased risk of NP, respectively (TBA = 0.622, CVC = 8/10, OR = 2. 39, CI = 0.2067-27.7643, P = .007; Figure 3).
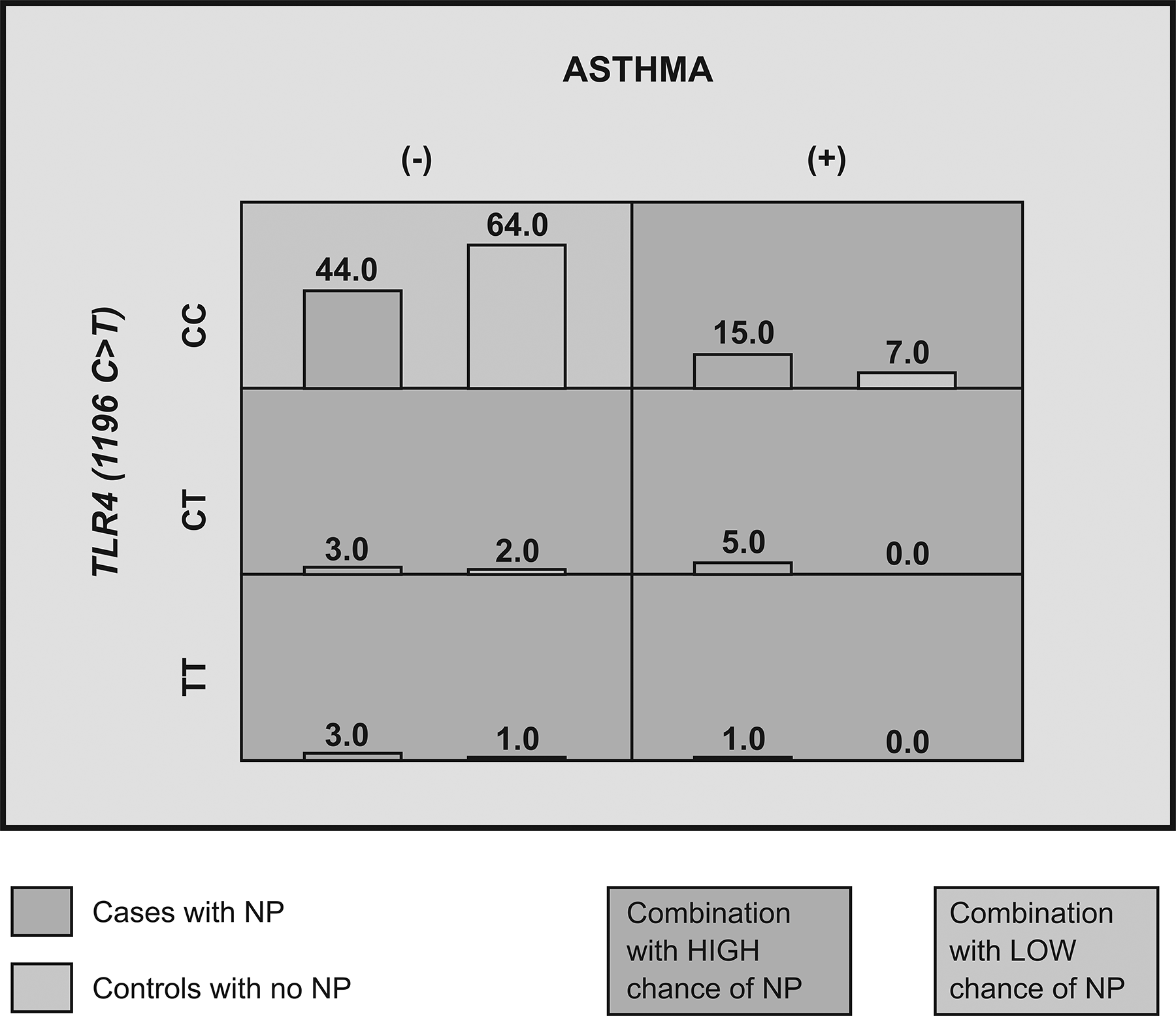
Distribution of the asthma and TLR4 (1196 C>T) genotype in cases and controls in application of MDR. The 2-factors model demonstrating effect of the asthma and TLR4 (1196 C>T) genotype which is able to correctly predict NP with 62.7% accuracy. The asthma+CC, asthma+CT, and asthma+TT had a 2-fold, ∞-fold, and ∞-fold increased risk of NP, respectively. For each combination, the number of cases with asthma (positive or negative) and TLR4 (1196 C>T) genotype is displayed in the histogram on the left in each cell, while the number of controls with asthma (positive or negative) and TLR4 (1196 C>T) genotype is displayed by the histogram on the right. Darker shade indicates the high-risk group (TBA = 0.622, CVC = 8/10, OR = 2. 39, CI = 0.2067-27.7643, P = .007). CI indicates confidence interval; CVC, cross-validation consistency; MDR, Multifactor Dimensionality Reduction; NP, nasal polyposis; OR, odds ratio; TBA, testing balanced accuracy.
Discussion
In current study, it was investigated the association between common TLR polymorphisms and nasal polyposis. It has been found that TLR4 (1196 C>T) polymorphism carry an increased risk for nasal polyposis. The pattern of high and low risk for the TLR2 (2258 A>G) and TLR4 (896 A>G) polymorphisms differ depending on the value of the TLR4 (1196 C>T). Our findings indicate that there was significant difference in genotype and allele frequencies of the TLR4 (1196 C>T) polymorphism between the nasal polyposis and control groups. There was a synergistic interaction between the TLR4 (896 A>G) and TLR4 (1196 C>T) polymorphisms.
Toll-like receptors are a receptor family that plays an important role in immune response. Toll-like receptor 2 (2258 A>G) polymorphism is associated with reduced response to infectious agents. 18 Two different mutations in TLR 4 polymorphisms (896 A>G and 1196 C>T) have been shown to reduce receptor response in macrophages, epithelial cells, and mononuclear cells. 19 Activation of TLRs in neutrophils, macrophages, and dendritic cells after acute inflammatory reactions in the nasal mucosa by viral or bacterial agents results in the production of inflammatory cytokines and chemokines. Chronic inflammation in the nasal mucosa may cause polyp formation. In a study, it was found that TLR2, TLR4, and TLR7 protein production were increased in the TLR pathway in nasal polyposis. 20 It can be considered that functional reduction in the TLR pathway may play a role in the etiology of nasal polyposis. It can be said that increase in expression of these proteins is result of overproduction of hypofunctional proteins. The existing polymorphisms in TLR4 and TLR2 may play a role in nasal polyposis because they cause hypofunction in the TLR pathway.
In the current study, it was observed that the probability of nasal polyposis was 62.7% in the presence of TLR4 (1196 C>T) polymorphism with asthma. Similar to current study in a study, it was determined that allergy and asthma were related to TLR4 polymorphism in children. 21 This study demonstrated that there is strong association between TLR4 (1196 C>T) polymorphism and asthma and nasal polyposis.
Different theories related to the etiology of nasal polyposis have been proposed. One of the suspected etiologies is Staphylococcus aureus infection. It was known that chronic rhinosinusitis with nasal polyps and asthma is associated with nasal S aureus colonization. Based on that knowledge, in a study repeated intratracheal exposure to S aureus-derived serine protease-like protein (SpID) was applied to TLR 4 knockout mice. 22 In that study, SpID-induced inflammatory response in the airways was in an interleukin (IL)-dependent but TLR4-independent manner. Toll-like receptor 2 receptors detect bacterial cell wall compartments such as peptidoglycan of S aureus. But in that study, SpID didn’t active TLR2 receptor cells in vitro. In the light of those results, it can be said that S aureus colonization was not playing role in nasal polyposis etiology and nasal polyposis is facilitating S aureus colonization.
In a study, significantly higher concentration of IL-4, IL-5, IL-6, and tumor necrosis factor-β were detected in nasal secretions of allergic nasal polyp patients than in nonallergic ones. 23 In our study, there was no significant association between atopy and TLR2 (2258 A>G), TLR4 (896 A>G), and TLR4 (1196 C>T) genotypes.
It was shown that there was no significant association between TLR 2 polymorphisms and chronic rhinosinusitis. 24 Similar to that study there was no difference between nasal polyposis and control group in terms of TLR 2 polymorphism in our study. But it has been found that TLR4 (1196 C>T) polymorphism carry an increased risk for nasal polyposis in current study.
Nasal polyposis is an inflammatory disease characterized by multiple polyps in the nasal cavity and paranasal sinuses, its etiology is multifactorial and it represents highly heterogeneous disease group. Nasal polyposis can be classified as eosinophilic and noneosinophilic nasal polyposis according to the rate of eosinophils in total inflammatory cells in the tissue. 25 In the light of results of the current study, the relationship between TLR and eosinophilic and noneosinophilic differentiation of nasal polyposis may be the subject of further studies.
As a conclusion, this study showed that TLR4 and TLR2 polymorphisms were predisposing factors for nasal polyposis. These genetic variations may shed light on showing inflammatory pathways involved in the pathogenesis of nasal polyposis. Further functional studies investigating the consequences of loss of TLR function are needed.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
