Abstract
Objective
Pseudogenes are often referred to as “junk DNA.” Although they have been well characterized in mammals, pseudogenes have been identified in only a few plant species. As an important traditional Chinese medicinal plant, the genome of
Methods
Based on the
Results
A total of 3156 pseudogenes were identified. DUP-type pseudogenes exhibited more insertion and frameshift mutations than PSSD-type pseudogenes; furthermore, a recent expansion of DUP-type pseudogenes was observed. Expression analysis detected 802 pseudogenes expressed in various tissues, primarily associated with plant defense functions according to Gene Ontology enrichment analysis. Among these, DUP-type pseudogenes were the most prevalent, with most arising recently. Additionally, 45 pseudogenes corresponding to gene family members involved in the biosynthesis of the medicinal compounds (indigo and indirubin) in
Conclusions
In this study, we successfully identified and characterized 3156 pseudogenes in
Introduction
Pseudogenes are DNA sequences that originate from functional genes but have lost their activity. Pseudogenization can result from various factors, such as polyploidization and cosymbiosis.7,8 The concept of pseudogenes was first proposed by Jacq et al. in 2017 when they cloned a 5S rRNA-related gene. 9 Pseudogenes can be classified into processed pseudogenes (PSSD) and unprocessed pseudogenes, the latter further divided into fragmented pseudogenes (FRAG) and duplicated pseudogenes (DUP). 10 During evolution, pseudogenes may accumulate mutations, including insertions, deletions, base substitutions, and translocations. Comparing pseudogene sequences with their parental genes provides valuable insights into evolutionary processes. 11 Although pseudogenes were once considered nonfunctional, recent studies have demonstrated that they may play significant roles in gene expression, regulation, and the generation of genetic diversity. 12 Pseudogenes can produce various noncoding RNAs (ncRNAs) that participate in diverse biological functions,5,13 and they have been identified as potential tumor markers that may influence tumor development and progression at the gene regulatory level. 14
Pseudogenes have been identified in various plants, including barley, rice, and
Materials and methods
Identification and screening of pseudogenes
The genome sequence, annotation file, and RNA-seq data (including both two developmental stages of stem (S1 and S2), leaf (L1 and L4), and root (R) samples of
Analysis of expression of pseudogenes
Based on the identification results of pseudogenes, we constructed a GTF file containing all pseudogenes. All RNA-seq data were preprocessed using fastp v0.23.1
19
with the parameters: -g -q 5 -u 50 -n 15 -l 150 –min_trim_length 10 –overlap_diff_limit 1 –overlap_diff_percent_limit 10. STAR
20
version 2.7.10a (–outFilterMultimapNmax 1 –quantMode GeneCounts) was used to map RNA-seq data (from samples of different developmental stages of
Finally, TBtools
22
and the cloud website bioinformatics.com.cn were used to construct circos plots and perform Gene Ontology (GO) and KEGG enrichment analyses. The detailed steps for constructing a circos plot using TBtools are as follows: First, prepare pseudogene (or gene) data by extracting the chromosome ID (chrID), start position, end position, and strand (plus or minus) from the pseudogene GFF3 file. Add a new column after the end position and fill it entirely with the value 1. Save this processed data as the pseudogene information file. Second, generate a color scheme by opening the “Discrete Color Scheme Generator” in TBtools. For input, use the sequence length information for each chromosome. Specify the output file path and click the “Start” button. This will generate a file containing random RGB color codes, which will serve as the input file for Advanced Circos. Third, visualize the chromosome skeleton by importing the generated color scheme file into Advanced Circos and clicking “Show My Circos Plot” to generate the colored chromosome skeleton Circos diagram. Finally, add pseudogene density by clicking “Show Control Dialog,” then clicking “Add” on the right panel and selecting the previously prepared pseudogene information file. Click “BIN Setting” and change the “Mean” option in BIN Mode to “Sum.” Click “Refresh Graph” to generate the Circos plot displaying pseudogene density. For GO and KEGG enrichment analyses, three files were used as input: plant GO or KEGG background data,
Results
Distribution of pseudogenes
A total of 3156 pseudogenes (Supplemental Table S1) were identified in the genome of
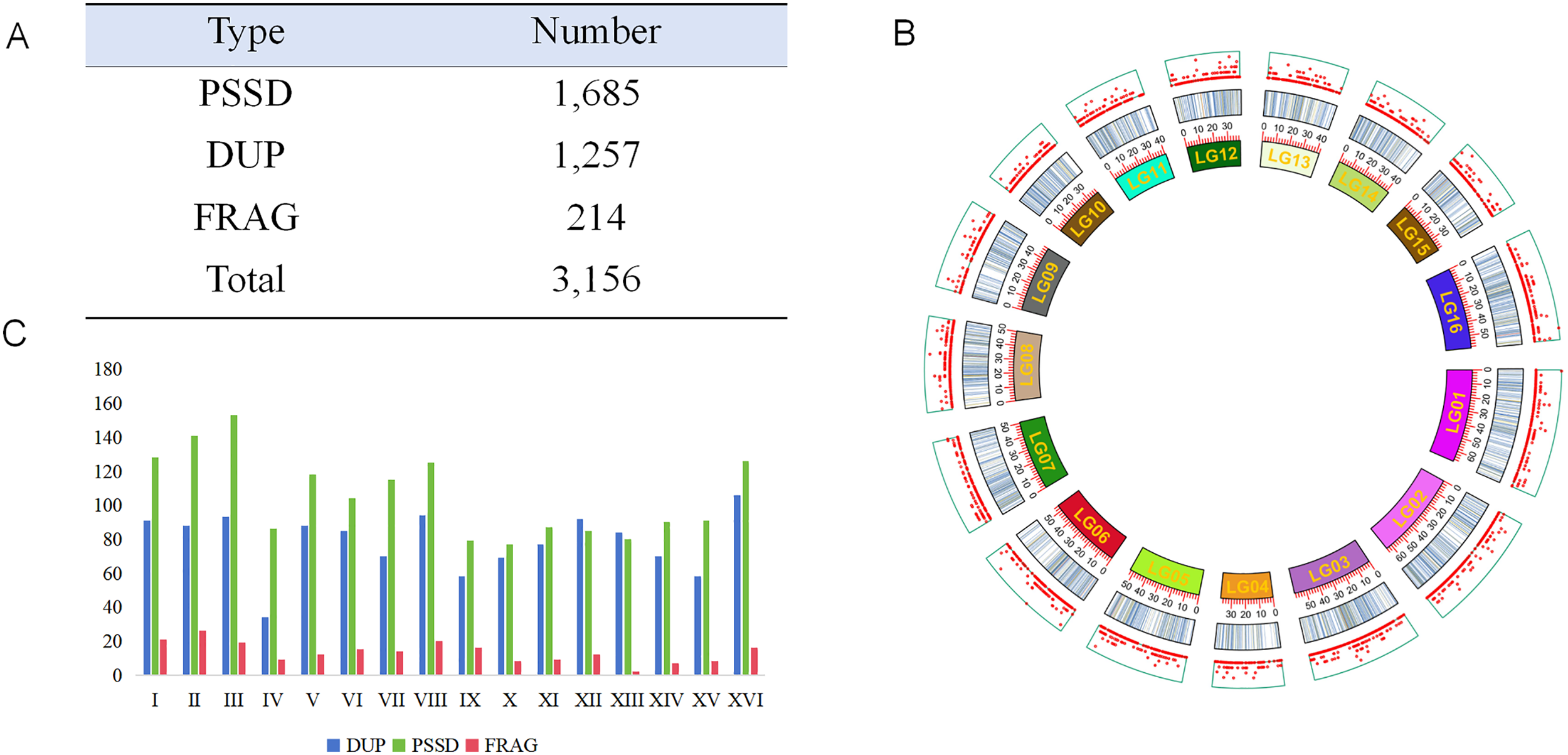
(A) Type and number of different pseudogenes, PSSD for processed pseudogenes, DUP for duplicated pseudogenes, FRAG means fragmented pseudogenes. (B) Distribution of pseudogenes and genes in different chromosomes: the outermost plate is genes, the middle one is pseudogenes, and the inner plate is chromosomes. (C) Distribution of different type of pseudogenes in different chromosomes.
The number of pseudogenes on each chromosome generally correlated with chromosome length (Figure 1C). Further analysis of the density and distribution of pseudogenes on each chromosome revealed that, although the density of pseudogenes was relatively consistent across chromosomes (ranging from 3 to 5 genes per Mb), there were significant differences in the distribution density of pseudogenes among different regions of the same chromosome (Figure 1B).
Functional enrichment analysis of pseudogenes
We carried out GO enrichment and KEGG pathway enrichment analyses based on the GO and KEGG terms associated with each pseudogene's parental genes. The results indicated that many pseudogenes were enriched in GO terms related to plant defense, including both abiotic and biotic stress responses (Figure 2A, Supplemental Table S2). DUP-type pseudogenes were also enriched in GO terms associated with pollen and stigma recognition (Figure 2B). According to the KEGG pathway enrichment analysis, pseudogenes were primarily enriched in pathways related to photosynthesis, signal transduction, and various metabolic processes. Additionally, some pseudogenes were enriched in pathways involved in terpenoid biosynthesis.

(A) GO enrichment of all pseudogenes; (B) GO enrichment of duplicated pseudogenes. From top to bottom are biological process, cellular component and molecular function.
Pseudogene mutation and evolutionary analysis
Following the development of most pseudogenes, their sequences may exhibit specific alterations due to the loss of function, external selective pressures, and other factors.
11
We then performed a statistical analysis of the different types of pseudogene mutations. As shown in Figure 3A and B, insertion and deletion mutations constitute most mutation types observed in PSSD and DUP pseudogenes. Although deletion mutations are relatively rare, both insertion and frameshift mutations in DUP pseudogenes occur at significantly higher rates than in PSSD pseudogenes (

Mutation statistics of DUP type (A) and PSSD type (B) pseudogenes (Mutation type: insertion, deletion, frame-shift (Shift), and premature stop codon mutations (Stop)).
We also examined the degree of sequence similarity between pseudogenes and their parental genes to estimate the formation time of pseudogenes. Overall, as shown in Figure 4A, pseudogenes exhibited two peaks and subsequent declines, with maxima at 0.4–0.5 and 0.8–0.9, indicating increases in pseudogene counts during these two-time intervals. The number of DUP pseudogenes initially increased and then decreased; the highest number was observed around 0.8–0.9 (Figure 4B), suggesting a burst of DUP pseudogene formation during this period. In contrast, the number of PSSD pseudogenes has been declining, with no recent increases observed (Figure 4C). Meanwhile, the number of FRAG pseudogenes has been growing, reaching a peak at 0.4 (Figure 4D).

Sequence identity between of pseudogenes and their parent genes. (A) All type of pseudogenes, (B) DUP type of pseudogenes, (C) PSSD type of pseudogenes, (D) FRAG type of pseudogenes.
Analysis of pseudogene expression patterns
We examined the expression of pseudogenes in two developmental stages of leaves (L1 and L4), two developmental stages of stems (S1 and S2), and roots (R) using transcriptome data from the genome study by Xu et al. (2020). 2 After screening, we found that 620 pseudogenes were expressed in L1; 603 in L4; 620 in S1; 576 in S2; and 603 in roots. In total, 801 pseudogenes (Supplemental Table S3) were expressed, including 391 DUP, 364 PSSD, and 47 FRAG pseudogenes. Among these, 421 pseudogenes were consistently expressed across all tissues at different developmental stages (Figure 5A). GO enrichment analysis of these pseudogenes indicated that their primary functions are related to plant defense.

(A) Venn diagram of pseudogenes expressed in different tissues (L, leaf; R, root; and S, stem) and stages (L1 and L4 represent two developing stages of leaf, and S1 and S2 mean two developing stages of stem). (B) Expression heatmap of pseudogenes related to gene family members involved in indigo and indirubin biosynthesis, all the samples have three replicates. The names outside the parentheses are detail information of each pseudogene (including chromosome number, pseudogene type, start position and end position), while the IDs inside the parentheses correspond to their parental function genes. (C) Expression profile of pseudogenes corresponding to genes family members involved in indigo and indirubin biosynthesis. The blue column represents all the gene family member-related pseudogenes, and the red column represents all the expression pseudogenes.
We also examined the expression profiles of pseudogenes belonging to gene families, including CYPs, UGTs, BGLs, and FMOs. Among the 27 pseudogenes identified as being expressed in various tissues (Figure 5B, Supplemental Table S4), only those related to the CYP and UGT gene families showed detectable expression. Three pseudogenes—pseudoEVM0012390-1 (CYP), pseudoEVM0012390-2 (CYP), and pseudoEVM0009100 (UGT)—were expressed in both stems and leaves but not in roots. PseudoEVM0002447 (UGT) and pseudoEVM0026966 (UGT) were detected exclusively in leaves, with pseudoEVM0002447 expression limited to the L1 leaf layer. PseudoEVM0012039 (UGT) and pseudoEVM0018899 (UGT) were not expressed in leaves but were present in roots and stems. Additionally, pseudogenes associated with indigo and indirubin synthesis located on chromosomes II, XII, and XIII showed no expression (Figure 5C).
Discussion
Here, we detected 1685 PSSD-type pseudogenes in
Although the overall chromosome size is proportional to the number of pseudogenes, there are a few exceptions. For example, Chromosome III is smaller than Chromosome II but contains more pseudogenes. A positional preference is observed within the same chromosome, as evidenced by the uneven distribution of pseudogenes, with some regions having fewer and others having more. Further analysis revealed that the ratio of PSSD-type pseudogenes to DUP-type pseudogenes on Chromosome IV is relatively high, reaching 2.52. The number of DUP-type pseudogenes on Chromosome IV is only 34, which is significantly lower than the average number on other chromosomes (average of 77, minimum of 58). In the pseudogene evolution analysis, there is a recent surge of DUP-type pseudogenes, indicated by a similarity of 0.8–0.9 between pseudogenes and their parental genes. Considering the distribution of DUP-type pseudogenes across chromosomes, it can be inferred that this recent surge mainly occurred on chromosomes other than Chromosome IV. Additionally, based on GO enrichment and KEGG pathway analyses, all three types of pseudogenes are closely associated with environmental response and defense against external stimuli, suggesting that stress plays a significant role in pseudogene formation in
Pseudogenes associated with the indigo and indirubin synthesis pathways are distributed on chromosomes other than Chromosome VII, indicating a preferential chromosomal distribution. The expression patterns of these pseudogenes generally consistent with the overall expression profile; however, spatial and temporal differences exist among individual pseudogenes. Additionally, expression of pseudogenes on certain chromosomes was not detected, suggesting chromosomal preference for the expression of these pseudogenes. Some pseudogenes exhibit significant differential expression among leaves, stems, and roots, implying a potential role in regulating the distinct pharmacological properties of
Conclusion
This study identified 3156 pseudogenes at the
Supplemental Material
sj-pdf-1-sci-10.1177_00368504261420981 - Supplemental material for Genome-wide identification and expression pattern analysis of pseudogenes in Strobilanthes cusia (Nees) Kuntze
Supplemental material, sj-pdf-1-sci-10.1177_00368504261420981 for Genome-wide identification and expression pattern analysis of pseudogenes in
Footnotes
Acknowledgments
Not applicable.
Ethical considerations
Not applicable.
Author contributions
Z Lin designed the projects and carried out most of the analysis, and Q Cai handled the statistical analysis of pseudogenes.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was funded by Fujian Provincial Natural Science Foundation Youth Project (Grant No. 2022J05254), Putian University scientific research launch project (Grant No. 2022051), and the Fujian Provincial Science and Technology Project (Grant No. 2022N5006).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Not applicable.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
