Abstract
Objective
Marine organisms such as sea cucumbers represent valuable sources of specialized metabolites with promising applications, making them relevant targets for exploring metabolite distribution and subsequent in-depth characterization. This study aimed to chemically and biologically characterize the parent methanolic extract and its derived fractions, obtained through sequential liquid–liquid partitioning, from the sea cucumber Isostichopus sp. aff. badionotus collected from the Colombian Caribbean Sea.
Methods
The extract and fractions were analyzed using ultra-high-performance liquid chromatography coupled with mass spectrometry and the Global Natural Products Social-based molecular networking platform, enabling dereplication and identification of metabolites. Additionally, the extract and fractions were evaluated against eight Gram-positive and Gram-negative bacterial strains using broth microdilution assays.
Results
The dichloromethane fractions exhibited the highest antibacterial activity against both Gram-positive and Gram-negative bacteria. For the first time, chemical characterization allowed the identification of a series of metabolites present in the extract and fractions.
Conclusion
This study provides the first report on antibacterial activity in I. sp. aff. badionotus, demonstrating that medium-polarity compounds in the dichloromethane fraction exhibited moderate bacterial inhibition, particularly against Gram-positive strains.
Keywords
Introduction
Bacterial drug resistance remains one of the most pressing challenges to global public health, largely driven by the overuse and misuse of antibiotics. This phenomenon results in a marked reduction in the efficacy of conventional treatments and a significant increase in patient morbidity and mortality rates. 1 In response to this urgent threat, searching for new bioactive compounds with antibacterial potential has become a global research priority. Considering this context, natural products have historically been a critical source of therapeutic agents, and marine-derived compounds, in particular, have gained prominence due to their chemical diversity, structural complexity, and unique bioactivity profile. 2 Marine ecosystems, with their vast and largely unexplored biodiversity, offer promising opportunities for discovering and developing novel antibacterial agents.
Sea cucumbers, marine invertebrates belonging to the Holothuroidea, have attracted noteworthy attention among marine organisms. Distributed in both deep and shallow coastal waters worldwide.
3
Sea cucumbers possess a diverse repertoire of metabolites linked to their detritivorous feeding habits and adaptation to complex ecological niches. In addition to their traditional use as a food source in many cultures, particularly in Asia, sea cucumbers have been the focus of numerous studies investigating their biological properties. From Stichopus vastus, Stichopus quadrifasciatus, Holothuria (Metriatyla) lessoni Massin, Holothuria (Mertensiothuria) leucospilota, and Bohadschia marmorata, volatile cyclic methyl siloxane compounds were identified. In Holothuria scabra, holothurin, holothurinosides, scabraside, and other similar compounds with antioxidant properties were detected.4–6 Regarding anti-inflammatory compounds, Actinopyga agassizi, Pentacta quadrangularis, and other holothuroids contain triterpene glycosides that have been identified as anti-inflammatory metabolites.6,7 Frondoside A–A5, isolated from Cucumaria frondosa, as well as protein hydrolysates from Actinopyga lecanora, exhibited anticancer activity in lungs (LNM35, A549, among others), breast (MDA-MB-435 and MCF-7), and liver (HepG2) cell lines.8,9 Protein hydrolysates from A. lecanora and Holothuria parva were also evaluated as bioactive peptides with ACE (angiotensin i converting enzyme) inhibitory and antihypertensive properties.10,11 Furthermore, several studies have reported the antimicrobial potential of different sea cucumber species, highlighting their promise as natural alternatives to combat bacterial resistance. In this context, cyclic siloxanes such as octamethyl, decamethyl, and dodecamethyl cyclohexasiloxane, identified in the muscular tissues of S. vastus, S. quadrifasciatus, H. lessoni Massin, H. leucospilota, and B. marmorata, demonstrated promising antimicrobial activity.
4
To better understand the antimicrobial effects of the lipophilic extract of H. leucospilota, the authors suggested that the presence of fatty acids enhanced both toxicity and antimicrobial activity.
12
In addition, the use of
In the present study, we aimed to expand the chemical and biological understanding of the Isostichopus genus by characterizing the chemical composition of the parent methanol extract and its derived fractions obtained through sequential liquid–liquid partitioning from I. sp. aff. badionotus, collected from the coastal waters of northern Colombia. Ultra-high-performance liquid chromatography coupled with mass spectrometry (UHPLC-MS), along with the Global Natural Products Social (GNPS) molecular networking platform, was employed for metabolite profiling. Additionally, we evaluated, for the first time, the antibacterial activity of the extract and fractions against eight Gram-positive and Gram-negative bacterial strains. This integrative approach enabled the identification of metabolites potentially responsible for the observed antibacterial effects and contributed to the growing body of knowledge on marine-derived antibacterial compounds (https://www.preprints.org/manuscript/202505.1052/v1).
Materials and methods
Collection and extraction of sea cucumber Isostichopus sp. aff. badionotus
Adult specimens of the sea cucumber I. sp. aff. badionotus were collected by hand between January and March of 2025. The sampling plan identified three sites along the coast of Santa Marta Bay in the Colombian Caribbean: Rodadero (11°13′22.73″ N, 74°13′32.59″ W), the airport (11°07′10″ N, 74°13′50″ W), and Taganga beach (11°16′04.6″ N, 74°11′46.9″ W). Ten specimens were selected from each site during collection and transferred to the aquaculture laboratory at the University of Magdalena. The specimens were stored in a polyethylene bag at 4 °C until processing. The voucher samples for taxonomic identification were deposited in the Biological Collections at the University of Magdalena (CBUMAG:ECH:00001, CBUMAG:ECH:00002, and CBUMAG:ECH:00003).
The animal tissue samples from the grouped specimens were cut into small pieces, frozen at −80 °C, and dried using a freeze dryer. Extraction was performed by cold maceration. First, 600 g of dry cucumber biomass were mixed with 6 L of solvent in a glass container. The solute–solvent ratio was 1 g/10 ml. The mixture was stirred at room temperature for 72 h. After 72 h, the extract was filtered with Whatman No. 1 filter paper (Massachusetts, USA), and the filtrate was concentrated under reduced pressure with a Heidolph Hei-VAP Value rotary evaporator (Schwabach, Germany) to obtain the crude methanol extract. The resulting extract was subjected to sequential liquid–liquid separation using solvents of increasing polarity. Each separation was performed in a separation funnel using equal volumes (1:1, v/v) of aqueous methanol residue and the respective organic solvent (Merck, Darmstadt, Germany): n-hexane (n-Hex), dichloromethane (DCM), n-butanol (BuOH), and methanol. The solvent separations were performed in a separation funnel with equal volumes (1:1, v/v) of the aqueous-methanol residue and the respective organic solvent. The resulting organic phases were collected individually, dried over anhydrous sodium sulfate, filtered, and evaporated under reduced pressure. This process yielded 3.6 g (7.6%) of the Hex phase, 6.6 g (14%) of the DCM phase, 8.6 g (18.2%) of the BuOH phase, and 28.5 g (60.2%) of the methanol phase. The products were then stored at 4 °C until they were used in determining chemical profiles and conducting biological assays.
Biological assay
Strains
Eight bacterial strains were utilized to assess the antibacterial activity of the fractions from I. sp. aff. badionotus. These selected pathogens are clinically significant and are included in the list of the prioritized bacteria by the WHO due to the complications they cause in human health. These included four Gram-negative strains: Escherichia coli ATCC 25922, Klebsiella pneumoniae ATCC 1705, Shigella flexneri ATCC 12022, and Pseudomonas aeruginosa ATCC 10195, and four Gram-positive strains: S. aureus ATCC 6538, Bacillus cereus ATCC 11778, Staphylococcus epidermidis ATCC 1228, and Staphylococcus warneri ATCC 27836. All strains were cultured in Mueller–Hinton broth and maintained under standard laboratory conditions.
Antibacterial activity assay
The initial antibacterial activity of each fraction separated by solvent was determined using the broth microdilution method, according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI) (version 2022). For this purpose, a stock solution of each fraction was prepared by weighing 100 mg of the dry product and dissolving it in 1 ml of pure dimethyl sulfoxide (DMSO) ([Extract] = 100.000 μg/ml; [DMSO] ∼ 100%). The working solutions of each extract were then prepared at 2.000 μg/ml by dilution with 1× buffered sodium phosphate ([DMSO] ∼ 2%). Next, 100 μl of each working solution was mixed with 100 μl of bacterial suspension and broth (adjusted to optical density [DO]600 = 0.05) in 96-well microplates, resulting in a final assay concentration of 1.000 μg/ml per well ([DMSO] ∼ 1%). Gentamicin (40 μg/ml) served as a positive control, while 1% DMSO and uninoculated broth were included as negative controls. In addition, broth alone was used as a sterility control. The plates were incubated at 37 °C for 150 min, and bacterial growth was monitored by measuring the optical density at 600 nm every 30 min. The percentage of bacterial inhibition was calculated according to the following equation:
In addition, the minimum inhibitory concentration (MIC) was determined for extracts that reduced bacterial growth kinetics at the screening concentration. In summary, the inhibition of bacterial growth by the active extracts was evaluated with serial dilutions from 1000 to 100 µg/ml, and the MICs were calculated using nonlinear regression, in which the experimental data were fitted using a modified Gompertz equation using GraphPad Prism 9.3. This procedure used the same methodology of initial screening. The positive control was gentamicin; the negative control was DMSO at concentrations between 0.1 and 1.
Statistical analysis
Each experimental condition was performed in triplicate, with four technical replicates per treatment (n = 12). Data reproducibility was assessed using Levene's test for homogeneity of variances (coefficient of variation < 20%). Statistical significance among groups was evaluated using one-way analysis of variance followed by Tukey's posthoc test (p < 0.05). All statistical analyses were carried out using GraphPad Prism software, version 9.3.0 (GraphPad Software, San Diego, CA, USA).
Ultra-high-performance liquid chromatography coupled with high-resolution mass spectrometry-based analysis
The extract and its fractions were subjected to metabolic profiling using ultra-high-performance liquid chromatography coupled with high-resolution mass spectrometry (UHPLC-HRMS) to understand better and relate the observed biological activity. Dried fractions were subjected to chemical characterization using UHPLC coupled with quadrupole time-of-flight mass spectrometry (UHPLC-QTOF-MS; Compact QTOF MS + Elute UHPLC, Bruker Daltonik GmbH, Bremen, Germany). The methanolic (MeOH) extract and its Hex, DCM, and BuOH fractions were dissolved and transferred into HPLC-grade vials. Chromatographic separation was performed on a Kinetex C18 column (2.1 mm×100 mm, 1.7μ particle size) (maintained at 40 °C), using 0.1% formic acid in 10% water (solvent A) and 0.1% formic acid in 90% acetonitrile (solvent B) as the mobile phases. A 5 μl injection volume was used, with a flow rate of 0.5 ml/min. The gradient elution program was as follows: 0 min, 88% A; 12.5 min, 70% A; 14.5 min, 40% A; 19 min, 88% A, for a total run time of 19 min. Mass spectrometric data were acquired in both negative and positive electrospray ionization (ESI) modes over an m/z range of 120–1800. The ESI parameters included a capillary temperature of 200 °C, a capillary voltage of 2.0 kV, a dry gas flow rate of 8 L/min, and a nebulizer pressure of 2 bar.
Compound annotation was achieved by integrating high-resolution monoisotopic mass measurements, MS/MS fragmentation patterns, and chromatographic retention times. These data were cross-referenced with the established spectral libraries, including MS-DIAL, MassBank of North America (MoNA), and LipidBlast, to ensure the accurate annotation of the detected metabolites. The confidence level for metabolite identification via HRMS was assigned according to the previously established reporting standards (levels 1–4). 15
Molecular networks
Raw chromatographic data were converted to the mzXML format and uploaded to the GNPS Molecular Networking platform (https://gnps.ucsd.edu/) for the dereplication and construction of molecular networks. 16 Spectral clustering was based on cosine similarity scores and compared with public GNPS libraries to facilitate metabolite annotation. Molecular networking allowed the visualization of structurally related compounds and the putative identification of bioactive metabolites, contributing to the observed antibacterial effects. Data processing and compound annotation were performed using MetaboScape 4.0 software (Bruker), referencing the MoNA spectral library.
Results
To evaluate the antibacterial potential of I. sp. aff. badionotus, specimens were collected from three coastal sites in the Santa Marta Bay (Colombia), and an MeOH crude extract was prepared and subsequently partitioned through liquid–liquid extraction to yield Hex, DCM, and BuOH fractions. The extract and its fractions were tested against eight clinically relevant bacterial strains using the CLSI broth microdilution method. In parallel, UHPLC-QTOF-MS-based metabolite profiling coupled with GNPS molecular networking was conducted to characterize the specialized metabolites in the fractions and explore potential correlations with the observed antibacterial activity.
Chemical characterization of extract and fractions from Isostichopus sp. aff. badionotus
The UHPLC-QTOF-MS-based chemical characterization aimed to reveal the profile of specialized metabolites present in each fraction and assess the effectiveness of the fractionation strategy. This approach was conducted in both positive and negative ionization modes to ensure comprehensive detection. High-resolution monoisotopic masses, tandem MS/MS fragmentation patterns, and chromatographic retention times were systematically compared against the spectral databases including MS-DIAL, MoNA, and LipidBlast for accurate compound annotation. Considering the nature of the compounds and the lack of in-house standards, the identification confidence was assigned to level 3, indicating annotation at the compound-class level with evidence supporting possible structures rather than complete structural elucidation. This classification aligns with the accepted confidence levels for communicating metabolite identification via HRMS. 16 With this clarification, subsequent analysis revealed a diverse array of lipid-type specialized metabolites. In the negative ionization mode, compounds were primarily associated with classes such as lysophosphatidylethanolamines (LPEs), lysophosphatidic acids (LPAs), lysophosphatidylcholines (LPCs), lysophosphatidylserines (Lyso-PS), phosphoethanolamines (PE-O), glycerophosphoinositols (GPI), and sulfated sterols (ST). Conversely, positive ionization mode predominantly identified lipids including lysodiacylglycerylcarboxyhydroxymethylcholines (LDGCC), sphingomyelins (SM), LPEs, phosphatidylcholines (PC), and LPCs. Figure 1 depicts representative molecular structures of the major specialized metabolites detected under each ionization condition, with panel A corresponding to those annotated in the negative mode and panel B to those in the positive mode.

Annotated compound classes in Isostichopus sp. aff. badionotus extract and fractions by UHPLC-MS/MS-based analysis. (a) Metabolites detected and annotated in negative mode. (b) Metabolites detected and annotated in positive mode. Lysophosphatidylethanolamines (LPE), lysophosphatidic acids (LPA), lysophosphatidylcholines (LPC), lysophosphatidylserines (Lyso-PS), phosphoethanolamines (PE-O), glycerophosphoinositols (GPI), sulfated sterols (ST), lysodiacylglycerylcarboxyhydroxymethylcholines (LDGCC), sphingomyelins (SM), phosphatidylcholines (PC). R1–R12 represent different substituents, varying according to the annotated compound class. UHPLC-MS/MS: ultra-high-performance liquid chromatography coupled with mass spectrometry-mass spectrometry.
The distribution and relative abundance of the annotated metabolites across the different extract and its derived fractions are intuitively visualized in the heatmap (Figure 2), which presents two subsets of detected compounds obtained under negative and positive ionization modes (Figure 2(a) and (b), respectively). Metabolite intensities were normalized and visualized using a color gradient, with red tones indicating higher abundance and blue tones representing lower abundance. Thus, hierarchical clustering of both subsets and fractions revealed distinct chemical profiles among the fractions, emphasizing the efficiency of the liquid–liquid partitioning in segregating metabolites based on solvent removal. The DCM fraction showed the highest diversity and intensity of detected metabolites (especially in positive mode), suggesting enrichment of moderately polar metabolites (mainly ST, LPE, LPC, LDGCC, and PC) in both subsets. The BuOH fraction also retained several polar and amphiphilic compounds (e.g. LPI and LPA), some overlapping with those in the DCM fraction (ST, LPE, and PC).
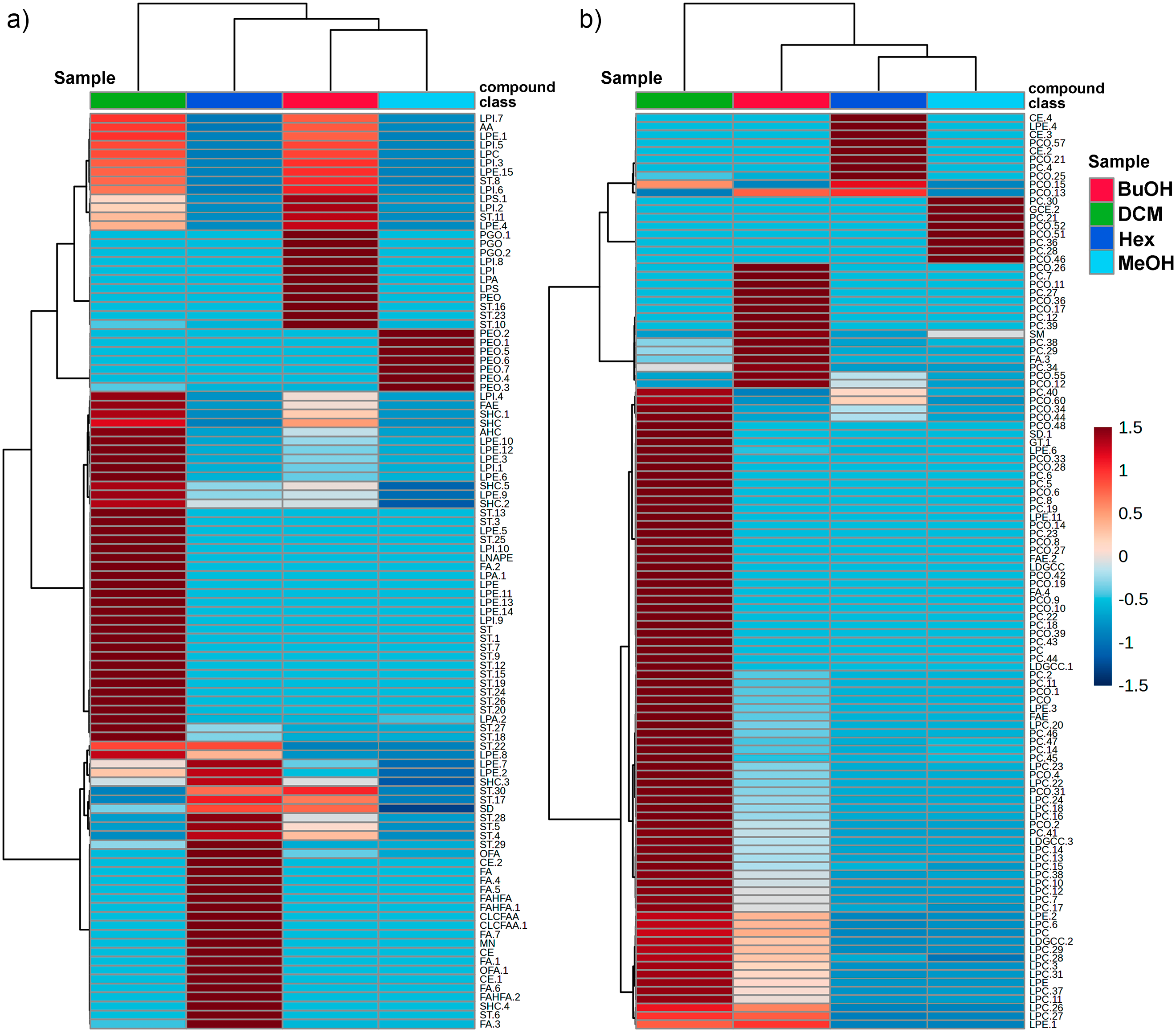
Heatmap of the relative abundance of metabolites detected in the methanolic extract (MeOH) and its solvent fractions—hexane (Hex), dichloromethane (DCM), and butanol (BuOH)—of Isostichopus sp. aff. badionotus. (a) and (b) represent the two subsets of metabolites from negative (a) and positive (b) ion modes, respectively, clustered based on their intensity patterns across the parent extract (MeOH) and its three fractions. The color scale indicates normalized intensity values, with red representing higher abundance and blue lower abundance. Hierarchical clustering highlights horizontally distinct chemical profiles among the fractions, reflecting the differential distribution of metabolites according to partitioning. Fractions per ionization mode are vertically organized according to hierarchical clustering. Class compounds are abbreviated according to Figure 1. Final numbers after a dot indicate sequential annotations within the same compound class.
In contrast, the Hex fraction exhibited a more restricted profile, enriched in a narrower set of nonpolar metabolites (fatty acids, terpenoids, and miscellaneous). The parent MeOH extract exhibited an unfractionated chemical profile, characterized by intermediate levels of many annotated metabolites, and served as a baseline for comparative analysis. This profile likely reflects the presence of residual compounds (e.g. PE, PC) that were not entirely removed during the partitioning process, consistent with its role as the original matrix encompassing a broader diversity and potentially higher abundance of metabolites. Consequently, the clustering patterns and differential distribution confirm that solvent-guided fractionation effectively separated metabolites according to their physicochemical properties. Notably, the BuOH and DCM fractions retained a higher abundance of structurally diverse metabolites, supporting their potential role in the observed biological activities and underscoring their relevance for pharmacological investigation (Figure 3).

PLS-DA score plots generated from the UHPLC-QTOF-MS datasets. Panel (a) corresponds to the negative ionization mode, while panel (b) corresponds to the positive ionization mode. Each point represents an individual sample from the methanolic extract (MeOH) and its fractions: hexane (Hex), dichloromethane (DCM), and butanol (BuOH). Colored ellipses indicate 95% confidence regions for each group. The percentage of variance explained by each principal component is displayed on the axes. UHPLC-QTOF-MS: ultra-high-performance liquid chromatography coupled with quadrupole time-of-flight mass spectrometry.
To identify the metabolites driving the separation observed in the PLS-DA (partial least squares discriminant analysis) score plots, the corresponding loading plots (Figure 4) were analyzed for both negative (panel A) and positive (panel B) ionization datasets. The compounds displayed represent the top contributors (highest loadings on component 1) to the discriminative power of the model. In the negative ion mode (Figure 4(a)), LPI (LPI 21:0) and several LPEs, such as LPE 22:4 and LPE 22:5, were among the most influential compounds. STs (e.g. ST 29:1;O;S and ST 27:1;O;S) also exhibited strong discriminative contributions.

PLS-DA loading plots displaying the top discriminant metabolites. (a) Metabolites contributing most to sample separation under negative ionization mode. (b) Metabolites contribute most under the positive ionization mode. Loadings on component 1 are shown on the x-axis. Heatmaps indicate the relative abundance of each metabolite across the four animal-derived mixtures (BuOH, DCM, Hex, MeOH), with a red-to-blue color scale representing high-to-low abundance. Compound classes are abbreviated as in Figure 1, and individual structures follow standard lipid abbreviated nomenclature.
On the other hand, the distribution of STs across the solvent fractions revealed distinct patterns according to ionization mode and fraction polarity. Under negative ionization, although some STs were most abundant in the DCM fraction, the higher ST total content was particularly found in the BuOH and DCM fractions, accounting for 36.52% and 17.46% of the total metabolite content, respectively. A lower proportion was detected in the Hex fraction (2.16%), while STs were observed at very low abundance in the methanol (MeOH) fraction. In contrast, the positive ionization mode revealed that PCs were the predominant lipid class detected, with their relative abundance varying across fractions: 10.64% in DCM, 5.88% in BuOH, 4.82% in Hex, and 3.28% in MeOH.
Molecular networks based on the GNPS molecular networking
To explore and connect the UHPLC-MS-based chemical diversity of the specialized metabolites present in I. sp. aff. badionotus across fractions, we further analyzed the chemical profiles through the GNPS molecular networking computational web platform, using a precursor and fragment ion tolerance of 0.02 Da, and edges were retained only when spectra shared a cosine score above 0.6 with at least five matched peaks. The resulting spectra were then compared against GNPS libraries, keeping only matches with a cosine score above 0.6 and at least five fragment matches. The mass spectral molecular networking job is accessible at: https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=ca5da72bba83495db3b6742f03a46027 and https://gnps.ucsd.edu/ProteoSAFe/status.jsp?task=adcb9f8e3140475b802cc629b64e9b4b. The generated molecular networks (Figure 5) visually represent the chemical space occupied by specialized metabolites in the four solvent fractions. In both negative (panel A) and positive (panel B) ionization modes, the clusters of structurally related metabolites were evident, indicating chemical diversity and specificity across fraction types. Based on mass and fragmentation patterns, molecular networks were then examined.

Molecular networks constructed from LC-MS/MS data of Isostichopus sp. aff. badionotus extract and fractions: (a) negative mode and (b) positive mode. Each node represents a distinct molecular feature (putative metabolite), and edges indicate structural similarity based on MS/MS fragmentation patterns. The node color reflects the origin of the metabolite across the different solvent extract/fraction (dark blue: methanol, MeOH) and fractions (light blue: dichloromethane, DCM; light green: hexane, Hex; dark green: butanol, BuOH). LC-MS/MS: liquid chromatography coupled with mass spectrometry-mass spectrometry.
The network obtained from the dataset in the negative mode (Figure 5(a)), which is particularly suitable for detecting acidic and sulfated compounds such as STs, reveals 13 densely interconnected clusters predominantly colored in light blue and dark green, indicative of the DCM and BuOH fractions, respectively.
These analyses revealed that cluster N1 involved PE-type compounds with different side chains and substitution patterns, which were mainly identified in the Hex and DCM fractions, with a lower proportion in the BuOH, confirming the richness of lipid-like compounds for the less polar fractions (Figure 6(a)). This well-defined cluster shows a clear pattern in mass (e.g. m/z 478.3, 490.2, 506.8), consistent with a PE-like lipid core modified by alkyl chain variations.
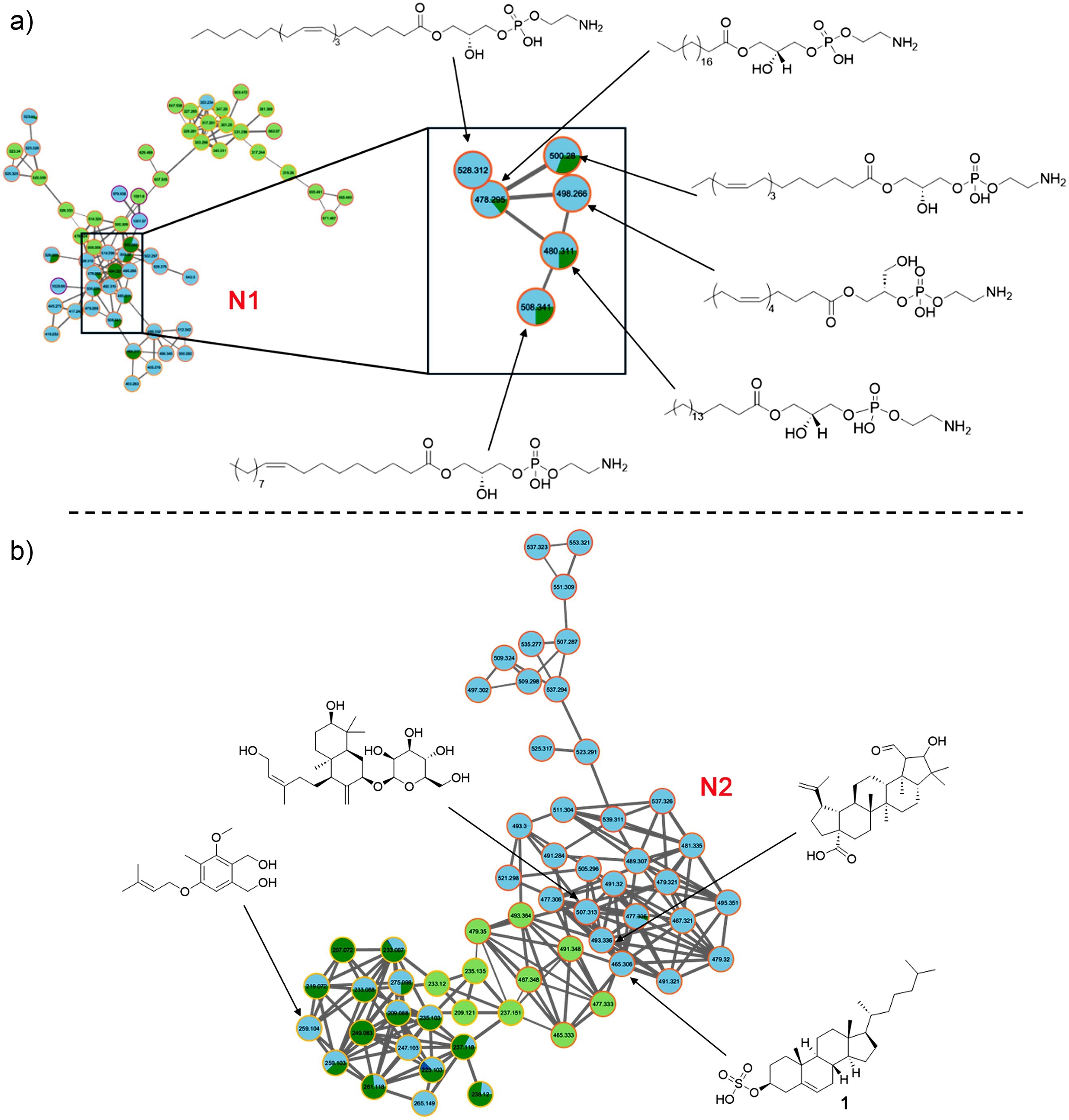
Molecular networks based on GNPS. Identified steroidal compounds and phosphatidylethanolamines (PE). (a) Cluster N1 and (b) cluster N2 of data analysis in negative mode. Light blue node: DCM fraction; dark green node: BuOH fraction; light green node: Hex fraction. GNPS: Global Natural Products Social.
Similarly, STs were grouped within the same cluster (N2) in their [M − H]− form (Figure 6(b)), suggesting structural similarity. This dense network is composed primarily of nodes with light blue and dark green borders, indicating the dominance of metabolites in the DCM and BuOH fractions.
The remaining identified clusters in the negative ion mode dataset (N3–N10) were associated with various lipid classes and other compound types. Cluster N3 (m/z 512–854, [M − H]−) primarily comprised ether-linked phosphatidylethanolamines (PE-O), ether-linked phosphatidylglycerols (PG-O), and LPE, while cluster N4 (m/z 830–928, [M − H]−) involved parkeol derivatives and holostane-type triterpene glycosides. 16 Clusters N5 and N6 (m/z > 564, [M − H]−) were attributed to LPIs and LPCs, respectively. Cluster N7 (m/z < 327, [M − H]−) included fatty acids, and cluster N8 (m/z < 339, [M − H]−) was dominated by sulfated long-chain hydrocarbons. Cluster N9 (m/z < 532, [M − H]−) was assigned to lysoneuroaminophosphatidylethanolamines. Finally, cluster N10 (m/z > 1082) comprised high-molecular-weight, yet unidentified, compounds that may represent complex or conjugated lipids.
A different network topology was observed in the positive mode (Figure 5(b)), where neutral and basic compounds such as PCs ionize more efficiently. The formed clusters (P1–P10) were more evenly distributed among the fractions, although DCM (light blue) and Hex (light green) still dominate, aligning with their richer and diverse chemical profiles. Additionally, more dark blue nodes (MeOH) appeared in this mode compared to the negative mode, suggesting that the methanolic extract contained metabolites more prone to ionization in positive polarity, likely including polar lipids, peptides, or even glycosides.
Detailed analysis of the molecular network obtained in the positive ion mode confirmed a high abundance of [M + H]+ adducts related to PCs, primarily in the DCM and BuOH fractions. Figure 7 displays four relevant clusters (P3, P5, P11, and P12), in which the identified compounds mainly belong to the PC family, with a smaller proportion corresponding to LPCs and phosphatidylinositols.

Molecular networks based on GNPS. Data analysis in positive mode. Light blue node: DCM fraction; dark green node: BuOH fraction; light green node: Hex fraction; dark blue node: MeOH extract. GNPS: Global Natural Products Social.
The compact and well-structured cluster P11 also included metabolites with strong DCM and BuOH representation involving phosphocholine-like species. The similarity between nodes and their consistent MS/MS fragmentation supported a common backbone structure with acyl variation. Their solubility in polar solvents again supported their amphipathic nature and roles in membrane dynamics or bioactive signaling. In addition, clusters P3 and P5 were dominated by light blue (DCM) and dark green (BuOH) node borders. They featured ions in the m/z 490–726 range (e.g. 538.4, 544.3, 550.3, 714.55, 716.57, 726.57), which align with midpolar compounds, for example, oxidized sterols and glycosylated terpene glycosides. The connectivity suggested a shared core structure modified by alkyl chain variations or oxidations. The BuOH preference reflected their higher polarity, while DCM's enrichment hints at a mix of amphipathic or lightly functionalized analogs. Finally, the sparse mini-cluster P12 contained larger molecules (e.g. m/z 763.6, 803.6) with mixed extract/fraction prevalence but notable Hex dominance. Their high mass range and limited network connectivity suggested the presence of complex glycosylated compounds, such as the annotated steroidal metabolite glycosylated 3,12-dihydroxycholan-24-oic acid (m/z 803.5612, [M − H]−) (Figure 7).
Antibacterial activity
In this study, the antibacterial activity of the MeOH extract and its fractions DCM, Hex, and BuOH from I. sp. aff. badionotus was evaluated at an initial concentration of 1000 µg/ml against eight clinically relevant Gram-positive and Gram-negative bacteria.17–19 Growth curve analysis revealed that all fractions effectively inhibited bacterial growth by delaying the growth rate of both Gram-positive and Gram-negative strains, with the DCM fraction exhibiting the most pronounced effect (Figure 8).

Growth curves of bacteria over time following exposure to Isostichopus sp. aff. badionotus extract (MeOH) and its fractions (Hex, DCM, BuOH) at 1000 µg/ml against various bacterial strains: Escherichia coli (a), Klebsiella pneumoniae (b), Shigella flexneri (c), Pseudomonas aeruginosa (d), Staphylococcus aureus (e), Bacillus cereus (f), Staphylococcus epidermidis (g), and Staphylococcus warneri (h). C+ = positive control (gentamicin, 40 μg/ml); NC− = negative control (1% DMSO).
The DCM fraction exhibited significant antibacterial activity, particularly against E. coli (Figure 9(a)), with an inhibition rate of 76.5 ± 2.51%, and against several Gram-positive bacteria: S. aureus (62.5 ± 3.56%, Figure 9(e)), B. cereus (67.0 ± 2.02%, Figure 9(f)), S. epidermidis (65.5 ± 6.39%, Figure 9(g)), and S. warneri (73.6 ± 6.23%, Figure 9(h)). These values were statistically higher than those obtained with the other fractions (p < 0.05). The Hex and BuOH fractions showed moderate activity for some bacterial strains, with the highest inhibition also observed against E. coli, at 59.8 ± 4.06% and 69.2 ± 4.28%, respectively (Figure 9(a)). For K. pneumoniae, the inhibition values were 38.1 ± 7.27% (Hex) and 46.3 ± 3.00% (BuOH) (Figure 9(b)); for S. flexneri, 45.6 ± 7.64% (Hex) and 42.7 ± 5.18% (BuOH) (Figure 9(c)); for S. aureus, 52.2 ± 3.13% (Hex) and 24.6 ± 2.54% (BuOH) (Figure 9(e)); and for B. cereus, 50.5 ± 3.48% (Hex) and 58.9 ± 5.78% (BuOH) (Figure 9(f)). In contrast, the MeOH extract showed only mild antibacterial activity overall. Still, its efficacy exceeded that of Hex against K. pneumoniae (Figure 9(b)), S. epidermidis (Figure 9(g)), and S. warneri (Figure 9(h)).
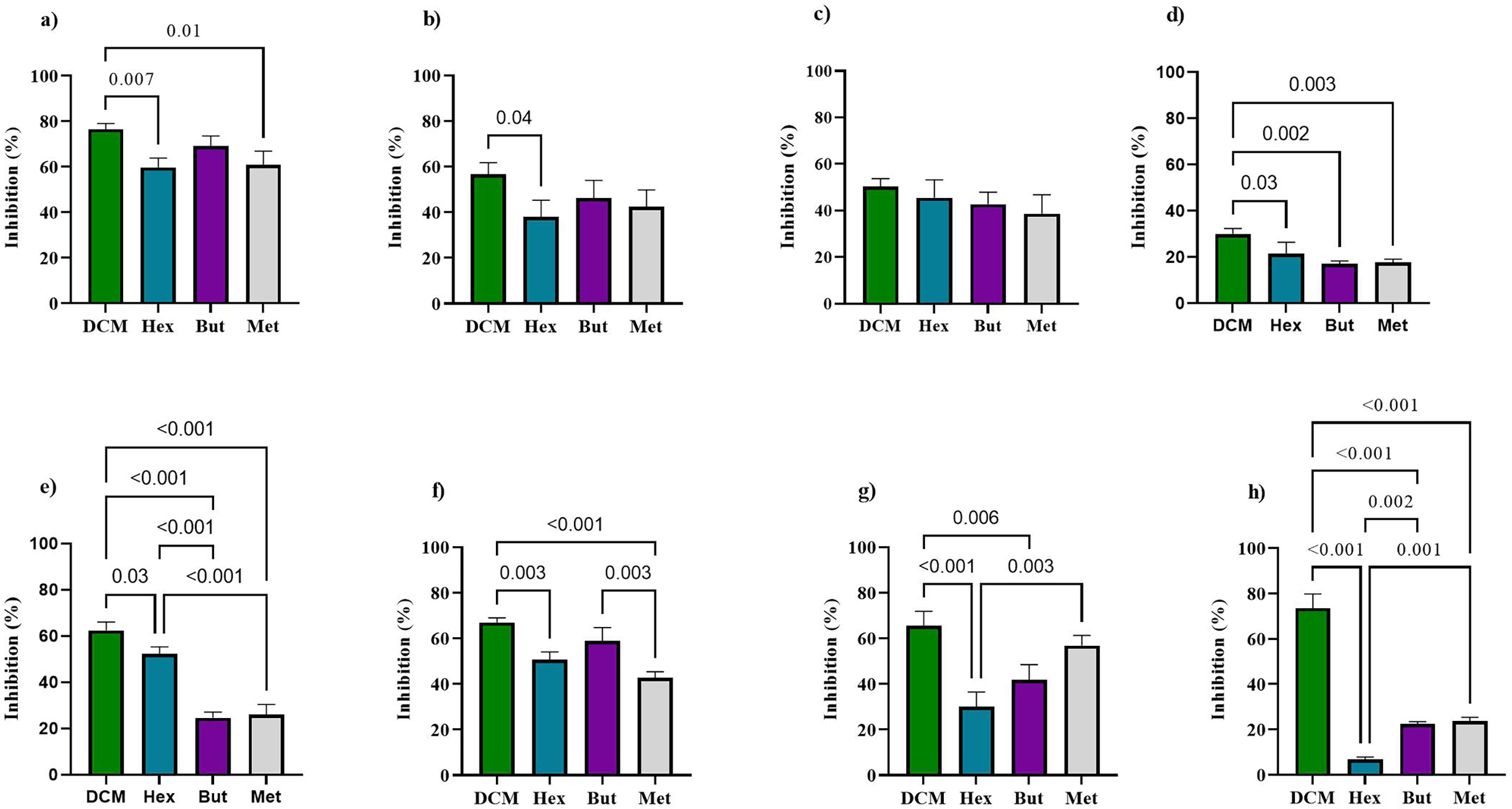
Antibacterial effect of parent extract (MeOH) and its derived fractions (Hex, DCM, BuOH) from Isostichopus sp. aff. badionotus evaluated at 1000 µg/ml against Escherichia coli (a), Klebsiella pneumoniae (b), Shigella flexneri (c), Pseudomonas aeruginosa (d), Staphylococcus aureus (e), Bacillus cereus (f), Staphylococcus epidermidis (g), and Staphylococcus warneri (h).
The MIC of the most active fractions was determined to fall within the 100–1000 µg/ml range. MIC values were calculated using nonlinear regression, fitting the experimental data to the modified Gompertz equation with GraphPad Prism 9.3. 20 The calculated MICs aligned with the observed inhibition percentages (Table 1), confirming the better activity of the DCM fraction.
Minimum inhibitory concentration (MIC) and R2 values obtained from fitting the Gompertz equation for MeOH extract and DCM, Hex, and BuOH fractions against the tested bacterial strains.
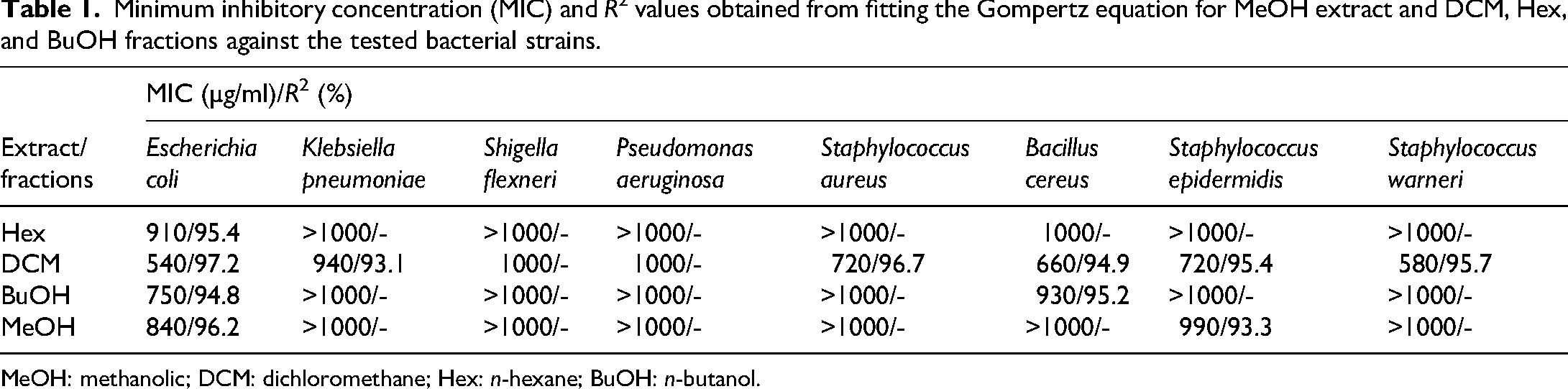

MeOH: methanolic; DCM: dichloromethane; Hex: n-hexane; BuOH: n-butanol.
Discussion
Chemical characterization of extract and fractions from Isostichopus sp. aff. badionotus
The results analyzed through multivariate analysis indicated that the methodological strategy of fractionation was optimal for the bioprospecting of the native sea cucumber, I. sp. aff. badionotus. The DCM and BuOH clusters contained, with high statistical confidence, the classes of metabolites (lysolipids and saponins) responsible for the bioactivity observed in other Holothuroidea. Our study confirms that the extract from this Caribbean species behaves similarly to other more extensively studied sea cucumbers, such as the genera Stichopus and Holothuria.14,17,18,21
The hierarchical clustering of both subsets and fractions revealed distinct chemical profiles among the fractions, emphasizing the efficiency of the liquid–liquid partitioning in segregating metabolites based on solvent removal. The DCM fraction showed the highest diversity and intensity of detected metabolites (especially in positive mode), suggesting the enrichment of moderately polar metabolites (mainly ST, LPE, LPC, LDGCC, and PC) in both subsets. The BuOH fraction also retained several polar and amphiphilic compounds (e.g. LPI and LPA), some overlapping with those in the DCM fraction (ST, LPE, and PC). In contrast, the Hex fraction exhibited a more restricted profile, enriched in a narrower set of nonpolar metabolites (fatty acids, terpenoids, and miscellaneous). The parent MeOH extract exhibited an unfractionated chemical profile, characterized by intermediate levels of many annotated metabolites, and served as a baseline for comparative analysis. This profile likely reflects the presence of residual compounds (e.g. PE and PC) that were not entirely removed during the partitioning process, consistent with its role as the original matrix encompassing a broader diversity and potentially higher abundance of metabolites. Consequently, the clustering patterns and differential distribution confirm that solvent-guided fractionation effectively separated metabolites according to their physicochemical properties. Notably, the BuOH and DCM fractions retained a higher abundance of structurally diverse metabolites, supporting their potential role in the observed biological activities and underscoring their relevance for pharmacological investigation.
The PLS-DA score plots distinguish the samples according to their chemical profiles in both negative and positive ionization modes. The score plot reveals a distinct clustering of all four extract/fraction types, indicating that each extract/fraction harbors a unique metabolite composition. The BuOH and DCM fractions form separate, tight clusters, suggesting well-defined chemical profiles enriched in polar and moderately polar compounds, respectively. The Hex fraction, dominated by nonpolar metabolites, is separated along component 1. Meanwhile, the MeOH extract occupies a unique position between the fractions, reflecting its broader chemical diversity as the parent extract.
As in the negative mode, the fractions form well-separated clusters with minimal overlap. Again, the BuOH and DCM fractions show strong clustering, highlighting their compositional consistency. The Hex fraction appears more distinct in this mode, and the MeOH extract is again positioned between the extremes, further supporting its role as a chemically rich and heterogeneous matrix. The PLS-DA-derived outcome demonstrated that solvent partitioning effectively segregated metabolite classes based on their polarity and ionization properties. The clear separation among extract/fraction types reinforces the utility of this approach in enriching and distinguishing specialized metabolite subsets for further bioactivity evaluation.
The heatmap presented in Figure 2 illustrates a pattern concerning the distribution of metabolites across fractions, suggesting that specific STs, LPIs, and LPEs were notably enriched in the DCM fractions, which reflects the polarity-dependent partitioning of these lipids.
In the positive ion mode, the distinguishing features comprised various PCs, in addition to LPE 20:4 and a glycosylated alkyl-substituted nor-terpenoid. These metabolites also showed strong associations with the DCM fractions, consistent with their intermediate polarity. In contrast, the MeOH extract exhibited moderate levels for most metabolites, underscoring its broad and unspecific metabolite composition as the parent matrix. Collectively, these findings highlight the selective enrichment of lipid subclasses in the DCM fraction. In this regard, particular STs and PCs play a pivotal role in differentiating the fractions, supporting the efficiency of the partitioning strategy to segregate lipid species and providing a chemical rationale for future targeted downstream bioactivity assays.
As noted in the Results, based on the distribution of STs across the solvent fractions, distinct patterns were revealed according to the ionization mode and the polarity of the fraction. These results demonstrate that DCM and BuOH fractions are richer in key specialized metabolites, while the MeOH extract contains lower proportions of both ST and PC families.
Within the organisms belonging to the family Stichopodidae, metabolites associated with the families of PC, PE, LPC, LPE, LPI, and SM were identified through mass spectrometry analysis of the lipid fraction of Apostichopus japonicus.19,20,22,23 By comparing our results with those reported in the literature, we can relate that I. sp. aff. badionotus contains a variety of phospholipids similar to the species grouped under the families Cucumariidae, Holothuriidae, and Stichopodidae. 24 This profile serves as a starting point for future research to focus on the purification and functional testing of these lipid extracts, confirming specific activities against relevant pathologies (e.g. infections or inflammation) in in vitro and in vivo models. This finding transforms a potentially underresearched local species (I. sp. aff. badionotus) into a strong candidate for biotechnological research by linking its chemical profile with species that have already demonstrated pharmaceutical value, such as those previously mentioned.
Molecular networks based on the GNPS molecular networking
The groups of structurally related metabolites indicated a significant chemical diversity and specificity among the types of fraction. The color distribution within clusters indicated that certain metabolite families are exclusive to specific fractions. In contrast, others are shared among two or more solvents, revealing the complementary nature of the fractionation. DCM and BuOH share many chemical features, likely due to overlapping polarity ranges, which might explain the shared presence of key sterols and lipids. On the other hand, Hex and MeOH show fewer shared nodes, reflecting their extraction of chemically distinct metabolite sets—nonpolar lipids in Hex and hydrophilic or polar compounds in MeOH. The network analysis confirms that DCM and BuOH fractions harbor the highest chemical diversity, particularly for bioactive specialized metabolites, while the MeOH and Hex fractions show lower complexity or narrower chemical space coverage. These insights reinforce the importance of using a polarity-gradient extraction strategy to capture the broadest spectrum of marine-derived metabolites.
The groups of structurally related metabolites indicated a significant chemical diversity and specificity among the types of extract. Based on the mass and fragmentation patterns, the molecular networks were examined. The results presented from the network based on the dataset in the negative mode corroborate the previous analysis, indicating a significant presence of STs within these two fractions (DCM and BuOH). The Hex fraction (light green) adds only a limited number of nodes, which aligns with its reduced extraction efficiency for polar or ionizable compounds. The metabolites present in the methanolic extract (dark blue nodes) are distributed sparsely, underscoring a relatively lower chemical diversity or ionization efficiency in this mode.
Similarly, the STs were categorized within the same group (N2) in their [M − H]− form, indicating structural similarity, with a notable predominance of metabolites in the DCM and BuOH fractions. The high connectivity suggests a family of structurally related compounds, potentially sterol-like lipids or sulfated triterpenoids, abundant in sea cucumbers and known for their bioactivity. The consistent m/z spacing and node proximity suggest sequential modifications such as hydroxylation or sulfation. In this regard, an abundant ST exhibited an MS2 fragment ion with m/z 96.960 ([M − H]– m/z 465.3061), matching the structure of cholesterol-3-sulfate (1) in the GNPS database, with a cosine score of 0.99. This ST 1 has been previously identified in other sea cucumber species and is known as the most abundant ST in Eupentacta fraudatrix, along with other cholestane derivatives and, to a lesser extent, sulfated stigmastanes. 25
Similarly, metabolite analysis of Psolus fabricii revealed the presence of the same sterols, although at significantly lower concentrations. 26 In the case of C. frondosa, it was reported that 51% of the pure fraction of STs corresponds to cholesterol sulfate. This evidence supports the identification of compound 1 as cholesterol sulfate. In comparison, the precursor ions of other ST-like compounds showed mass differences of 12 Da (+CH2/−H2) ([M − H]− m/z 477.3062) relative to compound 1, while others differed by 14 Da (+CH2) ([M − H]− m/z 479.3205) and 16 Da (+CH2/+H2) ([M − H]− m/z 481.3525), consistent with campesterol-type sterols. Both cholestane (compound 1) and campesterol analogs also exhibited hydroxylated variants (+16 Da, +OH). Additional STs displayed a 29 Da increase (+CH3CH2) ([M − H]− m/z 491.3203) relative to compound 1, indicative of stigmastane-type sterols. These STs have been previously reported in other sea cucumber species.27–29
The ions detected within the m/z range of 490–726 correspond to medium-polarity lipophilic compounds, such as oxidized sterols and glycosylated terpenoid glycosides, which are commonly found in sea cucumbers. In fact, in sea cucumbers, triterpene glycosides (saponins) play a key role in chemical defense. 30 These compounds typically consist of a triterpene aglycone conjugated with oligosaccharides containing xylose/3-O-methylxylose, quinovose, and glucose/3-O-methylglucose. Common forms include sterol glycosides (lanosterol, cholesterol, or parkeol derivatives) with masses of ∼500 Da, and saponins, ranging from biocide (∼850 Da) to hexaosides (∼1400 Da), which may be mono-, di-, or tri-sulfated. 30 Our findings confirm the defensive chemical signature of the Caribbean species (terpenoid glycosides and oxidized, glycosylated sterols), indicating that they utilize the same mechanism to repel their predators or pathogens, which is conserved throughout the Holothuroidea class (for instance, Holothuria and Stichopus).26,31,32 The presence of sugars (xylose, quinovose, glucose) and aglycones (lanosterol, cholesterol, parkeol) also confirms that the biosynthetic pathway in this species is typical of sea cucumbers, further validating the identity of the extract. The alignment of the revealed ions with those described in the databases for sea cucumbers, in addition to validating the UHPLC-MS technique, establishes a reference profile for the lighter and more abundant compounds of the species, which possess significant biological or pharmacological potential.
Antibacterial activity
The antibacterial activity of the MeOH extract and its DCM, Hex, and BuOH fractions from I. sp. aff. badionotus was evaluated at an initial concentration of 1000 µg/mL against eight clinically relevant Gram-positive and Gram-negative bacteria. The results were comparable to those previously reported for other sea cucumbers, with minimum bactericidal concentrations (MBCs) ranging from 0.500 to 10.000 µg/mL.5,14,33
The observed inhibition percentages confirmed a higher activity of the DCM fraction. Recent studies have shown that H. parva, H. scabra, and H. leucospilota exhibit antimicrobial properties. Furthermore, extracts from H. scabra have demonstrated activity against Aeromonas hydrophila, E. coli, Enterococcus sp., K. pneumoniae, P. aeruginosa, Salmonella typhi, S. aureus, V. harveyi, among others.27,33 Our outcome suggests that the synergistic effects of the constituent compounds of I. sp. aff. badionotus present a promising potential for the treatment of infections, particularly those caused by Gram-positive bacteria.
To understand our results, in general, sea cucumbers have increasingly been recognized as a promising source of antibacterial metabolites, with activity demonstrated across multiple species and tissue extracts. Methanol extracts from the gonads and muscle tissue of H. leucospilota inhibited Lactococcus garvieae and A. hydrophila, while Hex fractions were less effective. 34 Similarly, ethyl acetate, methanol, and aqueous extracts from different organs of Holothuria species collected in the Persian Gulf showed stronger activity against Saccharomyces cerevisiae and S. epidermidis, particularly in the methanolic extract. 35 In another comparative study, chloroform and Hex extracts of the body wall and intestine of H. leucospilota significantly reduced the growth of P. aeruginosa, while methanol extracts of the intestine and gonads exhibited MIC and MBC values ranging from 2.5 to 10 mg/ml. 36 Ethyl acetate extracts of Holothuria forskali were evaluated against Bacillus subtilis, B. cereus, E. coli, and P. aeruginosa, and demonstrated antibacterial activity particularly against E. coli and B. subtilis. 37 Furthermore, methanol:ethyl acetate extracts of H. scabra, H. atra, and Stichopus sp. from the Philippines exhibited broad-spectrum antibacterial activity, with MIC values ranging from 0.0008 to 0.0944 mg/ml against S. aureus, B. subtilis, E. coli, and P. aeruginosa. 38 More recently, saponins isolated from the cuvierian tubules of H. leucospilota have shown to inhibit A. hydrophila with an MIC of 30 µg/ml and an MBC of 80 µg/ml, while also disrupting quorum sensing and biofilm formation. 34
In our case, the observed differences in antibacterial activity may be attributed to structural differences between Gram-positive and Gram-negative bacterial membranes. Gram-positive bacteria possess a simpler cell wall composed primarily of a thick peptidoglycan layer, which facilitates the permeation of bioactive compounds. 20 This increased permeability enhances compound access to critical cellular targets, leading to disruption of essential bacterial processes and, ultimately, cell death. 22 In contrast, Gram-negative bacteria have a more complex cell envelope, consisting of both outer and inner membranes and an intervening layer rich in lipopolysaccharides (LPS). This multilayered structure provides a robust barrier, contributing to resistance against antibiotics, disinfectants, and other lytic agents in the environment.35–39
Structurally, the steroidal compounds identified in the most active DCM and BuOH fractions possess both hydrophilic and hydrophobic regions, imparting amphipathic properties. This structural polarity is associated with antimicrobial activity, particularly against Gram-positive bacteria, through mechanisms that include destabilization of membrane integrity and increased permeability. These disruptions lead to membrane rupture and leakage of intracellular contents, ultimately resulting in bacterial death.40–45 These factors may explain the differential antibacterial effects observed between the DCM fraction and the other fractions tested.
The antibacterial activity demonstrated by the test materials from the sea cucumber I. sp. aff. badionotus makes a valuable contribution to the field of natural product research. This work supports ongoing efforts to discover potent specialized metabolites from marine sources with activity against clinically relevant pathogens. In recent years, many studies have focused on exploring natural resources due to their diverse array of molecules with antibacterial potential.46–48 Such research has emphasized mechanisms of action targeting bacterial membrane proteins, efflux pumps, biofilm formation, and quorum sensing, all of which play crucial roles in developing antimicrobial resistance. Natural products may act independently or serve as complementary agents to enhance the efficacy of conventional antibiotic therapies.
Conclusions
This study underscores the potential of marine-derived natural products, particularly those from sea cucumbers, as valuable sources of antibacterial agents. The extract and fractions from I. sp. aff. badionotus, particularly the DCM fraction, exhibited reasonable antibacterial activity against both Gram-positive and Gram-negative bacteria. This bioactivity is primarily attributed to the presence of STs and triterpenes, which were found in high abundance in the DCM fraction. Complementary metabolomic profiling revealed a chemically diverse set of specialized metabolites, including STs and PCs, that may synergistically contribute to the observed antimicrobial effects. These results highlight the relevance of marine invertebrates as underexplored reservoirs of bioactive compounds and support their continued investigation as alternative solutions to address the global threat of antibiotic resistance. Further bioactivity-guided fractionation and mechanistic studies are warranted to assess their therapeutic potential fully.
Footnotes
Acknowledgments
YAR-N acknowledges Project FONDECYT No. 11241068. MB acknowledges the FONDECYT project No. 3220756. MET-M acknowledges Ministerio de Ciencias Tecnología e Investigación de Colombia (BPIN 2019000100048). EC-B acknowledges Universidad Militar Nueva Granada (Project EXT-CIAS-3854).
ORCID iDs
Ethical considerations
This research has the endorsement of the Research Ethics Committee of the University of Magdalena, Colombia created by rectoral resolution 427 of 2018 and by resolution 1861 of December 26, 2024, which provides access to Genetic Resources and Derived Products # 400 signed between the Ministry of Environment and Sustainable Development and the Universidad del Magdalena.
Author contributions
Conceptualization: YAR-N, MET-M, MB, and AR-F; investigation: YAR-N, MET-M, MB, EC-B, and FE-A; writing—original draft preparation: YAR-N, MET-M, MB, and FE-A; writing—review and editing: YAR-N, EC-B, AR-F, and MET-M. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministerio de Ciencia, Tecnología e Innovación, General System of Royalties BPN 2019000100048, Vice Rectory of Research at the Universidad del Magdalena, Universidad Militar Nueva Granada, Fondo Nacional de Desarrollo Científico y Tecnológico (grant numbers 256-2356987, EXT-CIAS-3854, 11241068, 3220756).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The original contributions presented in the study are included in the article. Further inquiries can be directed to the corresponding authors.

