Abstract
Objective:
Endometrial cancer (EC) is a malignant tumor with various histological subtypes and molecular phenotypes. The evaluation of drug resistance is important for cancer treatment. Progesterone resistance is the major challenge in EC. Knowledge of drug resistance in EC is important in the development of novel therapies.
Methods:
In this study, ten paracancerous and ten tumor tissues were collected to measure the expression of insulin-like growth factor 2 mRNA binding protein 3 (IGF2BP3) and high-mobility group protein 1 (HMGA1). AN3CA and Ishikawa cells were used to explore the effects of IGF2BP3 on EC.
Results:
The expression levels of IGF2BP3 and HMGA1 were higher in EC tumor tissues than in paracancerous tissues. IGF2BP3 and HMGA1 are highly expressed in cisplatin-resistant EC cells. IGF2BP3 knockdown decreased the growth of cisplatin-resistant EC cells. Knockdown of IGF2BP3 decreased the level of HMGA1 protein. HMGA1 knockdown decreased the growth of cisplatin-resistant EC cells.
Discuss and conclusions:
The findings demonstrate that IGF2BP3 is upregulated in EC and closely regulates the growth of drug-resistant EC cells via HMGA1. The findings will inform the development of novel therapies for EC.
Introduction
Endometrial cancer (EC) is a malignant tumor with various histological subtypes and molecular phenotypes. 1 EC is one of the main diseases that burden public health, affecting an estimated 142,000 women worldwide each year, with approximately 42,000 women dying annually due to EC. 1 The 5-year overall survival rate of patients with EC is approximately 80% at all stages. 1 The global incidence of EC continues to increase. 2 Although most patients are diagnosed at an early stage, the 5-year survival rate is quite low. 2 In the United States, the population with EC is doubling annually. 3
EC comprises types I and II, 4 caused by multiple factors, and they often involve excessive unopposed exposure of the endometrium to estrogen, including unopposed estrogen therapy, early menarche, late menopause, tamoxifen therapy, nulliparity, infertility or failure to ovulate, polycystic ovary syndrome, and others. 4 EC is induced by germline mutations, with Lynch syndrome displaying one of the strongest associations. 4 Postmenopausal bleeding is a common characteristic of EC. 4 Nonsurgical options can be used for low- to medium-risk endometrial hyperplasia. 4 For young patients, conservative medroxyprogesterone acetate (MPA) treatment is one of the optimal therapies; however, new therapies are required. 4
In cancer treatment, it is important to detect drug resistance. Progesterone resistance is a major challenge in EC. 5 Progesterone resistance occurs in patients with advanced and recurrent EC, which limits the effectiveness of treatment. 5 Cisplatin-based chemotherapy and radiotherapy, together with surgery used as first-line treatments, often result in resistance. The recurrence rate of advanced EC is relatively high, and the 5-year survival rate of patients with EC is low because of drug resistance. 6 However, the mode of action of drug resistance in EC remains unclear and requires further exploration.
As one member of the insulin-like growth factor messenger RNA-binding protein (IGF2BP) family, IGF2BP3 is considered essential for the functions of IGF2. 7 IGF2BP3 plays an important role in cellular development. 7 Accumulating evidence suggests that IGF2BP3 is closely involved in tumorigenesis, as the gene is highly expressed in a variety of tumor tissues compared to that in adjacent normal tissues, including different types of tumors.8-10 Thus, it is attractive to explore effects of IGF2BP3 on the development of EC.
High-mobility group A1 (HMGA1) is essential for important cellular activities such as chromatin remodeling. 11 Various diseases are induced by HMGA1 dysfunction. 11 The expression of HMGA1 is often abnormal in different types of cancer. 12 A strong correlation has been demonstrated between HMGA1 expression and the developmental stage and survival rate of colorectal cancer. 12 Thus, it is important to investigate the effects of HMGA1 on EC.
In this study, we determined the expressions of IGF2BP3 and HMGA1 in paracancerous and cancerous tissues of EC. In parallel, IGF2BP3 and HMGA1 expression were detected in cisplatin-resistant AN3CA and Ishikawa cell lines. These EC cell lines were used to explore the involvement of IGF2BP3 in the role of HMGA1 in EC cell growth using colony formation and wound healing assays. The findings will inform the development of novel treatments for EC.
Materials and methods
Paracancerous and tumor tissues
Ten paracancerous (paracancerous tissue was defined as the tissue less than 2 cm away from the tumor edge) and ten tumor tissue samples were obtained from the bio-bank of Hebei Medical University Third Hospital. All procedures involving human participants were performed following the ethical standards of the institutional and/or national research committee(s) and the 2013 Helsinki Declaration and its later amendments or comparable ethical standards. Approval was obtained from the ethical board of The Third Hospital of Hebei Medical University (approval no. S2021-022-1). Informed consent was obtained from all participants.
Cell culture
Ishikawa was purchased from Procell with STR authentication by manufacturer (catalog number: CL-0283, Wuhan, China, https://www.procell.com.cn/view/634.html) and AN3CA was purchased from Procell with STR authentication by manufacturer (catalog number: CL-0505, Wuhan, China, https://www.procell.com.cn/view/979.html) EC cell lines were maintained in Dulbecco's modified Eagle's medium (DMEM; cat. no. D6429-500ML; MilliporeSigma) supplemented with 10% fetal bovine serum (FBS; cat. no. MFCD00132239; MilliporeSigma), 1% l-glutamine (cat. no. 25030081; Invitrogen; Thermo Fisher Scientific, Inc.), and 1% penicillin-streptomycin solution (cat. no. V900929-100ML; MilliporeSigma) at 37˚C in a 5% CO2 incubator. The cells were subcultured when they were 80% confluent. Both cell lines were regularly tested for Mycoplasma using a MycoAlert Plus kit (Lonza Group, Ltd) to ensure that they were not contaminated with mycoplasma.
Generation of cisplatin-resistant cells
Ishikawa and AN3CA cells were cultured at 37˚C in a humidified 5% CO2 incubator in DMEM supplemented with 10% FBS for a period not exceeding 3 weeks. Cisplatin-resistant cells were generated by incubating wild-type cells in a medium with 2 μM cisplatin (DDP) for 4 weeks, followed by 10 μM cisplatin (DDP) for 4 weeks. Successful DDP resistance of AN3CA cells and Ishikawa cells was demonstrated as shown in supplementary Figure 1.

mRNA expression level of IGF2BP3 and HMGA1 is higher in EC tumor tissue compared to paracancerous tissue detected by qRT-PCR. (A) mRNA expression level of HMGA1 is significantly higher in EC tumor tissue compared to paracancerous tissue detected by qRT-PCR (n = 10, ****p < 0.0001). (B) mRNA expression level of IGF2BP3 is significantly higher in EC tumor tissue compared to paracancerous tissue detected by qRT-PCR (n = 10, ****p < 0.0001). NOTE: fold of change was normalized to paracancerous tissue.
Western blot analysis
RIPA cell lysis buffer (Beyotime) was used to cleave proteins for 30 min on ice. The lysed cells were centrifuged at 12,000 × g for 10 min at 4˚C. Protein in the supernatant was quantified using the BCA Protein Assay kit (Thermo Fisher Scientific, Inc.) before the addition of 5× SDS and boiling for 10 min. Thirty micrograms of total protein were added to 12% SDS-PAGE gels for electrophoretic separation. The resolved proteins were transferred to 0.45 µm Polyvinylidene Fluoride (PVDF) membranes (Amersham Hybond; GE Healthcare). The membranes were blocked with 1% bovine serum albumin (Amresco) for 2 h and then incubated overnight on a shaker at 4˚C with 1:1000 dilutions of antibodies to IGF2BP3, HMGA1, and β-actin (HPA002037, ZRB2154-25UL, and SAB5500001-100UL, respectively; all from Sigma-Aldrich). The membranes were washed three times for 10 min each time with Tris-buffered saline containing 0.1% Tween-20 (TBS-T) at room temperature, incubated in a 1:2000 dilution of horseradish peroxidase-conjugated goat anti-rabbit IgG H&L (ab7090; Abcam) for 1 h, and then washed. Finally, the membranes were exposed to an enhanced chemiluminescence substrate detection solution (Lulong Biotech).
Immunofluorescence staining
AN3CA and Ishikawa cells were plated following the procedure mentioned above for immunofluorescence staining. The cells were allowed to reach confluence within two days. Fixation was carried out by incubating the cells with 4% paraformaldehyde (PFA) (P6148, Sigma-Aldrich) diluted in 1X PBS (LM-S2041/500, Biosera) for 25 min at room temperature. Following aspiration of the 4% PFA, the cells were washed three times for 5 min each with 1X PBS.
A blocking solution was prepared using 5% normal goat serum and 0.3% TritonX (9002-93-1, Sigma-Aldrich) in 1X PBS, and it was applied for 2 h at room temperature in a humidified chamber. The primary antibody used was anti-P Glycoprotein (catalog number: ab129450), applied at a 1:200 ratio. After aspirating the blocking solution, the cells were washed three times for 5 min with 1X PBS, and the primary antibodies were then added and allowed to incubate at +4°C for 24 h. A negative control using 1X PBS was included. For the secondary antibody, goat-anti-rabbit Alexa Fluor 488 (ab150077, Abcam) was prepared in a 1:200 dilution solution. Following aspiration of the primary antibody, the cells were washed three times for 5 min with 1X PBS. The secondary antibody was added to the wells and incubated at room temperature in a humidified chamber for 2 h. After the incubation period, the cells were washed three times for 5 min with 1X PBS. Finally, a coverslip solution containing DAPI (00-4959-52, Invitrogen) was used, and cell images were captured using a laser scanning confocal microscope (Zeiss LSM 800).
Co-IP
Co-IP was performed to confirm protein-protein binding. Briefly, Ishikawa and AN3CA cells were extracted with ice-cold low-salt lysis buffer. The cell lysate was incubated with antibodies to IGF2BP3 (HPA002037; Sigma-Aldrich), HMGA1 (ZRB2154-25UL; Sigma-Aldrich), or IgG (#2729, Cell Signaling Technology) overnight on a turning wheel at 4°C. PureProteome™ Protein A/G Mix Magnetic Bead System (#LSKAGAG10, Thermo Fisher Scientific, Inc.) was used to purify the co-IP complex. Low-salt lysis buffer was used to wash the beads. The co-IP complex was analyzed using SDS-PAGE and western blotting.
RNA isolation and reverse transcription-quantitative (RT-q)PCR
TRIzol (Beyotime) was used to isolate total RNA from both cell lines (seeded in 96-well plates at a seeding density of 6000 cells/well) following the manufacturer's protocol. BeyoRT First-Strand cDNA Synthesis Kit (cat. no. D7166; Beyotime) was used for cDNA synthesis from the total RNA. RT-qPCR was performed using BeyoFast SYBR Green qPCR Mix (2X; cat. no. D7260-25 ML; Beyotime) on a 7500 Fast Real-time PCR System (Applied Biosystems, Thermo Fisher Scientific, Inc.). RT-qPCR was performed at 50˚C for 2 min, 95˚C for 2 min, 40 cycles of 95˚C for 15 s, 60˚C for 1 min, extension at 72˚C for 1 min, and a final extension step at 72˚C for 10 min. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the internal control. The relative gene expression levels were calculated using the 2−ΔΔCq method. 13 The primers used in this study are listed in Table 1. This experiment was repeated three times.
Primers of reverse transcription-quantitative PCR used in the present study.

Small interfering (si)RNA-based knockdown assay
Gene knockdown was performed using siRNAs. The sequences of siRNAs against IGF2BP3 and HMGA1 were designed using siRNA-Target-Finder (GenScript). These siRNAs were synthesized by and purchased from SynBio Technologies. The sequence of the empty vector siRNA-negative control (NC) was 5′-UUCUCCGAACGUGUCACGU-3′. The NC of siRNA-IGF2BP3 was the pooled sequence of two siRNAs: 5′-AAGACTGGCTACGCGTTCGTG-3′ and 5′-AAGCTAGACAAGCACTAGACA-3′. The siRNA for HMGA1 was the pooled sequence of two siRNAs: 5′-AAGATCCGCATTTGCTACCAG-3′ and 5′-AAGAAGATCCGCATTTGCTAC-3′.
The siRNAs (non-targeting control siRNA and target siRNA) were transiently transfected into Ishikawa and AN3CA cells using FuGENE HD Transfection Reagent (cat. no. E2311; Promega Corporation) according to the manufacturer's instructions at 37˚C in a 5% CO2 incubator. siRNA transfection was performed for 24 h before the subsequent experiments. The knockdown efficiency was evaluated using RT-qPCR and western blot assays, following the protocols described in this study.
Colony formation assay
Ishikawa and AN3CA cells (500 cells/well) were seeded in a 6-well plate and cultured for 14 days in a culture medium containing 10% FBS. The culture solution was discarded, and the cells were infiltrated with methanol for 10 min, stained with crystal violet, washed with water, dried, and counted.
Wound healing assay for colorectal cancer cell migration
Ishikawa and AN3CA cells were seeded in the wells of a 6-well plate. After the cells had grown to 100% confluence, the cell layer was scratched with a 20-µL pipette tip, and the medium containing 10% FBS was replaced with a serum-free medium. Cell images were captured at 0, 24, 48, and 72 h.
Statistical analyses
All data are represented as the mean ± standard error of the mean. One-way analysis of variance and Tukey's post-hoc test were used for the statistical analysis of continuous variables. Categorical variables were analyzed using Fisher's exact tests. Correlation analysis (Pearson) and statistical analyses were performed using GraphPad Prism software (version 5.0; GraphPad Software, Inc.). p < 0.05 was considered statistically significant.
Results
Expression levels of IGF2BP3 and HMGA1 are higher in EC tumor tissue than that in paracancerous tissue
IGF2BP3 and HMGA1 are closely associated with various cancers. Thus, it was interesting to explore the expression levels of IGF2BP3 and HMGA1 in EC tumor tissue compared to paracancerous tissue. The expression levels of HMGA1 (Figure 1(A)) and IGF2BP3 (Figure 1(B)) were higher in EC tumor tissue than that in paracancerous tissue.
Expression levels of IGF2BP3 and HMGA1 are higher in cisplatin-resistant EC cell lines
Since some literature have showed that IGF2BP3 and HMGA1 are involved in activities of different cancers, thus it is important to explore the effects of these two genes on cisplatin-resistant EC cell lines. To investigate the role of IGF2BP3 and HMGA1 in cisplatin-resistant EC cells, cisplatin-resistant AN3CA and Ishikawa cell lines were established, and cisplatin-resistant features were confirmed, respectively (Figure 2(A) and (B)). IGF2BP3 mRNA expression level was higher in cisplatin-resistant AN3CA cells (Figure 2(C)) and Ishikawa cells detected by qRT-PCR (Figure 2(D)). In parallel, HMGA1 mRNA expression level was higher in cisplatin-resistant AN3CA cells (Figure 2(E)) and Ishikawa cells detected by qRT-PCR (Figure 2(F)).
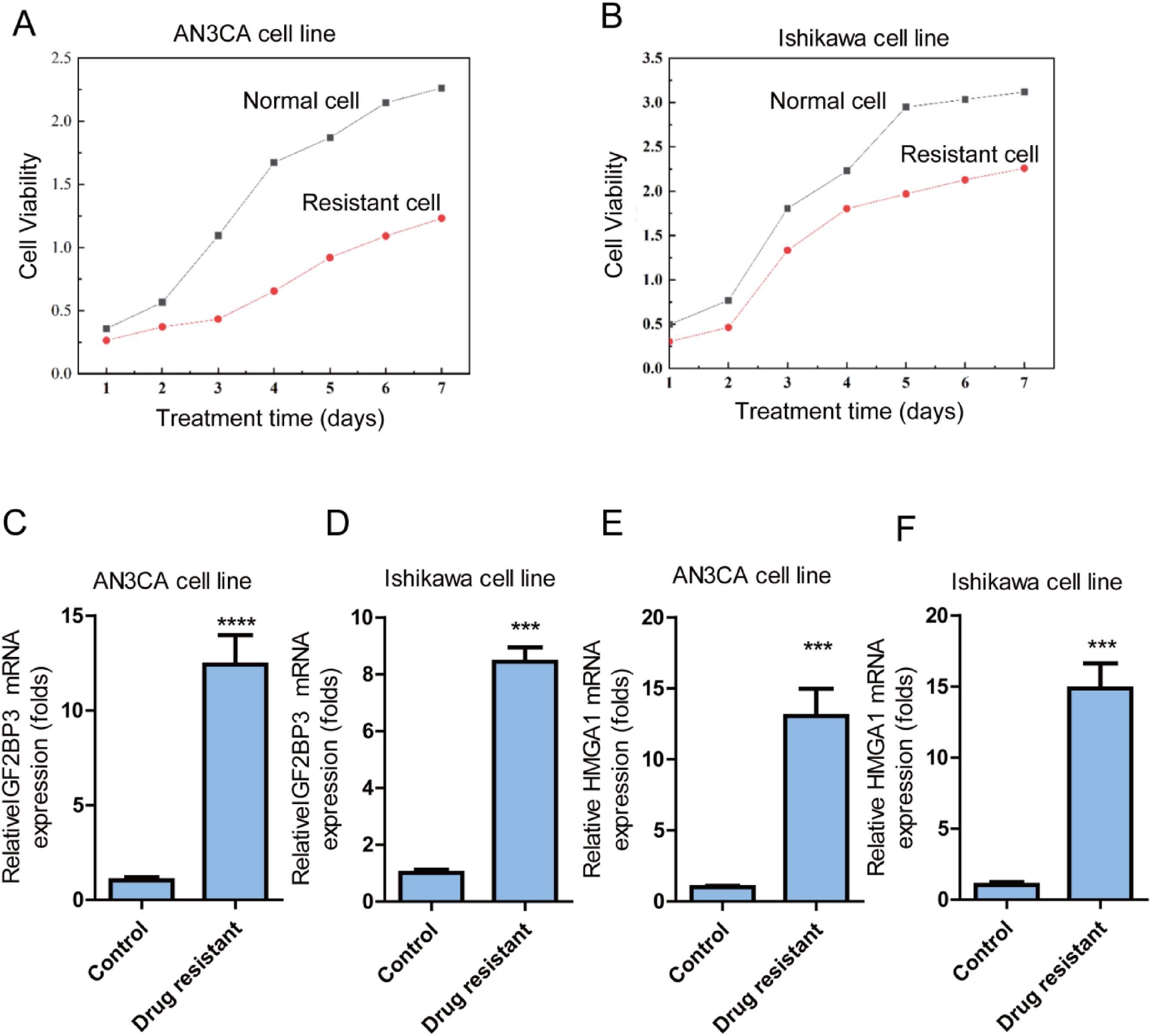
mRNA expression level of IGF2BP3 and HMGA1 is higher in cisplatin-resistant EC cell lines detected by qRT-PCR. (A) cisplatin-resistance was confirmed in AN3CA cell line; (B) cisplatin-resistance was confirmed in Inshikawa cell line; (C) mRNA expression level of IGF2BP3 is higher in cisplatin-resistant AN3CA cell line detected by qRT-PCR. (D) mRNA expression expression level of IGF2BP3 is higher in cisplatin-resistant Inshikawa cell line detected by qRT-PCR. (E) mRNA expression expression level of HMGA1 is higher in cisplatin-resistant AN3CA cell line detected by qRT-PCR. (F) mRNA expression expression level of HMGA1 is higher in cisplatin-resistant Ishikawa cell line detected by qRT-PCR.
Knockdown of IGF2BP3 potently inhibits the growth of cisplatin-resistant AN3CA cells
To evaluate the effects of IGF2BP3 on the growth of cisplatin-resistant AN3CA cells, the gene for IGF2BP3 was successfully knocked down in AN3CA cells (Figure 3(a)). IGF2BP3 knockdown significantly decreased colony formation in AN3CA cells (Figure 3(b) and (c)). Additionally, IGF2BP3 knockdown significantly decreased the migration of AN3CA cells (Figure 3(d) and (e)). These results demonstrate that IGF2BP3 knockdown potently inhibits the growth of cisplatin-resistant AN3CA cells.

Knockdown of IGF2BP3 by siRNA potently inhibited growth of cisplatin-resistant AN3CA cells. (A) IGF2BP3 was successfully knock down on AN3CA cells using siRNA against IGF2BP3 detected by western blot. (B) IGF2BP3 knockdown significantly decreased colony formation on AN3CA cell line. (C) quantification of colony formation from (B). (D) IGF2BP3 knockdown significantly decreased migration of cisplatin-resistant AN3CA cells detected by wound healing assay. (E) Quantification of distance of cellular migration in isplatin-resistant AN3CA cells.
Knockdown of IGF2BP3 potently inhibits the growth of cisplatin-resistant Ishikawa cells. To evaluate the effects of IGF2BP3 on the growth of cisplatin-resistant Ishikawa cells, IGF2BP3 siRNA was used to successfully knockdown the gene in cisplatin-resistant Ishikawa cells (Figure 4(A)), which was quantified as well (Figure B). IGF2BP3 knockdown significantly decreased colony formation by Ishikawa cells (Figure 4(C) and (D)). In addition, IGF2BP3 knockdown significantly decreased the migration of Ishikawa cells (Figure 4(E) and (F)). Collectively, these results demonstrate that IGF2BP3 knockdown potently inhibited the growth of cisplatin-resistant Ishikawa cells.

Knockdown of IGF2BP3 by siRNA potently inhibited growth of cisplatin-resistant Ishikawa cells. (A) IGF2BP3 was successfully knockdown by siRNA against the gene on Ishikawa cells detected by western blot. (B) IGF2BP3 knockdown significantly decreased colony formation on Ishikawa cell line detected by western blot. (C) Quantification of colony formation of IGF2BP3 knockdown affecting Ishikawa cell line. (D) IGF2BP3 knockdown by siRNA against the gene significantly decreased migration of Ishikawa cells measured by wound healing assay. (E) Quantification of migration of Ishikawa cells affected by IGF2BP3 knockdown (n = 4, *p < 0.05).
Knockdown of HMGA1 potently inhibits the growth of cisplatin-resistant AN3CA cells
The description that IGF2BP proteins may interact with HMGA proteins 14 prompted the assessment of whether IGF2BP3 could affect HMGA1. Knockdown of IGF2BP3 inhibited the production of HMGA1 protein (Figure 5(A) and (B)). To further investigate the effects of HMGA1 on the growth of cisplatin-resistant AN3CA cells, HMGA1 siRNA was used to knockdown the gene in AN3CA cells (Figure 5(C) and (D)). To investigate whether IGF2BP3 has interaction with HMGA1, a co-immunoprecipitation assay was performed using anti-IGF2BP3 antibodies, which indicated the interaction between IGF2BP3 and HMGA1 protein in AN3CA cells (Figure 5(E)). To further explore the interaction between IGF2BP3 and HMGA1, a co-immunoprecipitation assay was performed using anti-HMGA1 antibody, which indicated the interaction between IGF2BP3 and HMGA1 protein in AN3CA cells (Figure 5(F)). HMGA1 knockdown significantly decreased colony formation in AN3CA cells (Figure 5(G) and (H)). In addition, HMGA1 knockdown significantly decreased the migration of AN3CA cells (Figure 5(I) and (J)). Collectively, these results demonstrate that HMGA1 knockdown potently inhibits the growth of cisplatin-resistant AN3CA cells.

Knockdown of HMGA1 by siRNA potently inhibited growth of cisplatin-resistant AN3CA cells. (A) knockdown of IGF2BP3 inhibited protein level of HMGA1 detected by western blot assay. (B) quantification of protein level of HMGA1 detected by western blot assay. (C) siRNA against HMGA1 knockdown HMGA1 on AN3CA cells detected by western blot assay. (D) quantification of HMGA1 protein on HMGA1 knockdown AN3CA cells. (E) Co-IP study indicated that IGF2BP had interaction with HMGA1 on AN3CA cells with IGF2BP3 antibody. (F) Co-IP study indicated that IGF2BP had interaction with HMGA1 on AN3CA cells with HMGA1 antibody. (G) HMGA1 knockdown by siRNA significantly decreased colony formation on AN3CA cell line detected by colony formation assay. (H) Quantification of colony formation of HMGA1 knockdown affecting AN3CA cell line. (I) HMGA1 knockdown significantly decreased migration of AN3CA cells measured by wound healing assay. (J) Quantification of migration of AN3CA cells affected by HMGA1 knockdown (n = 4, **p < 0.01).
Knockdown of HMGA1 potently inhibits the growth of cisplatin-resistant Ishikawa cells
To evaluate the effects of HMGA1 on the growth of cisplatin-resistant Ishikawa cells, knockdown of IGF2BP3 inhibited the production of HMGA1 protein (Figure 6(A) and (B)). To further investigate the effects of HMGA1 on the growth of cisplatin-resistant Ishikawa cells, HMGA1 siRNA was used to knockdown the gene in Ishikawa cells (Figure 6(C) and (D)). To investigate whether IGF2BP3 has interaction with HMGA1, a co-immunoprecipitation assay was performed using anti-IGF2BP3 antibodies, which indicated the interaction between IGF2BP3 and HMGA1 protein in Ishikawa cells (Figure 6(E)). To further explore the interaction between IGF2BP3 and HMGA1, a co-immunoprecipitation assay was performed using anti-HMGA1 antibody, which indicated the interaction between IGF2BP3 and HMGA1 protein in Ishikawa cells (Figure 6(F)). HMGA1 knockdown significantly decreased colony formation in Ishikawa cells (Figure 6(G) and (h)). In addition, HMGA1 knockdown significantly decreased the migration of Ishikawa cells (Figure 6(I) and (J)). Collectively, these results demonstrate that HMGA1 knockdown potently inhibited the growth of cisplatin-resistant Ishikawa cells.

Knockdown of HMGA1 by siRNA potently inhibited growth of cisplatin-resistant Ishikawa cells. (A) Knockdown of IGF2BP3 inhibited protein level of HMGA1 detected by western blot. (B) quantification of protein level of HMGA1 detected by western blot assay. (C) HMGA1 was successfully knock down on Ishikawa cells detected by western blot. (D) quantification of HMGA1 protein on HMGA1 knockdown AN3CA cells. (E) Co-IP study indicated that IGF2BP had interaction with HMGA1 on Ishikawa cells with IGF2BP antibody. (F) Co-IP study indicated that IGF2BP had interaction with HMGA1 on Ishikawa cells with IGF2BP antibody. (G) HMGA1 knockdown by siRNA significantly decreased colony formation on Ishikawa cell line detected by colony formation assay. (H) Quantification of colony formation of HMGA1 knockdown affecting Ishikawa cell line (**p < 0.01). (I) HMGA1 knockdown significantly decreased migration of Ishikawa cells measured by wound healing assay. (J) Quantification of migration of Ishikawa cells affected by HMGA1 knockdown (n = 4, *p < 0.05).
HMGA1 over-expression compromised effects of IGF2BP3 on growth of EC cells
To further explore whether HMGA1 is downstream of IGF2BP3, the IGF2BP3 knockout AN3CA cell line was created and IGF2BP3 knockout was confirmed using western blot assay (Figure 7(A)). Then, HMGA1 over-expression was performed to top of IGF3BP3 knockout in AN3CA cell line, and HMGA1 over-expression was confirmed (Figure 7(B)). Importantly, it was found that HMGA1 over-expression compromised inhibitory effects of IGF2BP3 knockdown on AN3CA cell line (Figure 7(C)). The IGF2BP3 knockout Ishikawa cell line was created and IGF2BP3 knockout was confirmed using western blot assay (Figure 7(D)). HMGA1 over-expression was performed to top of IGF3BP3 knockout in Ishikawa cell line, and HMGA1 over-expression was confirmed (Figure 7(E)). It was found that HMGA1 over-expression compromised inhibitory effects of IGF2BP3 knockdown on Ishikawa cell line (Figure 7(C)). Taken together, it was demonstrated that HMGA1 over-expression compromised effects of IGF2BP3 on growth of EC cells.

HMGA1 over-expression compromised effects of IGF2BP3 on growth of EC cells. (A) Knockout of AN3CA cell line was confirmed using western blot assay. (B) HMGA1 over-expression on top of IGF3BP3 knockout in AN3CA cell line was confirmed. (C) HMGA1 over-expression compromised inhibitory effects of IGF2BP3 knockdown on AN3CA cell line. (D) Knockout of Ishikawa cell line was confirmed using western blot assay. (E) HMGA1 over-expression on top of IGF3BP3 knockout in Ishikawa cell line was confirmed. (F) HMGA1 over-expression compromised inhibitory effects of IGF2BP3 knockdown on Ishikawa cell line.
Discussion
The incidence and disease-associated mortality of EC continue to increase worldwide. 15 Clarification of the pathogenesis of EC is necessary to develop new therapies. We observed markedly elevated expressions of IGF2BP3 and HMGA1 in cancer tissues compared to that in paracancerous tumors. IGF2BP3 and HMGA1 were highly expressed in cisplatin-resistant EC cells. IGF2BP3 knockdown decreased the growth of cisplatin-resistant EC cells. Knockdown of IGF2BP3 decreased the level of HMGA1 protein. The knockdown of HMGA1 decreased the growth of cisplatin-resistant EC cells. The findings demonstrate that IGF2BP3 is potentiated in endometrial tumors and closely regulates the growth of cisplatin-resistant EC cells via HMGA1.
IGF2BP3 regulates tumorigenesis in various types of cancers. 16 The expression level of IGF2BP3 is potentiated in bladder cancer tissues compared to that in normal tissues and has been strongly correlated with disease progression. 17 Moreover, activation of IGF2BP3 was observed to significantly suppress apoptosis and proliferation of bladder cancer cells. 17 In lung cancer, overexpression of IGF2BP3 significantly compromises the effects of tyrosine kinase inhibitor treatment and induces drug resistance by promoting oxidative phosphorylation. 8 IGF2BP3 significantly potentiates the growth of breast cancer cells, induces drug resistance by binding to CD44, and increases IGF2 levels in fibroblasts. 18 Moreover, IGF2BP3 is reportedly inhibited by the let-7 family of miRNAs to regulate the activities of several types of tumors. 19 In this study, IGF2BP3 was highly expressed in EC tumor tissues (Figure 1(A)) and cisplatin-resistant AN3CA and Ishikawa cells (Figure 2). Knockdown of IGF2BP3 potently inhibited colony formation and migration of AN3CA cells (Figures 3 and 4). Overall, the findings demonstrate that IGF2BP3 plays an important role in cisplatin resistance and EC cell growth.
HMGA1 is important in several malignant cancers. 20 In colorectal cancer stem cells, programmed death-ligand 1 significantly activates HMGA1 expression to further activate phosphoinositide 3-kinase/Akt and mitogen-activated protein kinase kinase/extracellular signal-regulated kinase signaling to promote the growth of colorectal cancer stem cells. 20 Zhang et al. demonstrated the up-regulation of HMGA1 expression in gastric cancer and the enhanced cisplatin resistance of gastric cancer cells to the HCP5 gene, mediated by promoting HMGA1 expression by miR-519d-sponging microRNA. 21 Kim et al. demonstrated that a reduction in HMGA1 expression alleviated drug resistance to chemical anticancer drugs in spheroid cells, whereas activation of HMGA1 potentiated drug resistance in ovarian cancer cells. 22 In this study, HMGA1 expression was upregulated in EC tumor tissues (Figure 1) and cisplatin-resistant AN3CA and Ishikawa cells (Figure 2). Knockdown of HMGA1 potently inhibited colony formation and migration of AN3CA cells (Figure 5 and 6). Moreover, it was demonstrated that HMGA1 over-expression compromised effects of IGF2BP3 on growth of EC cells (Figure 7). Overall, these findings demonstrate that HMGA1 is essential for cisplatin resistance and EC cell growth.
The current study highlights IGF2BP3's significant up-regulation in endometrial cancer (EC) and its essential role in the growth regulation of drug-resistant EC cells, particularly through its modulation of HMGA1 levels. This research underscores the promise of targeting both IGF2BP3 and HMGA1 in the development of novel therapeutic strategies for addressing drug-resistant EC, potentially leading to more effective treatment options. However, several limitations of the study need to be addressed. One key consideration is the heterogeneity of endometrial cancer, which encompasses various subtypes with differing clinical behaviors and treatment responses. Investigating the role of IGF2BP3 across these distinct EC subtypes could uncover subtype-specific vulnerabilities that might inform tailored therapeutic approaches. Additionally, the study does not correlate the expression levels of IGF2BP3 and HMGA1 with patient outcomes, such as survival rates and responses to therapy. Establishing these correlations is crucial for understanding the clinical relevance of IGF2BP3 and HMGA1 and their potential as biomarkers in patient stratification. Future research should delve into the molecular pathways involving IGF2BP3 and HMGA1 to elucidate their roles further and explore the feasibility of targeted therapies aimed at these proteins within clinical settings. By understanding IGF2BP3's broader implications across various EC subtypes, researchers could significantly contribute to the personalization of treatment strategies, ultimately improving patient outcomes in endometrial cancer management.
Conclusion
Overall, the expression levels of IGF2BP3 and HMGA1 were higher in EC tumor tissues than that in paracancerous tissues. IGF2BP3 and HMGA1 are highly expressed in cisplatin-resistant EC cells. IGF2BP3 knockdown decreased the growth of cisplatin-resistant EC cells. Knockdown of IGF2BP3 decreased the protein level of HMGA1. HMGA1 knockdown decreased the growth of cisplatin-resistant EC cells. These findings demonstrate that IGF2BP3 is upregulated in EC and is closely regulated by the growth of drug-resistant EC cells via HMGA1. Our findings should inform the development of novel therapies for EC.
Supplemental Material
sj-pptx-1-sci-10.1177_00368504251315008 - Supplemental material for IGF2BP3 is upregulated in endometrial cancer and tightly regulates the growth of drug-resistant endometrial cancer cells via HMGA1
Supplemental material, sj-pptx-1-sci-10.1177_00368504251315008 for IGF2BP3 is upregulated in endometrial cancer and tightly regulates the growth of drug-resistant endometrial cancer cells via HMGA1 by Yiwei Zhang, Yanlai Xiao, Xiangzhai Zhao, Jie Xu, Huan Zhao, Zhaojun Guo, Jun Zhao, Yajing Zhang, Ruoxi Wang and Jian Wang in Science Progress
Footnotes
Authors’ contributions
Y.Z., Y.X., X.Z., J.X., H.Z., Z.G., J.Z., Y.Z., R.W., performed the experiments. Y.Z. and J.W. designed the research. Y.Z. and J.W. wrote the manuscript, and J.W. supervised the project. Y.Z. and J.W. confirm the authenticity of all the raw data. All authors read and approved the final manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Hebei Province Natural Science Foundation Basic Research Project - Precision Medicine Joint Fund Cultivation Project, (grant number H2021206367).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
