Abstract
Peripheral nerve and large-scale muscle injuries result in significant disability, necessitating the development of biomaterials that can restore functional deficits by promoting tissue regrowth in an electroactive environment. Among these materials, graphene is favored for its high conductivity, but its low bioactivity requires enhancement through biomimetic components. In this study, we extrusion printed graphene-poly(lactide-co-glycolide) (graphene) lattice scaffolds, aiming to increase bioactivity by incorporating decellularized extracellular matrix (dECM) derived from mouse pup skeletal muscle. We first evaluated these scaffolds using human-induced pluripotent stem cell (hiPSC)-derived motor neurons co-cultured with supportive glia, observing significant improvements in axon outgrowth. Next, we tested the scaffolds with C2C12 mouse and human primary myoblasts, finding no significant differences in myotube formation between dECM-graphene and graphene scaffolds. Finally, using a more complex hiPSC-derived 3D motor neuron spheroid model co-cultured with human myoblasts, we demonstrated that dECM-graphene scaffolds significantly improved axonal expansion towards peripheral myoblasts and increased axonal network density compared to graphene-only scaffolds. Features of early neuromuscular junction formation were identified near neuromuscular interfaces in both scaffold types. These findings suggest that dECM-graphene scaffolds are promising candidates for enhancing neuromuscular regeneration, offering robust support for the growth and development of diverse neuromuscular tissues.
Introduction
While skeletal muscle is inherently capable of self-repair from minor injuries, large-scale injuries can overwhelm the physiological repair capabilities of the body. Efforts to repair are often inhibited by the destruction of satellite cell microenvironments which lead to fibrotic cascades that develop into function-limiting scar tissue. 1 Functional autologous tissue transfer is the current gold standard of treatment, but donor site availability and morbidity are limiting factors. Other engineering approaches currently being investigated fail to address the complex management of cell types that enable functionality in skeletal muscle tissue. These studies investigate the efficacy of material systems by probing the response of an isolated cell type but fail to consider the support needed by a variety of cell types that must thrive in a shared tissue ecosystem. 2 Large-scale damage to skeletal muscle requires a formative solution that can provide a foundation to support a neuromuscular environment.
One notable shortcoming of existing materials for supporting peripheral nerve and muscle cell expansion and interaction is their limited electroactivity. The electronic properties of materials are highly relevant to both cell types. Neurons and myocytes rely extensively on bioelectronic signals to communicate and manage repair, yet many existing materials do not efficiently facilitate these cues. Several different motifs of electroactive components such as conductive polymers3,4 or metallic nanoparticles have been incorporated into composite materials with varying degrees of success.
Among these bioelectronic materials, graphene has rapidly emerged as a promising compound for promoting migration and regeneration in electrogenic tissue, such as skeletal muscle.5–8 Graphene's high electronic conductivity, structural stability, and form factor diversity all contribute to its success in various bioelectronic applications. 9 Despite these advantages, graphene exhibits relatively low bioactivity lacking physiochemical biomimicry and an ability to initiate specific regenerative cascades.
One possible solution to increase bioactivity is to combine a graphene-based material with decellularized extracellular matrix (dECM) components. 10 dECM-based solutions have the unique ability to simulate growth promoting environments by improving upon cell migration and viability.11,12 ECM-bound growth factors render dECM-based materials with bioactivity that can effectively drive cellular processes towards promising outcomes. 13 Notable drawbacks of current dECM-based scaffolds include their mechanical strength14,15 and electrically inert nature. For these reasons, dECM-based materials fail to promote cell alignment and limit overall skeletal muscle repair. 16
Recent literature has explored combining dECM with graphene to promote cardiac muscle, bone, and neural regeneration.17–19 Notably, a recent study found that doping polydopamine-functionalized reduced graphene oxide onto adipose-derived dECM scaffolds promoted neural stem cell differentiation. 19 However, the use of dECM-graphene scaffolds in neuromuscular co-cultures has yet to be investigated.
In this study, we hypothesized that the inclusion of dECM in graphene-based composites will lead to structurally robust and physiologically potent scaffolds with the capability to drive co-repopulation of motoneurons and myocytes. We incorporated dECM into graphene-based ink to generate 3D extrusion printed dECM-graphene composite scaffolds, evaluated mechanical and electronic properties of the scaffolds, and demonstrated motoneuron and myoblast growth and differentiation. Therefore, we employed 2D motor neuron (MN) cultures for their ease in quantifying live/dead assays, which is essential for evaluating cell viability on the scaffolds, and complemented these with 3D motor neuron spheroids (hMNS) co-cultured with myoblasts to better simulate peripheral nerve growth, where axons grow together in bundles, providing a more physiologically relevant assessment of the scaffold's potential to support neuromuscular growth.
Materials and methods
Preparation of decellularized ECM
Extracellular matrix was acquired from mouse pup muscle tissue under approval from Northwestern's Institutional Animal Care and Use Committee (Study #IS00010080). Muscle tissue was collected from P0-P1 C57BL/6J mouse pups (Jackson Laboratory, Bar Harbor, ME, USA). Tissues were placed in a beaker containing 0.5% sodium dodecyl sulfate (SDS) (Bio-Rad) solution again under constant agitation. The SDS solution was changed every 24 h for 4 to 7 days until the muscle turned completely translucent. Decellularized muscle was washed in DI every 2 to 4 h then overnight to remove residual SDS. All steps involving decellularization were conducted at 4 °C. Decellularized muscle was lyophilized and cryomilled using a 6775 Freezer/Mill Cryogenic Grinder (SPEX SamplePrep LLC, Metuchen, NJ, USA). The powder was resuspended in dichloromethane (DCM), triturated until homogenous, and finally filtered using a 10 μm PTFE filter (LCWP02500, Millipore Sigma, Burlington, MA, USA).
Bioink preparation and 3D printing
Poly-lactide-co-glycolic acid (PLGA) (P2191; Sigma Aldrich, St Louis, MO, USA; 82:18 poly-lactide to poly-glycolide; ρ 1.15 g cm−3) was dissolved in DCM (270997; Sigma Aldrich, St Louis, MO, USA). Graphene (A-12-5G; Graphene Supermarket, Ronkonkoma, NY, USA) was added into a 10:2:1 by mass mixture of DCM, ethylene glycol butyl ether (537551; Sigma Aldrich, St Louis, MO, USA) and dibutyl phthalate (524980; Sigma Aldrich, St Louis, MO, USA). For dECM-graphene inks, dECM was weighed, resuspended in DCM, and then added to the bioink base so that a 2.5% concentration was achieved. The new suspension was sonicated at 40 °C to allow the excess DCM to evaporate. Both of the graphene constructs were fabricated using a 3D BioPlotter (EnvisionTEC, Germany). The printing parameters were set to 30–40 mm/s, 4–5 Bar pressure, and a 200 μm clear tip was used (Nordson Corporation, Westlake OH, USA) with a starting needle offset of 0.45 mm. All graphene constructs were printed onto a smooth glass substrate at room temperature. Each layer was deposited to form 90° angles by offsetting each previous layer by 400 μm in the x and y planes. Previous work has shown that scaffolds with PLGA alone have vastly different properties compared to those that incorporate a powder. 20 Thus, we did not utilize a plastic control in this study and chose only to compare graphene and graphene-dECM scaffolds.
Mechanical testing
Graphene 3D printed scaffolds were cut into 10 × 5 mm strips and loaded into an RSA-G2 Solids Analyzer (Mettler Toledo, Columbus, OH) at a loading gap of approximately 8 mm. Tensile testing was executed with a HENCKY strain rate of 5% for about 120 s. The linear portion of the stress–strain curve was defined using a first degree polynomial fitting (best fit linear regression line) onto a minimum of 30 consecutive points that produced an R2 value of 0.99. The elastic modulus was calculated from the slope of this regression. This process was done in triplicate using three separately produced scaffolds per each scaffold type.
Zeta potential
About 5 mg of each scaffold variant was swollen in DI water to equilibrium state and then frozen in liquid nitrogen. The samples were then pulverized into smaller particles using a Cryomill and resuspended in 5 ml of a 0.01 M NaCl electrolyte solution with 7.4 Ph and filtered using a 70 μm strainer (CLS431751; Corning, NY, USA). Samples were loaded into a Malvern DTS1070 capillary cell and measured on a Malvern Zetasizer NanoZSP, applying the Smoluchowski equation for large particles.21,22
Resistivity measurements
The conductivity of dry graphene and graphene-dECM scaffolds was measured via electrochemical impedance spectroscopy. Contact with the scaffolds was made with pogo pins. Frequency was varied from 0.1 to 106 Hz with an AC amplitude of 10 mV. The average impedance over the frequency range was used to calculate material conductivity.
Scanning electron microscopy
Graphene scaffolds were directly mounted onto pin stubs with carbon tape and coated with 5 nm of osmium tetroxide with a Filgen Osmium Coater OPC-60A prior to imaging. Scaffolds were imaged by scanning electron microscopy (SEM, Hitachi 4800-S). Images were captured using Hitachi S-4800 PC-SEM software.
Human and C2C12 myoblast culture
The C2C12 mouse myoblast cell line (CRL-1772; ATCC, Manassas, VA, USA) was cultured following the manufacturer's protocol. Briefly, C2C12 myoblasts were seeded at a density of 5.0 × 103 cells/cm2 onto 0.5% gelatin-coated dishes (100 mm × 20 mm) containing 10 mL of growth medium, containing Ham's F10 (Corning) with 20% Fetal Bovine Serum (from Gibco), 10 mg/ml streptomycin and 100 units/ml penicillin (Gibco). Growth medium was changed every other day. Primary myoblasts from human hamstring muscle (passage 5–6) were seeded at the same density and cultured using the same growth medium complemented with human recombinant bFGF (5 ng/ml) (BD). Primary human myoblasts were a gift from Dr Andrea A. Domenighetti (Shirley Ryan AbilityLab, Chicago, IL, USA), who previously isolated them from surgical biopsy specimens under approval from Lurie Children's Hospital's Institutional Review Board (IRB 2016-186). 23 Cells were maintained at 37 °C, 5% CO2, and passaged at 75% confluency.
Quantification of myotube formation
C2C12 and human myoblasts were seeded onto scaffolds (0.5 mm × 0.5 mm) at a density of 5.0 × 103 cells/cm2 and cultured until approximately 80% confluency was reached. Myoblast differentiation, fusion and myotube formation were promoted by switching from the high-serum growth medium to a low-serum DMEM medium (Gibco) containing 2% Horse Serum (HS, Gibco), 4.5 g/L d-Glucose, L-Glutamine, 10 mg/ml streptomycin, 100 units/ml penicillin (Gibco), and Insulin (50 nM) (Sigma-Aldrich). Differentiation was promoted for a duration of 10 days for human myoblasts and 15 days for C2C12 cells. Total cell and myotube counts were determined from immunofluorescent-labeled cells using Nikon software (NIS Elements; Nikon; Melville, NY, USA). Analysis was performed on three separate images (10x, Nikon Eclipse Ti2 Series, Nikon, Melville, NY, USA) from three biological replicates corresponding to each scaffold variant. The total cell numbered was calculated as the sum of multi- and mononucleated cells distinguished using Hoechst 33258 (1:300; 94403; Sigma-Aldrich, St Louis, MO, USA). To assess the differentiation of C2C12 and human myoblast cells into myotubes, we counted the number of desmin and α-actinin positive myotubes containing more than four nuclei.
In-vitro biocompatibility test
A live/dead assay was conducted using confocal microscopy. C2C12 mouse myoblasts were seeded onto a 12-well plate containing both graphene scaffold variants at a density of 5.0 × 103 cells/cm2. Following 3 days of culture, cells were incubated with green-fluorescent Calcein, AM (1:2000; C3099; Thermo Fisher Scientific, Waltham, MA, USA) and red fluorescent propidium iodide (PI; 1:20; 75002; Stemcell Technologies, Vancouver, Canada) for 30 min prior to imaging. Cell nuclei were stained using Hoechst 33258 (1:300; 94403; Sigma-Aldrich, St Louis, MO, USA). Images were captured using a Nikon Eclipse Ti2 Series confocal microscope (Nikon, Melville, NY, USA) and quantified using Nikon software (NIS Elements; Nikon; Melville, NY, USA). Binary masks were created for each stain and overlapped to distinguish live cells (Hoechst + Calcein AM) from dead cells (Hoechst + PI).
Culturing of human stem cell-derived motor neurons
MNs were derived from the commercially available male control iPSC line BJFF.6 obtained from the Washington University Genome Engineering and iPSC Core (GEiC) facility. MN differentiation was conducted following the previously reported method by Ziller et al. with minor modifications.24,25 iPSC colonies were dissociated using ReLeSR (5872; Stemcell Technologies, Vancouver, Canada) and plated at a density of 74,000 cells/cm2 in mTeSR Plus (5825; Stemcell Technologies, Vancouver, Canada) containing 10μM ROCK inhibitor (129830-38-2, DNSK International, Hamden, CT, USA) for 24 h. Differentiation was initiated once cells reached 80–90% confluency by changing the base medium (N2B27 medium) to one containing 47.5 vol % DMEM:F12, 47.5 vol % Neurobasal, 1 vol % NEAA, 1 vol % Glutamax, 1 vol % N2 and 2 vol % B27. From day 0 to day 5, cells were fed daily using the N2B27 medium supplemented with 10μM SB431542 (DNSK International), 100 nM LDN193189 (DNSK International), 1 μM Retinoic Acid (RA, Sigma-Aldrich) and 1μM of Smoothened-Agonist (SAG, DNSK International). Starting on day 6, the N2B27 culture medium supplements were changed to 1 μM RA, 1 μM SAG, 5 μM DAPT (DNSK International) and 4 μM SU5402 (DNSK International). Cells were fed daily until day 14 of differentiation. The resulting differentiated MNs were then dissociated using TrypLE Express (GIBCO, Life Technologies) supplemented with DNase I (Worthington) and further dissociated to single cells with Accutase.
Glia and motor neuron seeding and culture on graphene scaffolds
Graphene scaffolds with and without 2.5% dECM were cut into square pieces 1 mm × 1 mm in dimension. Prior to cell seeding, scaffolds were incubated in 0.1 μg/ml of poly-D-lysine (PDL) overnight at 4 °C. Glia isolated from mouse pup cortices (P1) was then seeded at a density of 1.0 × 104 cells/well onto each scaffold type. The glial cells were cultured in MEM medium containing 10% horse serum, 1.3% D-glucose, and 1% P/S for 2 days before MNs were added. On day 3 following the addition of glia, MNs were then seeded onto each scaffold type at a density of 2.5 × 104 cells/well in neurobasal medium containing 1% non-essential amino acids, 1% Glutamax, 1% N2, 2% B27, and 1% P/S supplemented with ascorbic acid (1:1000),26,27 BDNF (1:10,000), CNTF (1:10,000), GDNF (1:10,000), and rock inhibitor (1:1000).
Motor neuron immunofluorescence staining and axon length quantification
Four days after initial seeding, cells were then fixed in 3–4% paraformaldehyde in PBS (pH 7.4) for 15 min at room temperature, followed by two rinses with ice cold PBS. Samples were incubated for 10 min in PBS + 0.25% Triton X-100 to improve antibody penetration, followed by three rinses with PBS (5 min per rinse). This was followed by incubation with 1% BSA in PBST for 30 min, to block unspecific binding of the antibodies. Samples were then stained with chicken anti-Nf-H (1:1000; ab72996, Abcam, Cambridge, UK), mouse anti-islet-1 (1:100; 15661-1-AP, Proteintech Group, Rosemont, IL, USA), and mouse anti-GFAP (1:20; DSHB, 8-1E7) and incubated for 1 h at room temperature. This solution was then removed, and the cells were washed three times with PBS. Samples were then stained with secondary antibodies donkey anti-chicken-Alexa488 (1:500; 703-545-155; Jackson ImmunoResearch Laboratories, Inc., West Grove, PA, USA), donkey anti-rabbit-Alexa555 (1:500; A-31572; Thermo Fisher Scientific, Waltham, MA, USA), and goat anti-mouse-Alexa647 (1:500; A-21235; Thermo Fisher Scientific, Waltham, MA, USA), then incubated in 1% BSA for 1 h at room temperature in the dark. The secondary antibody solution was removed, and the cells were rinsed three times in PBS (5 min per rinse) in the dark. To counter stain, cells were incubated with 0.1–1 μg/mL Hoechst (Sigma-Aldrich, 94403) for 1 min, then rinsed with PBS. Following fixation and immunostaining, tile scan images of the surfaces of each scaffold were acquired using a Leica DMi8 inverted microscope. Isl1 + MN nuclei were quantified using the Multi-point Tool in ImageJ. The Simple Neurite Tracer plugin in ImageJ was used to record paths of Nf-H expressing neurites extending from MN cell bodies and to quantify neurite lengths. The sum of neurite path lengths per number of nuclei was calculated for each image.
Formation of hiPSC-derived 3D motor neuron spheroids
Differentiation of human-induced pluripotent stem cells (hiPSCs) into hMNS was carried out as previously described27–29 with modifications applied to the iPSC line BJFF, which were obtained from a healthy control subject from the Washington University GEiC. Briefly, pluripotent stem cell colonies were dissociated with Accutase (AT 104-500, Innovative Cell Technologies, San Diego, CA, USA) and seeded into a 96-well low adherence plate (day 0) at a 400,000/mL density in N2B27 medium containing 0.1 vol % ß-mercaptoethanol (55 mM; 21985023; Thermo Fisher Scientific, Waltham, MA, USA) supplemented with ascorbic acid26,27 (AA, 0.2 μg/mL; 1:2000; A4403-100MG, Sigma Aldrich, St Louis, MO, USA), ROCK inhibitor (10 mM; 1:2000), laminin (1.6 μg/mL; 1:3200; 23017-015; Thermo Fisher Scientific, Waltham, MA, USA), and 3 µM Chir99021 (4423, Tocris Bioscience, Bristol, UK), 0.2 µM LDN193189 (1062368-24-4, DNSK International, Hamden, CT, USA), and 40 µM SB431542 (301836-41-9, DNSK International, Hamden, CT, USA) for 24 h. On day 2, embryoid bodies (EBs) were formed and the culture medium was further supplemented with all-trans retinoic acid (RA; 10 mM; 1:100,000; R2625, Sigma Aldrich, St Louis, MO, USA) and smoothened agonist (SAG; 10 mM; 1:20,000; 364590-63-6, DNSK International, Hamden, CT, USA). On day 4, the differentiation medium was changed to include only RA and SAG supplementation. DAPT (10 mM; 1:1000; 208255-80-5, Fisher Scientific, Hampton, NH, USA) was then added as an additional supplement to the medium starting on day 9. On day 11, BDNF (100 μg/ml; 1:5000; 248-BDB; R&D Systems, Minneapolis, MN, USA) and GDNF (100 μg/ml; 1:10,000; 212-GD; R&D Systems, Minneapolis, MN, USA) were added to form the final maturation medium which was used throughout all experiments involving hMNSs. All resulting hMNSs were used during their period of highest functionality between days 30–50 of formation. Cultures were maintained at 37 °C in 5% CO2 and maturation medium was changed every 3 days.
Human motor neuron spheroid co-culture with human myoblasts
Scaffolds were cut into 15 mm × 5 mm segments and prepared for cell culture in the manner described above. Human myoblasts were distributed onto either side of each scaffold type equally at a density of 5.0 × 103 cells/cm2 and were cultured in growth medium for 3 days to allow increase in confluency. The human myoblasts were then cultured for 5 days in low-serum differentiation medium to allow for myotube formation. On day 6 of the myoblast differentiation stage (day 9 from seeding), 32-day-old hMNSs prepared as described previously were seeded onto the center of each scaffold 1:1. The hMNSs and newly formed myotubes were co-cultured in final maturation medium for a duration of 14 days.
Immunofluorescence staining of hMNS-myoblast co-culture
Following day 14 the co-cultures were fixed and prepared for immunocytochemistry as previously described. The co-cultures were then incubated with chicken anti-Nf-H (1:1000; ab72996, Abcam, Cambridge, UK), mouse anti-islet-1 (1:100; 15661-1-AP, Proteintech Group, Rosemont, IL, USA), and anti-α-actinin (1:500; A7811; Sigma-Aldrich, St Louis, MO, USA) for 1 h. Samples were incubated with secondary antibodies donkey anti-chicken-Alexa488 (1:500; 703-545-155; Jackson ImmunoResearch Laboratories, Inc., West Grove, PA, USA), donkey anti-rabbit-Alexa647 (1:500; ab150075; Abcam, Cambridge, UK), goat anti-mouse-Alexa647 (1:500; A-21235; Thermo Fisher Scientific, Waltham, MA, USA), and α-Bungarotoxin-Alexa555 conjugate (α-Btx) (1:100; B35451; Thermo Fisher Scientific, Waltham, MA, USA) for 1 h at room temperature in the dark. For nuclei staining, cells were incubated with 0.1–1 μg/mL Hoechst (Sigma-Aldrich, 94403) for 1 min, then rinsed with PBS.
Sholl analysis
The fluorescent threshold of z-stack co-culture images was adjusted with ImageJ to identify all axons around the periphery of the hMNS. The neuroanatomy plugin for ImageJ was used to perform a sholl analysis. The radius of the hMNSs was assumed to be 500 µm and the step size was set as 10 µm.
Statistical analyses
All data sets are presented as mean ± SE (standard error of the mean). Statistical significance was determined by a one-tailed, unpaired t-test as appropriate. P < 0.05 was considered to be statistically significant.
Results
Graphene and dECM-graphene scaffolds are structurally comparable but electronically distinct
Using the method previously described by Jakus et al. 30 (Figure 1(a)), graphene scaffolds were extrusion printed with and without dECM. Brightfield images were captured to determine whether the incorporation of dECM led to gross macrostructure changes (Figure 1(b) and (d)). SEM imaging further corroborated the structural similarities (Figure 1(c) and (e)). In both cases, flakes could be identified throughout the material, revealing similar topographical profiles at the microscale. We then evaluated the Young's modulus of graphene and dECM-graphene scaffolds to determine if dECM inclusion caused bulk scaffold mechanical properties (Figure 1(f) and (g)). The nominal tensile modulus for pristine graphene scaffolds was 23.57 ± 6.72 MPa and 28.78 ± 13.45 MPa for dECM-graphene scaffolds (Figure 1 (g)). No statistical difference was detected between these data sets, demonstrating that the mechanical properties of dECM-graphene scaffolds were comparable to the scaffolds comprised of graphene alone.

Characterization of graphene and dECM-graphene scaffolds. (a) Schematic of scaffold printing concept. dECM-graphene scaffolds are structurally feasible and comparable to pristine graphene. Brightfield images of (b) graphene and (d) dECM-graphene show grid scaffold morphology, and (c,e) SEM images show that the microstructure is not significantly affected by the addition of dECM. (f,g) Tensile testing and translation into Young's Modulus. Data are presented as a mean ± SE for n = 3 scaffolds per group. (h) Zetapotential analyses were also performed to evaluate differences in material parameters following the addition of dECM. Data are presented as a mean ± SE for n = 6 technical replicates. Statistical significance was determined using a one-tailed unpaired t-test, (****) P < 0.0001. (i) Conductivity measurements for each scaffold type. Data are presented as a mean ± SE for n = 3 scaffolds per group. Statistical significance was determined using a one-tailed unpaired t-test, (*) P < 0.05.
After evaluating the mechanical properties of the scaffolds, electrochemical changes were analyzed by zetapotential and conductivity measurements (Figure 1(h) and (i)). Zetapotential was performed to evaluate differences in surface charge between the graphene and graphene-dECM scaffolds and a statistical difference (****P < 0.0001) was found (Figure 1(h)). Zetapotential values were measured to be −18.3 ± 0.4 mV in graphene scaffolds and −22.2 ± 0.6 mV in dECM scaffolds (Figure 1(h)). Further examination of the scaffold electronic properties revealed dECM-graphene materials demonstrated a two-fold increase in electrical conductivity from 68.97 ± 7.92 S/m in pristine graphene to 140.66 ± 6.75 S/m in dECM (*P < 0.05, Figure 1(i)).
Graphene and dECM-graphene scaffolds are equivalent in driving human and mouse myoblast viability and differentiation
A live/dead assay performed using mouse myoblasts on graphene and dECM-graphene scaffolds demonstrated favorable viability on both scaffolds (Supplemental Figure 1). We then screened graphene and dECM-graphene scaffolds for myogenic properties with both human and mouse myoblasts to determine whether there would be species-dependent differences in cellular outcomes, considering the incorporated dECM was mouse muscle-derived. To observe cell morphology on graphene and dECM-graphene scaffolds, histology staining for α-actinin and desmin was performed. 31 On both scaffolds, myotubes were highly elongated and multinucleated, which is indicative of cell maturation and demonstrates that the scaffolds were effective in driving cell differentiation. Between scaffold variants, however, we were unable to discern any qualitative differences in myoblast cell phenotype.
In addition to histological analysis, immunofluorescent images were analyzed to determine the quantity of myotubes, which were classified as cells containing more than four nuclei, within pristine graphene and dECM-graphene scaffolds. No statistical significance was found in comparing myotube formation derived from human and mouse myoblasts between the two scaffold types. 72.9 ± 2.4% of mouse myoblasts contributed to myotube formation in pristine graphene scaffolds; in graphene scaffolds containing dECM, 79.8 ± 4.7% of all cells contributed towards the formation of myotubes (Supplemental Figure 2). With respect to the total amount of human primary cells observed in each scaffold, 22.8 ± 5.3% contributed to myotube formation on graphene scaffolds; in comparison, 24.7 ± 8.6% of all cells on graphene scaffolds containing dECM were found within the myotubes present (Figure 2(a) to (d)).
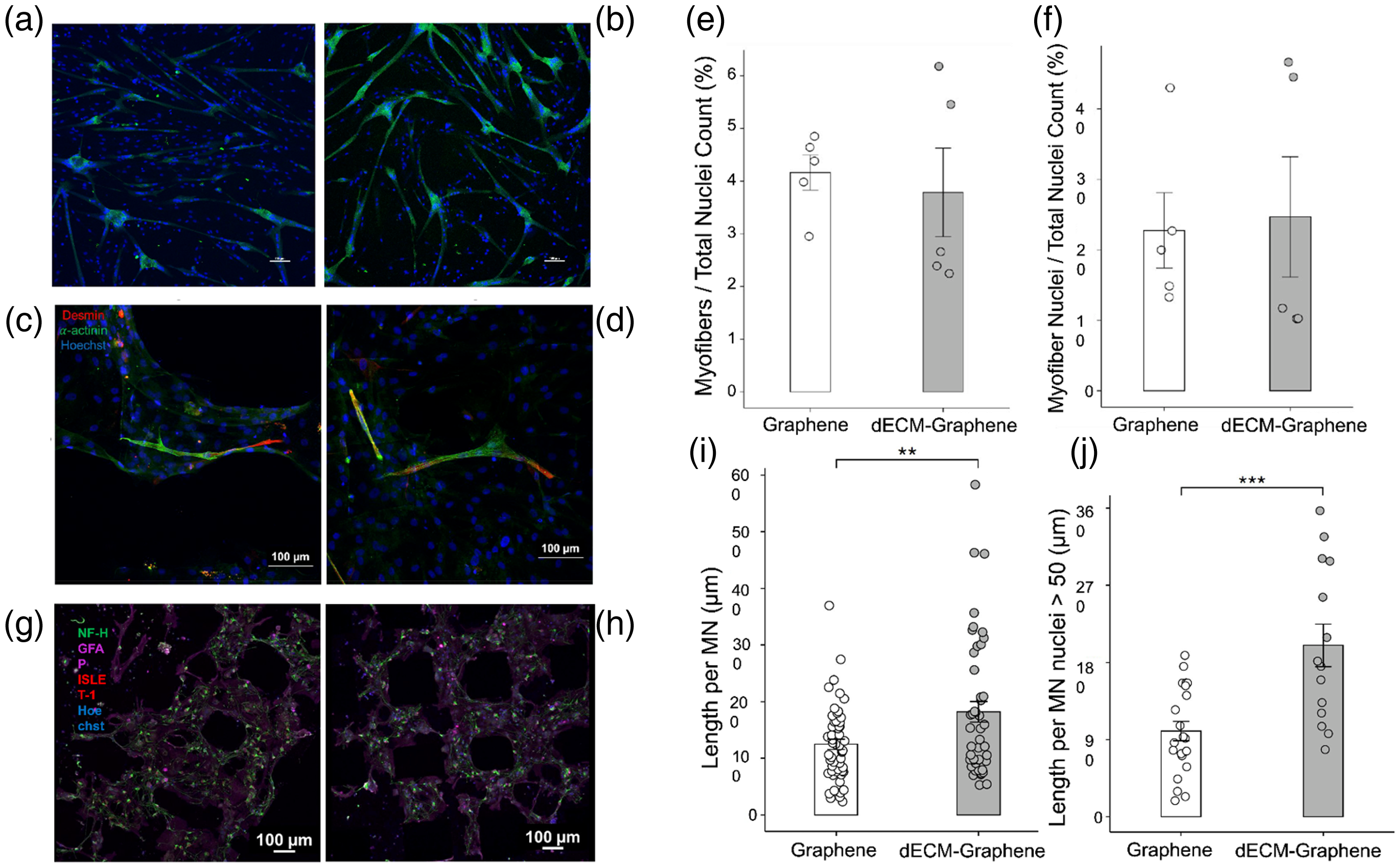
2D in vitro models of human derived muscle and neural cells on graphene and dECM-graphene scaffolds. Representative images of human myoblasts seeded on (a) graphene and (b) dECM-graphene scaffolds, highlighting presence of myofibers at lower magnification (scale bars 100 μm). Representative images of human myoblasts seeded on (c) graphene and (d) dECM-graphene scaffolds at higher magnification and stained to highlight myofiber maturity (red = desmin, green = ɑ-actinin, blue = hoechst), quantified with respect to (e) the amount of myofibers formed from the total population present and (f) the number of participating cells involved in myofiber formation. Data are presented as a mean ± SE for n = 5 scaffolds per group. dECM-graphene scaffolds significantly improve axonal extension. Representative confocal microscopy images of MNs seeded with supporting glial cells on (g) graphene and (h) dECM-graphene scaffolds were captured and used to perform analysis (green = NF-H, pink = GFAP, red = ISLET-1, blue = hoechst). (i) Axonal outgrowth with respect to the total nuclei count. Data are presented as a mean ± SE for n = 40–50 quantified images from 3 scaffolds per group. Statistical significance was determined using a one-tailed unpaired t-test, (*) P < 0.05. (j) Axonal length per nuclei considering only tile scan sets containing more than 50 nuclei. Data are presented as a mean ± SE for n = 13–15 quantified images from 3 scaffolds per group. Statistical significance was determined using a one-tailed unpaired t-test, (**) P < 0.01.
dECM-graphene scaffolds significantly improve axonal outgrowth
Considering the physiologically synergistic nature of muscle and nervous tissue, we next evaluated the response of neural-derived cells. MNs derived from human iPSCs, along with glial supportive cells, were seeded onto each construct. Neurons were distinguished from glial cells using nucleic marker Islet-1 A, and the total axon lengths per nuclei were measured. MNs seeded on dECM-graphene constructs had average axon/nuclei lengths of 189.3 ± 21.2 µm, while axon growth on graphene scaffolds averaged 124.8 ± 8.5 µm in length (Figure 2(e) to (g)).
Incorporation of dECM improves systematic outcomes in neuronal-muscular co-culture
To explore whether the incorporation of dECM in the graphene scaffolds could lead to systematic improvements, we applied an hMNS-myoblast coculture system (Figure 3). MN formation from hiPSCs and formation into hMNSs was carried out following previously described methods with minor modifications (Figure 4).27,29 Myoblasts were first seeded onto each scaffold type and were cultured in isolation until myotube fusion was initialized (Figure 3(a), left). hMNSs were seeded on the scaffolds after the myoblasts fused into myotubes, at which point the cells reach a state of lower metabolic activity (Figure 3(a), middle). After hMNS seeding, the cell medium was changed to accommodate the organoids (Figure 3(a), right).
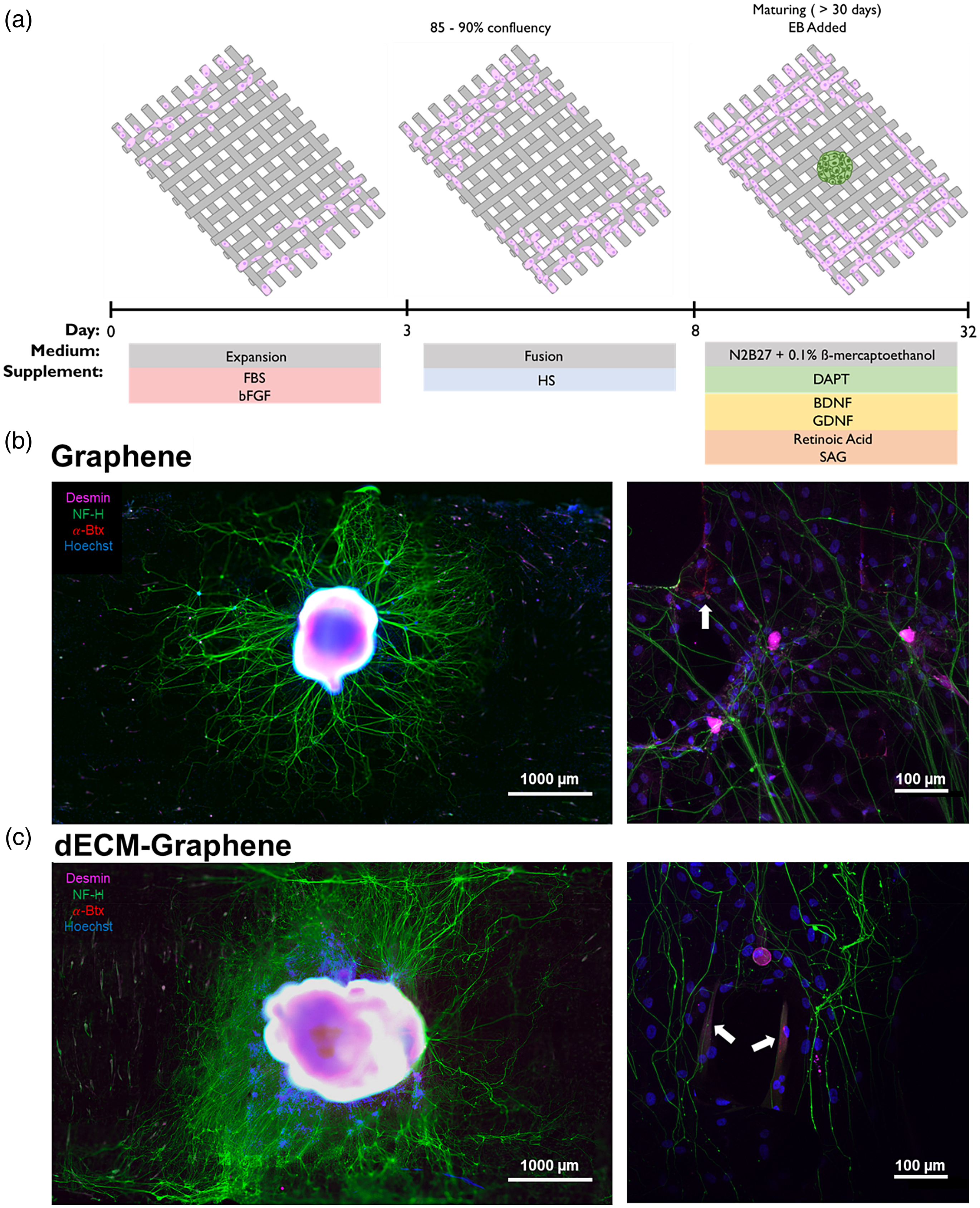
Advanced co-culture with both human derived myoblasts and 3D motor neuron spheroids for evaluation of systematic regeneration. (a) The neuro-muscular co-culture timeline is depicted. To develop co-cultures, myoblasts were first seeded and allowed to fuse on the scaffolds. After allowing eight days of myoblasts fusing into myofibers, the hMNSs made from EBs (32 day old) were also placed on the scaffolds with the pre-seeded myotubes. Neuro-muscular co-cultures on both extruded graphene scaffolds demonstrate complex cell-cell interactions. Axons of hMNS seeded on (b, left) graphene and (c, left) dECM-graphene scaffolds can be seen terminating at the interface of mature myotubes. (b,c, right) For both scaffolds, acetylcholine receptor clustering can be seen at various sites along regions of myotubes (white arrows).

Differentiation and culture of motor neurons in 2D and 3D formats. Panel A illustrates the 2D culture of motor neurons, which is ideal for quantification of live/dead assays and initial assessments of cell viability on the scaffolds. Panel B demonstrates the 3D human motor neuron spheroids (hMNS) made from embryoid body (EBs) co-cultured with myoblasts, a model that mimics peripheral nerve growth by allowing axons to grow together in bundles, as compared to mostly the single neurites observed in 2D cultures. This approach provides a more physiologically relevant environment to evaluate the scaffold's potential for supporting neuromuscular growth. Summary of small molecules used in differentiation: Ascorbic Acid (AA), BDNF (Brain-Derived Neurotrophic Factor), CHIR99021 (GSK-3 inhibitor), DAPT (γ-Secretase inhibitor), GDNF (Glial cell line-Derived Neurotrophic Factor), Laminin (Extracellular matrix protein), LDN193189 (BMP inhibitor), N2B27 (Media Supplement), Retinoic Acid (RA), ROCK inhibitor, SB431542 (SMAD inhibitor), Smoothened-Agonist (SAG), SU5402 (FGF receptor inhibitor).
After hMNS seeding, outwards axonal branching towards myotubes was observed on both graphene and dECM-graphene scaffolds (Figure 3(b) and (c), left). We stained the co-cultures for alpha-bungarotoxin (α-Btx) to screen for acetylcholine receptors, which are the first indication of neuromuscular junction (NMJ) formation.32,33 We observed acetylcholine receptor clustering at the neuromuscular interfaces in both graphene and dECM-graphene scaffolds (Figure 3(b) and (c), right).
A Sholl analysis was performed to quantitatively evaluate the degree of hMNS branching between scaffold types (Figure 5). From the Sholl analysis, we see a higher number of axon intersections stemming from hMNSs seeded on dECM-graphene scaffolds (Figure 5(c)). We also analyzed differences in hMNS radius, and despite being comparable in size at initial seeding (Supplemental Figure 3), hMNSs seeded on dECM-graphene scaffolds presented significant size increases compared to their counterparts seeded on pristine scaffolds (Figure 5(d)).

dECM-graphene enhances hMNS radial growth and axon branching compared to pristine graphene. (a,b) Co-cultures stained with anti-Nf-H (green), anti-islet-1 (magenta), anti-α-actinin (magenta), α-Btx conjugate (red), and counterstained with Hoechst (blue). Isolation of Nf-H channel and axonal morphology of hMNS on graphene and dECM-graphene scaffolds is shown. Threshold levels were adjusted to highlight proximal and distal axonal projections. The Sholl plot is also shown with 10 μm intervals starting at radial distance 500 μm from center. (c) Sholl analysis was performed between scaffolds to compare the degree of axonal branching and networking on the different types. Data are presented as a mean ± SE from n = 3 experiments. (d) The average co-culture hMNS radii at the conclusion of the experiment were quantified and compared. Data are presented as a mean ± SE from n = 3 co-culture hMNS. Statistical significance was determined using a one-tailed unpaired t-test, (**) P < 0.01.
Discussion
The mechanical similarities between these materials indicate that any observed functional improvements in cellular processes are not owed to altered scaffold mechanics, rather they can be attributed to biochemical changes in the scaffolds from the dECM inclusion. dECM-based materials are typically associated with weak structural integrity, yet in this case, the dECM-graphene scaffolds had a tensile modulus showing no significant difference from the tensile strength of pristine graphene. 15 This maintenance of favorable scaffold mechanics also demonstrates that in the future, dECM may be incorporated into other graphene-based composites with more complex architectures without detrimentally affecting structural integrity. Overall, it is important that dECM preserves the mechanics of the pristine graphene, considering that graphene's structural characteristics are an especially attractive motivator for its use in tissue engineering. 5
Scaffolds containing dECM molecules overall exhibited a more negative surface charge than their graphene-only counterparts. The decreased zeta potential of the scaffold particles is in line with other studies, which have previously shown a direct relationship between the quantity of dECM incorporated into the composite and decreasing zeta potential. 34 In the previous study, they attributed the decrease in zeta potential to the increased amount of “electrostatically neutral” molecules that results from increasing levels of dECM. The combined zetapotential and conductivity analysis demonstrate that there are distinct changes in the electroactive properties of the scaffolds following the addition of dECM. dECM incorporation elevates scaffold conductivity, which may have critical downstream influence on cellular processes.
To overcome the electronegative nature and resulting hydrophobicity of the two scaffold types, cationic PDL was first applied prior to any cell seeding. Initial attempts in culturing cells onto the graphene and dECM-graphene scaffolds without PDL resulted in position instabilities from the constructs themselves as well as low cellular adherence in all cell types tested. When assessing for myotube formation, there was no significant advantage gained by incorporating dECM into the scaffold supports in either mouse or human-derived muscle cell lines. Although the dECM was specifically muscle-derived, the cells seeded on these substrates did not demonstrate measurably higher degrees of myoblast maturation. For future studies, we may explore the use of dECM derived from a variety of tissues to determine whether dECM from different sources influences differentiation. Interestingly, the species of dECM may not be significant in bolstering scaffold bioactivity, since we observed no differential response for either human or mouse myoblasts to dECM-graphene.
Although we did not see significant differences between cell response of muscle-derived cells seeded on pristine graphene scaffolds compared to dECM-graphene scaffolds, the dECM-graphene scaffolds significantly bolstered neuronal cell axon outgrowth compared to the graphene scaffold. Previous studies have demonstrated that cell response to ECM composition is highly specific, 35 and this result not only corroborates those findings, but it also points to the importance of examining novel biomaterials holistically. Diverse incorporations of dECM components from various lineages may elucidate previously unrealized potential for improving regenerative results in complex systems. Future investigations may further explore the mechanisms by which different cell types respond to dECM compositions made from a variety of tissue types with differing age origins.
Apart from augmenting the physiochemical bioactivity with dECM, we also note composite material parameters are affected. As previously mentioned, dECM incorporation led to reduced scaffold surface charge and increased conductivity. One possible explanation for the differential outcomes observed in neuronal and muscular cell types may be that changes in electronic properties are more potent in neuronal lineages compared to muscular lineages. Delineating the biochemical influences of dECM incorporation from the electronic changes of the material may be subject of future investigation.
In our combined 3D and 2D culture model we see neural hMNS branching towards peripheral muscle cells in a physiologic manner; furthermore, we are providing a more accurate reflection of the compatibility of dECM-graphene and graphene scaffolds with functional skeletal muscle repair. This culture system allows us to observe systematic interactions, similar to those which occur during innervation, which is critical for skeletal muscle development and repair.36,37 For future experiments, we may perform additional long-term cultures to further analyze if there are significant differences in the mode in which the hMNSs influence NMJ formation.
Previous studies have corroborated the findings here, where initial acetylcholine receptor clustering can be observed with an hMNS-myoblast system. 38 In the case where hMNS size may be correlated to neuronal number, we expect dECM-graphene scaffolds to demonstrate more axonal outgrowth and intersections. Interestingly, we found that with both scaffolds, the maximum number of axon intersections occurred at the same radial distance, regardless of the scaffold type. One explanation could be that branch formation is reliant on the stability provided by the main organoid body. Therefore, if smaller hMNSs were used then we would expect neuronal branching to maximize at smaller radial intervals. We also observed that the number of intersections decreased at the same rate, with respect to increasing radial distance. Although there could be a spatial element which limits neuronal outgrowth due to the boundaries of the scaffolds, peripheral muscle cells may also occupy space that could serve to further extend axonal reach and interconnections.
With this co-culture system, we were able to observe initial signs of NMJ formation in the form of acetylcholine receptor clustering in both scaffold variants, which are highly critical in skeletal muscle development and repair. Although Sholl analyses had been previously demonstrated for individual cell systems, this is the first demonstration of such an approach to quantify differences among complex hMNS architectures.39,40 Using this application, neuronal networking was tracked, and the degree of pathway formation was quantified. With this analysis, we found that dECM-graphene scaffolds significantly promoted axonal outgrowth and branching. We did not, however, observe any qualitative differences in myotube morphology or quantity. In future studies, we may extend co-culture duration or utilize in vivo models to assess functional differences for myotubes seeded on varying scaffold types.
Limitations
Several limitations were encountered during this investigation that warrant acknowledgment. First, the extrusion printing method can lead to variability in the strength of each strut of the scaffolds, resulting in larger variance in Young's modulus than what would typically be expected from more conventional geometric forms. Additionally, in the maintenance of the co-culture models, the supplements added to the medium and the frequency of their replacement were not adjusted in response to the evolving cellular population and environment. As cells mature and begin to emit their own signaling molecules, there is a potential for supplemental factors to unfavorably influence cellular interactions. Moreover, frequent medium changes could disrupt molecular signaling environments that are crucial during specific time periods. Another limitation involves the use of mixed human and animal cell types in our study, particularly in the context of hiPSCs. While this approach allowed us to assess the scaffold's performance across a variety of cellular environments, it also introduces complexities and potential variability due to species differences. Generating all relevant cell types solely from hiPSCs is technically challenging and resource-intensive, but this should be a priority for future studies to create more homogeneous and controlled cellular models. Despite these limitations, the insights gained from this study contribute valuable information regarding the scaffold's versatility and potential applications in neuromuscular regeneration.
Conclusion
This study determined that PLGA graphene scaffolds treated with PDL were capable of sustaining neuronal and muscular cell cultures. Incorporation of dECM into these scaffold constructs resulted in improved neuronal network formation in MNs co-cultured with glial cells as a monolayer and in those cultured as a 3D hMNS in the presence of human myofibers. The presence of acetylcholine clustering observed in both scaffold variants present them as promising candidates as substrates for establishing NMJs. The successful support of multiple neuromuscular cell types from different sources, including MNs, C2C12 myoblasts, and hMNS, highlights the scaffolds’ robustness as versatile substrates for the growth and development of neuromuscular tissues, further enhancing its potential for clinical applications.
Supplemental Material
sj-docx-1-sci-10.1177_00368504241281469 - Supplemental material for Incorporation of decellularized-ECM in graphene-based scaffolds enhances axonal outgrowth and branching in neuro-muscular co-cultures
Supplemental material, sj-docx-1-sci-10.1177_00368504241281469 for Incorporation of decellularized-ECM in graphene-based scaffolds enhances axonal outgrowth and branching in neuro-muscular co-cultures by Carlos Serna, Kirtana Sandepudi, Rebecca L Keate, Sophia L Zhang, Kristen Y Cotton, Alberto De La Isla, Matias Murillo, Yasmine Bouricha, Andrea A Domenighetti, Colin K Franz and Sumanas W Jordan in Science Progress
Supplemental Material
sj-docx-2-sci-10.1177_00368504241281469 - Supplemental material for Incorporation of decellularized-ECM in graphene-based scaffolds enhances axonal outgrowth and branching in neuro-muscular co-cultures
Supplemental material, sj-docx-2-sci-10.1177_00368504241281469 for Incorporation of decellularized-ECM in graphene-based scaffolds enhances axonal outgrowth and branching in neuro-muscular co-cultures by Carlos Serna, Kirtana Sandepudi, Rebecca L Keate, Sophia L Zhang, Kristen Y Cotton, Alberto De La Isla, Matias Murillo, Yasmine Bouricha, Andrea A Domenighetti, Colin K Franz and Sumanas W Jordan in Science Progress
Footnotes
Acknowledgements
R.K. was supported in part by the National Institutes of Health Training Grant (Grant No. T32GM008449) through Northwestern University's Biotechnology Training Program. We thank the Alvin J. Siteman Cancer Center at Washington University School of Medicine and Barnes-Jewish Hospital in St Louis, MO., for the use of the Genome Engineering & Stem Cell Center, which provided the reprogrammed human iPSCs used in our experiments. The Siteman Cancer Center is supported in part by an NCI Cancer Center Support Grant #P30 CA091842. This work made use of the EPIC facility of Northwestern University's NUANCE Center, which has received support from the SHyNE Resource (Grant No. NSF ECCS2025633), the IIN, and Northwestern's MRSEC program (Grant No. NSF DMR-1720139). This work made use of the DMA at the MatCI Facility supported by the MRSEC program of the National Science Foundation (Grant No. DMR-1720139) at the Materials Research Center of Northwestern University. Zetapotential measurements were performed in the Analytical bioNanoTechnology Core (ANTEC) Facility of the Simpson Querrey Institute at Northwestern University. ANTEC is currently supported by the Soft and Hybrid Nanotechnology Ex-perimental (SHyNE) Resource (Grant No. NSF ECCS-2025633). C.K.F. acknowledges funding from the Belle Carnell Regenerative Neurorehabilitation.
Author contributions
CSIII: contributed to project conception, conducted most experiments, prepared all the materials/culture systems, and main contributor to data collection and analysis. RLK: contributed significantly to figure generation and manuscript writing, conducted some experiments (mechanical/conductivity testing), and assisted with data collection and analysis. KS: assisted with cell culture, data/image collection, manuscript editing, and final submission. SLZ: assisted with cell culture, fabrication of bioink, and with data collection/processing. Also contributed to manuscript editing. KYC: assisted with some data collection and cell culture, as well as manuscript edits. ADLI: maintained many of the cell cultures. Also contributed to image/data collection and manuscript editing. MM: assisted with maintenance of cell culture and manuscript editing. YB: assisted with cell cultures and manuscript editing. AAD: assisted with manuscript review and provided consultation in matters pertaining to the culturing and maintenance of muscle cells. CF: advised on all co-culture systems and assisted with image/data collection and processing. Reviewed and approved final manuscript. SWJ: project conception, overall management of experimental design, data collection and analysis, and manuscript writing. Reviewed and approved final manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Science Foundation, National Cancer Institute, National Institutes of Health (grant numbers DMR-1720139, ECCS2025633, P30 CA091842, T32GM008449).
Ethical considerations
ECM was acquired from mouse pups under approval from Northwestern's Institutional Animal Care and Use Committee (Study #IS00010080) and human myoblasts were isolated from surgical biopsy specimens under approval from Lurie Children's Hospital's Institutional Review Board (2016-186).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
