Abstract
The brain regulates every physiological process in the body, including metabolism. Studies investigating brain metabolism have shown that stress can alter major metabolic processes, and that these processes can vary between regions. However, no study has investigated how metabolic pathways may be altered by stressor perception, or whether stress-responsive brain regions can also regulate metabolism. The basolateral amygdala (BLA), a region important for stress and fear, has reciprocal connections to regions responsible for metabolic regulation. In this study, we investigated how BLA influences regional metabolic profiles within the hippocampus (HPC) and medial prefrontal cortex (mPFC), regions involved in regulating the stress response and stress perception, using optogenetics in male C57BL/6 mice during footshock presentation in a yoked shuttlebox paradigm based on controllable (ES) and uncontrollable (IS) stress. RNA extracted from HPC and mPFC were loaded into NanoString® Mouse Neuroinflammation Panels, which also provides a broad view of metabolic processes, for compilation of gene expression profiles. Results showed differential regulation of carbohydrate and lipid metabolism, and insulin signaling gene expression pathways in HPC and mPFC following ES and IS, and that these differences were altered in response to optogenetic excitation or inhibition of the BLA. These findings demonstrate for the first time that individual brain regions can utilize metabolites in a way that are unique to their needs and function in response to a stressor, and that vary based on stressor controllability and influence by BLA.
Introduction
The brain, while only representing around 2% of the total body weight in an adult human, is the most energy-demanding organ, accounting for about 20% of total energy consumption in a resting state. The mammalian brain relies heavily on glucose as its main energy source. In humans, the brain consumes nearly 20% of all glucose in the body, 1 with most being devoted to neurons. The brain can also utilize ketone bodies as a secondary energy source, but these are not relied on significantly, except during times of depleted glucose levels. 2
Stress can disrupt energy homeostasis by altering major metabolic processes such as hormone signaling and glucose uptake, 3 as well as alter gene expression related to glucose metabolism and reduce glucogenic and neuroactive amino acids. 4 Other evidence has shown that stress 5 increases consumption of fat-derived ketone bodies in the brain that varies across regions. 5 However, these studies focused on stressed-induced alterations of brain metabolism in the context of the general effects of stress on metabolic processes, and the subjects in these studies were passive recipients of stressful manipulations. It is well established that the medial prefrontal cortex (mPFC), hippocampus (HPC), and basolateral amygdala (BLA) form a neurocircuit that regulates many components of the stress response. 6 Our work, 6 and that of others, 7 have demonstrated that giving animals the ability to engage in behaviors that can modify stressor perception and/or stress outcomes can also alter stress-induced inflammatory responses in a region dependent manner, 6 and that these are modulated by the BLA. Interestingly, the immune system and metabolism are intimately related, as immune responses rely heavily on energy 8 and metabolism can influence immune cell function. 9 The BLA likely also has connections to both the hypothalamic and brainstem regions responsible for metabolic regulation via influences on the central nucleus of the amygdala, the principal output region of the amygdala. 10 Therefore, it is reasonable to postulate that individual brain regions, such as HPC and mPFC, can also have unique metabolic profiles that may also be modulated by BLA during the stress response. Currently, we are unaware of any previous evidence that suggests BLA has the capacity to influence metabolic signaling and pathway regulation.
In this study, we investigated the effects of controllable and uncontrollable stress (modeled by escapable [ES] and inescapable [IS] footshock, 6 respectively) on metabolic pathway regulation within the HPC and mPFC. We also assessed whether these pathways were influenced by BLA using optogenetics to selectively activate or inhibit glutamatergic neurons in BLA during stress.
Materials and methods
Subjects
Male C57BL/6 mice, 8–9 weeks of age and 20–25 g on arrival, were obtained from Charles River Laboratories (Wilmington, MA) and individually housed for the duration of the study. Food and water were available ad libitum. Housing rooms were kept on a 12:12 light:dark cycle and ambient temperature was maintained at 24.5° ± 0.5°C. All procedures were conducted in accordance with the National Institutes of Health Guide for the Care and Use of Experimental Animals and were approved by Eastern Virginia Medical School's Institutional Animal Care and Use Committee (Protocol#: 17-015).
Mice were randomly divided into three ES treatment groups (ES, ES with excitatory light [ESOe] or ES with inhibitory light [ESOi]), three IS treatment groups (IS, ISOe, and ISOi), and two home cage (HC or HC with eYFP control construct [CHCO]) control groups (n = 4–6 per group). Previously, we have reported that stress-related measures, such as sleep or inflammatory-related gene expression, do not differ greatly between HC and a control group which was placed in the shuttlebox chamber on each experimental day but not shocked (mock trained). 11 Therefore, in this study, we primarily focused on how metabolic pathway regulation differs between ES/IS groups relative to the HC group, and how this may be modulated with optogenetic manipulations of BLA.
Virus vector construct
Purified adeno-associated virus preparations containing CaMKII that enables selective targeting of a subpopulation of glutamatergic neurons were used in this study as previously described. 6 The final viral concentration of each construct injection was 1.5 × 1012 virus molecules/ml in 350 mM NaCl, 5% D-Sorbitol.
Surgery
All surgical procedures were conducted as described in detail previously. 6 Briefly, microinjection cannulae (26-guage) connected to a syringe pump (BSP-99M, Braintree Scientific Inc., Braintree, MA) were stereotaxically placed bilaterally above BLA (−1.5 mm AP, ±2.9 mm ML, −4.7 mm DV) for administration of CaMKII constructs. Custom-made optic probes (200 µm, conical tip, mated to metal ferules) were then implanted directly above the injection sites and secured to the skull using dental cement. The animals were given at least four weeks to recover and to allow viral transduction. During that period, the animals were kept undisturbed except for postsurgical monitoring and weekly bedding changes.
Training procedures
On experimental days 1 and 2, animals were shock trained (ST) with ES or IS as previously described. 6 Briefly, the ES groups had the ability to learn they could behaviorally terminate the footshock by moving to the opposite shuttlebox chamber; the yoked IS groups could not control the shock. Termination of shock for an ES mouse also terminated the shock to its yoked IS mouse in a separate shock chamber (Coulbourn Instruments, Model E10-15SC), ensuring each yoked set of mice received the same duration of shock. Thus, a pair of mice received identical shock, but it was characterized as either controllable or uncontrollable based on ability to escape. For optogenetic manipulations of BLA, mice were connected bilaterally to optic fibers and placed in a shock chamber for ST as previously described. 6 Inhibitory light stimulation started prior to ST and continued for the duration of training. Excitatory light stimulation occurred at the start of shock presentations during ST and continued until the last shock presentation. HC and CHCO groups did not experience the shuttlebox or shock, nor did the CHCO group experience the hookup to optic fibers.
RNA extraction
All groups were euthanized following ST on the second training day and RNA isolated from whole HPC and mPFC as described previously. 6 Samples from each region were loaded into NanoString® Mouse Neuroinflammation Panels which provides a broad view of multiple underlying aspects of neuroinflammation, including an assessment of neurotransmission, neuron–glia interactions, neuroplasticity, cell integrity, as well as metabolism. Results from the panels were uploaded to the nSolver database (Version 4.0.70; NanoString Technologies; Seattle, WA) to assess relative levels of metabolic markers within the panels. Gene expression and pathway profiles related to metabolism were compiled for each group to assess expression levels relative to the stress response following ST.
Statistical analyses
Data were normalized to NanoString's internal positive and negative controls to account for slight differences in assay efficiencies. The normalized gene counts for each gene in each assay were then divided by the appropriate normalization factor and averaged for the samples of each mRNA type to generate counts normalized to the internal reference genes. Fold changes in gene transcript levels were determined relative to basal levels detected in the HC (or CHCO) group. Relative fold changes in transcript levels for each determined gene were compared between groups. The data were first internally analyzed within nSolver's automatic analysis function that used multiple t-tests with Benjamini–Yekutieli correction to control for the false discovery rate to determine significance for each individual normalized gene. When comparing overall Pathway Regulation Scores between Groups (ES versus IS, ESOi versus ISOi, and ESOe versus ESOi), further two-way analyses of variance (ANOVAs) with Treatment (ES or ESOi/e and IS or ISOi/e) as between factors and Pathway Regulation Scores as a within-subject's factor where appropriate. Further one-way ANOVAs were used to compare differences in individual Pathway Regulation Scores between Treatments (no/excitatory/inhibitory optogenetic manipulations) within each Group (ES and IS). Tukey's post hoc analysis guided by the ANOVA (alpha = 0.05) on group and related gene expression was performed where appropriate. The following comparisons were evaluated: ES compared to HC, IS compared to HC, ES versus IS, ESOi compared to CHCO, ISOi compared to CHCO, ESOi versus ISOi, ESOe compared to CHCO, ISOe compared to CHCO, ESOe versus ISOe, comparisons between ES/ESOi/ESOe, and comparisons between IS/ISOi/ISOe.
Results
Signaling pathways related to metabolism
Pathway Regulation Scores were determined using the nSolver database via directed global significance scores of overlaid differential gene expression data for sets of genes grouped by biological function relative to HC (or CHCO). This analysis measures the extent to which genes within a given set are up- or downregulated with the independent variable. We examined overall regulatory changes in three major metabolic pathways based on changes in the expression levels of preselected genes incorporated in the NanoString® Mouse Neuroinflammation Panels, including: carbohydrate metabolism (10 total genes: Akt1, Akt2, Creb1, Igf1, Igf1r, Pik3ca, Pik3cb, Pik3r1, Sirt1, and Slco2b1), insulin signaling (27 total genes: Akt1, Akt2, Atp6v0e, Atp6v1a, Braf, Csk, Dlg4, Dusp7, Eef2k, Egfr, Erbb3, Fyn, Grin2a, Grin2b, Igf1, Igf1r, Il2rg, Kit, Mafb, Map2k1, Nefl, Pik3ca, Pik3cb, Pik3r1, Pik3r2, Psmb8, and Tcirg1), and lipid metabolism (18 total genes: Apoe, Crebbp, Ep300, Fa2h, Gal3st1, Gba, Gdpd2, Hpgds, Lsr, Pik3ca, Pik3cb, Pik3r1, Pla2g4a, Prkaca, Prkacb, Pten, Ptgs2, and Ugt8a).
Metabolic pathway regulation immediately following stress exposure
No amygdalar manipulation
In HPC, the ES and IS groups displayed distinct metabolic regulatory pathway profiles associated with carbohydrate metabolism, insulin signaling, and lipid metabolism compared to HC (Figure 1a). In general, the ES group exhibited downregulation of all three metabolic pathways, while the IS group exhibited an upregulation of these pathways. Analysis of variance found significant differences on Treatment (F1,12 = 31.52; p = 0.0001) but not on Pathway (p = 0.94) The greatest fold change was in the lipid metabolism pathway (p < 0.01), although both the carbohydrate metabolism and insulin signaling pathways yielded significant differences (p < 0.05).

Metabolic pathways are differentially regulated between hippocampus (HPC) and medial prefrontal cortex (mPFC). Graphs plotting the regulation of metabolic pathway scores in (a) HPC and (b) mPFC. * p < 0.05; ** p < 0.01.
In mPFC, ANOVA found no significant differences (p = 0.86). Interestingly, metabolic pathway regulation seemed to have a greater expression change, whether up or down, in HPC than in mPFC compared to the HC group (Figure 1A and B).
Amygdalar inhibition
In HPC, both the ESOi and ISOi groups displayed a suppression of pathway profiles associated with carbohydrate metabolism, insulin signaling, and lipid metabolism. Analysis of variance found significant differences on pathway (F2,12 = 4.001; p < 0.05) and on treatment (F1,14 = 8.607; p = 0.01); however, there were no significant interactions found between them (p = 0.92). In general, the ISOi group exhibited a greater downregulation of these pathways compared to the ESOi group, and this was found to be significant for all pathways (p < 0.05) (Figure 2A).

Optogenetic inhibition of basolateral amygdala (BLA) differentially modified metabolic pathway regulation between hippocampus (HPC) and medial prefrontal cortex (mPFC). Graphs plotting the regulation of metabolic pathway scores in (a) HPC and (b) mPFC. * p < 0.05.
Analysis of variance found no significant differences in metabolic Pathway Regulation Scores in the mPFC (p = 0.82). Once again, metabolic pathway regulation seemed to have a greater expression change in HPC compared to mPFC, whether up or down, in the ESOi and ISOi groups compared to the baseline of the CHCO group (Figure 2A and B).
Amygdalar activation
In HPC, ANOVA found significant differences on Treatment (F1,11 = 12.29; p = 0.006) but not on Pathway (p = 0.49). The ESOe and ISOe groups displayed opposite regulation of the three metabolic pathways profiles associated with carbohydrate metabolism, insulin signaling, and lipid metabolism. In the ESOe group, there was a significant increase in the Pathway Regulation Scores of both carbohydrate metabolism (p < 0.05) and lipid metabolism (p < 0.01) compared to ISOe, which had a decrease in these Pathway Regulation Scores. The inverse was true for the insulin signaling pathway, as the ESOe displayed a downregulation of this pathway, while the ISOe displayed an upregulation (p < 0.05) (Figure 3A).

Optogenetic activation of basolateral amygdala (BLA) differentially modified metabolic pathway regulation between hippocampus (HPC) and medial prefrontal cortex (mPFC). Graphs plotting the regulation of metabolic pathway scores in (a) HPC and (b) mPFC. * p < 0.05; ** p < 0.01.
In mPFC, there was surprisingly minimal change in the regulation of the carbohydrate metabolism pathway in both the ESOe and ISOe groups compared to the baseline of the CHCO group and ANOVA found no significant differences in metabolic Pathway Regulation Scores in the mPFC (p = 0.82) (Figure 3B). Again, metabolic pathway regulation seemed to have a greater expression change in HPC compared to mPFC, whether up or down, in the ESOe and ISOe groups compared to the baseline of the CHCO group.
Between groups
In HPC, there were significant differences between groups in all three metabolic pathways observed. In the carbohydrate metabolism pathway, ANOVA found significant differences on Treatment (F5,22 = 3.257; p = 0.02). In the ESOe group, the carbohydrate metabolism pathway was significantly upregulated compared to the ES (p < 0.05) and ESOi (p < 0.01) groups (Figure 4A

Carbohydrate metabolism pathway scores between groups are modified by optogenetic manipulation of the basolateral amygdala (BLA). Graphs plotting the regulation of carbohydrate metabolism pathway scores in (a) Hippocampus (HPC) and (b) medial prefrontal cortex (mPFC). * p < 0.05; ** p < 0.01.
Regarding the insulin signaling pathway, ANOVA found significant differences on Treatment (F5,22 = 3.433; p = 0.0002), with the ISOe group having a higher Pathway Regulation Score compared to the IS (p < 0.001), ESOi (p < 0.01), and ISOi (p < 0.01) groups. Additionally, the ISOi group had a decrease in their insulin signaling Pathway Regulation Score compared IS (p < 0.05) (Figure 5A). Once again, ANOVA found no significant differences in insulin signaling Pathway Regulation Scores in the mPFC (p = 0.82) (Figure 5B).
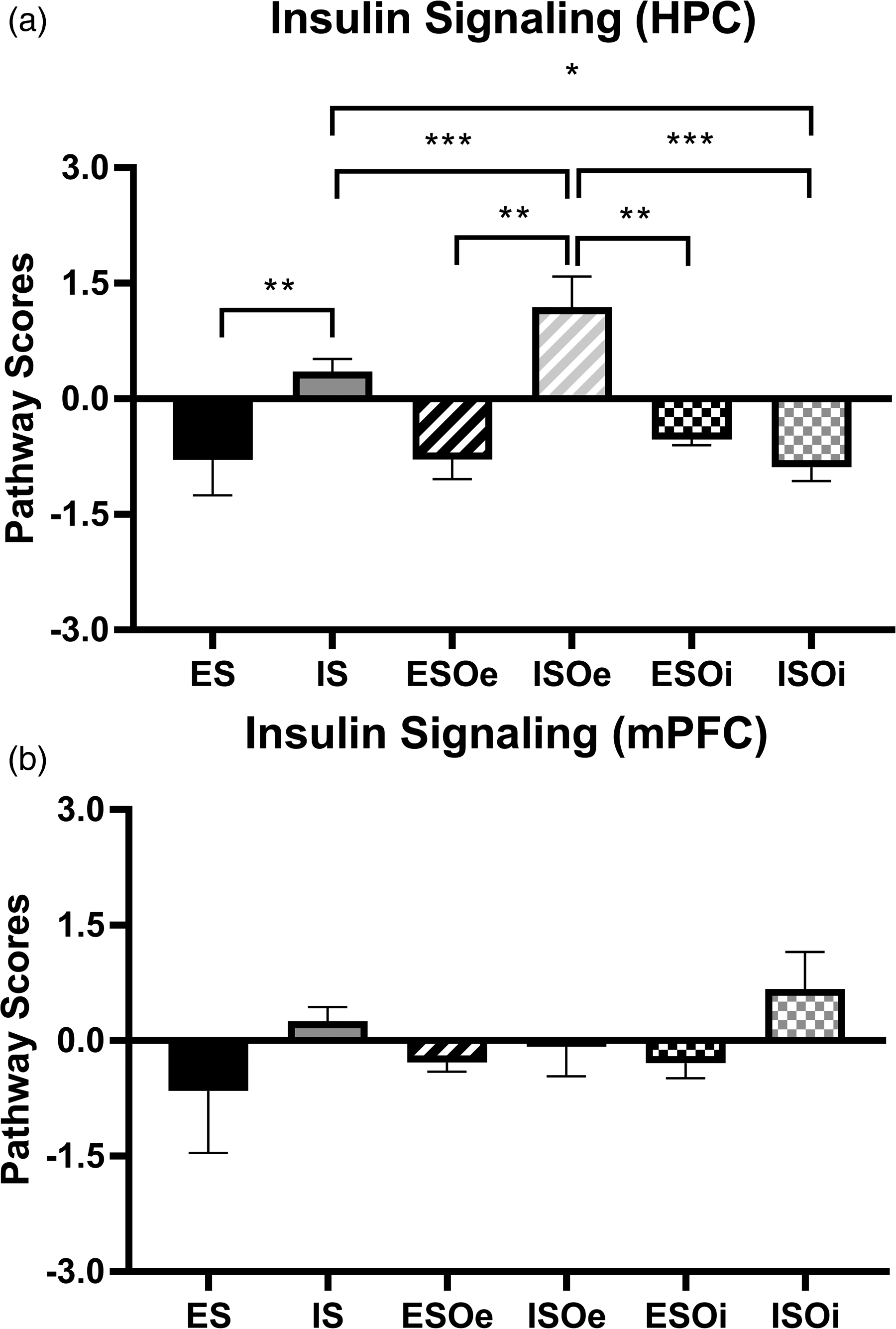
Insulin signaling pathway scores between groups are modified by optogenetic manipulation of the basolateral amygdala (BLA). Graphs plotting the regulation of insulin signaling pathway scores in (a) Hippocampus (HPC) and (b) medial prefrontal cortex (mPFC). * p < 0.05; ** p < 0.01; *** p < 0.001.
Regarding lipid metabolism, ANOVA found significant differences on Treatment (F5,22 = 1.9543; p = 0.002). Lipid metabolism in the ES group was significantly reduced compared to the ESOe group (p < 0.05), while the IS group had significantly increased lipid metabolism compared to the ISOe group (p < 0.05) (Figure 6A). Analysis of variance once again reveals that there were no significant differences in the lipid metabolism Pathway Regulation Scores in the mPFC (p = 0.82) (Figure 6B). Overall, the fold change in all of these metabolic pathways is greater in the HPC than the mPFC.

Lipid metabolism pathway scores between groups are modified by optogenetic manipulation of the basolateral amygdala (BLA). Graphs plotting the regulation of lipid metabolism pathway scores in (a) Hippocampus (HPC) and (b) medial prefrontal cortex (mPFC). * p < 0.05.
Discussion
Our study shows that the context of fear (escapable versus inescapable) can play a significant role on the metabolic pathways involved in brain responses. When comparing escapable groups (ES, ESOi, and ESOe) to their inescapable counterparts (IS, ISOi, and ISOe), we saw significant differences across carbohydrate metabolism, insulin signaling, and lipid metabolism pathways in the HPC. Furthermore, our study shows that optogenetic manipulation of the BLA significantly alters Pathway Regulation Scores between groups in the HPC.
The regulation of metabolism by the brain is well accepted, and many factors that regulate metabolism also regulate neurological processes. The brain receives and integrates multiple metabolic inputs via tightly regulated pathways that control food intake as well as energy allocation and homeostasis. 12 Of the genes analyzed across three pathways, seven were in more than one panel. Both Akt1 and Akt2 genes were included in the carbohydrate metabolism and insulin signaling panels. AKT can process glucose by activating glucose transporter 4 or phosphorylating hexokinase and also regulates insulin signaling by interacting with phosphoinositide 3-kinase (PI3K). 13 Our data show a trend where both ES and IS groups exhibited upregulation of Atk1 and Atk2 in carbohydrate metabolism and insulin signaling pathways in the HPC (with the ES group exhibiting more robust upregulation than the IS group) (Figures S1 and S2). This difference in upregulation may be a result of decreased carbohydrate metabolism and insulin signaling in the ES group (Figure 4). In addition to the Akt genes, insulin growth factor (Igf1 and Igf1r) genes also impact carbohydrate and insulin signaling. IGF genes increase insulin signaling, which in combination with growth hormone, could potentially alter carbohydrate metabolism. 14 Our data show a trend of negligible difference in Igf1 gene expression, but a notable upregulation in Igf1r expression (Figures S1 and S2). This upregulation was more robust in the ES group, which may again be a result of decreased carbohydrate metabolism and insulin signaling in the ES group (Figure 1). Finally, genes implicated in the PI3K signaling cascade (Pik3ca, Pik3cb, and Pik3r1) were included in all three panels. Phosphoinositide 3-kinase forms a signaling cascade with AKT to impact both glucose homeostasis and lipid metabolism. In addition, homologs of PI3K have been shown to play a prominent role in insulin signaling. 15 Interestingly, all these genes are linked through a shared signaling cascade or their ability to interact with insulin. Across all three pathways, the ES group had an upregulation of Pik3cb and a downregulation of Pik3ca and Pik3r1, while the IS groups had an upregulation of Pik3ca and Pik3r1, but a downregulation of Pik3cb (Figures S1–S3). These differences could be attributed to the downregulation of these pathways in the context of ES and their upregulation in IS (Figure 1). Regarding fold changes of individual genes, there were genes in all three metabolic pathways that had differential expression depending on both optogenetic manipulation and the ability to escape the stressor. The top five notable fold changes for each metabolic pathway in each treatment condition—as well as their log2 fold changes, standard deviations, and p-values—can be found in Supplementary Material (Figures S4, S5 and Tables S1–S6).
Overall, our studies attempted to assess this connection by examining differences in gene expression within the neurocircuitry of the mPFC, HPC, and BLA. It is well established that the mPFC, HPC, and BLA form a neurocircuit with one another and regulate many components of the stress response. 6 Projections from the BLA to the HPC help regulate the fear response, as disruption of amygdalar signals to the HPC result in fear generalization. 16 Hippocampus projections to the mPFC are necessary for working, episodic and emotional memory consolation and retrieval, 17 and mPFC-BLA projections are important for social context evaluation and emotional processing. 18
Previous research has already shown that the metabolic rates in the mPFC, HPC, and BLA can be altered by differences in ion receptor availability 19 and regional excitation. 20 However, this is the first study to our knowledge that shows the same stressor (footshock) eliciting different changes in metabolic pathway regulation in the same brain region, depending on the context of the stressor (escapable versus inescapable). This study found that BLA can regulate both carbohydrate and lipid metabolism, as well as insulin signaling, since optogenetic inhibition or excitation of the BLA modified stress-induced ES and IS outcomes on metabolic pathway regulation. A key example of this is in the insulin signaling pathway, which is significantly upregulated in the HPC of the IS group, and increased even more when the BLA is optogenetically activated (ISOe group). However, inhibiting the amygdala during inescapable stress (ISOi group) resulted in a decrease in insulin signaling. Because increased insulin is associated with increased glucose production and utilization, 21 we would expect to see alterations in insulin signaling as a reflection of carbohydrate metabolism. The groups that did not receive optogenetic manipulation (ES and IS) and those that received optogenetic inhibition (ESOi and ISOi) followed this pattern, where carbohydrate metabolism decreased as insulin signaling decreased; however, optogenetic excitation of the BLA produced an interesting difference, where carbohydrate metabolism decreased as insulin signaling increased, and carbohydrate metabolism increased as insulin signaling decreased. In addition to its role in glucose utilization, insulin can also function as a hormone and produce an anti-inflammatory effect. 22 Furthermore, previous studies in our lab have shown that optogenetic excitation of the BLA results in an upregulation of various genes associated with inflammatory signaling pathways. 6 Therefore, amygdalar activation and the inflammatory response could be interacting with one another, and this interaction could be interfering with different metabolic pathways in the brain regions that receive BLA projections (e.g., HPC and mPFC).
The amygdala also appears to modulate lipid metabolism. The ES group displays a decrease in the lipid metabolism Pathway Regulation Score, and the IS group shows an increase in this Score. When the BLA is optogenetically activated, the escapable group (ESOe) inversely has an increase in lipid metabolism, and the inescapable group (ISOe) has a decrease in lipid metabolism. This reversal of the lipid metabolism Pathway Regulation Scores was likely caused by optogenetic manipulation, as inhibiting the BLA resulted in minimal change compared to CHCO in the escapable group (ESOi) and a slight downregulation in the inescapable group (ISOi).
In addition to the differences seen within groups, there were also regional differences in pathway regulation. For example, Pathway Regulation Scores in the mPFC had little difference from the Control group in all three metabolic pathways observed, while Pathway Regulation Scores in the HPC expressed multiple differences based on context and optogenetic manipulation. Previous studies have noted these differences in the amygdala, cortex, and HPC of other neurocircuits, 23 but there is currently no explanation as to why these differences occur.
This preliminary work has limitations that should be addressed. While Nanostring® panels revealed significant differences in genes associated with metabolic pathways, the small number of animals in some groups may not have provided sufficient statistical power to reveal all potential differences. Additionally, while the Nanostring® panels revealed differences between treatment groups, the panels are specifically designed to investigate neuroinflammation signaling and only provide a broad view of multiple underlying aspects of neuroinflammation, including metabolism. Additional work is needed to elucidate specific metabolic processes. It would also be useful to examine subregions of HPC and mPFC which could reveal differences not observed when the whole area was examined.
Conclusion
This study provides important preliminary evidence identifying unique metabolic profiles within regions that are regulated by BLA during, and after, the stress response. Based on our findings, carbohydrate metabolism, insulin signaling, and lipid metabolism in HPC may be regulated by the BLA. Future work will be needed to reveal the precise mechanisms of these alterations, as well as to further examine the temporal course of BLA regulation of stress-induced metabolic regulation. Work will also be needed to assess other regions (e.g., mPFC) to understand circuit-level regulation of metabolism during the stress response.
Supplemental Material
sj-docx-1-sci-10.1177_00368504241253692 - Supplemental material for Evidence for a role of the basolateral amygdala in regulating regional metabolism in the stressed brain
Supplemental material, sj-docx-1-sci-10.1177_00368504241253692 for Evidence for a role of the basolateral amygdala in regulating regional metabolism in the stressed brain by Austin M Adkins, Zachary N M Luyo, Woong-Ki Kim, Laurie L Wellman and Larry D Sanford in Science Progress
Supplemental Material
sj-pdf-2-sci-10.1177_00368504241253692 - Supplemental material for Evidence for a role of the basolateral amygdala in regulating regional metabolism in the stressed brain
Supplemental material, sj-pdf-2-sci-10.1177_00368504241253692 for Evidence for a role of the basolateral amygdala in regulating regional metabolism in the stressed brain by Austin M Adkins, Zachary N M Luyo, Woong-Ki Kim, Laurie L Wellman and Larry D Sanford in Science Progress
Footnotes
Acknowledgements
This project was supported by an EVMS Institutional Multidisciplinary Research Award. The authors thank Dr Karl Deisseroth for sharing the optogenetic constructs with our lab and the EVMS Biorepository for running the NanoString assays.
Authors' contribution
Conceptualization, L.D. Sanford, L.L. Wellman, and W.K. Kim; methodology, L.D. Sanford and L.L. Wellman; software, A.M. Adkins; validation, A.M. Adkins; formal analysis, A.M. Adkins; investigation, L.D. Sanford, L.L. Wellman, A.M. Adkins, and Z.N.M. Luyo; resources, L.D. Sanford and L.L. Wellman; data curation, A.M. Adkins; writing original draft preparation, A.M. Adkins and Z.N.M. Luyo; writing—review and editing, L.D. Sanford, L.L. Wellman, W.K. Kim, A.M. Adkins, and Z.N.M. Luyo; visualization, A.M. Adkins and Z.N.M. Luyo; supervision, L.D. Sanford and L.L. Wellman; project administration, L.D. Sanford and L.L. Wellman; funding acquisition, L.D. Sanford, L.L. Wellman and W.K. Kim. All authors have read and agreed to the published version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, author ship, and/or publication of this article: This project was supported by an EVMS institutional Multidisciplinary Award.
Supplemental material
Supplemental material for this article is available online.
Author biographies
Austin M Adkins is a postdoctoral fellow specializing in the developmental origins of health and disease. His expertise is focused on the context of the biological basis of behavior, and physiology.
Zachary N M Luyo is a Biomedical Sciences PhD student. His research in the Sanford lab investigates the role of spaceflight stressors on behavior, immunology, and brain morphology.
Woong-Ki Kim is a neuroimmunologist with focus on neuroHIV and Alzheimer's disease. He has been studying microglia, CNS border-associated macrophages, and pericytes.
Laurie L Wellman is a professor in Neuroscience who specializes in behavior and neural structure and function. Her research focuses on the amygdala and fear circuitry as well as the role of sleep on cognitive and emotional processing.
Larry D Sanford is a professor in Neuroscience who specializes in sleep and behavior. His research focuses on the interactions between learning and memory, stress, sleep, and the immune system.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
