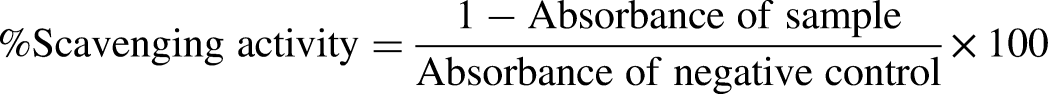Abstract
Termites cause a serious menace to wooden structures all over the world. They rely mostly on entozoic fauna residing in their hindgut for the digestion of cellulosic and hemicellulosic materials. One of the ways to control termites is through their gut symbionts. The present study was designed to characterize the hindgut bacteria isolated from Odontotermes obesus and Heterotermes indicola. Furthermore, the growth inhibitory effect of eight tropical plant extracts was investigated to find out potential control agents for these bacterial isolates. The characterization of bacteria was carried out based on their morphology, Gram staining, biochemical and amplification of 16SrRNA gene. Amplified products were sequenced to confirm their relationship with bacterial isolates from termites of other regions. The growth inhibitory effect of ethanolic leaf extracts of eight plants was evaluated in an invitro agar well diffusion method. Qualitative and quantitative phytochemical analysis of the most effective plant was carried out to learn about bioactive agents. The results confirmed the presence of five bacteria from each termite species. The Bacillus cereus, Escherichia coli, and Lysinibacillus fusiformis were common to both termites whereas Lysinibacillus xylanilyticus and Lysinibacillus macrolides were found in O. obesus only and H. indicola harbor Bacillus subtilis and Shigella sonnei in addition to common three ones. Among the plant extracts of Carica papaya, Eucalyptus camaldulensis, Osmium basilicum, Grevillea robusta, Eucalyptus globulus, Pongamia pinnata, Mentha longifolia, and Melia azedarach, the G. robusta > E. camaldulensis > O. basilicum were found to have growth inhibitory effects with increasing concentrations from 100 to 2000 µg/mL. The biodiversity of the bacterial fauna is important for the biological control of termites. Leaf extracts of these medicinal plants can be used to control termite infestation in an environment-friendly manner to save huge economic loss.
Keywords
Introduction
Termite infestation caused a significant economic loss all over the world by damaging wood and wooden materials except Antarctica. Along with their own digestive enzymes, they harbor a variety of symbiotic microorganisms including archae, bacteria, and eukarya in their hindgut which not only form a mutualistic relationship with each other as well as between microbes and their termite host. 1 The microbes are the source of nitrogen, carbon, and energy for their host. About 74–99% lignocellulose and 65–87% hemicellulose digestion is accomplished by termites. The effective breakdown of these chemical compounds is carried out by the synergistic activity of microbial fauna of termite gut along with the wide variety of cellulolytic enzymes. 2 These organic materials are initially converted into simple sugars and then finally into ethanol by the action of the microbial community present in the hindgut. 3 Bacterial fauna in termite hindgut consists of both obligate and facultative anaerobes. For the survival of facultative anaerobes, oxygen penetrates the gut through the body wall. They are not only the symbionts of termites but also live as ecto- and endosymbionts of flagellates found in their gut, a main source of cellulose digestion. In Mastotermes darwiniensis approximately 90% of bacteria were found to be associated with flagellates as symbionts. 4
Previous studies revealed that high bacterial diversity is present in hindgut as well as termite-specific bacterial lineages. 5 Further, knowledge about genes involved in reductive acetogenesis, fermentation, lignocellulose digestion, and nitrogen fixation is also elaborated.1,6 As termites are very diverse, the knowledge about the evolutionary relationship between termites and their endosymbionts is insufficient. Here the main aim of the present study is the isolation, characterization, and identification of bacterial symbionts from the hindgut of two commonly found termites in Pakistan; a higher Odontotermes obesus and a lower Heterotermes indicola by using morphological, biochemical, and molecular techniques and finding their evolutionary relationships with other bacteria and host termites. Moreover, ethanolic leaf extracts from eight tropical plants, that is Eucalyptus camaldulensis, Osmium basilicum, Grevillea robusta, Carica papaya, Eucalyptus globulus, Mentha longifolia, Pongamia pinnata, and Melia azedarach were evaluated for their antibacterial activity against identified bacterial isolates to identify most effective one. So these plants derived natural products could be an effective alternate for termite control by targeting their bacterial fauna.
Materials and methods
Sample collection
Workers of two termite species, that is H. indicola and O. obesus, were collected from dead wooden logs and mud mounds with the help of collection trap units as described by Sornnuwat et al. 7 and identified by using the key prepared by Akhtar. 8 The termite species were kept in petri dishes with moistened filter paper at 25°C for 1 week before the experiment to make their hindgut contents clear from debris and other wood particles.
Dissection and inoculum preparation
The exterior surface of about 100 termite workers was washed at least 10 times with distilled water and the gut was expelled from the entire body with a fine tip of sterilized forceps in 0.08% normal saline solution. The gut was opened to release its contents and 1 mL was serially diluted to get discreet bacterial colonies. Cultures were prepared on nutrient agar plates by incubating at 37°C for 24 h and transferred to fresh media plates after every week to get pure colonies. 9 Pure cultures were preserved in nutrient agar slants at 4 °C for further investigations.
Bacterial identification
Isolated bacteria were identified and characterized by the following methods.
Colony morphology Gram staining Biochemical identification Molecular identification
Colony morphology and gram staining
The morphological identification of bacteria was performed based on visual differences in size, shape, pigmentation, elevation, margin, opacity and appearance in bacterial colonies. Gram staining was performed to discriminate between gram-negative and gram-positive bacteria based on difference in their cell wall composition. 10
Biochemical characterization
Biochemical characterization of isolated bacterial cultures was performed 11 and following biochemical tests were used (1) triple sugar iron test for lactose/glucose fermentation; (2) Indole and H2S test; (3) citrate utilization test; (4) nitrate reduction test; (5) catalase test; (6) Urase test; and (7) methyl red Vogas-Proskauer test.
Molecular identification
DNA extraction and PCR amplification of bacterial 16SrRNA gene
Bacterial DNA was extracted by using the CTAB method with some modifications. 12 Polymerase chain reaction (PCR) amplification of the 16SrRNA gene was performed by using the following set of primers UNR-5′-GGACTACCAGGGTATCTAAT-3′ and UI-5′-CCAGCAGCCGCGGTAATACG-3′. PCR reaction was conducted by using a Thermo cycler (96 universal Gradient Peq Star Lab Peq UK) having a reaction mixture of 50 μL, comprising PCR water (33.3 μL),Taq buffer (5 μL), MgCl2 (4 μL), dNTPs (1 μL), forward and reverse primer (2 μL), and Taq DNA polymerase (0.7 μL). At the end, 2 μL of DNA template was added. Cyclic conditions involve, initial Denaturation (1 cycle) at 95°C for 4 min, denaturation (35 cycles), annealing and extension at 95°C for 1 min, 60°C for 2 min and 72°C for 2 min then final extension (1 cycle) at 72°C for 10 min. The amplified PCR products were visualized by using 1% agarose gel. Samples were loaded, stained with ethidium bromide. Bands were observed under UV light and images were taken with a Dolphin gel documentation camera (Wealtec, USA). The gene jet purification kit (Invitrogen) was used for the separation of a particular PCR product according to the manufacturer's protocol. Amplified PCR products were sequenced according to the protocol provided by ABI Prism Big Dye Terminator Cycle Sequencing Ready Reaction Kit v. 3 (Applied Biosystems) using ABI 373A automated sequencer.
Plants selected for bioassay
Leaves of eight plants including C. papaya Linn. (Papaya), E. camaldulensis Dehnh (River red gum), O. basilicum Linn.(Sweet basil), G. robusta Cunn. (Silver oak), E. globulus Labill (Blue gum), P. pinnata Linn.(Indian beech tree), M. longifolia Linn.(Horse mint)and M. azedarach Linn. (Chinaberry tree) were selected to evaluate their antibacterial activity against bacterial isolates from hindgut of O. obesus and H. indicola. The leaves were collected from the vicinity of Quaid-i-Azam University Islamabad, Pakistan and identified with the help of a key described by Nasir and Ali.
13
The leaves were then washed, shade dried at room temperature (27°C–37°C) and powdered mechanically with an electric blender (Daigger Scientific®, USA). Thirty grams of plant powder were subjected to obtain extract in 250 mL of ethanol solvent by using Soxhlet apparatus at 40°C–60°C (two cycles per hour) for 8 h. The extract was concentrated by evaporating the solvent with a rotary vacuum evaporator (R-300, Rotavapor®, Germany). The stock solution was prepared by dissolving 1 g of crude extract in 100 mL of distilled water and different concentrations (100, 500, 1000, 1500, and 2000 µg/mL) were prepared by applying the formula C1V1 = C2V2.
Antibacterial bioassay
The agar well diffusion method was used for antibacterial activity of different plant extracts against bacterial strains. 14 Nutrient agar plates were prepared and 6 wells were pinched using sterile 5 mm cork borer. Using sterile cotton swab bacterial inoculum was uniformly spread on sterile agar plates. 150 µL of each plant extract (100, 500, 1000, 1500, and 2000 µg/mL) was poured into the wells while using water as negative control and left for diffusion at room temperature. Plates were then incubated for 24 h at 37°C ± 1 °C. After incubation antibacterial activity of the plant extract was evaluated and zone of inhibition was measured in millimeter.
Phytochemical analysis of G. robusta leaf extracts
Qualitative phytochemical screening
Gas chromatograph coupled mass spectroscopy (GC-MS) was used to qualitatively characterize the ethanolic leaf extract of G. robusta by gas chromatograph coupled with a mass spectrophotometer (GC-2010, Shimadzu). The mass spectrum was identified by the National Institute of Standards and Technology (NIST) database having more than 62,000 patterns. The comparison between spectra of known and unknown compounds helps in the interpretation of the mass spectrum. 15
Quantitative phytochemical screening
Total phenolic content
Total phenolic content was estimated by using Folin-Ciocalteu (FC) as a reagent while gallic acid was a positive control. About 20 µL of crude plant extract (4 mg/mL DMSO) was transferred to 96-well microtiter plate then 90 µL of FC reagent was added. After incubation for 5 min at room temperature, 90 µL of Na2Co3 (6% w/v) was added and absorbance was recorded at 630 nm. The total phenolic content was represented as µg of gallic acid equivalent (GAE)/mg of dry weight (DW). 16
Total flavonoid content estimation
Total flavonoid content was estimated by using Quercetin as standard. About 20 µL of sample (4 mg/mL DMSO) was added in 96-well plates along with 10 µL of potassium acetate (1 M) and 10 µL of AlCl3 (10% w/v), and 160 µL of distilled water. The mixture was left at room temperature for 30 min and absorbance was recorded at 415 nm. Quercetin concentration was 2.5–40 µg/mL and the resultant total flavonoid content was represented as µg of Quercetin equivalent (QE) per mg of DW.
Antioxidant assay
DPPH radical scavenging activity
The crude leaf extract of G. robusta may be comprised of potential radical scavengers that enhance their toxic effect. The DPPH (1,1-diphenyl-2-picryl-hydrazyl) free radical scavenging activity of extract was evaluated according to Brand-Williams (1995). The 20 µL aliquot of plant extract (4 mg/mL DMSO) was added to DPPH solution (9.2 mg/100 mL methanol) and incubated at 37% for 60 min. The absorbance of the sample and standard (ascorbic acid) was recorded at 517 nm. Percent radical scavenging activity was calculated by the following formula 16 ;
Ferric reducing antioxidant potential
The total reducing power of G. robusta was estimated by adding 200 µL of the sample with 400 µL of phosphate buffer (0.2 M, pH 6.6) and 500 µL of potassium ferric cyanide (1% w/v). After incubating at 30°C for 50 min, 400 µL of trichloroacetic acid (10% w/v) was added and centrifuged at 3000 rpm for 10 min. Finally, 150 µL of supernatant was mixed with 50 µL of FeCl3 (0.1% w/v) and absorbance was recorded at 700 nm. Ascorbic acid was used as positive control. The total reducing power of the sample was taken as µg of ascorbic acid equivalent per mg of DW. 17
Total antioxidant capacity
The total antioxidant capacity of G. robusta leaf extract was carried out by adding 1 mL of reagent solution (ammonium molybdate (4 mM), sodium phosphate (28 mM), sulfuric acid (0.6 M)) with 100 µL of plant sample and incubated for 90 min at 90°C. Ascorbic acid was used as the standard solution and absorbance was recorded at 645 nm. The total antioxidant capacity of the sample was taken as µg of ascorbic acid equivalent per mg of DW. 18
Results
Colony morphology
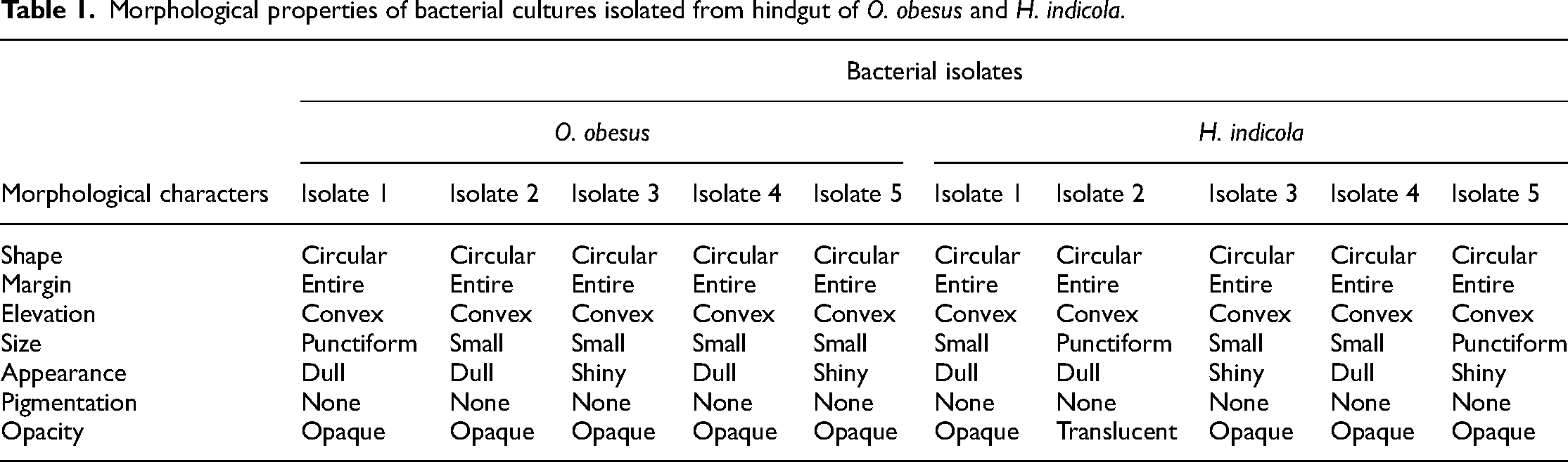
The colony morphology of pure cultures isolated from the hindgut of O. obesus and H. indicola was observed under a magnifying glass and the results are given in Table 1.
Morphological properties of bacterial cultures isolated from hindgut of O. obesus and H. indicola.

Gram staining identification and biochemical identification
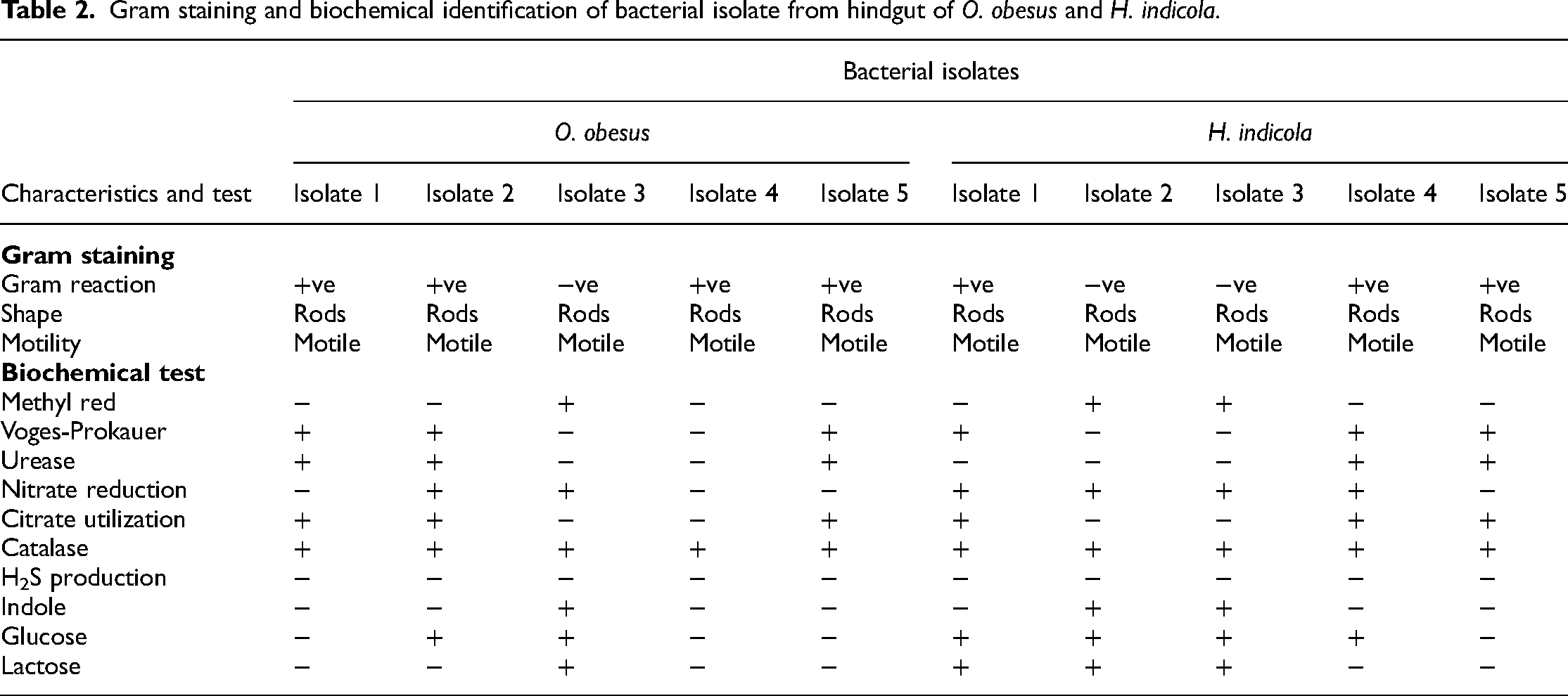
Gram staining was performed on pure bacterial isolates of both termite species to discriminate between gram-negative and gram-positive bacteria. Similarly, biochemical characterization of bacterial isolates was carried out by using different biochemical tests. Results of gram staining and biochemical identification are given in Table 2.
Gram staining and biochemical identification of bacterial isolate from hindgut of O. obesus and H. indicola.
Molecular identification
PCR amplification of 16SrRNA gene of bacterial isolates from the hindgut of O. obesus and H. indicola
The amplification of the 16SrRNA gene was confirmed by performing a gel electrophoresis assay and results revealed that bacterial strains isolated from the hindgut of O. obesus and H. indicola had nucleotide sequence of 193 bp (Figure 1).

PCR amplification of 16SrRNA gene of (a) O. obesus and (b) H. indicola.
Phylogenetic analysis
The NCBI (http://blast.ncbi.nlm.nih.gov/) blast search results for these isolated samples showed the highest sequence similarity with the different bacteria represented in Tables 3 and 4. Moreover, the neighbor-joining tree based on the 16SrRNA gene sequence was constructed to show the relationship between different isolates. Bayesian probability together with bootstrap values for Niebuhr Joining, Parsimony and Maximum Likelihood analysis strongly supported our phylogenetic tree and its clades. Evolutionary analyses were conducted in MEGA6. 19 These results also confirmed the results of NCBI-BLAST for bacterial isolates, the phylogenetic tree based on 16SrRNA gene sequence showed the relationship between the different isolates and the representative species of the family Enterobacteriaceae and Bacillaceae as shown in Figures 2 and 3.

Molecular phylogenetic tree of bacterial isolates (1–5) from hindgut of O. obesus.

Molecular phylogenetic tree of bacterial isolates (1–5) from hindgut of H. indicola.
Accession number of different bacterial species from NCBI used for constructing a phylogenetic tree of isolates 1–5 from hindgut of O. obesus.

Accession number of different bacterial species from NCBI used for constructing a phylogenetic tree of isolate 1–5 from hindgut of H. indicola.

Antibacterial activity of plant extracts
Potential of plant extracts against bacteria isolated from the hindgut of O. obesus
The potential of eight tropical plant extracts in ethanol solvent was tested against bacterial isolates from hindgut of O. obesus. All plants showed differential antibacterial activity at all concentrations (100, 500, 1000, 1500, and 2000 µg/mL). P. pinnata and C. papaya were found least effective plants having activity against two isolates only at higher concentrations, that is 1500 and 2000 µg/mL. P. pinnata showed a zone of inhibition against B. cereus (4.3 ± 0.03, 7.8 ± 0.67 mm) and L. macrolides (8.6 ± 0.45, 9.5 ± 0.89 mm). Similarly, C. papaya also has a zone of inhibition against B. cereus (9.8 ± 0.05, 12.0 ± 0.45 mm) and E. coli (5.7 ± 0.06, 9.3 ± 0.5 7 mm). M. azedarach showed zone of inhibition against L. fusiformis at all above mentioned concentrations, that is 3.50 ± 0.09, 4.90 ± 0.57, 4.90 ± 0.02, 5.30 ± 0.03, and 6.90 ± 0.88 mm, respectively. However, it was active against L. macrolides and L. xylanilyticus only at 1500 and 2000 µg/mL having zones of inhibition 8.06 ± 0.92, 11.40 ± 1.22, 7.06 ± 0.31, and 9.56 ± 0.43 mm, respectively. M. longifolia was found active against B. cereus, L. macrolides, and L. xylanilyticus at higher concentrations (1000, 1500, and 2000 µg/mL). E. globulus showed activity against B. cereus and E. coli at all concentrations while active against L. macrolides only at 1500 and 2000 µg/mL having zone of inhibition 11.50 ± 1.32 and 14.40 ± 1.53 mm, respectively. E. globulus has the largest zone of inhibition (21.40 ± 0.95 mm) at 2000 µg/mL against E. coli and the smallest zone of inhibition (11.33 ± 0.16 mm) against B. cereusat 100 µg/mL (Table 5). However, it does not have any growth inhibitory effect on L. fusiformis and L. xylanilyticus.
Zone of inhibition (mm) measured against five bacterial isolates from hindgut of O. obesus.

The third most effective plant was E. camaldulensis having activity against all isolates. Its maximum zone of inhibition was 24.83 ± 1.08 mm at 2000 µg/mL against L. macrolides and its minimum zone of inhibition was 5.00 ± 0.65 mm at 100 µg/mL against L. fusiformis. O. basilicum was the second most active plant at all concentrations except 100 µg/mL against B. cereus, E. coli, L. macrolides, and L. xylanilyticus while against L. fusiformis it was active only at 1500 and 2000 µg/mL having a zone of inhibition 5.65 ± 1.21 and 9.31 ± 1.87 mm, respectively. Its maximum zone of inhibition was recorded against L. macrolides (14.89 ± 1.32, 20.16 ± 2.12, 22.89 ± 0.87, and 23.33 ± 1.18 mm) at all active concentrations as compared to the rest of the isolates. G. robusta was the most effective among all the tested plants having activity against all isolates at all concentrations. Its maximum zone of inhibition was recorded at 1500 and 2000 µg/mL against L. macrolides and L. xylanilyticus, that is 23.33 ± 1.56, 25.26 ± 3.01, 26.33 ± 0.87, and 27.50 ± 1.02 mm, respectively. Similarly, its smallest zones of inhibition were recorded against L. fusiformis, all above the mentioned concentrations having values 5.4 ± 0.03, 8.52 ± 1.74, 12.26 ± 1.36, 13.50 ± 1.20, and 15.00 ± 1.44 mm, respectively (Table 5).
Potential of plant extracts against bacteria isolated from the hindgut of H. indicola
The potential of plant extracts was also evaluated against bacterial isolates of H. indicola. All plants showed differential antibacterial activity at all concentrations (100, 500, 1000, 1500, and 2000 µg/mL). P. pinnata was least effective among all tested plants and had a zone of inhibition against B. subtilis at 2000 µg/mL and L. fusiformis at 1500 and 2000 µg/mL. The second least effective plant was C. papaya which has a zone of inhibition against B. subtilis at 1500 and 2000 µg/mL (9.0 ± 0.32, 10.3 ± 0.08 mm) and against S. sonnei at all concentrations with a zone of inhibition of 2.8 ± 0.01, 5.0 ± 0.01, 5.0 ± 0.57, 11.3 ± 0.08 and 11.8 ± 0.02 mm, respectively. Similarly, M. azedarach, M. longifolia, and E. globules were also active against only two isolates. M. azedarach was active against B. subtilis and L. fusiformis at above all mentioned concentrations having zones of inhibition 00 ± 00, 6.83 ± 0.72, 9.93 ± 1.05, 12.00 ± 0.76, 15.66 ± 1.20 mm and 5.30 ± 0.57, 7.09 ± 0.12, 11.83 ± 1.48, 15.06 ± 0.31, 19.90 ± 0.57 mm, respectively. M. longifolia has activity only against B. subtilis and B. cereus at all concentrations except 100 µg/mL with zone of inhibition 2.9 ± 0.35, 4.9 ± 0.57, 8.0 ± 0.12, 10.3 ± 0.08 mm and 4.57 ± 0.54, 5.80 ± 2.30, 7.16 ± 0.72, 7.50 ± 0.57 mm, respectively. E. globules was also effective against B. subtilis and E. coli. It showed a zone of inhibition at 1000 (11.83 ± 1.48 mm), 1500 (14.06 ± 1.31 mm), and 2000 µg/mL (18.90 ± 0.57 mm) against B. subtilis. While it was found active with all the above mentioned concentrations against E. coli having a zone of inhibition 5.50 ± 0.98, 6.67 ± 2.12, 7.72 ± 0.06, 9.3 ± 0.57 and 12.30 ± 0.01 mm, respectively (Table 6).
Zone of inhibition (mm) measured against five bacterial isolates from hindgut of H. indicola.

O. basilicum showed antibacterial activity against B. subtilis, E. coli, and B. cereus. The largest zone of inhibition (22.10 ± 0.18 mm) of O. basilicum was reported against B. cereus at 2000 µg/mL and smallest zone of inhibition (9.30 ± 1.01 mm) was against B. subtilis at 1500 µg/mL. The second most effective plant was E. camaldulensis against all isolates except E. coli at all the above mentioned concentrations. It was found least effective at 100 µg/mL having no zone of inhibition against all isolates except B. cereus (7.67 ± 0.56 mm) while antibacterial activity was increased with increase in concentration having the largest zone of inhibition at 2000 µg/mL with 19.54 ± 0.57, 20.73 ± 0.88, 22.75 ± 0.56, and 21.66 ± 2.68 mm against B. subtilis, S. sonnei, B. cereus, and L. fusiformis, respectively. G. robusta was found most effective among tested plants and showed a zone of inhibition against all isolates at all concentrations except B. subtilis at 100 and 500 µg/mL where no zone of inhibition was recorded (Table 6).
Phytochemical analysis of G. robusta leaf extracts
Gas chromatograph coupled mass spectroscopy
The GC-MS analysis of G. robusta leaf extracts indicated 15 peaks which revealed the presence of 15 phytochemical compounds. Compound name, retention time, area percentage, molecular weight, and molecular formula were identified by comparison with the NIST library. Coumarin, 7-hydroxy (30.64%), methyl hexofuranoside (27.63%), and 9,12,15-octadecatrienoic acid (Z,Z,Z) (8.97%), are reported as major compounds while 2,3-dihydrobenzofuran (1.93%), 3-hydroxy-2-methylbenzaldehyde (3.07%), 2-coumaranone (2.55%), 1,2,3,4,5-cyclohexanepentol (1.77%), 5-hydroxy-1-tetralone (0.69%), beta-d-mannofuranoside, methyl (3.34%), 5,5, 8a-trimethyldecalin-1-one (0.73%), n-hexadecanoic acid (6.30%), phytol (1.37%), 9,12, 15-octadecatrienoic acid (2.16%), bis(2-ethylexyl)phthalate (3.08%) and 4-hexyloxyphenol (5.77%) are present as minor constituents. The GC-MS chromatograms of identified compounds are presented in Figure 4 and their reported biological activities are presented in Table 7.
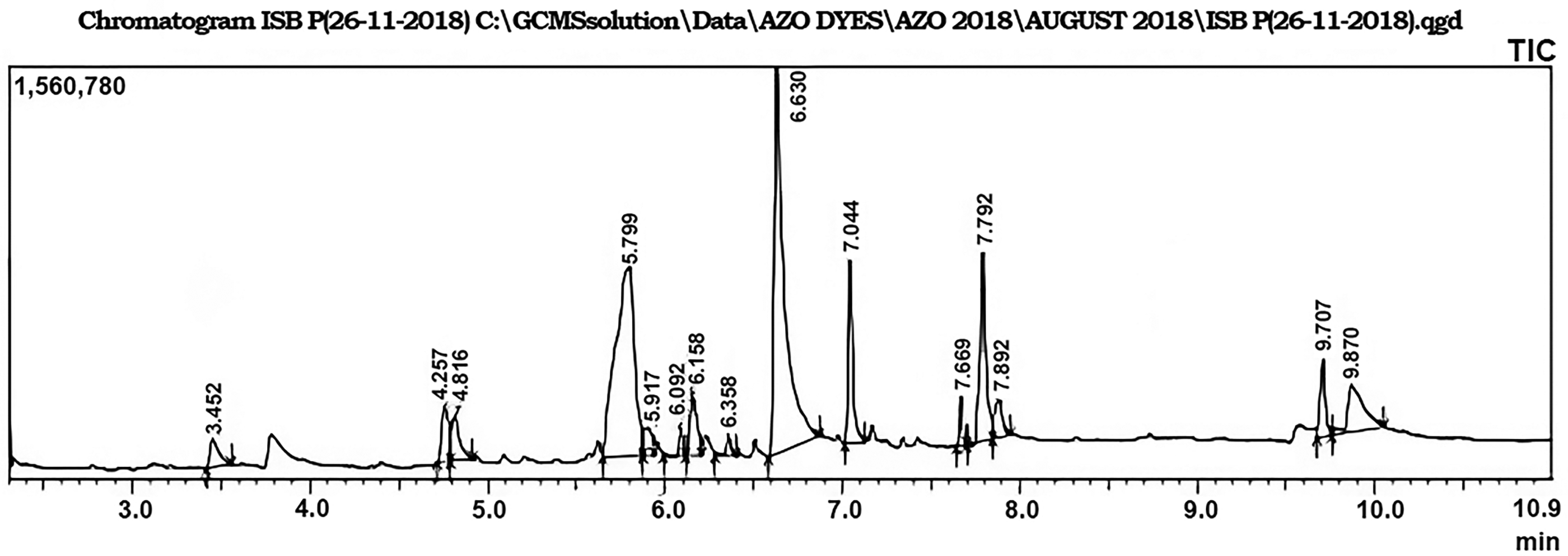
GC-MS chromatogram of ethanolic leaf extract of G. robusta. GC-MS: gas chromatograph coupled mass spectroscopy.
Reported biological activities of phytochemical compounds isolated from ethanolic leaf extract of G. robusta.

Total phenolic and flavonoid content
Quantitative phytochemical analysis showed that the total phenolic content of G. robusta leaf extract was 7.34 µg GAE/mg of DMSO. Similarly, total flavonoid content is expressed as Quercetin equivalent (µg GAE/mg DW), that is 9.38 ± 1 µg QE/mg DW and total flavonoid content was 5.29 µg QE/mg DW.
Antioxidant assay
Free radical scavenging activity (DPPH assay)
The percent DDPH free radical scavenging activity was detected based on the discoloration of the methanolic DPPH purple colored solution. The G. robusta leaf extract exhibited the most potent radical scavenging activity having an IC50 value of 5.25 μg/mL.
Ferric reducing antioxidant potential
The ferric reducing antioxidant potential of the crude plant extract was detected based on the conversion of the ferric ion (Fe+3) to the ferrous ion (Fe+2). The reducing potential of G. robusta was 28.69 μg AAE/mg of the extract.
Total antioxidant capacity
The total antioxidant capacity was evaluated by phosphomolybdenum-based method. The results revealed the total antioxidant capacity of 27.02 μg AAE/mg of the extract.
Discussion
Termite hindgut microbiota carry out different physiological functions, that is degradation of hemicellulose, cellulose, methanogenesis, hydrogenesis, acetogenesis, nitrogen fixation, and sulfate reduction. 10 The breakdown of these biopolymers into simplest sugars and then into ethanol is accomplished by these bioreactors of termite hindgut. 23 Scientist investigated different species of symbiotic microbial communities and revealed their promising role in cellulose digestion. Several studies are available on the identification and isolation of hindgut bacteria of different termites, for example Methanobrevibacter genus belongs to lower termites while Methanomicrococcus is associated with higher termites.24,25 Similarly,16SrRNA clones analysis of Reticulitermes falvipes showed the presence of bacteriodales (family Porphyromonadaceae) and Spirochetes (genus Treponema). 26 The presence of these bacteria suggests the unique lineages of bacteria residing in termite hindgut. Vikramet al. 27 also investigated that symbiotic microbiota associated with the termite gut wall are different from each other depending upon feeding habits of the different termite species. The present study has been conducted for the isolation and characterization of bacterial fauna from the hindgut of O. obesus and H. indicola. The absence of protozoan fauna from hindgut of higher termite species makes them more interesting subjects for the cellulose containing food digestion process.
Batool et al. 28 isolated two bacterial strains belonging to genus Bacillus from the digestive tract of Microtermes obesi which carry out xylanolytic and cellulolytic activity. Bacteroides, Parabacteroides, Dysgonomonas and phylotypes of genus Alistipes are abundantly present in Odontotermes than Microtermes sp. Bacteriodes abundance in Odontotermes sp. is a key finding in other fungus-growing termites, promoting their significant role in the termite hindgut. 29 The bacteria present in the termite gut play an important role in biofuel production in term of plant decomposition and are involved in sugar metabolism and polysaccharide breakdown. 30 In the present study bacterial isolates belonging to different families Enterobacteriaceae and Bacillaceaewere investigated from peripheral hindgut contents of O. obesus and H. indicola which play a significant role in maintaining the internal hindgut environment by secreting different enzymes. The presence of these microbes helps in nitrate fixing from the food chain and acts as a resistant barrier against microbial pathogens to protect microbial fauna of termite hindgut. B. subtilis reported in the present study, has also been investigated previously as aerobic, gram-positive, rod-shaped bacteria that help in nitrate fixation, inhibit fungal pathogens and are resistant to them. 31 Yuki et al. 32 investigated that bacteria belonging to the genus Bacillus produce alkaline enzymes like cellulases, hemicellulases, proteases, and amylases.
Different aerobic, anaerobic, and facultative bacteria have been isolated from termites belonging to Enterobacteriaceae, Enterobactor, Acinetobactor, Yersiniaceae, Citobactor, TrabulsiellaKluyvera, and Moraxellaceae. 33 Similar results were reported in the present study. All the bacterial isolates from the hindgut of O. obesus (higher termite) and H. indicola (lower termite) belong to the family Enterobacteriaceae and Bacillaceae. As termites feed on a variety of food sources that have cellulose and hemicellulose as major components of their diet which may contribute to the presence of plant associated bacteria in their hindgut. Diverse bacterial species belonging to different genera like Bacillus, Pseudomonas, Delftia, Serratia, Cedecea, Viridbacillus, and Lysinibacillus have been isolated from bryophytes of naturally raised swamp ecosystems. Serratia, Bacillus, and Pseudomonas were found frequently with plants suppressing the growth of bacterial and fungal pathogens, for example Erwinia sp. and Alernaria alternaria. 34 The presence of genus Lysinibacillus has also been confirmed from the hindgut of O. obesus and H. indicola in the present study suggesting the feeding habit of termites on various food sources.
In the present study antibacterial activity of crude ethanolic leaf extracts from eight medicinal plants was assessed against bacterial isolates from the hindgut of two termite species (O. obesus and H. indicola). All plants exhibit differential antibacterial activity at all concentrations (100, 500, 1000, 1500, and 2000 µg/mL), that increased with an increase in dose level. The difference in antibacterial activity of ethanolic leaf extracts of these plants might be due to the difference in their bioactive compounds. The results of the present study are supported by Rabe and Staden who reported that plant extracts are more effective against gram-positive bacteria as compared to the gram-negative bacteria due to the difference in their cell wall composition. 35 G. robusta, E. Camaldulensis, and O. basilicum were found to have maximum antibacterial activity against almost all bacterial isolates of both species at all concentrations. Most promising activity was observed at higher dose levels, that is 1500 and 2000 µg/mL. These findings are supported by previous studies that antibacterial activities are dose dependent. Stumpf et al. 36 reported the growth inhibitory concentrations of the tannins against Staphylococcus aureus and found that these values increase with increasing concentration in the growth medium. These findings are also supported by a study conducted by Kolenc et al. 37 that antibacterial activity of plants extracts against S. aureus and L. acidophilus was also dose dependent. Similarly, when the activity of tannins and their extracts was evaluated against E. coli similar results were observed that by increasing the dose, growth inhibitory effects also increased. 38 Maximum zone of inhibition of G. robusta was 24.54 ± 1.32 and 27.50 ± 1.02 mm against L. fusiformis isolated from H. indicola and O. obesus, respectively. Similarly, maximum inhibition of E. camaldulensis and O. basilicum were 22.75 ± 0.50, 22.10 ± 0.18 and 24.83 ± 1.08, 22.33 ± 1.18 mm, respectively against isolate four of both species. However, remaining plants were least effective and showed activity against one or two isolates at higher concentrations (Tables 5 and 6). The results of the present findings are supported by Ghalem and Mohamed. 39 They reported that ethanolic leaf extract of E. camaldulensis was a more potent antibacterial agent as compared to the E. globulus against S. aureus. Similar findings were also reported by Akin et al. 40 that crude leaf extract of E. camaldulensis in ethanol has strong antibacterial activity against B. subtilis, S. epidermidis, E. coli, S. marcescens, and S. aureus. O. basilicum is one of the most important plants among 60 species of Ocimum.41,42 Methanol, petroleum ether, and chloroform extract of O. basilicum were used against gram-positive bacteria (C. defficile, B. subtilis, and S. aureus) and gram-negative bacteria (E. coli, S. typhi, K. pneumoniae). These extracts were also evaluated for their antifungal activities against A. falvus, A. niger, and C. albicans. Results showed that O. basilicum has promising antimicrobial activities against tested pathogens.43,44 However, contradictory results were observed against leaf extracts of C. papaya and Eu. globulus in the present study as they were found as least effective against bacterial isolates. Antitumor, antibacterial, antifungal, and wound healing activities of C. papaya were reported previously. 45 Comparison between antibacterial and antifungal activities of C. papaya leaf extract showed that C. papaya has good antibacterial and antifungal activities.45,46 However, the present results are very different from the previous studies as C. papaya exhibits very little antibacterial activity against isolate 2 of O. obesus at 2000 µg/mL with a zone of inhibition of 12.0 ± 0.45 mm. Similar results were observed for E. globulus which was not more effective in present studies but previously reported to have moderate antibacterial activities. The effectiveness of these plant extracts may be due to their hydrophobic nature which helps them to penetrate deep into bacterial cells, destroying their cellular organelles, and releasing essential molecules leading to bacterial cell death. 47 GC-MS analysis of G. robusta leaf extract characterized 15 compounds (Figure 4). Among all the bioactive compounds identified in the present study (n-hexadecanoic acid, phytol, 12,15-octadecatrienoic acid (Z,Z,Z) and 2, 3-dihydrobenzofuran) have been previously reported for their various biological and antimicrobial activities (Table 7). Inside biological systems, phenolics, flavonoids, and polyphenols are oxygenated derivatives and have been reported to be associated with definite antioxidant action having free radical and single oxygen quenching ability. 48 The presence of hydroxyl, methoxy, and ketonic functional groups might be attributed to the antioxidant properties of phenols. Plant phenolic have reported antioxidant properties which make them imperative in fighting different conditions like oxidative stress, cell death, and cytotoxicity by their radical scavenging and trace elements chelating potential. 49 Free radical scavenging and inhibition of lipid peroxidation are considered as the most useful traits that make these compounds pharmacologically important. A linear correlation exists between the total contents of polyphenols and free radical scavenging. Therefore, it can be inferred that various polyphenols in the extract of G. robusta are largely responsible for radical scavenging mediated antioxidant activity. Similarly, the ferric reducing power has been reported to be associated with phenolic contents. 50 The significant reducing potential of ethanolic crude extracts in the present study affirms the presence of phenolic and flavonoid compounds. The polarity-based increase in results is in accordance with the previously reported studies where a different solvent range of varying polarity was employed. The results also indicated the presence of appreciable antioxidant capacity in the ethanolic extract of G. robusta. 51 The complete phytochemical screening revealed that the ethanolic extracts of G. robusta could be potential candidates for natural antioxidant preparations.
Conclusion
It is concluded from the present study that termite hindgut (O. obesus and H. indicola) harbors the bacteria belonging to the families Enterobacteriaceae and Bacillaceae. The isolated bacteria are related to their feeding habitats but their function is somewhat similar in all termite species including acetogenesis and cellulase production thus providing them the ability to digest wood. These termites are detrivorous, feeding on plants, litter, soil, and other substances which may contribute to the varying degree of microbial fauna in hindgut. Furthermore, the plant extracts of G. robusta, E. camaldulensis, and O. basilicum having highest antibacterial activity indicate the effectiveness of these plants as a potential candidate for the control termites by targeting their gut bacteria.
Footnotes
Acknowledgements
We are thankful to the Department of Animal Sciences, Quaid-i-Azam University, Islamabad, Pakistan for providing laboratory facilities to carry out this research work. We are also thankful to Mr Abdur Rehman Azam and Dr Sobia Abid for their help and support in refining the manuscript.
Author's contribution
AA and NAQ designed the study. AA carries out laboratory experiments. SQ and SU analyzed and interpreted the data. MA and NS helped in sampling of plant leaves and termites. MFN helped in bacterial culturing and identification. AA helped in the drafting of manuscript and NAQ proofread the manuscript. All authors approved the final manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Higher Education Commission, Pakistan (grant number 2291).
Author biographies
Asma Ashraf is an assistant professor of Parasitology. Her areas of interest are parasitology, entomology, microbiology, insect pest management and vector borne diseases.
Saima Qadeer is an assistant professor of Developmental Biology. Her areas of interest are developmental biology, endocrinology and teratology.
Sana Ullah is an assistant professor of Fisheries. His areas of interest are aquaculture and fisheries, toxicology.
Muhammad Asad is an assistant professor of Fisheries. His areas of interest are aquaculture and fisheries toxicology.
Huma Fatima is an assistant professor of Parasitology. Her areas of interest are parasitology, entomology, molecular biology and genetics.
Muhammad Farhan Nasir is an assistant professor of Fisheries. His areas of interest are aquaculture and fisheries toxicology.
Nargis Shaheen is a Postdoctoral fellow of Parasitology. Her areas of interest are parasitology, entomology, molecular biology and genetics.
Naveeda Akhtar Qureshi is an associate professor of Parasitology. Her areas of interest are parasitology, entomology, insect pest management and molecular biology.