Abstract
The search for 3R-relevant information is a prerequisite for any planned experimental approach considering animal use. Such a literature search includes all methods to replace, reduce and refine (3Rs) animal testing with the aim of improving animal welfare, and requires an intensive screening of literature databases reflecting the current state of knowledge in experimental biomedicine. We developed SMAFIRA, a freely available online tool to facilitate the screening of PubMed/MEDLINE for possible alternatives to animal testing. SMAFIRA employs state-of-the-art language models from the field of deep learning, and provides relevant literature citations in a ranked order, classified according to the experimental model used. By using this classification, the search for alternative methods in the biomedical literature will become much more efficient. The tool is available at https://smafira.bf3r.de.
Literature search as a legal prerequisite
Scientists are always looking for the best model to achieve the best possible result to answer a specific research question. For the biomedical field this means that researchers in principle weigh up whether an animal experiment or a non-animal approach is the most suitable model. This balancing is particularly required within the EU, because researchers have to justify the use of animals when applying for project authorisation (Article 37, Directive 2010/63/EU). 1 Such justification includes the consideration of methods to replace, reduce and refine (3Rs) the use of animals and has to be accompanied by an evaluation of the current state of scientific knowledge.
MEDLINE is a prominent database of biomedical and life sciences journal citations and currently provides over 30 million citations. 2 Researchers routinely access the database via PubMed, the associated search interface, to fulfil the legal requirements. This task, however, can prove arduous, since scientists add new scientific knowledge at a tremendous rate: on average, 3,750 citations were added to MEDLINE per day in 2022. 2 Current bibliographic classifications like MeSH (Medical Subject Headings) marginally support a literature search that specifically targets 3R-relevant contents. 3
Large language models to speed up the information retrieval process
We developed the SMAFIRA (Smart Feature-based Interactive Ranking) tool to support a 3R-relevant information retrieval from PubMed. SMAFIRA employs state-of-the-art language models from the area of deep (machine) learning, that is, artificial neural network-based word embeddings4,5 and BioBERT. 6 Our rationale was to speed up the information retrieval process by providing citations retrieved from PubMed in a prioritised and filtered hit list.
Searching is based on a reference abstract
The first version of our tool is now freely accessible at https://smafira.bf3r.de. To use the tool, researchers have to enter the PubMed Identifier (PMID) of a reference abstract that corresponds with the animal experiment the user wishes to replace with an alternative method.
Using the given PMID, SMAFIRA retrieves similar articles (citations and attached abstracts) from PubMed. The retrieved articles are classified by pre-defined categories of experimental models and are tagged accordingly: ‘vertebrate in vivo’, ‘vertebrate tissue/biopsy’, ‘vertebrate primary cell’, ‘vertebrate immortal cell’, ‘invertebrate’, ‘human’, and ‘virtual model’. Non-experimental, observational research is assigned with the label ‘others’. Because we recognised that the terms
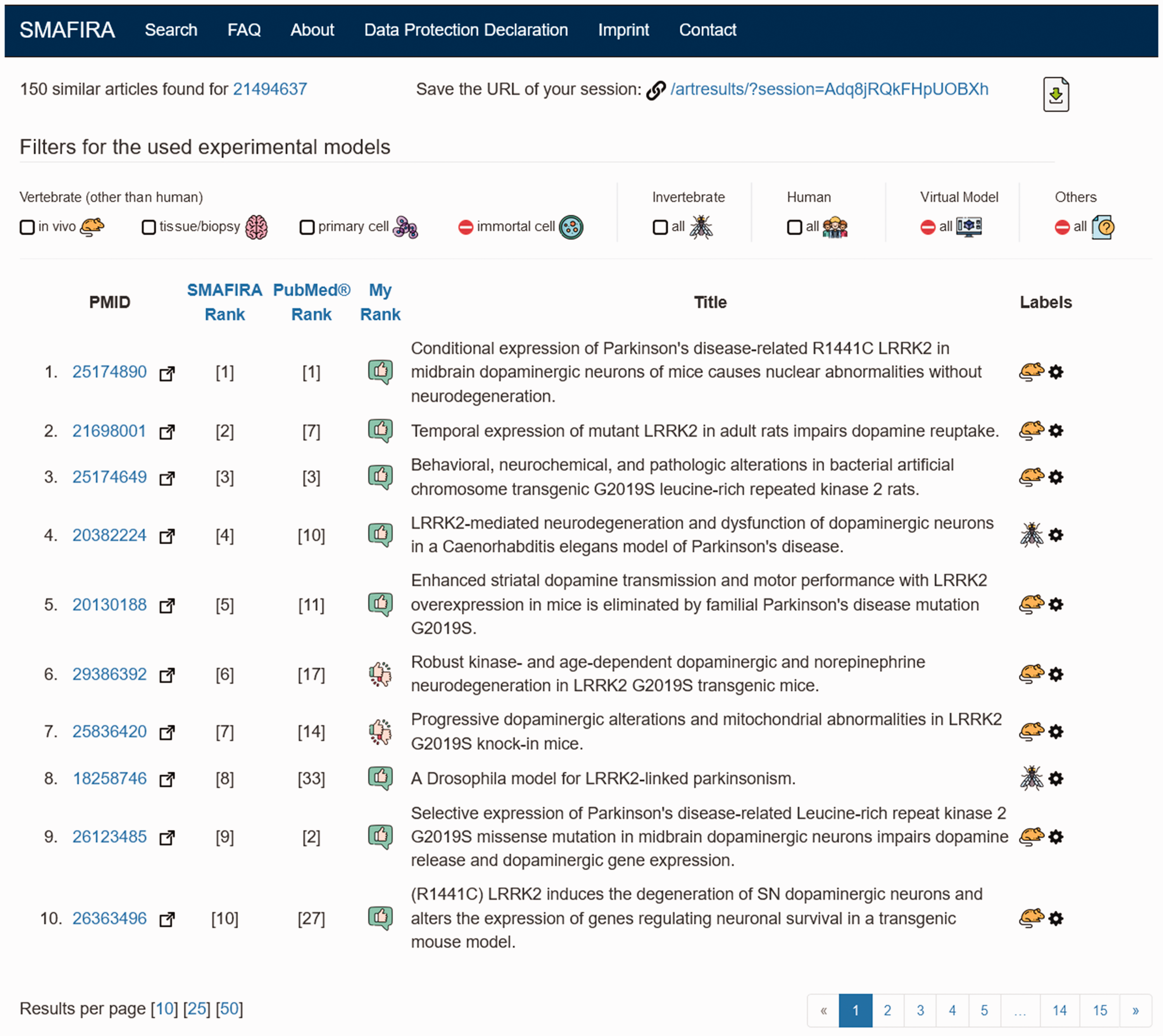
Screenshot of SMAFIRA results page using PMID 21494637 as the reference. Left: three rankings of retrieved literature are provided: 1) SMAFIRA ranking, 2) native PubMed ranking, and 3) My Rank based on the user’s judgement of similarity. When clicking on the PMID, the user is directed to the user opinion page (see Figure 2). Right: the category of the respective experimental model is depicted with icons (e.g. mouse for ‘vertebrate in vivo’). Top: the user can choose any category by clicking on the respective checkbox in the filter section. Selection of more than one filter is possible, the Boolean operator ‘OR’ is used as the connector. The URL of the results page can be saved for later recall of the session.
One of three rankings can be activated by clicking on the respective designator. The SMAFIRA ranking employs word embeddings generated via the word2vec algorithm4,5 in combination with cosine similarity. The PubMed ranking employs the pmra algorithm. 7
The icons under My Rank are only available if the user has submitted an own judgement regarding the similarity of two abstracts. To enable this judgement, a side-by-side comparison of the reference and listed abstract is provided on the user opinion page (Figure 2), which can be opened by clicking on the respective PMID in the hit list. Users can indicate their opinion on similarity by choosing among three icons. The default rankings provided by SMAFIRA and PubMed are not affected by this input. Furthermore, the automatic classification of the abstract according to the experimental model used can be overridden. All changes are automatically saved by the tool and can be recalled within 90 days by saving and calling up the URL on the results page. To support a further, user-oriented development of SMAFIRA, users are asked to forward the URL that links to their case study and changes to the developers (
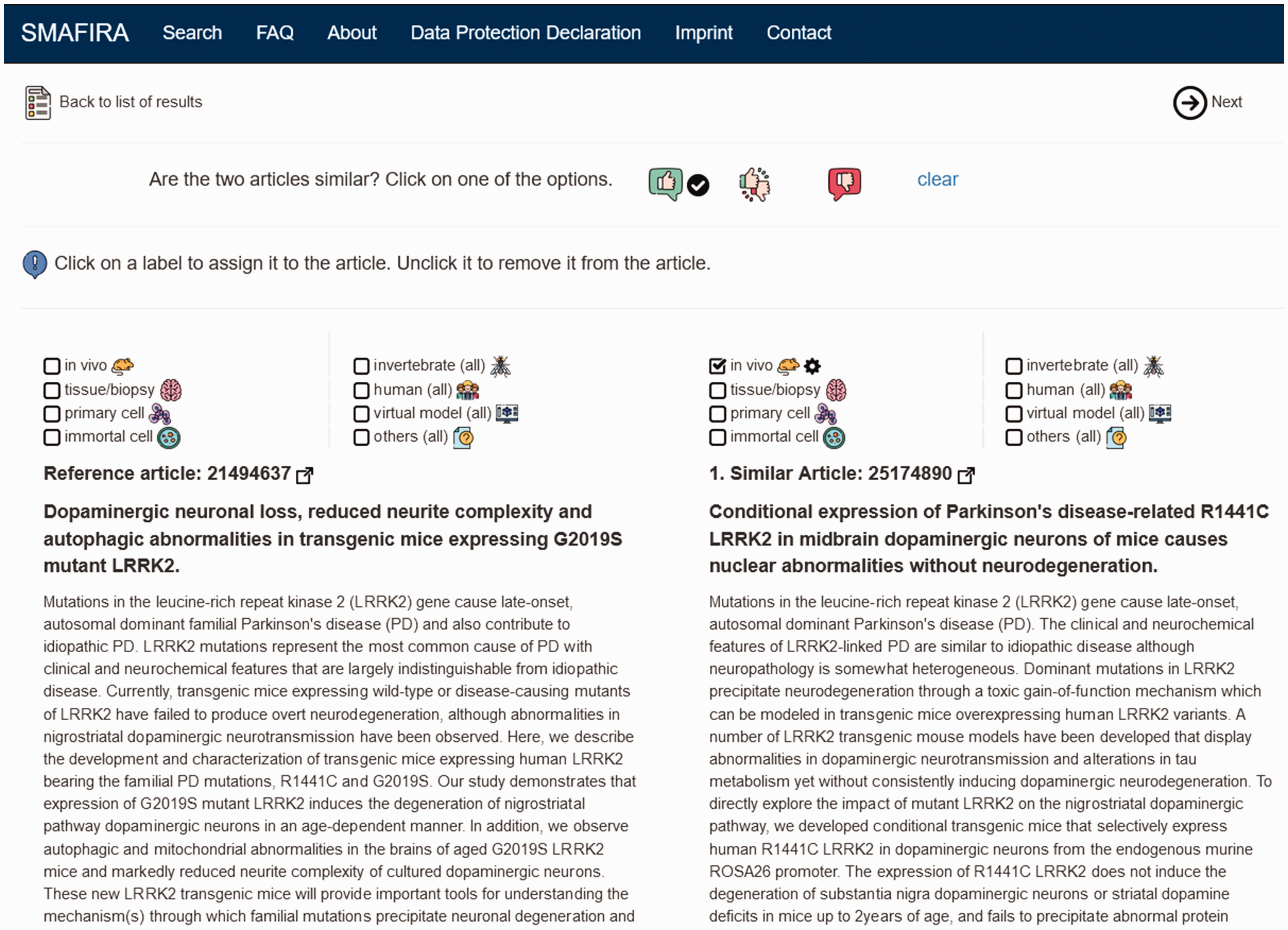
Screenshot of the user opinion page of SMAFIRA. The reference abstract (left) and the chosen similar article (right) are displayed for side-by-side comparison. Immediately above these articles, the experimental models predicted by the machine learning approach are indicated by an activated checkbox and a small cogwheel next to the icon. The automatically applied labels can be overridden by simply activating the relevant checkboxes. The similarity perceived by the user can be indicated by activating the respective icon (thumbs up = similar, mixed thumbs = uncertain, thumbs down = not similar).
Strategies for a 3R-relevant information retrieval using SMAFIRA
To retrieve possible replacement methods for a reference animal experiment, the user should choose between vertebrate tissue/biopsy, vertebrate primary cell, vertebrate immortal cell, invertebrate, human, and virtual model. Replacement options can then be identified by evaluating the top-ranked citations. However, SMAFIRA does not judge on the suitability of an alternative. This judgement is reserved for scientific experts. Thus, researchers have to evaluate the full text of a suggested citation to examine whether it really describes a candidate alternative method.
Reductions or refinements of an animal experiment can be retrieved indirectly by choosing the filter vertebrate in vivo. The top-ranked citations then need to be checked for the most similar vertebrate animal experiment. Subsequently, by inspecting the full text, 3R-relevant variations of the planned experimental procedure may be identified in the methods sections – if such variations are described.
A new instrument in the 3R-toolbox
SMAFIRA complements existing toolboxes to support application of 3R measures to animal experiments, for example, Experimental Design Assistant, 9 SWIFT-Review to support systematic reviews, 10 or 3Ranker, which was added only recently. 11 Apart from the ranking of semantically similar content, classification and filtering for specific experimental models is the chief added value of SMAFIRA. We hope that our tool will be a great support for experimental researchers and animal welfare officers in their search for alternative methods, and for members of ethical committees in their evaluation of animal projects.
Footnotes
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical statement
Our study did not require an ethical board approval because it did not contain human or animal trials.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
