Abstract
The extra-hospital epidemiology of Acinetobacter infections is a subject of debate. In recent years, the prevalence of animal multidrug-resistant Acinetobacter infections has increased considerably. The goal of the present study was to specify Acinetobacter species isolated from laboratory mice and to test them for their antimicrobial susceptibility. During routine microbiological monitoring of laboratory mice, 12 Acinetobacter spp. were isolated. By means of 16S rRNA and rpoB gene sequencing, seven of the isolates were identified as Acinetobacter radioresistens, three isolates belonged to Acinetobacter genomospecies 14BJ, one isolate was classified as Acinetobacter pitii and one as Acinetobacter sp. ANC 4051. The distribution of the minimal inhibitory concentration (MIC) values was uniform for 21 of the 23 antimicrobial agents tested, whereas a broad MIC distribution was recorded for tulathromycin and streptomycin. The MIC values recorded were low for the majority of the antibiotics tested. Nevertheless, very high MIC values, which will probably render a therapeutic approach using these substances unsuccessful, were recorded for florfenicol, tiamulin, tilmicosin and cephalothin in most of the isolates. In conclusion, we document colonization of laboratory mice with different Acinetobacter species, displaying similar antibiotic susceptibility profiles, with possible implications in the Acinetobacter epidemiology as well as in the husbandry and experimentation of the colonized animals.
Acinetobacter species are regarded as a major infectious problem in human medicine, particularly in immunocompromised patients. 1 The Acinetobacter strains involved in nosocomial infections are frequently multidrug-resistant. 2 The extra-hospital epidemiology of human Acinetobacter infections is a subject of debate. 3 Acinetobacter species are ubiquitous within the environment and can be found in soil, water, on surfaces or in food products.4,5 Moreover, they seem to persist for long periods of time in the environment.6,7 During recent years, the prevalence of animal multidrug-resistant Acinetobacter infections has increased considerably. Acinetobacter isolates have been collected over time mainly from cats and dogs, but also from horses, donkeys, cattle, goats, birds and even from rabbits and guinea pigs.3,8–10 Although the presence of Acinetobacter species in laboratory rodents is known, to date there is only one study reporting the isolation and phenotypical identification of Acinetobacter strains using the Vitek-2 system. 11 Animal Acinetobacter isolates not only present concerns regarding infections of humans that are in contact with animal carriers, but can also produce serious animal diseases ranging from localized infections to septicemia.10,12 Concerns for animal welfare or pathology, as well as for experimentation and the husbandry of Acinetobacter spp. infected groups of animals are aspects that should not be neglected. The FELASA recommendations for health monitoring imply examination for infectious agents other than those listed in the recommendations, whenever considered relevant for a microbiological unit, thus granting flexibility in the design of the microbiological surveillance programs of a facility. 13 The goal of the present study was to specify Acinetobacter spp. isolates obtained from laboratory mice and to investigate them for their antimicrobial susceptibility.
Materials and Methods
Bacterial isolation
Identification of mice Acinetobacter species isolates.
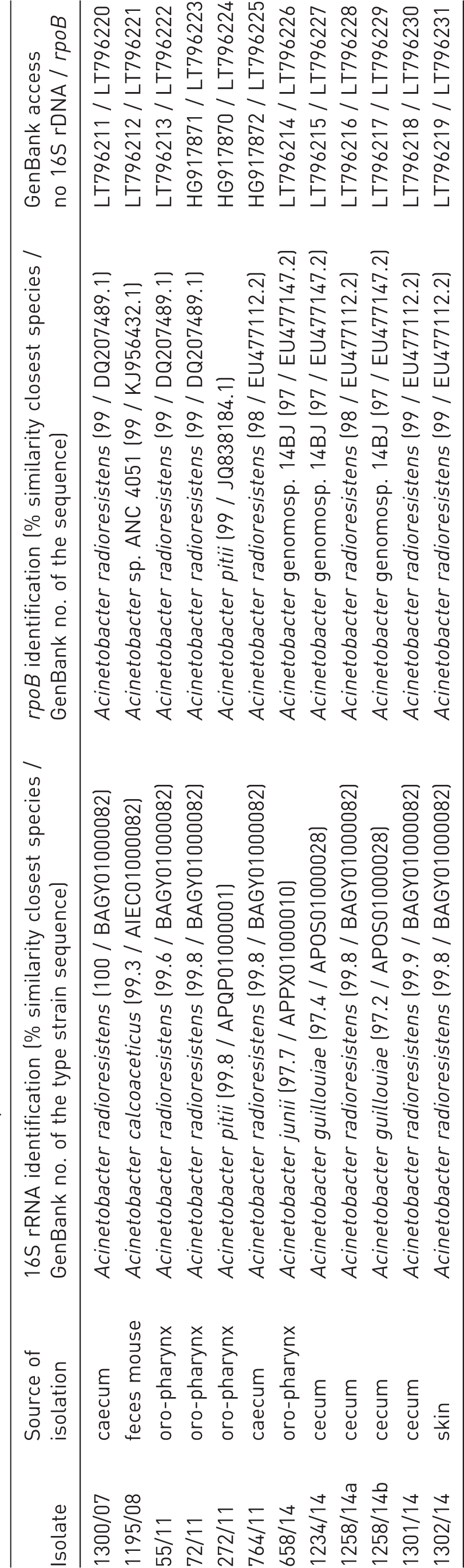
For health monitoring, the mice were euthanized by the CO2 method, in accordance with the directive 2010/63/EU, and cecum and/or feces and skin samples as well as swabs of the nasal cavities, oro-pharynx, and genital mucosa were cultured on Columbia blood and MacConkey agar plates (bioMérieux, Nürtingen, Germany) for approximately 48 h at 37℃ in aerobic and anaerobic conditions. This is to examine the samples for the presence of the bacteria listed in Table 3 of the FELASA recommendations that are able to grow on the aforementioned agar media. 13
Bacterial species identification
Single isolated colonies were sub-cultured and tested for purity and then subjected to phenotypic identification by colony morphology, ability to grow on special media, cell morphology after Gram staining, spore forming ability, respiratory type, motility, oxidase and catalase tests. The assignment to the genus Acinetobacter was based presumptively on the growth on MacConkey agar, Gram staining, a negative oxidase test, and the results of the API 20 NE (bioMérieux, Marcy l’Étoile, France) test. Further identification to species level was performed by partial sequencing of the 16S rRNA and rpoB genes. DNA for PCR amplification of the two genes was extracted from 24-h-old blood-agar-grown cultures as described previously. 15 Amplification of the 16S rDNA and analysis of the sequences was performed as described previously. 16 A 16S rDNA sequence similarity of ≥97% with that of an Acinetobacter species type strain was considered as sufficient for identification at genus level.17,18, and a prerequisite for further species identification based on rpoB gene sequencing, since 16S rRNA gene sequence variation are not sufficient to differentiate Acinetobacter species. 19 Amplification and analysis of the rpoB gene sequences were performed using a procedure described previously. 4 An rpoB sequence similarity of ≥95% to a reference strain was considered as sufficient for identification at species level. 4
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing of the 12 Acinetobacter isolates was performed by broth microdilution according to the human-specific Clinical & Laboratory Standards Institute (CLSI) document M100-S. 20 The decision to use the human-specific document for testing the Acinetobacter isolates from mice was taken because there is no testing method for Acinetobacter spp. available in the veterinary-specific documents VET06 21 and VET08 . 22 Moreover, neither mouse-specific nor veterinary-specific clinical breakpoints for Acinetobacter spp. are available.21,22 Therefore, the human-specific clinical breakpoints—when available—were used for categorization of the minimal inhibitory concentration (MIC) values as susceptible, intermediate, or resistant. 20 Custom-made microtiter plates (MCS Diagnostics, Swalmen, The Netherlands) were used to test the Acinetobacter isolates for their susceptibility to 23 antimicrobial agents/combinations (ampicillin, amoxicillin-clavulanic acid, ceftiofur, cefquinome, cephalothin, cefotaxime, cefoperazone, imipenem, tetracycline, doxycycline, gentamicin, neomycin, streptomycin, nalixic acid, ciprofloxacin, enrofloxacin, marbofloxacin, florfenicol, tilmicosin, tulathromycin tiamulin, colistin, trimethoprim-sulfamethoxazole). The test panel was based on the microtiter plates used for Gram-negative bacteria in the German National Resistance Monitoring program of veterinary pathogens GERM-Vet (https://www.bvl.bund.de/EN/09_Laboratories/01_Tasks/03_national_resistance_monitoring/resistance_monitoring_node.html). For antimicrobial susceptibility testing, fresh overnight cultures on blood agar plates were used, and a density of McFarland 0.5 was adjusted in 0.85 % saline. The inoculum was prepared by adding 5 µL of the bacterial suspension per 1 mL of cation-adjusted Mueller-Hinton broth. From this suspension, 50 µL was then pipetted into each well of the microtiter plate and the results were read after 20–24 h of incubation at 35℃ ± 2℃. 20 The Escherichia coli reference strain ATCC® 25922 served as quality control.
Results
Identification of Acinetobacter isolates
Direct culture of animal samples on agar plates yielded 12 isolates, which were presumptively identified as belonging to the genus Acinetobacter after phenotypic analysis of colony morphology, Gram staining, spore forming ability, respiratory type, motility, oxidase, catalase and API 20 NE tests (Table 1). Overall, Acinetobacter spp. was found in approximately 0.5% of the mice examined per year. They were not detectable in all microbiological units of the facility by using routine health monitoring procedures described above. Nevertheless, the isolates described here were found in both breeding and experimental barriers. The strains, isolated mostly from cecum or the oropharynx, were subjected to 16S rDNA and rpoB gene sequencing. Nine of the isolates possessed a 16S rDNA sequence similarity ≥ 99% to that of a bacterial type strain deposited in GenBank, whereas the remaining three strains showed a sequence similarity ≥ 97% but < 99% to that of a prototype strain (Table 1). The similarity level of 16S rRNA genes allowed the classification of all isolates into the genus Acinetobacter. The further rpoB gene analysis permitted identification at species level for all isolates. Overall, seven isolates belonged to the species Acinetobacter radioresistens, three isolates were Acinetobacter genomosp. 14BJ. The remaining two isolates were classified as Acinetobacter pitii and Acinetobacter sp. ANC 4051, respectively (Table 1).
Antimicrobial susceptibility of the Acinetobacter isolates
Antimicrobial susceptibility of mice Acinetobacter species isolates.
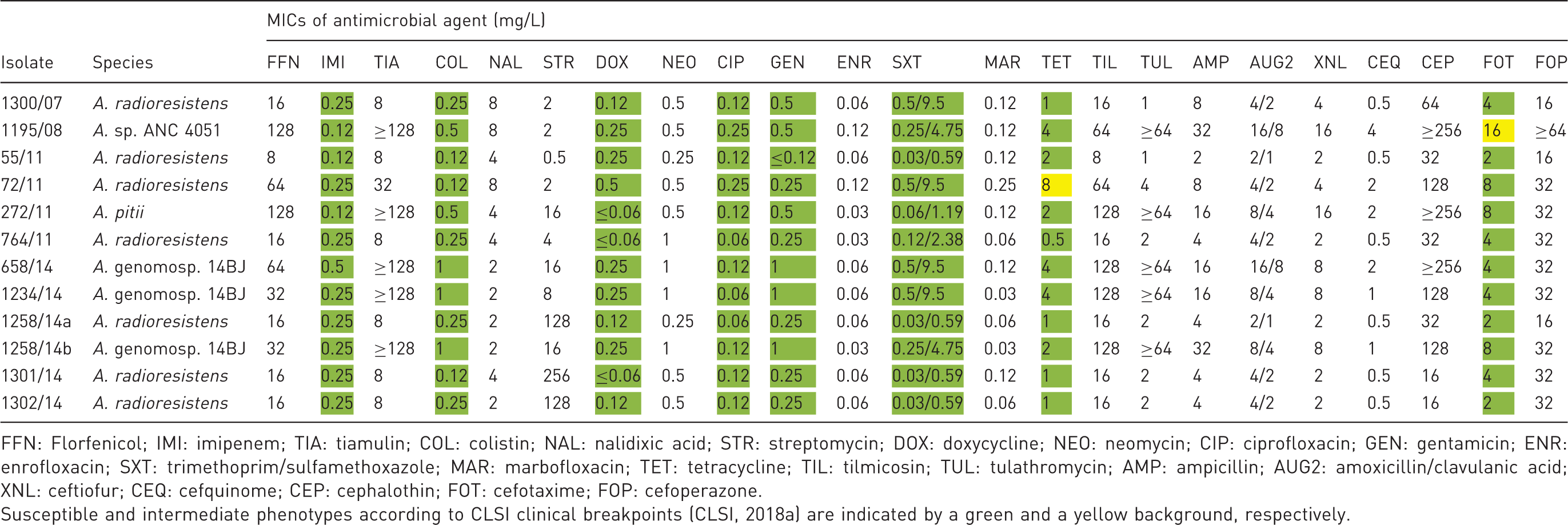
FFN: Florfenicol; IMI: imipenem; TIA: tiamulin; COL: colistin; NAL: nalidixic acid; STR: streptomycin; DOX: doxycycline; NEO: neomycin; CIP: ciprofloxacin; GEN: gentamicin; ENR: enrofloxacin; SXT: trimethoprim/sulfamethoxazole; MAR: marbofloxacin; TET: tetracycline; TIL: tilmicosin; TUL: tulathromycin; AMP: ampicillin; AUG2: amoxicillin/clavulanic acid; XNL: ceftiofur; CEQ: cefquinome; CEP: cephalothin; FOT: cefotaxime; FOP: cefoperazone.
Susceptible and intermediate phenotypes according to CLSI clinical breakpoints (CLSI, 2018a) are indicated by a green and a yellow background, respectively.
For eight of the tested antimicrobial agents, namely cefotaxime, imipenem, colistin, tetracycline, doxycycline, ciprofloxacin, gentamicin, and trimethoprim-sulfamethoxazole, the CLSI-approved human-specific clinical breakpoints were applied. 20 However, it should be noted that these breakpoints might not be appropriate to predict the clinical outcome for the treatment of laboratory mice, due to possible differences in the pharmacokinetic and pharmacodynamic parameters between humans and mice. Nevertheless, they allow a prediction in case of an infection of humans due to transmission of these Acinetobacter strains from the laboratory mice. Using these human clinical breakpoints, all isolates were susceptible to imipenem, colistin, doxycycline, gentamicin, trimethoprim/sulfamethoxazole and ciprofloxacin. 20 The low MIC values (0.03–0.25 mg/L) seen for enrofloxacin and marbofloxacin are in accordance with the susceptibility to ciprofloxacin, since resistance to fluoroquinolones is commonly mediated via the same mechanism. 23 The MICs for nalidixic acid ranged between 2 mg/L and 8 mg/L.
In comparison to doxycycline, to which all isolates were classified as susceptible, one isolate, 72/11, was classified as intermediate to tetracycline. 20 Only strain 1195/08 showed intermediate susceptibility to cefotaxime, while the remaining isolates were classified as susceptible. 20 However, the MIC values for other β-lactams such as cephalothin (8– ≥ 256 mg/L) and cefoperazone (16– ≥ 64 mg/L) were comparatively high. The MICs for neomycin ranged from 0.25 mg/L to 1 mg/L and were relatively low. However, the MIC values for other substances, such as florfenicol (8–128 mg/L), tiamulin (8– ≥ 128 mg/L), and tilmicosin (8–128 mg/L) were comparatively high for most of the isolates tested, suggesting that a therapy using these antimicrobial agents might be unsuccessful.
Discussion
In the present investigation, we analyzed the species diversity of Acinetobacter spp. isolates obtained during the routine health monitoring program of laboratory mice from our institution. The prevalence of about 0.5% Acinetobacter spp. assumed in our study might be underestimated, since the methods used here did not specifically target the isolation of Acinetobacter spp., but rather all FELASA-listed bacteria. The presence of Acinetobacter spp. in laboratory rodents is known among people performing microbiological monitoring in rodent facilities. Despite this fact, to date there is only one study reporting the isolation of Acinetobacter strains from laboratory mice and rats. 11 In that study, all strains isolated were classified as A. baumannii by phenotypical methods. 11 However, phenotypical species identification among Acinetobacter isolates remains challenging. 24 The genotypic identification procedure used in our study allowed an exact identification to species level. Most of the species found in our study belonged to A. radioresistens or Acinetobacter genomosp. 14BJ, but also isolates of A. pitii or Acinetobacter sp. ANC 4051, belonging to the A. calcoaceticus-baumannii complex, which constitutes the most common causal agents of Acinetobacter infections in hospitals worldwide, were documented (Table 1). The finding of these bacterial species points towards a potential risk of transfer of these bacteria between animals and humans. The Acinteobacter strains were isolated from several parts of the bodies of the animals, including skin, upper respiratory tract, cecum and feces. We have recently shown that survival of A. radioresistens on animal bedding in a mouse room easily exceeded 6 weeks, 6 suggesting that this group of bacteria is capable of persisting over long periods of time in the mouse room environment, representing a risk of infection for humans and animals. This finding is in accordance with the findings that Acinetobacter can survive several weeks on abiotic surfaces under dry conditions due to biofilm formation and the development of dormant cells. 24 Nevertheless, all materials used in the routine husbandry procedures in the mouse room, such as cage, bedding, feed, water, are sterilized before use, suggesting that the Acinetobacter isolates found were more probably transferred from the mouse into the environment rather than being picked up from environmental sources.
A further valuable finding is the antimicrobial susceptibility data of the Acinetobacter isolates (Table 2). By using the human-specific clinical breakpoints for Acinetobacter spp., none of the isolates was classified as resistant to imipenem, colistin, doxycycline, ciprofloxacin, gentamicin, trimethoprim/sulfamethoxazole, tetracycline and cefotaxime. 20 Moreover, only single isolates were classified as intermediate to tetracycline and cefotaxime, respectively. For 15 out of 23 (70%) antimicrobial agents used, no CLSI-approved clinical breakpoints were available. For antimicrobial agents such as neomycin, enrofloxacin, and marbofloxacin, the MIC values recorded were comparatively low (≤1 mg/L), whereas for other substances such as florfenicol, tiamulin, tilmicosin, cephalothin, and cefoperazone, they were comparatively high (up to ≥ 64 mg/L to ≥ 256 mg/L) (Table 2). Such high antimicrobial concentrations would be hard to achieved in vivo and, thus, render a therapeutic approach using these substances most likely unsuccessful. It should be noted that Acinetobacter are intrinsically resistant to certain antimicrobial agents, and the CLSI document M100 20 stated that Acinetobacter baumannii and the Acinetobacter calcoaceticus complex are intrinsically resistant to penicillin, ampicillin, amoxicillin, amoxicillin-clavulanic acid, first generation cephalosporins (cephalothin, cefazolin), second generation cephalosporins (cefuroxime), cephamycins (cefoxitin, cefotetan), clindamycin, daptomycin, fusidic acid, glycopeptides (vancomycin, teicoplanin), linezolid, macrolides (erythromycin, azithromycin, clarithromycin), quinupristin-dalfopristin, rifampicin, ertapenem, trimethoprim, chloramphenicol and fosfomycin. Due to the known intrinsic resistance of Gram-negative bacteria to some of these antimicrobial agents, some of them, such as erythromycin, clindamycin, vancomycin, linezolid and quinupristin-dalfopristin, were not included in the test panel used. The comparably high MICs observed for ampicillin and amoxicillin-clavulanic acid, as well as the high cephalothin MICs can be explained by intrinsic resistance. 20 In addition, the finding of high florfenicol MICs, pointing towards a possible resistance, are in accordance with the intrinsic resistance reported for the related substance chloramphenicol. 20 Multidrug-resistant Acinetobacter isolates are common among hospital isolates. 4 Resistance to antimicrobial agents of Acinetobacter animal isolates is widespread, covering a broad resistance spectrum, including gentamicin, tetracycline, and enrofloxacin, among others. 10 Laboratory animals are not usually exposed to antibiotic treatments, which would trigger the selection of resistant strains. Nevertheless, therapy with antimicrobial agents might be indicated in order to save valuable unique mice strains until rederivation by other methods is possible. Goo and co-workers tested the resistance of several rodent Acinetobacter isolates to a restricted group of antimicrobial agents and found them resistant to cefazolin, cefoxitin and nitrofurantoin, but susceptible to other commonly used antibiotics. 11 However, the use of the human-specific clinical breakpoints also allows a prediction of a therapeutic success in the case of human infection with such a mouse-associated Acinetobacter isolate, which might occur after transmission to an immunocompromized person. The finding that these strains were comparably susceptible to most of the antimicrobial agents for which clinical breakpoints are available seems promising, since resistant Acinetobacter isolates are a global problem in human medicine. 24 Especially resistance to last resort antimicrobial agents such as carbapenems is a major threat for the treatment of nosocomial infections.24,25.
In conclusion, we document that the laboratory mouse can be involved in the extra-hospital epidemiology of Acinetobacter infections. Moreover, the colonization or infection of laboratory rodents with this group of bacteria can be problematic, especially for immunocompromized laboratory animal breeds.
Footnotes
Acknowledgments
We gratefully acknowledge Vivian Hensel, Isabel Schäfer, and Manuela Stockhausen for excellent technical assistance.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
