Abstract
Pigs are used with increased frequency to model different kinds of orthopedic surgical conditions. In order to show the full potential of porcine models in orthopedic research, it is therefore required to examine the expression of bone regulatory genes in pigs affected by orthopedic surgery and compare it to the expression in humans and mice as mice, are one of the most applied animal species in orthopedics today. In the present study, the local molecular response to drilling of a tibial implant cavity, and the subsequent insertion of a steel implant was examined in a porcine model. Pigs were euthanized five days after drilling of the bone. The molecular response of 73 different genes was analyzed using a high-throughput quantitative polymerase chain reaction platform and compared to histopathology. Histologically, it was found that bone remodeling was initiated on day 5 after surgery and was associated with upregulation of several genes involved in bone degradation and formation (CTSK, ACP5, IBSP, RANK, RANKL and COL1A1). Interleukin-6 and several acute-phase proteins (C3, SAA and ITIH4) were significantly upregulated, indicating their importance in the initial process of healing and osseointegration. All tested bone morphogenic proteins (BMP2, -4 and -7) including their inhibitor noggin were also significantly upregulated. Surprisingly, vascular endothelial growth factor A was not found to be regulated five days after surgery while several other vascular growth factors (ANGPT1, ANGPT2 and PTN) were upregulated. The pig was found to be a useful model for elucidation of bone regulatory genes in humans.
Introduction
Anatomically and physiologically, pigs are similar to humans 1 and have many advantages as a model for orthopedic conditions when compared to rodents. 2 The size of pigs makes them suitable for using orthopedic devices and procedures similar to those applied in humans, and in contrast to mice, it is possible to sample large quantities of blood and tissue for analysis. Most important, the immune system and the bone matrix composition of humans are more similar to pigs than to mice.3–7 However, there is no argument for a bone genetic superiority of a porcine orthopedic model over a mouse model, as the local expression of bone regulatory genes is non-characterized in pigs. Increasingly more pigs are used to model different kinds of orthopedic surgical conditions, and recently several studies have been published highlighting the advantages of porcine models of osseointegration of orthopedic implants.8,9 Therefore, it is necessary to examine the expression of bone regulatory genes in pigs after orthopedic surgery and compare them to the expression in humans and mice in order to evaluate the full potential of pigs as experimental animal models in orthopedic research.
Reports on the local molecular response toward bone damage including orthopedic implant insertion originate mainly from fractures studied in murine models. 10 The best way to obtain insight into the human physiology is obviously to examine humans; however, for ethical reasons this is not always feasible. The systemic and local response in patients with impaired fracture healing have been examined in a few studies using blood samples or specimens of bone tissue, respectively.11–13 However, in order to fully understand the healing process to bone damage, the local molecular and morphological response is optimally examined in discriminative animal models. Both the local histology and gene expression after bone damage are well characterized in rodents.14–17 However, molecular characterization in large animal models like pigs and in humans8,18,19 are limited even though orthopedic conditions like joint replacement and osteosynthesis are standard procedures.20,21 A major issue related to orthopedic surgery is osteonecrosis caused by trauma and high temperatures during drilling in bones, 22 as well as the response to a foreign body, 23 which depends on the material and surface characteristics.8,9,18
Understanding the local impact of bone damage both morphologically and at the molecular level is required in order to improve the procedures related to insertion of orthopedic devices and to prevent implant-associated osteomyelitis. A detailed examination of the molecular changes and the cellular environment adjacent to bone damage might provide a platform for identifying novel agents with therapeutic potential, such as bone morphogenic proteins (BMPs), which already have been used clinically to reduce healing time in tibial fractures. 24
The aim of the present study was to analyze the genetic expression of normal bone tissue and bone tissue affected by drilling of an implant cavity with insertion of a steel implant in a porcine model. Based on the obtained results, the study aimed to describe the potential of porcine models in orthopedics on a bone molecular level. Overall, we demonstrate that the expression of bone regulatory genes in pigs affected by orthopedic surgery is comparable to results obtained in mice and humans.
Animals
Six 3-month-old (∼30 kg) female Danish Landrace pigs from a specific pathogen-free herd were used. At arrival, the animals were allowed to acclimatize for one week before entering the trial. The animals were housed in the same pen (3 × 3 m) and fed a commercial pig diet ad libitum and had free access to tap water. Control pigs previously used to characterize and show reproducibility of a porcine implant-associated osteomyelitis model were used in the present study. 25 Therefore, with reference to reduction, refinement and replacement (the 3Rs), the present study fulfills the expectation of reduction, as the material is collected from previously used pigs, instead of using new pigs to generate similar results.
Materials and methods
Experimental design
The surgical procedure has recently been described in detail 25 and was approved by the Danish Animal Experiments Inspectorate (license No. 2013/15-2934-00946) according to the Danish proclamation of law on animal experimentation (LBK No. 1306,23/11/2007). All animals were cared for in accordance with the principles laid down by the European Union Directive 2010/63/EU for Protection of Animals used for Scientific Purposes. Briefly, a 4 × 20 mm implant cavity was drilled with a steel k-wire in the trabecular bone at the proximal end of the right tibia approximately 10 mm distal to the growth plate in anesthetized pigs. Saline (10 µl) was injected into the cavity, and a stainless steel implant (2 × 15 mm) was inserted. Five days after the operation, the pigs were euthanized by an overdose of 20% pentobarbital given intravenously and the implants were removed. Hereafter, the right tibial bones were sagittal-sectioned through the implant cavity. One half was used for histology and placed in formalin for fixation, while trabecular bone tissue surrounding the implant cavity from the other half was placed in RNAlater for molecular analysis. From each animal, the left tibial bone was likewise split in two and placed in formalin and RNAlater, respectively, as controls.
Anesthetic and analgesic protocol
Premedication and induction of anesthesia was performed in the stable with intramuscular injections (1 ml/10 kg body weight (BW)) of a solution containing a mixture of 125 mg zolezepam, 125 mg tiletamine, 6.25 ml xylazine (20 mg/ml), 1.25 ml ketamine (100 mg/ml) and 2 ml butorphanol (10 mg/ml). Two catheters were then inserted into lateral ear veins. One catheter was used for surgical anesthesia with propofol (10 mg/ml) 1 ml/kg BW, and one catheter was used for intraoperative analgesia with Fentanyl (0.5 mg/h). At the end of surgery, animals received an intramuscular injection 0.1 mg/kg BW of Buprenorphine (0.3 mg/ml) resulting in postoperative analgesia for the following six hours. Thereafter, and throughout the experiment, animals received daily oral analgesic treatment with Meloxicam 0.3 mg/kg BW. The animals recovered from anesthesia under observation in individual pens. Throughout the postoperative period, the pigs were followed up daily including inspection of surgical wound and evaluation of gait. Impaired ability to walk and stand, and anorexia were set as human endpoints.
Histology
After one week of fixation in formalin the bone tissue was decalcified in formic acid for four weeks. Blocks of bone tissue, representing the area surrounding the implant cavity and normal trabecular bone tissue, were cut from the right and left tibial bones, respectively. All blocks of bone tissue were then passed through graded concentrations of alcohol and embedded in paraffin wax. Finally, tissue sections (4–5 µm) were stained with hematoxylin and eosin in order to examine the morphology and with Safranin O to study chondroid tissue.
Assay design
A literature search was conducted to find genes likely to be involved in bone healing and osseointegration. Selected genes for the expression analysis can be divided into three main groups: (1) markers of bone remodeling, (2) inflammatory genes and (3) growth factors.
Primer3 software (http://bioinfo.ut.ee/primer3-0.4.0/) was used to design primers with a mean temperature (Tm) between 59℃ and 61℃ and amplicon lengths from 70 to 150 bp. Primers were designed to span introns and to target most or all splice variants of the gene of interest. Sequences, amplicon lengths, polymerase chain reaction (PCR) efficiencies, fold changes and p values for all primers and primer assays can be found in the supplementary data (Table S1). Nucleotide blast searches were performed to test the specificity of all primers (https://blast.ncbi.nlm.nih.gov/Blast.cgi) before synthesis (Sigma-Aldrich).
RNA extraction
Tissue samples preserved in RNAlater were transferred to 1 ml Qiazol (Qiagen) in M-tubes (Miltenyi Biotec) and homogenized using a gentleMACS Dissociator (Miltenyi Biotec). RNA was isolated using an miRNeasy Mini kit (Qiagen) and treated with RNase-Free DNase set (Qiagen) according to the manufacturer’s specifications. The concentration and purity of the extracted RNA was determined with a NanoDrop ND-1000 ultraviolet spectrophotometer (Thermo Fisher Scientific). The purity was evaluated based on the A260/280 and A260/230 ratios and accepted for further analysis. The RNA concentration ranged from 218 ng/µl to 685 ng/µl with a mean of 433 ng/µl. The quality of the RNA was determined using Agilent RNA 6000 Nano chips, Agilent RNA 6000 Nano reagents and Agilent 2100 Bioanalyzer (Agilent Technologies). An RNA integrity number (RIN) was assigned to each sample. Mean RIN was 5.9 ± 1.4. The extracted RNA was stored at –80℃.
Reverse transcription, pre-amplification and exonuclease treatment
Complementary DNA (cDNA) was synthesized from 500 ng extracted total RNA using the QuantiTect Reverse Transcription Kit (Qiagen) according to the manufacturer’s specifications but with a 25% reduction in the volume of buffers, primer mix and reverse transcriptase. For genomic DNA elimination, the RNA was mixed with 1.5 µl Wipeout buffer and RNase-free water to a final volume of 14 µl and incubated on a TProfessional TRIO thermocycler (Biometra). The cDNA synthesis was performed by adding 6 µl QuantiTect Reverse Transcription master mix (0.75 µl Reverse Transcriptase, 0.75 µl RT primer mix, 3 µl RT buffer and 1.5 µl RNase-free water) to the DNase-treated RNA, reaching a final reaction volume of 20 µl. Two technical replicates of cDNA were made for each RNA sample, except for three samples with an RIN under 6 for which three replicates were completed. Non-reverse transcriptase controls were also made and included in the following pre-amplification and quantitative polymerase chain reaction (qPCR). The cDNA was diluted 1:10 in low-ethylenediaminetetraacetic acid (EDTA) TE-buffer (Panreac AppliChem) before pre-amplification. One µl TaqMan PreAmp Master Mix (Applied Biosystems), 2.5 µl 200 mM mix of all primers used subsequently for qPCR, 4 µl low-EDTA TE-buffer (Panreac AppliChem) and 2.5 µl diluted cDNA was mixed and incubated at 95℃ for 10 minutes followed by 19 or 21 cycles of 95℃ for 10 seconds and 60℃ for four minutes. The pre-amplified cDNA was treated with 16 U Exonuclease I (New England Biolabs) for 30 minutes at 37℃. The product was stored at –20℃. Before qPCR the cDNA was diluted 1:10 in low-EDTA TE-buffer (Panreac AppliChem).
High-throughput qPCR
qPCR was performed using 96.96 Dynamic array integrated fluid circuit (IFC) chips (Fluidigm) in the BioMark HD System (Fluidigm) combining 96 primer sets with 96 samples. Each primer mix contained 3 µl 2X Assay Loading Reagent (Fluidigm) and 3 µl of 20 µM forward and reverse primers suspended in low-EDTA TE-buffer (Panreac AppliChem). The sample mixes were prepared using 3 µl 2X TaqMan Gene Expression Master Mix (Applied Biosystems), 0.3 µl 20X DNA Binding Dye Sample Loading reagent (Fluidigm), 0.3 µl 20X Evagreen (Panreac AppliChem), 0.9 µl low-EDTA TE-buffer (Panreac AppliChem) and 1.5 µl pre-amplified diluted (1:10) cDNA. Before loading the samples and primers, the chip was primed in an HX IFC controller (Fluidigm). After priming 5 µl of each sample mix and primer mix were distributed into the appropriate compartments and loaded into the chip in the HX IFC controller. Thereafter the chip was inserted in the BioMark real-time PCR instrument (Fluidigm) and the following program was used: two minutes at 50℃ and 10 minutes at 95℃, next 35 cycles with 15 seconds at 95℃ and one minute at 60℃. After qPCR, melting curves were generated from 60℃ to 90℃. Non-template controls and non-reverse transcriptase controls were included to indicate problems with contamination, non-specific amplification or genomic DNA. Standard curves constructed from three separate dilution series of pooled cDNA of all samples were used to determine the efficiency of each primer assay.
Data processing and statistics
Raw data were inspected using Fluidigm Real-Time PCR Analysis software (v. 4.1.3). GeneEx5 (v. 5.4.4.119) (MultiD) was used for pre-processing, normalization and relative quantification. Data were corrected for PCR efficiency for each primer assay and thereafter normalized with four reference genes (β-2-microglobin, glyceraldehyde-3-phosphate dehydrogenase, peptidylprolyl isomerase A and TATA-box binding protein). Out of six tested reference genes, these four were found to have the most stable expression using both GeNorm 26 and NormFinder. 27 After normalization, low-quality RNA samples were carefully inspected to ensure valid data. The cDNA replicates were averaged after normalization. Inconsistent primer assays (defined as >17% of cDNA replicates varying >1.5 Cq) were excluded or validated by a second qPCR run. For each primer assay the Cq values were converted to relative quantities. For visualization, the data were scaled to give the healthy bone a mean relative expression of 1. Data were log2-transformed before paired t tests, which were carried out using Excel (2010) (Microsoft). Genes were considered significantly regulated if at least a two-fold difference was found between the means of healthy and operated-on bones and if the p value was below 0.05.
Results
All animals were healthy prior to surgery. After recovery from anesthesia, all pigs were lame on the inoculated leg. However, they were able to use the leg and walked freely around in the pens. During the first day the lameness diminished and disappeared two to three days after surgery. All pigs ate and drank normally throughout the experiment. 25
Gene expression analysis
After data processing and removal of duplicate primers, valid expression data from 73 genes were obtained. Of these, 36 had a fold-change difference between the mean of the drilled and the healthy bone of at least 2 and were significantly different. Many of the molecular differences could be linked to morphological changes. Owing to limited space, only selected genes of importance are described and discussed in this paper, but a full list is available in the supplementary data (Table S1).
Bone remodeling
Toward the implant cavity an interrupted thin layer of elongated fibroblasts and leukocytes were seen lining compressed and osteonecrotic trabecular bone tissue showing a faint staining with empty osteocytic lacuna (Figure 1(a)). This layer was sporadically intermingled with single neutrophils and osteoclasts. Ruffle-bordered osteoclasts were also seen (Figure 1(b)) and their activity was confirmed by upregulation of the genes encoding cathepsin K (CTSK) and tartrate resistant acid phosphatase 5, also known as TRAP (ACP5) (Figure 2(a)), which are both central for bone resorption.
Necrotic trabecular bone and newly formed osteoid adjacent to a drilled implant cavity five days after surgery in a porcine model. Hematoxylin and eosin-stained (a) necrotic bone (NB) with trabeculae depleted of osteocytes leaving the lacunae empty. Bar 80 µm. (b) Osteoclast (arrow), resorbing trabecular bone tissue. Bar 350 µm. (c) Mineralized bone lined by layers of stimulated osteoblasts with a cuboidal shape (arrow). A mixed population of inflammatory cells is present around the bone structure. Bar 80 µm. (d) New mineralized osteoid (O) produced by surrounding cubical osteoblasts (arrows). Bar 80 µm. Relative expression (RQ) of genes related to (a) bone remodeling and (b) regulation of osteoclast activity in a porcine model of implant insertion five days after surgery. Dots represent a single sample and bars the mean for the group (n = 6); left unharmed tibia (•/solid bar) and the right operated-on bone (○/dashed bar). *p value < 0.05 in a paired t test.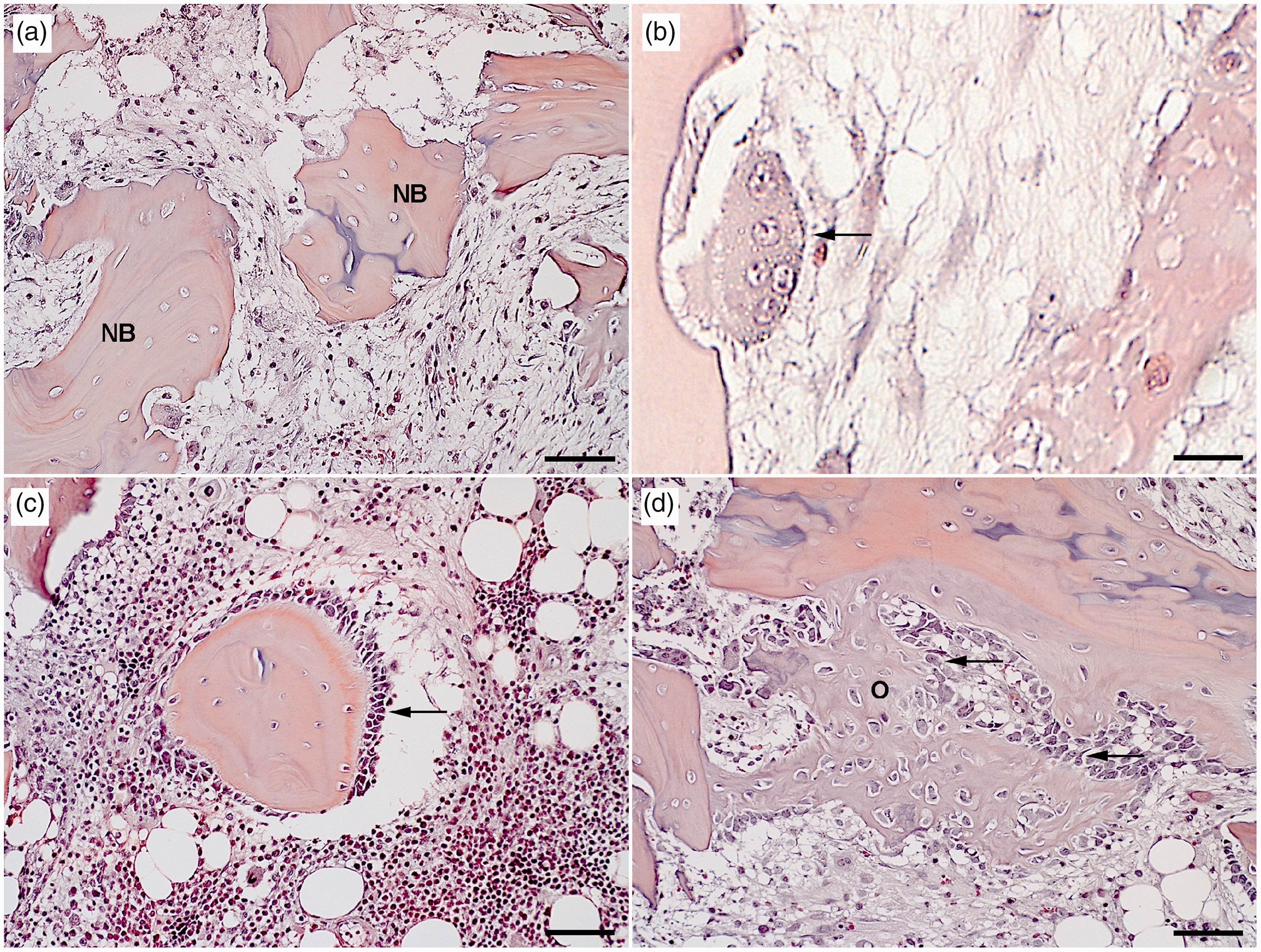
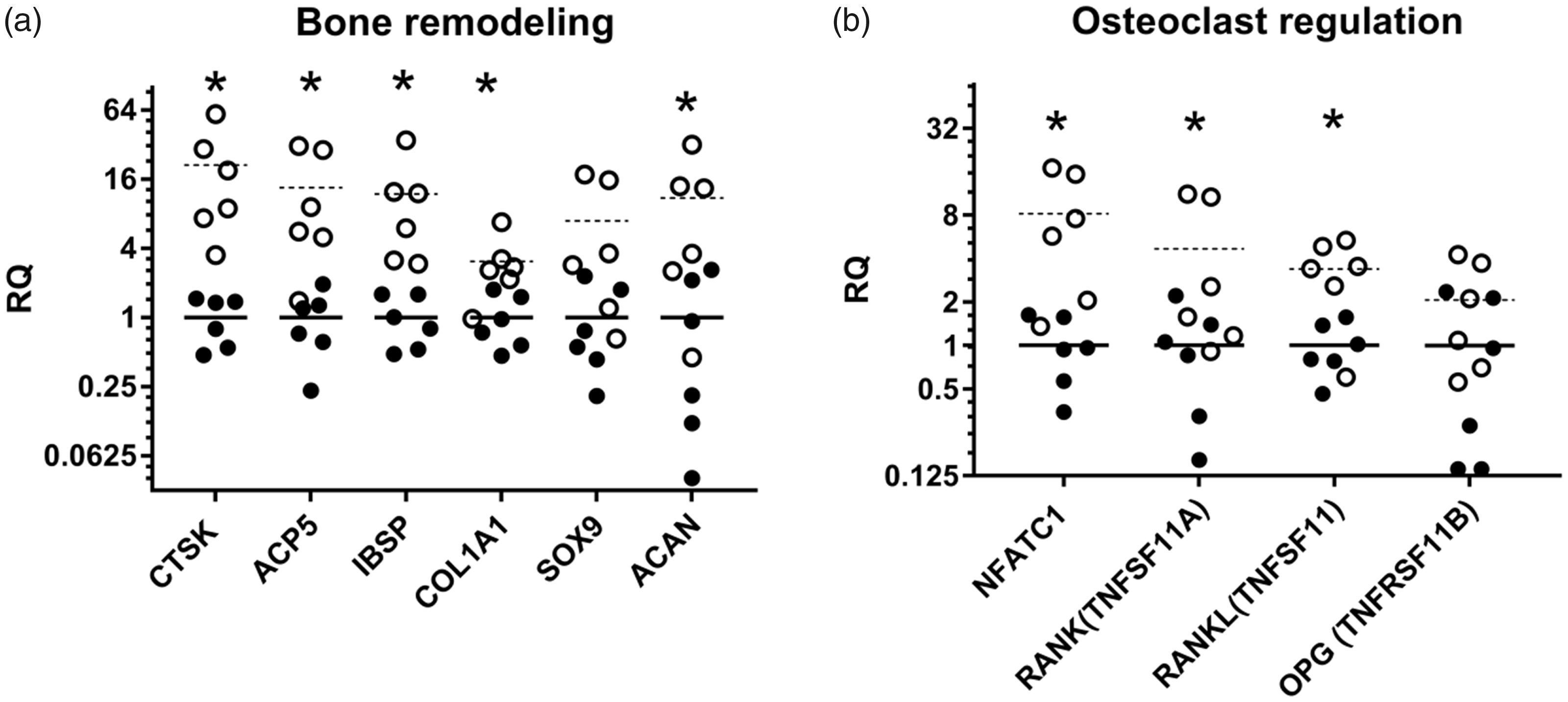
Receptor activator nuclear factor kappa B (RANK, encoded by TNFRSF11A) and the transcription factor nuclear factor of activated T cells-1 (NFATc1), both essential for osteoclast activation, 28 were both significantly upregulated. The same applied to receptor activator nuclear factor kappa B ligand (RANKL, encoded by TNFSF11), while there was no significant change in the expression of the decoy receptor osteoprotegrin (OPG, encoded by TNFRSF11B) (Figure 2(b)).
In the periphery of the tissue surrounding the implant cavity, edema and proliferation of fibroblast and osteoblast blended into the adjacent normal bone tissue and bone marrow. The production of new osteoid by proliferating cubic osteoblasts was also observed (Figure 1(c) and (d)), indicating intramembranous osteogenesis. New bone formation was confirmed by molecular analysis of the bone matrix proteins bone sialoprotein (IBSP) and collagen type 1 alpha 1 (COL1A1), which were both significantly upregulated (Figure 2(a)).
No cartilage formation could be determined by histology but aggrecan (ACAN), a matrix protein in cartilaginous tissue, showed a significant almost 11-fold upregulation and SOX9, an important transcription factor for chondrocytes, showed a borderline significant seven-fold upregulation with a p value of 0.059 (Figure 2(a)).
Inflammation
Histologically, infiltrating mononuclear cells were seen (Figure 1(c)) to aggregate in the damaged bone tissue. No regulation of interleukin-1 (IL-1) (Figure 3) or tumor necrosis factor (TNF) (below detectable levels (data not shown)) was found, while interleukin-6 (IL-6) was highly and significantly upregulated (more than 40-fold) (Figure 3). Likewise several acute-phase proteins (APPs) were found to be upregulated after surgery. The complement component 3 (C3), which has a central role in activating all three complement cascades,
29
was found to be significantly upregulated (Figure 3). The positive APPs serum amyloid A (SAA) and inter-α-trypsin inhibitor H4 (ITIH4, also known as pig major acute-phase protein (pigMAP)), were also found to be highly upregulated after implant insertion (Figure 3).
Relative expression (RQ) of inflammatory genes in a porcine model of implant insertion five days after surgery. Dots represent a single sample and bars the mean for the group (n = 6); left unharmed tibia (•/solid bar) and the right operated-on bone (○/dashed bar). *p value < 0.05 in a paired t test.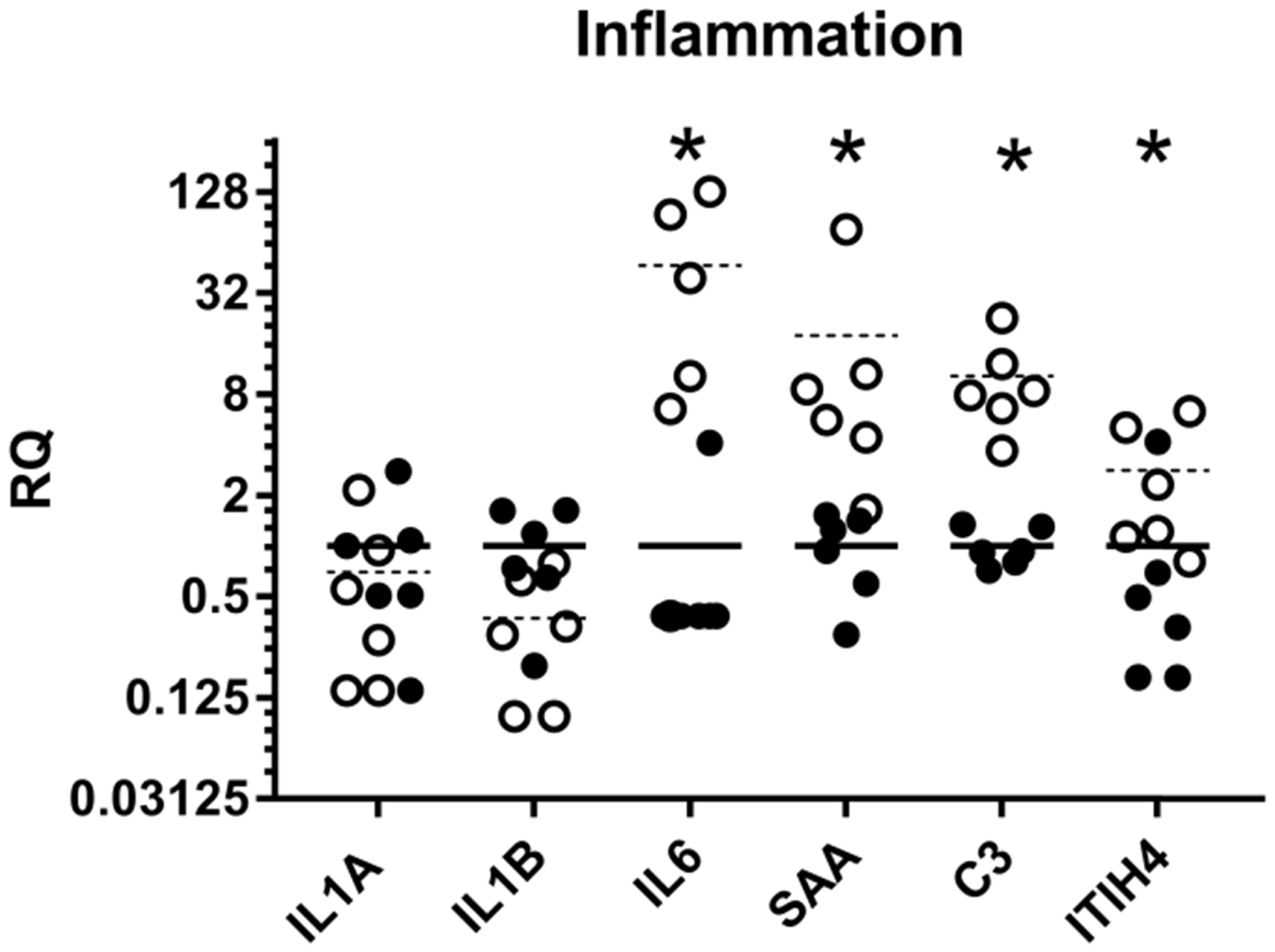
Growth factors
All the tested members of the transforming growth factor (TGF)-β superfamily were significantly upregulated after surgery (Figure 4(a)). With a four-fold upregulation, TGFB1 had the highest fold change, while bone morphogenic protein (BMP)2, -4 -7 and their inhibitor noggin (encoded by NOG) showed two- to three-fold upregulations.
Relative expression (RQ) of (a) bone morphogenic proteins and (b) vascular growth factors in a porcine model of implant insertion five days after surgery. Dots represent a single sample and bars the mean for the group (n = 6); left unharmed tibia (•/solid bar) and the right operated-on bone (○/dashed bar). *p value < 0.05 in a paired t test.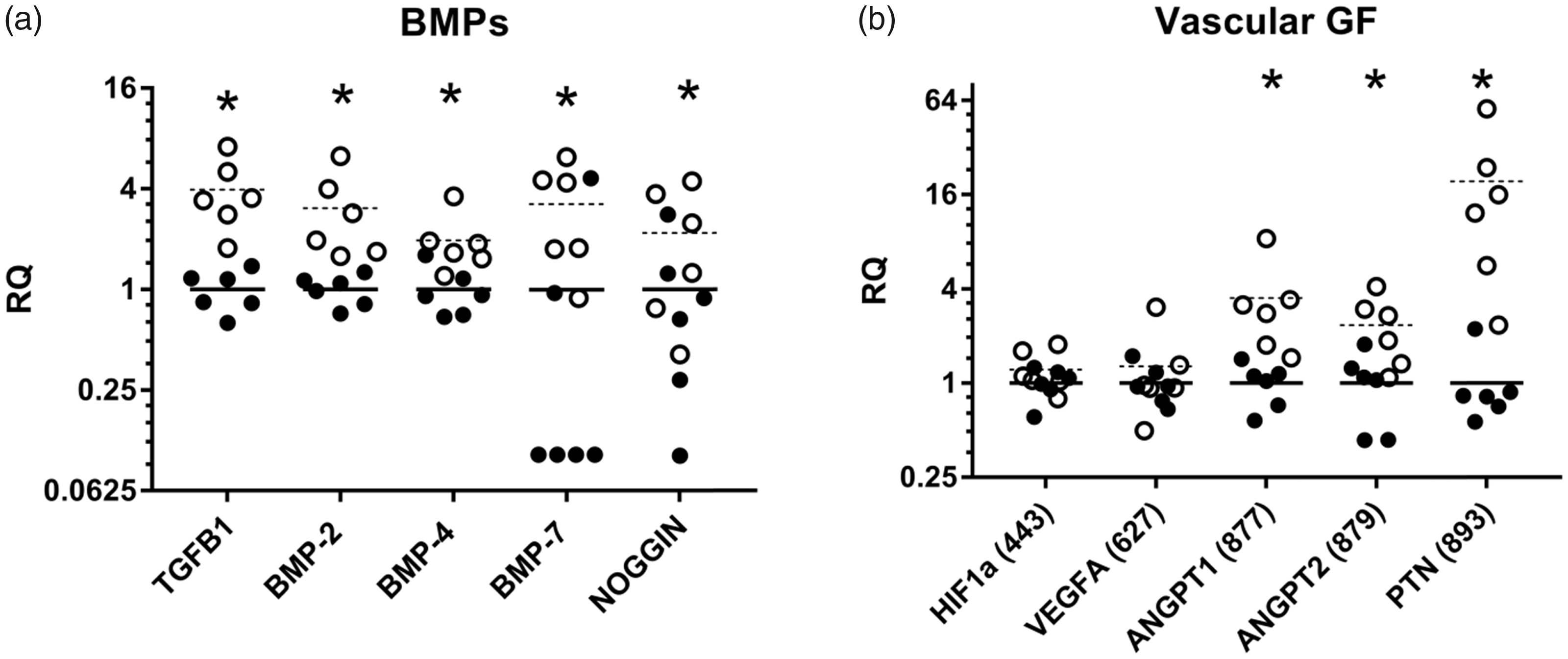
Vascular endothelial factor A (VEGFA) and its transcription factor hypoxia inducible factor-1α (HIF1A) were not regulated in the present study (Figure 4(b)), while the vascular growth factors angiopoietin-1 (ANGPT1), angiopoietin-2 (ANGPT2) and pleiotrophin (PTN also known as osteoblast-specific factor 1) were all significantly upregulated three-, two- and 19-fold, respectively (Figure 4(b)).
The tested fibroblast growth factors and insulin-like growth factors, also considered important in bone repair, 30 did not show any regulation five days after surgery. Neither did platelet-derived growth factor (PDGF)-B but PDGF-A was upregulated (Table S1).
Discussion
Bone remodeling
The upregulation of genes involved both in bone formation (IBSP and COL1A1) and bone degradation (CTSK and ACP5) confirmed that both osteoclasts and osteoblasts were actively degrading and regenerating bone tissue, respectively, already five days after surgery (Figure 2(a)).
Osteoclast activity and therefore bone resorption is influenced by the ratio between RANKL and its decoy receptor OPG. 28 Both molecules are secreted by osteoblasts and have also been shown to play a role in human fracture healing. 12 In the present study, RANKL was upregulated while no significant change in OPG expression was seen (Figure 2(b)), indicating that the RANKL/OPG ratio was increased. This is in agreement with the observed increased osteoclast activity and an increase in the ratio on day 7 in a murine fracture model. 16
Interestingly, SOX9 and ACAN, both markers of chondrogenesis, were upregulated (Figure 2(a)), while the histology did not reveal any formation of cartilage. This could imply that on day 5, pre-chondrocytes are differentiating and starting the production of cartilage, but an extracellular chondroid matrix has not yet been established. In a mouse fracture study, SOX9 has also been found to be highly upregulated. 15 However, in another murine model of osseointegration, no cartilage was detected histologically with Safranin O staining during a 28-day long time course. 14
Inflammation
The pro-inflammatory cytokines IL-1, IL-6 and TNF are especially important for initiation of inflammation and the repair process after injury by redirecting immune cells, inducing APP production and promoting bone resorption. 31 Messenger RNA coding for IL-6 was more than 40-fold upregulated after the implantation procedure (Figure 3). In agreement with these results, the expression of IL-6 was found to be elevated at day 4 in a murine fracture model, but only with a 3.4-fold upregulation. 16 However, IL-1 (Figure 3) and TNF (data not shown) were not regulated. We speculate that IL-1 and TNF were upregulated immediately after the surgical procedure but had returned to basal levels at day 5. This is in line with observations in mice, in which both IL-1 and TNF rapidly declined three days after a fracture. 32 In humans as well IL-1 has been reported to be downregulated from day 3 to 7 after insertion of titanium bone implants. 19
APPs are circulating plasma proteins found to be highly regulated by inflammation. 33 The expression of SAA and complement factors have not previously been characterized in porcine bone tissue. C3, SAA and the main porcine APP ITIH4 were all significantly upregulated (Figure 3). Previously, osteoclasts have been shown to produce C3, and the cleaved product C3a stimulates osteoblast differentiation in mice. 34 In addition, complements C3 and C5 have been linked to fracture healing in mice. 35 SAA is mainly produced and secreted by the liver, but expression has also been reported in human bone tissue, 36 and it has been shown to affect the differentiation of osteoblasts and osteoclasts in vitro. 37 In humans, downregulation of serum ITIH4 concentrations has been reported in patients with nonunion fractures, 38 and it has been suggested that fragments of the protein can be used as serum biomarkers for osteoporosis. 39
Growth factors
TGF-β is important for bone remodeling; it recruits mesenchymal stem cells, fibroblast and immune cells, stimulates the proliferation of osteoblasts and chondrocytes and increases the production of extracellular matrix proteins. 40 A four-fold upregulation of TGFB1 was found in response to the present implant insertion (Figure 4(a)). In accordance with our results, TGF-B1 was upregulated in bone tissue after fracture in mice 41 and elevated concentrations in serum and fracture hematomas of humans with long bone fractures have also been reported. 13
Bone morphogenic proteins (BMPs) are important for bone and cartilage development and repair; in addition, they are promising therapeutics for treatment of bone defects. 42 BMP2 and BMP7 have been used clinically for treatment of open fractures and nonunions to reduce healing time and complications. 24 BMP2, -4 and -7 were all significantly upregulated after surgery (Figure 4(a)). Cho et al. found that BMP2 and BMP4 were upregulated in mice immediately after fracture while BMP7 expression increased after seven days. 41 Based on the temporal expression of all the BMPs, Cho et al. speculated that BMP2 is responsible for triggering the healing cascade and expression of other BMPs. 41 This hypothesis is supported by the fact that the healing process is not initiated in fractures of BMP2 knockout mice. 43 The ratio between BMP4 and the BMP inhibitor noggin has also been speculated to be important in fracture healing. We found a two-fold upregulation of both BMP4 and noggin (Figure 4(a)), indicating that the BMP4/noggin ratio is not significantly changed by the implant insertion, but that both molecules play a role in the response to the insertion procedure. Kloen et al. showed that BMPs and their inhibitors are co-expressed and found a difference in the ratio of BMPs and their inhibitors in callus of human fractures compared to tissue from nonunions, indicating their importance in bone repair. 44
Following implantation, revascularization is important for the surrounding healing process, since angiogenesis is essential for ossification. 45 VEGFA is described as one of the most important growth factors for angiogenesis, also in relation to bone repair, 46 and it has been reported to be upregulated both in mouse and rat fracture models.15,47 However, in the present model, no regulation was seen of either VEGFA or its transcription factor HIF1A (Figure 4(b)). Instead the vascular growth factors ANGPT1, ANGPT2 and PTN were all significantly upregulated (Figure 4(b)). This indicates that VEGF-dependent angiogenesis pathways are not as important in the osseointegration of implants as in fracture healing. A difference in the role of VEGFA in fracture and implant insertion is supported by Schmid et al., who found that VEGFA was significantly upregulated both at one and nine days post-fracture in a murine model, while the gene was upregulated only at day 1 in controls without fractures but with a steel pin inserted in the tibia. 15 ANGPT1, -2 and PTN have also been reported to be upregulated in murine fracture models.17,48
The relevance of porcine models
Descriptive animal models are a necessity to advance modern medicine by increasing our knowledge of the complex reactions the body elicits in response to injury and disease. The most important parameter in choosing an animal model should not be low cost and easy handling, but how well the model mirrors human conditions. Porcine bone closely resembles human bone in regards to anatomy, mineral composition, rate of remodeling and microstructure3,7 and compared to mice the immune system of the pig is also more similar to the human counterpart.2,5,6 Therefore, results of bone inflammation attained in pigs have high translational potential.6,49 In today’s bone research rodents are often preferred over large animal models because they are so well characterized at the molecular level. This article contributes to alleviate the deficiency in molecular studies in porcine bone as the use of porcine models in biomedical research is increasing.1,49,50
Conclusion
The present molecular analysis gives a comprehensive understanding of how bone drilling and implant insertion changes the local gene expression in bone tissue five days after surgery in a porcine model. In conclusion, the porcine model revealed similarities to results from both human and murine studies. Therefore, pigs are useful and relevant, also on a local molecular level, as a large animal model in orthopedic surgery. Appropriate to the 3Rs, increased background knowledge of experimental animals will increase their reliability, which might lead to a reduction in the number of animals used. In the future, the present publication will serve as a strong argument for the use of porcine models in preclinical orthopedic studies, but also as a basis for further research of the transcriptional bone response related to infections and different kinds of local medical treatments.
Supplemental Material
Supplemental material for Pigs are useful for the molecular study of bone inflammation and regeneration in humans
Supplemental material for Pigs are useful for the molecular study of bone inflammation and regeneration in humans by Freja Lea Lüthje, Kerstin Skovgaard, Henrik Elvang Jensen and Louise Kruse Jensen in Laboratory Animals
Footnotes
Acknowledgments
We thank Betina Andersen, Elizabeth Petersen and Nicole Lind Henriksen for their excellent assistance with the histology and Karin Tarp for her technical assistance with the gene expression analysis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the European Union’s Horizon 2020 research and innovation program under the NoMorFilm project (grant agreement no. 634588), and by the Danish Research Council (grant no. 4005-00035B).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
