Abstract
The aim of this study was to document the pathogenic role of biovar Heyl of [Pasteurella] pneumotropica in mouse colonies. Fifty-three isolates associated with mastitis and orbital, cutaneous and vaginal abscesses as well as isolates from the nose and vagina of healthy mice were investigated. According to phenotypic characteristics and rpoB sequencing, the isolates were identified as [P.] pneumotropica biovar Heyl. Pulsed-field gel electrophoresis (PFGE) revealed five closely related profiles separated by only one to four fragments. The outbreak strains diverged from epidemiologically unrelated strains with the same rpoB sequence type, as shown by the PFGE profiles. The investigation documented that members of biovar Heyl of [P.] pneumotropica caused disease outbreaks in mouse colonies since the clonality indicated a primary role of [P.] pneumotropica biovar Heyl in the infections observed.
The most well-known opportunistic pathogen among members of the Pasteurellaceae family involved in rodent infections is [Pasteurella] pneumotropica. 1 [P.] pneumotropica has been separated into two biovars, Jawetz and Heyl.2,3 Biovar Heyl appears to represent a homogeneous taxon that is frequently isolated from mice and rats and is more rarely isolated from other rodent species (e.g. Syrian hamsters and mastomys).3,4 However, biovar Jawetz is considered to be a heterogeneous taxon with two different phenotypes that is usually isolated from mice, and is less common in rats.4,5 To separate the two biovars, acid formation from L-arabinose, melibiose, and raffinose in addition to lysine decarboxylase have been recommended.2,5,6 Biovar Heyl is positive in the lysine decarboxylase test and produces acid from L-arabinose and melibiose, whereas biovar Jawetz is negative in these tests. 2 Although separation of biovars is possible based on biochemical tests, the use of molecular techniques, such as 16S rRNA and partial rpoB gene sequencing, as well as PCR including 16S-23S rRNA internal transcribed spacer detection and high resolution melting curve analysis, have improved the differentiation of the biovars.6,7–15
[P.] pneumotropica has mainly been associated with latent infections of apparently healthy rodents, and it has been difficult to investigate its pathogenicity without serial passage. Thus, [P.] pneumotropica has been regarded as a secondary invader with a low pathogenicity in rodents.16–20 [P.] pneumotropica has been mainly associated with the respiratory tract of laboratory rodents since its original description.3,17 However, orbital infections have also been commonly associated with [P.] pneumotropica, and the most common clinical manifestations are reported as conjunctivitis, dacryoadenitis and panophthalmitis.4,21–23 In addition to the respiratory tract and orbital infections, mastitis, urogenital and subcutaneous infections have also been described.4,21,24–27
Most reports on disease conditions, such as orbital, vaginal, and subcutaneous abscesses and mastitis, do not refer to a specific biovar of [P.] pneumotropica.18,20,22–27 Artwohl et al. 28 reported that orbital abscesses are caused by biovar Heyl and confirmed this observation by 16S rDNA gene sequencing. The aim of the present study was to investigate the role of [P.] pneumotropica biovar Heyl in disease outbreaks by comparing the clonality of isolates involved since clonal outbreaks indicate a primary rather than secondary role in the infection. 29
Materials and methods
The specimens were collected from breeding colonies of outbred and inbred strains of mice (Bom:NMRI [outbred], C57BL/6JBom, BALB/cJBom, DBA/2JBom, and C3H/HeJBom [inbred], and Bom:NMRI-Foxn1nu [immunodeficient outbred] and B6.Cg/JBom-Foxn1nu [immunodeficient inbred]) that were produced by a commercial vendor (Bomholtgaard Ltd., Ry, Denmark) (approved by the Danish Experimental Animal Inspectorate) as part of the company’s health monitoring program of barrier-raised mice. All procedures were performed in accordance with the Danish guidelines for the care and use of laboratory animals.
Mice were housed in open standard plastic cages (Tecniplast, Buguggiate, Italy) in rooms under controlled temperature (20 ± 2℃) and relative humidity (55 ± 10%), with lights on from 06:00–18:00 h. All mice were bred and maintained in barriers with HEPA filtered air and chemical lockers using glutaraldehyde as a disinfectant. Immune-deficient mice were housed in open standard cages in laminar flow hoods. The animals were provided with autoclaved food (breeding diet, catalogue No. 01; Bomholtgaard Breeding Centre, Ry, Denmark) and acidified water (HCl at pH 3.5) ad libitum. Autoclaved bedding (Hahnflock® H9 granulate; Hahn & Co, Bredenbek-Kronsburg, Germany) and paper towels were used in the cages. Animal caretakers entered the barriers through lockers where they had to shower and changed into autoclaved clothes.
Description of the outbreaks
Investigated strains from the [Pasteurella] pneumotropica biovar Heyl outbreak.
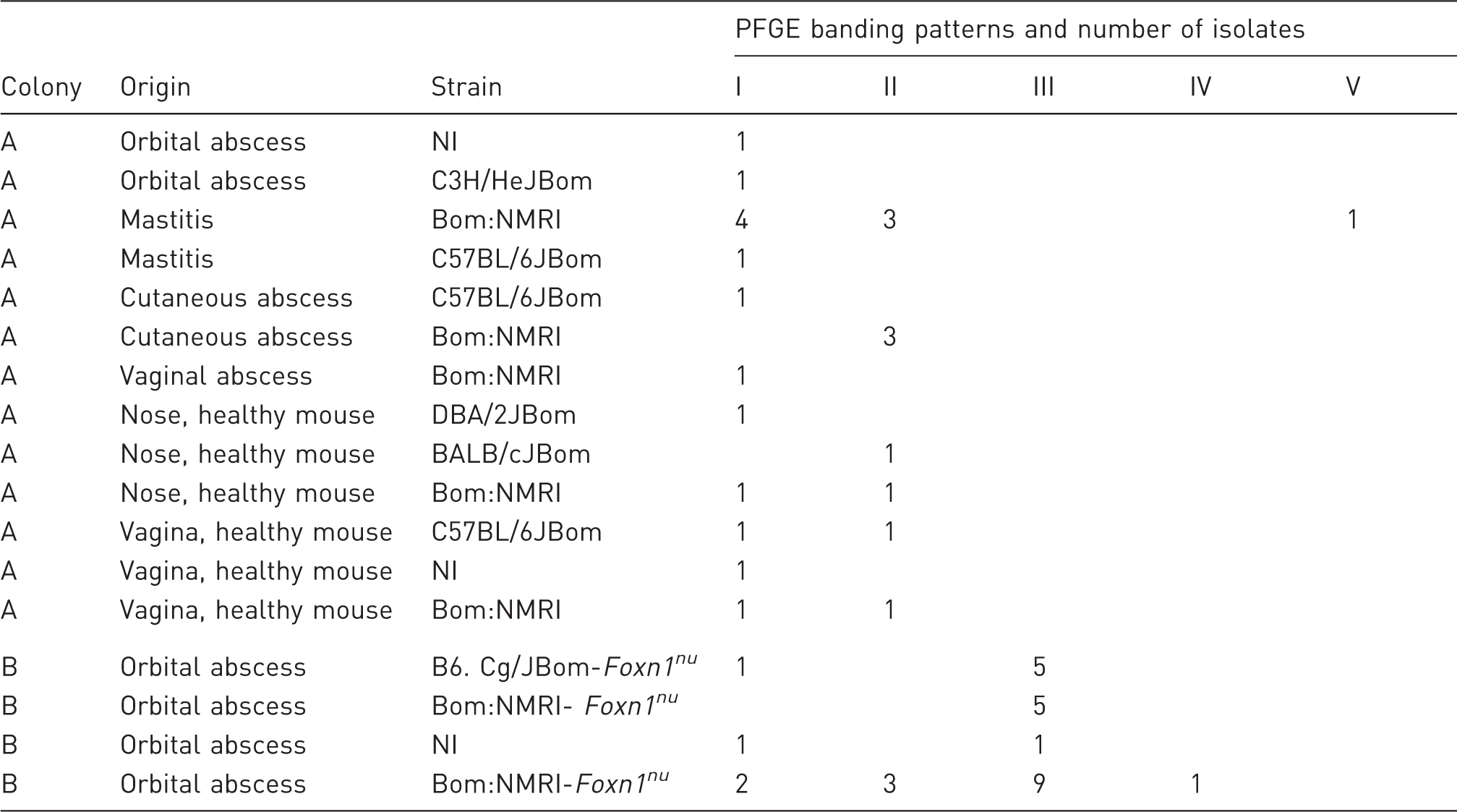
Bom:NMRI, Naval Medical Research Institute, outbred; DBA/2JBom, inbred; C57BL/6JBom, inbred; C3H/HeJBom, inbred; BALB/cJBom, inbred; B6.Cg/JBom-Foxn1nu, inbred; Bom:NMRI-Foxn1nu, outbred; NI, no information about strain. PFGE: pulsed-field gel electrophoresis.
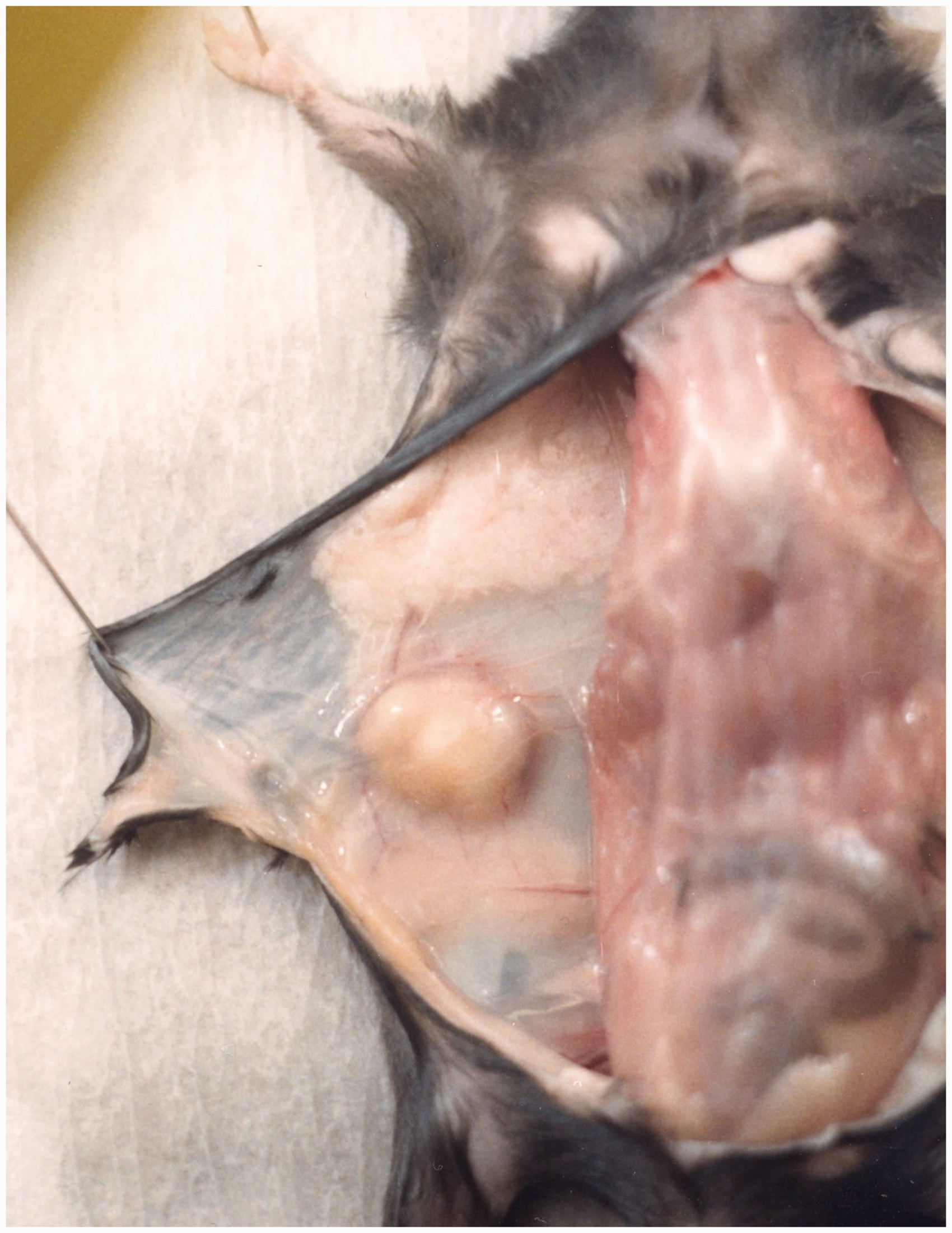
Example of a subcutaneous abscess investigated in an inbred C57BL/6JBom mouse from colony A.
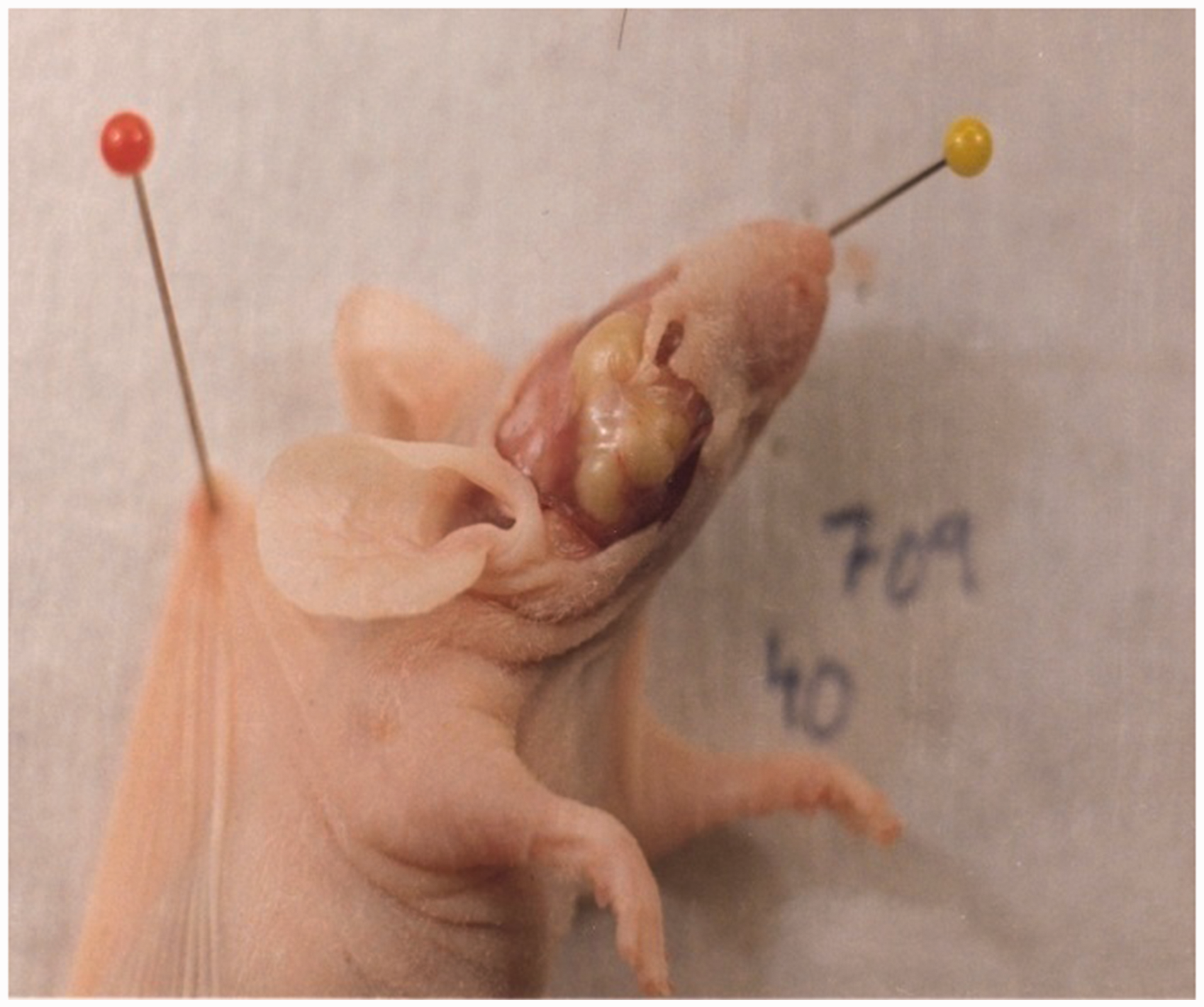
Example of an orbital abscess investigated in a mouse of the immune deficient outbred stock Bom:NMRI-Foxn1nu.
Isolation of bacteria
Live affected mice selected for examination were removed from the barrier and killed by cervical dislocation in the laboratory. They were then examined by post-mortem analysis. Sterile swabs were used to aseptically collect samples post-mortem. Samples were collected from the infected site and from the nose and vagina of non-infected mice. All samples were plated onto 5% calf blood agar plates and incubated at 37℃ for 24 h. Small non-hemolytic, grey or yellow, circular, low convex, even margined, non-transparent, smooth and glistening colonies with a butyrous consistency, with pin-point appearance sometimes, were isolated in pure culture. Isolated representative colonies in pure culture were preserved by freezing at −80℃. Preliminary identification was performed with API 20 NE (bioMérieux, Marcy l’Etoile, France). Full phenotypic identification was based on ‘classical’ biochemical testing. 7 For comparison with outbreak strains, the epidemiologically unrelated strains Ppn181, Ppn290 and Ppn264 isolated from mice in The Netherlands 30 were included, while strains P323 and MusIII were isolated from mice in Denmark and strain B7152/3 was isolated from a mouse in Australia.
Pulsed-field gel electrophoresis (PFGE)
PFGE was performed according to Pors et al. 31 with the following modifications. The cells were suspended in 2 mL of cell suspension buffer (100 mmol/L Tris/HCl, 100 mmol/L EDTA) to adjust the optical density (610 nm) of the bacterial suspension to 1.0. To make agarose plugs, SeaKem Gold Agarose (Lonza, Rockland, ME, USA) was used. Four hundred microliters of the bacterial suspension were mixed with 20 µL proteinase K (20 mg/mL), which was then mixed with an equal amount of agarose solution and the final mixture was embedded into plug moulds. The solidified agarose plugs were submerged in 5 mL of cell lysis buffer (50 mmol/L Tris-HCl, pH 8.0, 50 mmol/L EDTA, 1% N-lauroylsarcosine) containing 20 mg/mL of proteinase K and incubated in a water bath at 55℃ for 2 h. After washing in deionized water, the agarose plugs were stored at 4℃ until use. The plugs were cut into slices and placed in a buffer mix containing ultraclean water (AnalaR Normapur; VWR Chemicals, Leuven, Belgium) and Cut Smart (New England Biolabs [NEB], Ipswich, MA, USA) and incubated at room temperature for 15 min. After removing the buffer mix, the slices were digested by ApaI or NotI (NEB) using enzyme specific buffers. Subsequently, the plug slices were loaded into the wells of a 1% SeaKem Gold Agarose gel. The Low Range PFG Marker (NEB) was run in parallel with the samples. Strain H9812A of Salmonella enterica serotype Braenderup was also included as a size standard for PFGE as recommended by PulseNet. 32 Electrophoresis was run on a CHEF-DRII system (Bio-Rad Laboratories, Copenhagen, Denmark) at 6 V/cm with ramping times of 1–30 s for 18 h. After electrophoresis, the gel was stained with ethidium bromide, photographed and analysed using the program GelComparII (version 3.5; Applied Maths, Sint-Martens-Latem, Belgium). A dendrogram was drawn by cluster analysis based on the Dice coefficient and unweighted pair-group method using arithmetic averages (UPGMA) algorithm.
Partial DNA sequencing of the rpoB gene
Five strains were selected based on PFGE comparison for partial sequencing of the rpoB gene (560 bp fragment) to represent the different PFGE clusters observed. For comparison, a strain of biovar Jawetz isolated from a third colony during the same outbreak was also included, as well as the six reference strains previously stated. Sequencing was performed according to previously described protocols.8,9,33 A multiple alignment of DNA sequences was constructed by ClustalX2 34 and BioEdit. 35 Phylogenetic analysis of the rpoB gene sequences was performed by neighbor-joining using Jukes–Cantor correction, and included calculation of bootstrap support. MEGA6 was used for graphical representation of the tree. 36 Searches for sequences in public databases were performed by BLAST. 37
Results
Standard phenotypical tests allowed for the classification of all 53 isolates as [P.] pneumotropica biovar Heyl, the only deviations being a few isolates demonstrating small/pin-point colonies. Isolates from lesions in both colonies and isolates from the nose and vagina of healthy mice demonstrated the same phenotype. Identification of 16 out of 53 isolates investigated by API 20 NE did not result in any taxa recognized by the database (data not shown).
PFGE analysis after ApaI digestion of all 53 isolates resulted in five related PFGE profiles that differed in one to four fragments (Figure 3). Profiles I and III only differed in the size of the heaviest fragment, whereas profiles I and II also differed in one medium-sized fragment. Profile IV differed in three fragments from profile II and in four fragments from profiles I and III. Profile IV differed from profile V in two fragments (Figure 3). With the exception of the reference strain MusIII, which shared profile I of the outbreak strains, the other five reference strains diverged in five or more fragment positions. The PFGE profiles obtained by ApaI were confirmed by NotI digestion (data now shown). Cluster analysis showed that profiles I, II, III and V demonstrated 93% similarity (Figure 3). These profiles made up 43 out of 44 isolates from lesions and all nine isolates from apparently healthy mice. If the reference strains were included, the overall similarity decreased to 73% (Figure 3).
Unweighted pair-group method using arithmetic averages (UPGMA) dendrogram showing the five pulsed-field gel electrophoresis (PFGE) patterns of representative strains of [Pasteurella] pneumotropica biovar Heyl by ApaI restriction and comparision to six reference strains. The scale bar indicates cluster depth in the dendrogram based on the Dice similarity coefficient.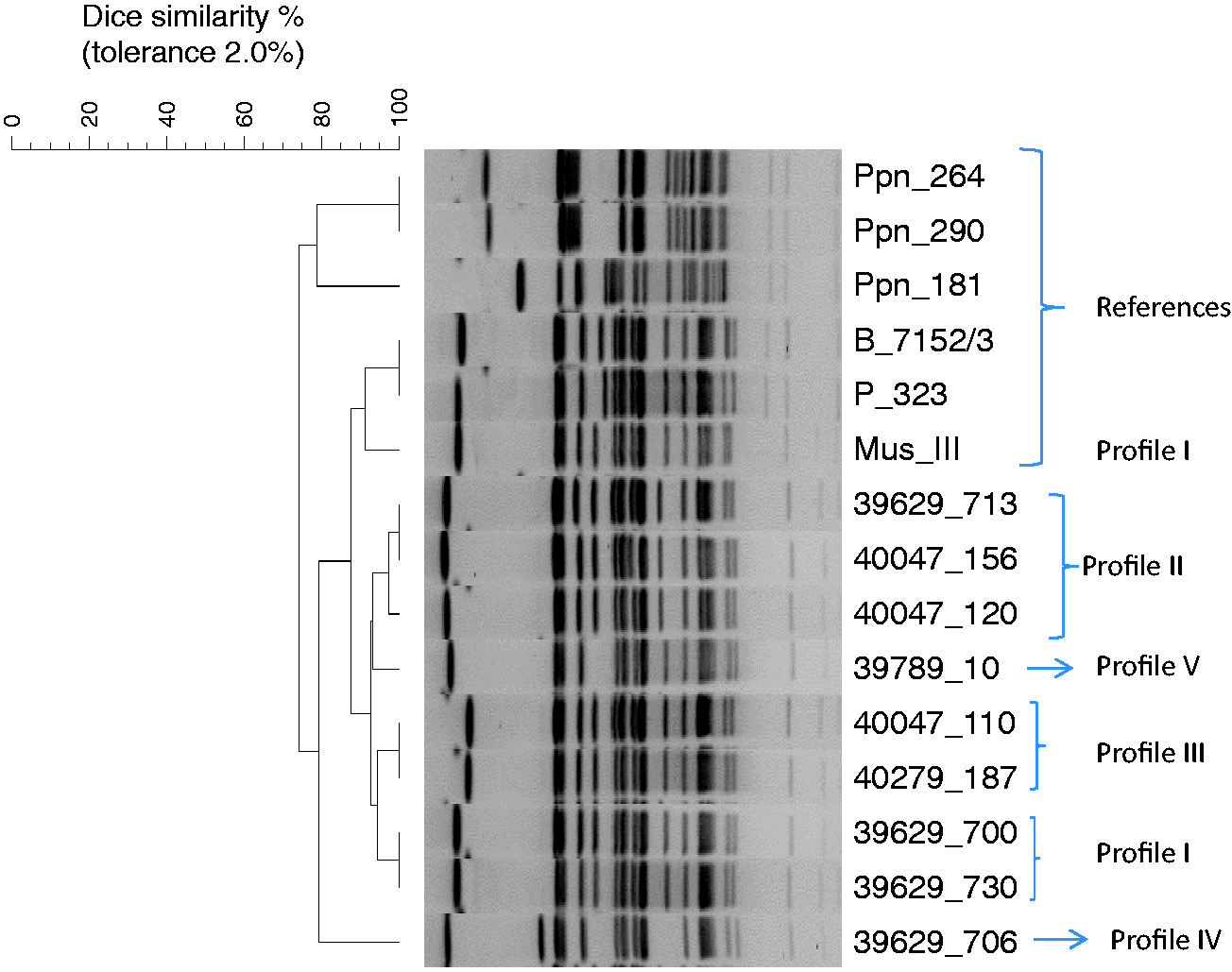
The association between PFGE types and lesion types in colonies A and B is shown in Table 1. While lesions in colony A were mainly associated with PFGE types I and II, lesions in colony B were mainly caused by PFGE type III. An association between lesion type and PFGE type was not demonstrated; however, isolates from apparently healthy mice in colony A belonged to the same PFGE type as isolates from lesions in this colony. All six rpoB sequenced isolates had identical sequences for the region of the gene analysed and showed 100% similarity with the reference strains CR30 and CR32
10
deposited in the GenBank database (Figure 4). This included one strain 39629-700 1-2-89 with pin-point colony morphology. All strains clearly differed from the type strain of [P.] pneumotropica biovar Jawetz, demonstrating only 88% similarity. Strain 40047-111 of biovar Jawetz was included as a control and shared the rpoB sequence with the type strain of [P.] pneumotropica (type Jawetz).
Phylogenetic tree of [Pasteurella] pneumotropica biovar Heyl based on rpoB sequence analysis of outbreak strains selected based on pulsed-field gel electrophoresis (PFGE) analysis. Strain 40047/111 of [P.] pneumotropica biovar Jawetz was included for comparison. Strain names are indicated with accession number in the parenthesis. Bar indicates 0.5% nucleotide difference.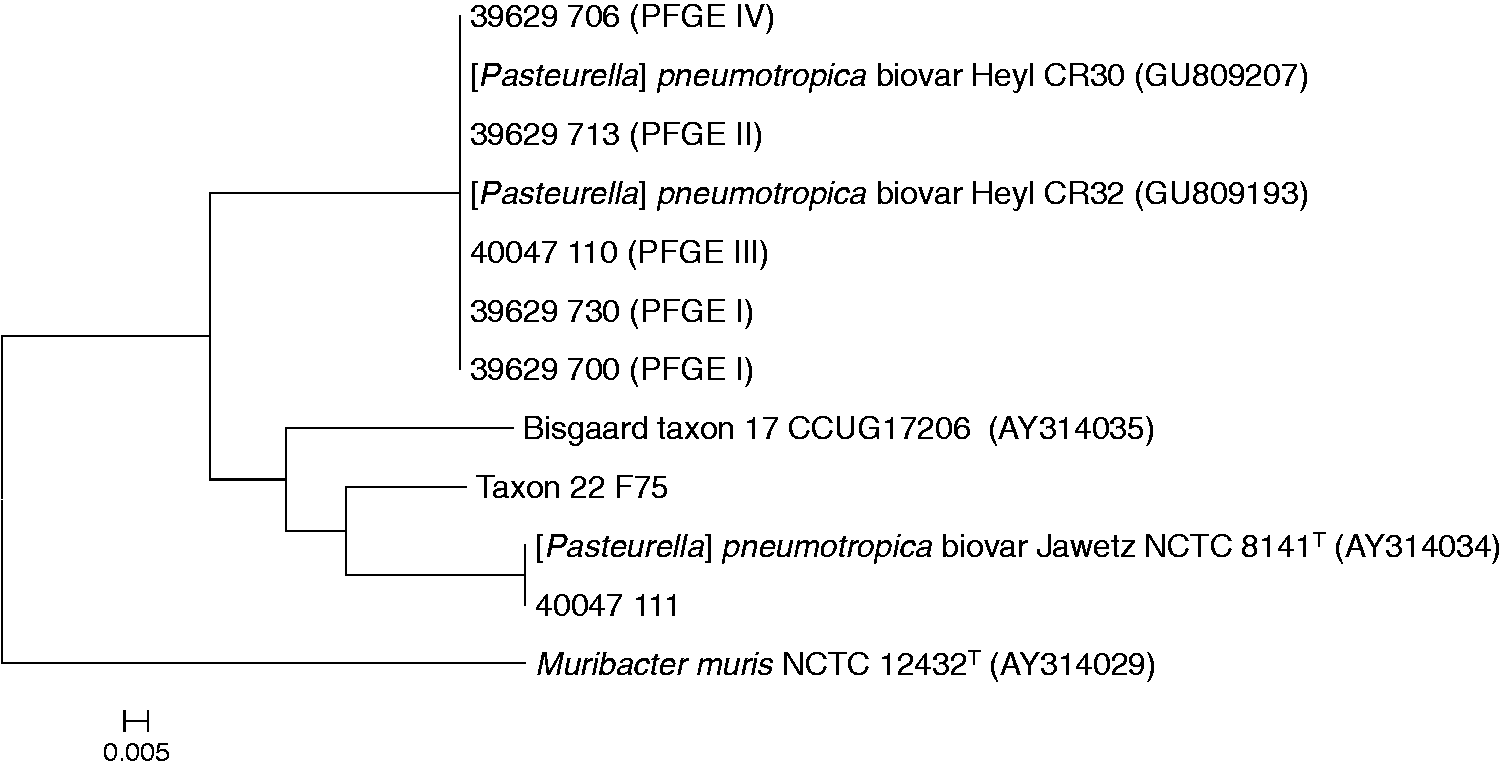
Discussion
Classical phenotypic tests for identification demonstrated similar phenotypic characters as reported previously for [P.] pneumotropica type Heyl.1,2 Phenotypic identification was confirmed by rpoB sequencing of selected isolates that demonstrated 100% similarity with reference strains. 10 The API 20 NE kit commonly used in animal diagnostic laboratories has been designated to be an appropriate tool for identification of Pasteurellaceae in rodents and rabbits. 38 Nevertheless, the strains investigated in this study were not recognized by the API database. The fastidious nature of these organisms might explain the diagnostic problems observed. The API 20 NE kit has also been found to be less reliable compared with other commercial test kits (such as ID HN-20 Rapid; Nissui Pharmaceutical, Tokyo, Japan) in identifying [P.] pneumotropica. 39 Similar problems have been reported for other members of Pasteurellaceae. 40
The five PFGE profiles observed for the 53 isolates investigated differed at most by four fragments. According to the ‘Tenover criteria’, two or three fragment differences between strains can be explained as a single point mutation or insertion or deletion of a DNA fragment, and such strains are considered to be members of the same outbreak clone, whereas strains with four to six fragment differences are considered to be possibly related. 41 According to these criteria, all strains from the outbreak were linked with at most four fragment differences. The strains were sampled over a period of five months, which might explain the genetic changes observed for some strains. The reference strains showed five or more differences in fragment positions compared with the outbreak strains, and were interpreted as unrelated (Figure 3). A single reference strain, MusIII, however, shared profile I of the outbreak strains. This isolate was obtained 10 years later from the same facilities and might have been maintained in the same breeding line. The reference strains had identical rpoB gene sequences as the outbreak strains, which explained the relationship with the outbreak strains in the PFGE analysis.
Despite the clonality of the outbreak strains, different lesions were demonstrated in the affected colonies. However, only local infections were observed, and the lack of systemic infections seems to indicate that the clone only has limited disease potential. Virulence factors of [P.] pneumotropica have not been studied in detail. A homologue of the RTX gene has been found in the type strain of [P.] pneumotropica (biovar Jawetz) with two paralogs pnxIA and pnxIIA. 42 Further screening of isolates showed a predominance of pnxIA in the strains investigated. Later, a third homologue of the RTX gene, pnxIIIA, was found in the type strain of [P.] pneumotropica (biovar Jawetz) and expressed less cytotoxic activity compared with PnxIA and PnxIIA. This homologue may play a role in hemagglutination. 43 Subsequently, a draft genome sequence of [P.] pneumotropica (biovar Jawetz) has been published, and additional candidate major virulence associated genes, such as filamentous hemagglutinin, type VI secretion system proteins and a novel RTX-like toxin coding gene, have been identified. 44 The ability of biofilm formation of both biotypes of [P.] pneumotropica has also been reported. 45 Recently, the presence of the RTX gene, as well as other virulence associated genes, in biovar Heyl has been investigated by comparative genome analysis of both biovars. Among the RTX genes, only pnxIII was identified in biovar Heyl, whereas the other genes were found to be common in both biovars. A YadA-like protein was considered to be a common virulence gene in both biovars, and its expression was related to autoagglutination and cell adhesion, which further indicates the influence of biofilm formation of [P.] pneumotropica. 46 Further studies are required to explore the mechanisms underlying the virulence factors and their distribution within the population, based on the present findings.
Previous investigations have not related to the transmission by carrier animals to the biotype of [P.] pneumotropica. 4 The current investigation indicates a high carrier rate of biovar Heyl. Transmission is also thought to be dependent on population density with inhalation of infectious droplets, dust or fomites as major vehicles. Additionally, vaginal colonization might represent an important possibility for horizontal and vertical transmissions within colonies.4,18 Self-scratching of external ears and the adjacent tissue of the head, as well as severe self-mutilation, has been reported in relation to otitis. 47
The isolates from the orbital abscess and nose of healthy mice showed PFGE pattern I in the same colony (A). It is possible that the infection may have ascended to the eye from the nasal cavity via the lacrimal duct. The nasolacrimal duct is present at an early age in mice, several days before their eyes are opened. 22 It is also stated that [P.] pneumotropica may be adapted to the Harderian gland and conjunctiva, suggesting that the orbital abscess had its origin in the Harderian gland. 21 Nicklas 4 has also mentioned that bedding particles can penetrate the conjunctiva due to the absence of eyelashes in nude mice, and that this may be the starting point of an infection. In colony B, all strains were isolated from orbital abscesses with some divergence in PFGE profiles (I–IV).
Three closely related PFGE profiles (I, II and V) were associated with mastitis in the same colony. This clinical condition may have had its origin in a contaminated environment. Mastitis has previously been reported in rats in barrier protected colonies. 48 Isolates from the cutaneous abscess also showed PFGE profiles I and II. Multiple subcutaneous abscesses caused by [P.] pneumotropica have been reported in rats. 26 PFGE patterns I and II were observed in the isolates of the vagina of healthy mice, whereas only pattern I was found in the vaginal abscess cases analysed. It is difficult to state with certainty whether or not the infection originates from the uterus. Previous studies49–51 have shown that uterine infections in laboratory rodents are possible and that [P.] pneumotropica in particular is recovered from infected uteri in mice.
Each clinical condition was recorded in different strains of mice. However, specificity for different strains of mice has been found. Wilson 52 reported an abscess in the masseter muscle that was only found in one strain of mice, although three different strains of mice were housed in the same environmental conditions. McGinn et al. 53 reported higher susceptibility to otitis in CBA/J mice compared with CBA/CaJ mice, which was probably related to differences in their degree of [P.] pneumotropica infection.
This report of a natural outbreak involving different clinical conditions caused by the same clone provides evidence for the pathogenicity of [P.] pneumotropica biovar Heyl. The clinical lesions were different in the two colonies, and this might be due to the different immune statuses of the mice in the two colonies. The immune-deficient animals were more prone to orbital abscesses, while immune-competent mice also suffered from abscesses at other locations. The finding of the same clone in both clinically affected and apparently healthy mice points to the opportunistic nature of this pathogen. Further investigations are needed to identify potential virulence factors that turn the [P.] pneumotropica biovar Heyl into a pathogen, and to investigate if diversity in virulence exists between subpopulations of this pathogen.
Footnotes
Acknowledgements
We would like to thank Ms Katrine Aagaard for excellent technical assistance in the laboratory, and Dr Louise Ladefoged Poulsen is thanked for her help with the GelCompar data analysis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
