Abstract
In this observational retrospective study, an outbreak of
Keywords
Introduction
Staphylococci are Gram-positive cocci inhabiting the skin and mucous membranes of animals, including humans. Staphylococci are considered opportunistic pathogens with
In mice,
The Federation of European Laboratory Animal Science Associations (FELASA) has been issuing guidelines for the health monitoring of mouse colonies since 1994.11–14 These recommendations include a list of additional agents for which testing is optional and thus are not tested except when the infectious agent may be relevant for the research programme. In the institution’s case, one of these relevant infectious organisms is
Here an observational retrospective study of a
Case description
A new rodent breeding area was opened in October 2012 at Trinity College Dublin (TCD) to centralise and facilitate rodent breeding and experimental animal work. At the time, the policy was to allow direct imports of live animals from trusted vendors with a health screening negative for the FELASA recommended infectious agents for laboratory mice 13 and embryo rederive all other rodents into the new area. This included mouse strains coming from an old breeding area within TCD’s campus.
All animals housed for experimental purposes are kept according to the European Union (Protection of Animals Used for Scientific Purposes) Regulations 2012 (S.I. No. 543 of 2012) 16 following the Commission Recommendation of 18 June 2007 on Guidelines for the Accommodation and Care of Animals Used for Experimental and Other Scientific Purposes. 17 At the time, mice were housed in individually ventilated cages (GM500, Tecniplast, Buguggiate, Varese, Italy), on poplar bedding (Eco-pure Poplar Chips3, Datesand Ltd, Manchester, UK), with two pieces of nesting material (Bed-r’Nest, Datesand Ltd, Manchester, UK), a shelter (Mouse House, Tecniplast, Buguggiate, Varese, Italy) and a raised polysulfone plastic platform (Figure 1; Mouse Loft, Tecniplast, Buguggiate, Varese, Italy) as standardised environmental enrichment per cage. Water was supplied through a reverse osmosis filtered automatic watering (Triple Red, Buckinghamshire, UK). Mice were all fed a standard irradiated pelleted diet (LabDiet PicoLab Rodent Diet 20 5053, LabDiet, Missouri, USA) within a range of 25.0 to 50.0 kGy. Mice were separated by sex at weaning and group housed up to a group size of five. The environmental conditions were 20–24°C, 45–65% relative humidity and a 12:12 light cycle. Animals were screened every three months according to FELASA recommendations. 13

(a) Mouse loft from Tecniplast used as environmental enrichment; (b) different mouse lofts representing undamaged (1 and 5) and different grades of damage (2, 3 and 4) and (c) to (e) detail of the damaged edges.
During the period June 2015 to January 2016, a total of 60 clinical cases were presented for veterinary examination showing a lump on the facial, periorbital and retroorbital area (Figure 2) in this new rodent breeding unit. Of the 60 cases, 57 were observed in C57BL/6 background (C57BL/6J, B6;129S4-
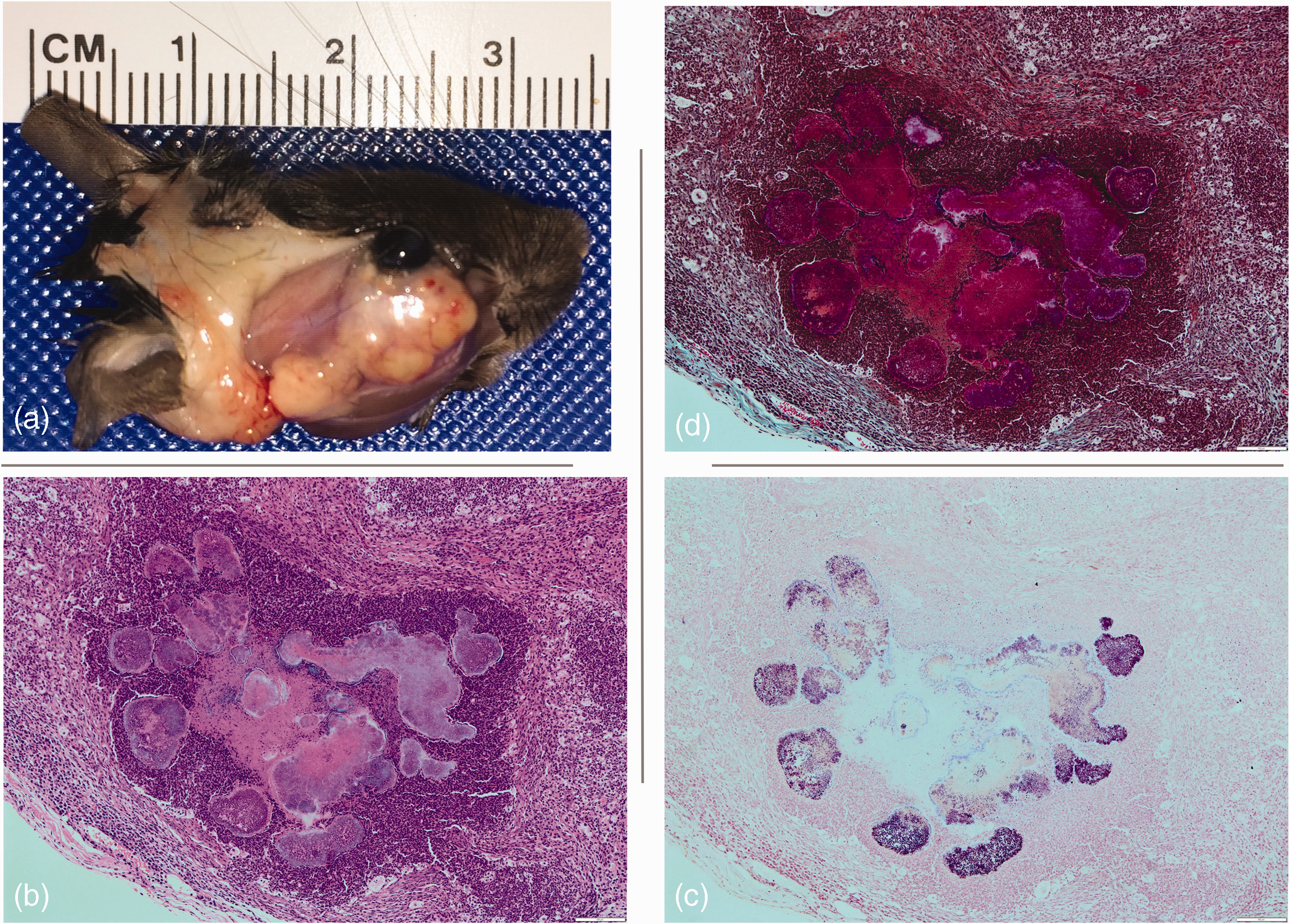
(a) Facial abscess; (b) to (d) micrographs showing characteristic
Concurrently, sharp edges were observed on our plastic mouse lofts used for environmental enrichment (Figure 1). All affected animals were housed with intact environmental enrichment that then became damaged by the mice. The mouse lofts are made from polysulfone plastic. Polysulfones are transparent, amorphous polymers with high strength and high heat resistance. 18 Lofts were damaged by gnawing, most probably through plastic stress and debilitation because of autoclaving (Figure 1). After three years of use, and approximately 78 washing and autoclave cycles, the polysulfone plastic environmental enrichment started to become fragile and the mice were able to gnaw through the plastic, creating these sharp ends. The institution stopped using this type of environmental enrichment in January 2016.
All clinically affected mice were euthanised by carbon dioxide inhalation using 20% volume per minute rate to fill the home cage, added to the existing air in the chamber before unconsciousness with minimal distress to the animals.
19
The lumps observed were abscesses in all cases. A differential diagnosis on pyogenic bacteria in mice includes streptococci,
1
Abscess, and sometimes also the mouth, were swabbed (eSwab, COPAN Italia, Brescia, Italy) and 100 µl of liquid plated on each of the following culture media: Columbia 5% Sheep Blood agar (reference 43041, Biomerieux, Hampshire, UK), MacConkey agar (reference 43141, Biomerieux, Hampshire, UK) and chromID® S. aureus Elite agar (reference 419042, bioMérieux UK Limited, Hampshire, UK) at 37°C in an atmosphere of 5% CO2; and on Chocolate agar PolyViteX (reference 43101, Biomerieux, Hampshire, UK) at 37°C in a jar (Oxoid AnaeroJar 2.5l, reference AG0025A, Thermo Scientific, Hampshire, UK) anaerobically (Oxoid™ AnaeroGen™ 2.5l sachet, reference AN0025A, Thermo Scientific, Hampshire, UK).
Pure bacteria culture was grown from the lesions in all cases. The bacteria recovered were Gram-positive cocci, β haemolytic on Columbia 5% Sheep Blood agar, catalase positive (Catalase Test Protocol, slide drop method, 3% H2O2), 22 oxidase negative and coagulase positive, except for isolate identified as number 16, which was coagulase negative (Staphylase Test Kit DR0595, Oxoid Limited, Hampshire, UK).
Twenty-one isolated colonies were grown on Brain Heart Infusion (BHI) broth (reference CM1135, Oxoid Microbiology Products, Thermo Scientific, Hampshire, UK) overnight and frozen using 50% v/v glycerol (reference 444482V, Prolabo, VWR International Ltd, Dublin, Ireland) in a cryotube (reference CRY8042, Nalgene Barcoded Cryovial, Sparks Lab Supplies, Dublin, Ireland) that was placed in a 4°C pre-equilibrated cooling rack (Labtop Mini-Cooler –20°C, reference 479-0642, VWR International Ltd, Dublin, Ireland) on dry-ice and left for 3–5 min before being transferred to a –80°C freezer for long-term storage. Cultured BHI broth was harvested, and DNA was extracted from pelleted bacteria using the Qiagen QIAamp Kit (Qiagen, West Sussex, UK) according to the manufacturer’s instructions. The DNA was dissolved in 200 µl of elution buffer (AE buffer, Qiagen, West Sussex, UK) and stored at −20°C. PCR products for sequencing were cleaned using the QIAquick kit (Qiagen, West Sussex, UK) according to the manufacturer’s instructions.
PCR amplification of the 16S rRNA gene was carried out as previously described 23 on the first seven different clinical isolates (Table 1 24 ) to ascertain initial genomic information on these isolates. Briefly, PCRs were carried out as follows: 50 ng of template was added to a PCR mix containing 2 mM dNTPs, 10 pmol of each primer and 2.5 U of HotStartTaq DNA polymerase (Qiagen, West Sussex, UK) in PCR buffer containing 15 mM MgCl2. PCR was performed using the C1000 Touch Thermal Cycler (Bio-Rad Laboratories Ltd, Watford, UK). Amplified DNA was resolved using gel electrophoresis in 1% agarose gels using Midori Green Advance DNA Stain (Nippon Genetics, Dueren, Germany). Amplified fragments were sequenced (Source BioScience, Cambridge, UK).
Isolate identification table.
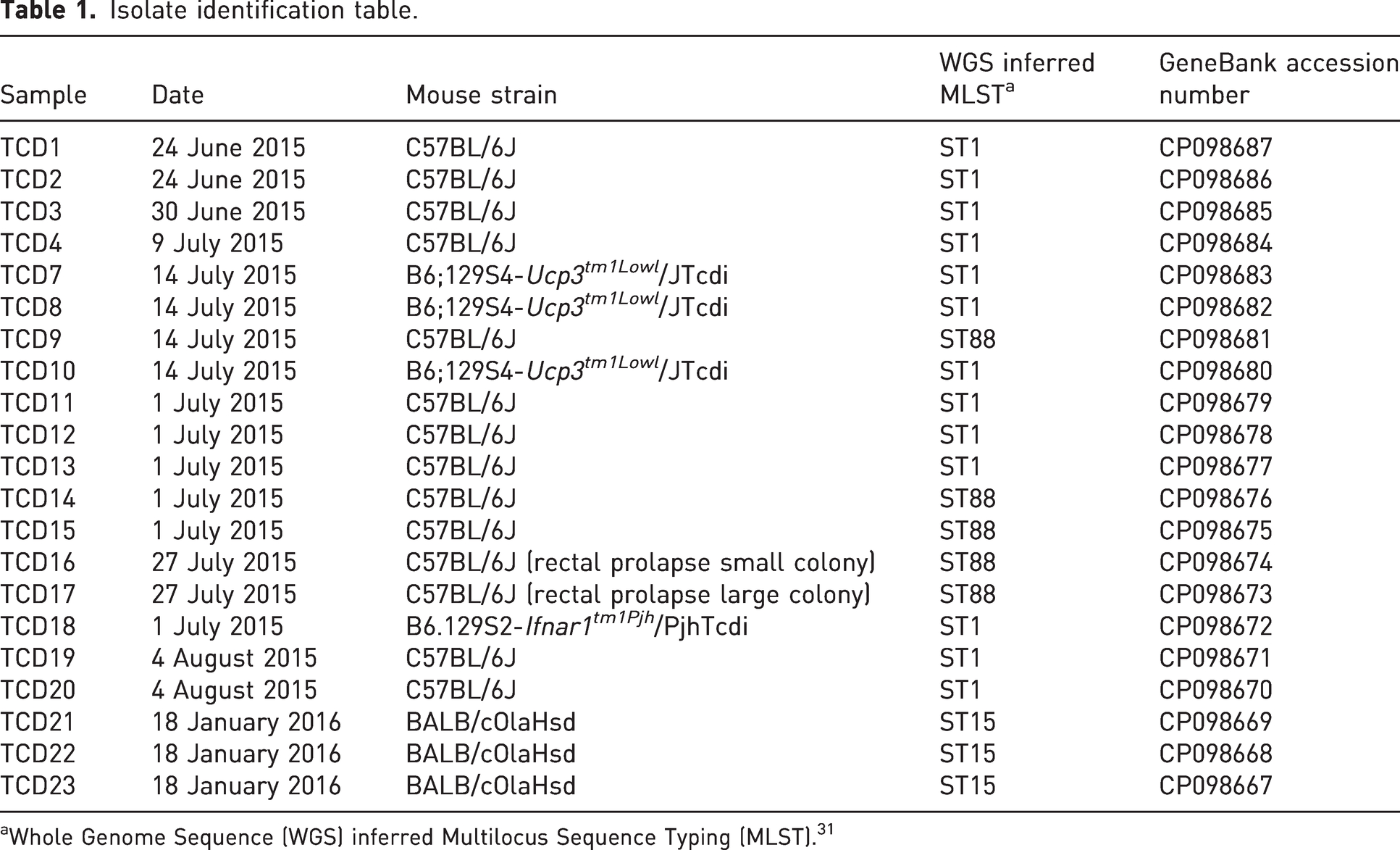
aWhole Genome Sequence (WGS) inferred Multilocus Sequence Typing (MLST). 31
Sequences were edited and analysed using EditSeq and SeqMan of Lasergene programs (DNAStar, Konstanz, Germany). Sequences were identified using BLAST from the National Center for Biotechnology Information (NCBI; http://www.ncbi.nlm.nih.gov/BLAST/). 16S sequences were also analysed using the ribosomal database project.25,26 Upon 16S rDNA sequencing, the isolates were identified as
Tissues for histological examination were taken in 10% neutral buffered formaldehyde (HistoTainer, Simport Scientific, Quebec, Canada). The microscopic lesions were characteristics of
To further characterise the outbreak, whole genome sequencing was done from the 21 clinical isolates. Briefly, the extracted DNA was cleaned using magnetic beads (Beckman Coulter AMPURE XP 5 ml Kit, Thermo Scientific, Hampshire, UK), and DNA eluted in molecular biology grade water (PCR Water, Qiagen, West Sussex, UK). DNA was quantitated (Qubit 4 Fuorometer, Thermo Scientific, Hampshire, UK) and the manufacturer-suggested protocol for generation of Nextera XT libraries followed (Nextera XT DNA Sample Preparation Kit, Illumina, Cambridge, UK). Samples were multiplexed using Illumina-supplied barcodes (Nextera XT Index Kit, Illumina, Cambridge, UK). Libraries’ concentrations were normalised to 4 nM, and when required the sample concentrated using vacuum (Savant SpeedVac, Thermo Scientific, Hampshire, UK). Paired-end sequencing was performed on the Illumina MiSeq using the 600-cycle (MiSeq Reagent Kit v3, Illumina, Cambridge, UK). Sequences were identified by contig alignment using BLAST from the NCBI (http://www.ncbi.nlm.nih.gov/BLAST/). Sequences were initially assembled using SPAdes
28
and a phylogenetic analysis performed using Harvest
29
(CosmosID, Germantown, Maryland, USA) (Figure 3). Final genome assembly for each of the isolates was carried out by SeqMan NGen through the whole genome SeqMan Ultra Variant Analysis/Resequencing (DNAStar, Konstanz, Germany) using
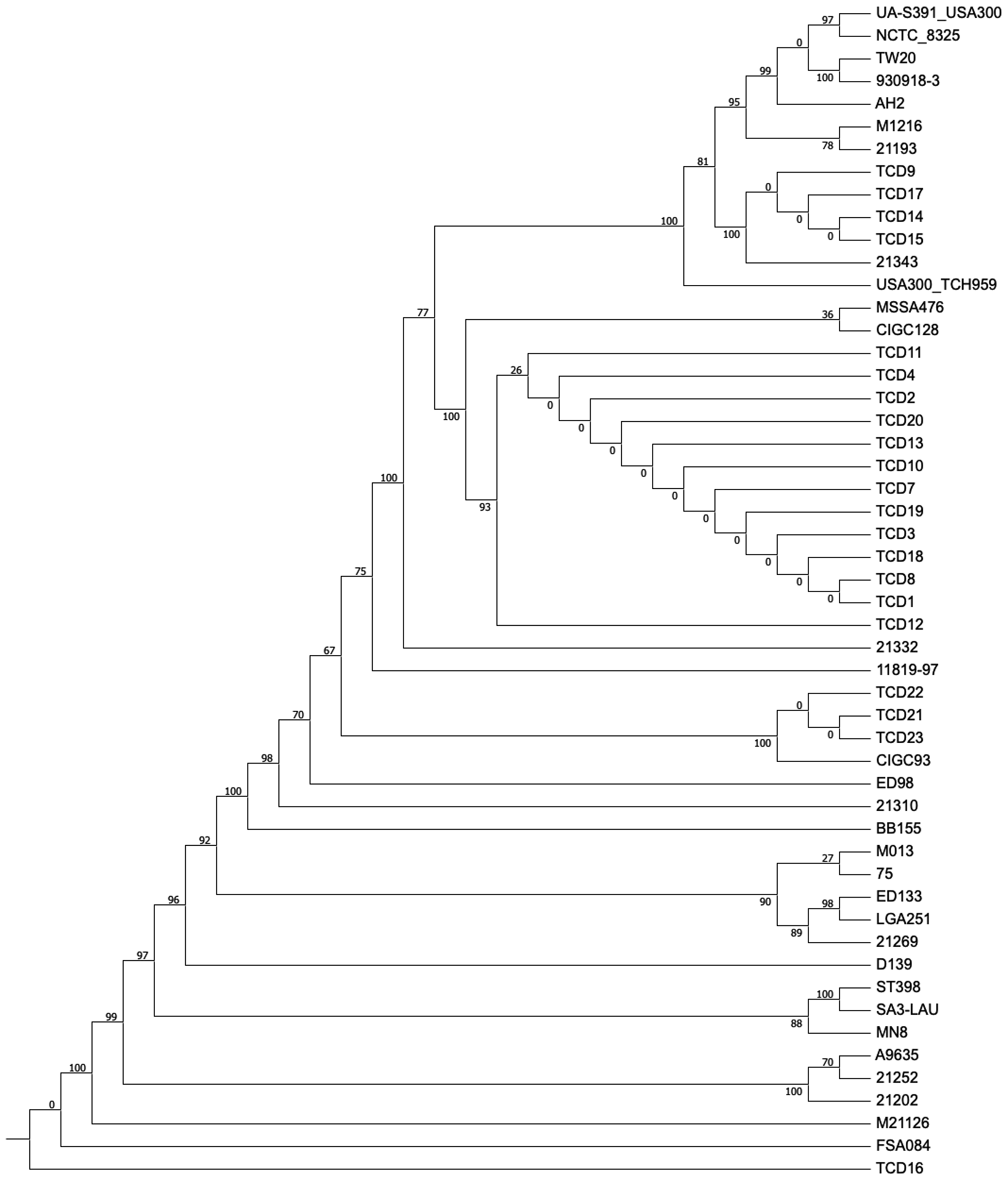

Whole genome analysis phylogenetic tree. The name of the
The personnel working in the area were screened to check for infection transmission from humans to mice through exposure to the hands of staff after they have been transiently colonised with staphylococci.
30
Nose samples from all people working in the area and mouse bacterial isolates were sent to the National MRSA Reference Laboratory (St James’s Hospital, Dublin, Ireland) for epidemiological typing by DNA sequence analysis of the X region of the protein A gene (spa).
31
Five out of 11 samples came positive for
Discussion
In this observational retrospective study, an outbreak of
The three clinical cases in late January 2016 occurred in BALB/cOlaHsd mice brought directly from a vendor into the experimental area. These mice came with a FELASA report negative for all listed organisms that unfortunately did not check for any of the additional agents, including
Here, I propose an additional aetiopathogenesis for
Data access
All data presented in this manuscript is accessible through GeneBank accession numbers – 16S rRNA gene: MH496641, MH496642, MH496643, MH496644, MH496646, MH496647 and MH496648; Genome: CP098667, CP098668, CP098669, CP098670, CP098671, CP098672, CP098673, CP098674, CP098675, CP098676, CP098677, CP098678, CP098679, CP098680, CP098681, CP098682, CP098683, CP098684, CP098685, CP098686 and CP098687.
Footnotes
Acknowledgement
I would like to thank Professor Timothy Morris for critical reading of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics statement
Legal and ethical requirements have been met with regard to the humane treatment of animals.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
