Abstract
Immunohistochemical markers for proliferation (bromodeoxyuridine, BrdU) and apoptosis (in situ terminal deoxynucleotide transferase dUTP nick end-labeling, TUNEL) were localized within glutathione S-transferase (GSTP)-positive hepatic foci in rats. Using the TechMate Automated Staining System (BioTek Solutions; Santa Barbara, CA), formalin-fixed, paraffin-embedded sections were run through a double-label avidin–biotin–immunoperoxidase protocol in less than 10 hr. Steam heat-induced epitope retrieval and/or proteolytic digestion preceded each labeling procedure. Color development was achieved using diaminobenzidine (DAB) with nickel enhancement for BrdU and TUNEL and VIP for GSTP. Results illustrate clear staining, brown-black BrdU-positive nuclei or TUNEL-positive apoptotic bodies within purple GSTP-positive hepatocytes. This automated procedure provides a method to easily identify and quantitate proliferating or apoptotic cells within foci of altered hepatocytes in rat liver and may have general applications for studies of cell or tissue kinetics during development, differentiation, and various pathological conditions in animals and humans.
Keywords
D
The control of cell numbers in tissues or specific cell populations is highly dependent on the balance between cell birth and cell death, or apoptosis (Pitot and Sirica 1980; Wyllie et al. 1980; Alison et al. 1995). Measurements of cell proliferation and apoptosis in normal or neoplastic cell populations can be informative indicators of the carcinogenic nature of many drugs or chemicals (Schulte–Hermann et al. 1993; Luebeck et al. 1995; Goldsworthy et al. 1996; Jones et al. 1996). Recent studies have evaluated proliferation and apoptosis within the target cell (i.e., preneoplastic) population to determine net growth, which may relate to neoplastic progression (Goldsworthy et al. 1996). Several laboratories are measuring cell proliferation and apoptosis within hepatic foci of rodents, the putative preneoplastic liver lesions that are used as markers in medium-term assays to estimate liver tumor risks (Pitot and Sirica 1980; Pitot 1990).
Methods for detecting cell proliferation in rodent liver have been reviewed (Goldsworthy et al. 1995, 1996; Eldridge and Goldsworthy 1996). Immunohistochemical markers of cell proliferation, such as antibodies to 5-bromo-2-deoxyuridine (BrdU) or proliferating cell nuclear antigen (PCNA), have recently been identified within rat hepatic foci that stain positive with various immunohistochemical markers, such as glutathione S-transferase (placental form; GSTP) (Tiwawech et al. 1991; Buchmann et al. 1994; Dragan et al. 1994).
There are several methods for detecting apoptosis in rat hepatocytes (Oberhammer and Roberts 1994; Wheeldon et al. 1995; Goldsworthy et al. 1996). One popular method useful for quantifying apoptosis in paraffin-embedded tissue is in situ end labeling (ISEL) (Gold et al. 1993; Wijsman et al. 1993). ISEL relies on exogenous enzymes, such as DNA polymerase I for in situ nick translation (ISNT) or terminal deoxynucleotidyl transferase (TdT) in nick end-labeling (TUNEL) to incorporate labeled nucleotides to the 3′-hydroxyl (3′OH) recessed termini of DNA breaks. TUNEL is considered more sensitive and specific than ISNT (Gold et al. 1993), and the former was chosen for two commercially available in situ apoptosis detection kits, ApoTag and ApoTag Plus (Oncor; Gaithersburg, MD) and one from Boehringer (Mannheim, Germany). Although apoptosis detection by ISEL techniques in normal or neoplastic rat liver has been described (Kong et al. 1995; Roberts et al. 1995; Wheeldon et al. 1995), a double labeling approach has not.
Our objective was to develop an automated double labeling technique to detect both proliferating and apoptotic cells in GSTP+ hepatic foci. BrdU- or TUNEL-labeled focal (GSTP+) and nonfocal (GSTP-) hepatocytes of rats were identified and quantitated following an initiation–promotion protocol. This automated double labeling technique provides advantages of minimal effort in an accurate, consistent, and expedient manner and may have applications for studies of proliferation or apoptosis within a wide variety of tissues and cell types.
Materials and Methods
Animal Treatment, Tissue, and Slide Preparation
In an initiation–promotion protocol, eight male Sprague–Dawley rats were given 150 mg/kg diethylnitrosamine (Sigma Chemical; St Louis, MO) by
First Label: BrdU or TUNEL
BrdU. Reagents for this assay, excluding the primary antibody, were purchased as a kit (BioTek Solutions). Slides were steamed (Handy Streamer Plus; Black and Decker, Shelton, CT) for 20 min in heat-induced epitope retrieval (HIER) buffer. The HIER procedure was performed manually but automation of this technique has been described (Myers 1995). Slides were incubated in 1 N HCl at 43C for 1 hr and processed for BrdU immunohistochemistry on the BioTek Solutions' TechMate Automated Staining System. A 15-min incubation in 3% H2O2 in buffer blocked endogenous peroxidase and a 15-min proteolytic digestion in prediluted proteinase K (Dako; Carpenteria, CA) exposed antigen binding sites. Nonspecific binding was blocked using goat serum in PBS buffer for 15 min, followed by a 45-min incubation in the first primary antibody, a 1:50 dilution of mouse monoclonal anti-BrdU (Becton–Dickinson; Mountain View, CA). After the initial primary antibody incubation, slides were placed in biotinylated secondary anti-mouse (rat absorbed) antibody for 30 min. This antibody complex was then detected using a 30-min incubation in an avidin–biotin–horseradish peroxidase complex (ABC) and a series of three 7-min 3,3′-diaminobenzidine (DAB) chromagen steps. To intensify color development, 1 μl of 8% nickel chloride per 1 ml of DAB was mixed 5–10 min before use (Hsu and So-ban 1982).
TUNEL. ISEL using the ApopTag detection kit (S7100 kit; Oncor) was used to detect cells undergoing apoptosis. Sections were processed in accordance with kit instructions, with modifications as noted below. The TechMate system automated protocol began with a 15-min incubation in 3% H2O2 in buffer, followed by a 10-min proteolytic digestion in prediluted proteinase K. Slides were incubated in equilibration buffer for 2 min, followed by incubation in working strength terminal deoxynucleotidyl transferase (TdT) enzyme at 37C for 70 min. The TdT working strength enzyme consists of TdT enzyme and digoxigenin-11-dUTP for end extension of the 3′OH ends of double- or single-stranded DNA of fragmented DNA. This was followed by stop/wash and anti-digoxigenin–peroxidase steps, which were identical to kit instructions. Color development with DAB and nickel chloride was identical to that described with BrdU, as above.
Second Label: GSTP
Reagents for this assay, excluding the primary antibody and the VIP chromogen (Vector Labs; Burlingame, CA) were purchased as a kit (BioTek Solutions). After automated staining of the first label, slides were removed from the TechMate machine and steamed manually in HIER for 20 min before automated staining for GSTP. Residual biotin from the previous BrdU staining procedure was blocked using an avidin–biotin blocking kit (Vector Labs) and peroxidase used in the first procedure was blocked with 3% H2O2 for 15 min. Slides were incubated in rabbit anti-human GSTP (Dako) in 1:100 dilution in PBS buffer for 45 min, followed by biotinylated anti-rabbit and ABC–peroxidase for 30 min each. Slides were developed using the VIP chromogen for 3–10 min. The VIP chromogen development was visually monitored until desired staining density was achieved. Slides were counterstained with hematoxylin, dehydrated, cleared, and coverslipped using a xylene-based mounting medium.
Controls
Positive controls included collection, sectioning, and staining for BrdU or TUNEL incorporation in duodenum along with liver in the same slide from each rat. Additional tissue slides were stained separately for BrdU, TUNEL, or GSTP to verify that staining sensitivity was not decreased by double labeling. Negative controls for BrdU or GSTP immunohistochemical staining included slides that were incubated with normal mouse or rabbit serum in place of BrdU or GSTP primary antibodies, respectively. To provide a negative control for TUNEL, we omitted TdT from the labeling mix. Specificity was controlled by comparing TUNEL with morphology as observed in H&E-stained slides.
Quantitation of Proliferation and Apoptosis
An electronic imaging program (Image Pro Plus; Media Cybernetics, Silver Springs, MD) was used as an aid in counting by projecting the electronic image onto a monitor for identification and tagging of BrdU- or TUNEL-positive hepatocytes. BrdU and TUNEL labeling indices of nonfocal (GSTP-) hepatocytes were determined by dividing the number of BrdU-positive hepatocyte nuclei or TUNEL-positive hepatocytes, respectively, by the total number of hepatocytes scored and multiplying by 100. A minimum of 1500 or 4000 hepatocytes were scored in random fields for BrdU or TUNEL labeling, respectively, from the left liver lobe of eight rats. BrdU and TUNEL labeling indices of focal (GSTP+) hepatocytes were determined by dividing the number of BrdU-positive hepatocyte nuclei or TUNEL-positive hepatocytes, respectively, by the total number of hepatocytes within each GSTP+ focus and multiplying by 100. BrdU and TUNEL labeling indices were determined for 20 random hepatic foci per section and were averaged to determine the focal BrdU or TUNEL labeling index for each rat. Mean ± SEM focal and nonfocal BrdU and TUNEL labeling indices were calculated. Statistical analysis was conducted to determine significant differences between focal and nonfocal labeling indices. Focal and nonfocal labeling indices were compared by a Student's t-test if the data passed a test for normality (ANOVA) or Mann–Whitney Rank Sum test if the data failed the normality test.
Results
The combination of dark brown-black DAB staining of BrdU or TUNEL and the purple VIP chromagen staining for GSTP gave excellent contrast and staining quality (Figures 1A and 1B). BrdU-labeled nuclei of normal hepatocytes were easily distinguished from labeled nonhepatocytes by their large, spherical contour and their central cellular location. Staining intensity of GSTP+ hepatocytes varied throughout each section, which is consistent with the variable phenotypic expression of this marker within rat hepatic foci. BrdU-labeled hepatocyte nuclei were easily identified within purple GSTP+ hepatocytes and were readily distinguished from unlabeled hepatocyte nuclei within the same focus.
We first examined H&E-stained liver sections for apoptosis because the current standard for recognition and quantitation of apoptosis in tissue sections is based on morphological features observed with this stain (Goldsworthy et al. 1996). The earliest visible stage of apoptosis by H&E is an apoptotic cell, characterized by a compact hepatocyte with condensed chromatin marginated to the nuclear membrane (Wyllie et al. 1980). This stage was rarely observed in H&E-stained sections, probably because this stage lasts only a few minutes and therefore the probability of detection is low (Goldsworthy et al. 1996). Apoptotic bodies, which are fragmented from cell surface pseudopodia, were seen more commonly than apoptotic cells because this stage is visible for a few hours (Goldsworthy et al. 1996). Most apoptotic bodies appear as discrete, eosinophilic membrane-bound bodies within surrounding hepatocytes and were occasionally identified in eosinophilic hepatic foci in H&E-stained liver (Figure 2). Apoptotic bodies exhibit diverse forms and sizes and can be categorized into chromatin- and nonchromatin-containing bodies (Goldsworthy et al. 1996). Most apoptotic bodies have been engulfed by surrounding hepatocytes and consequently are visualized as intracellular bodies.
In TUNEL/GSTP double labeled slides, apoptotic cells and bodies stained dark brown and hepatic foci stained purple (Figures 3A and 3B), which enabled easy identification of various stages of apoptosis. TUNEL-labeled apoptotic bodies were observed as discrete, membrane-bound, chromatin- or nonchromatin-containing bodies of variable size within the cytoplasm of GSTP+ hepatocytes (Figure 3B) or as diffuse light or dark brown cytoplasmic staining that covered the entire cell (Figure 3B). Occasional bodies did not stain positive with TUNEL (Figure 3B). TUNEL-labeled apoptotic cells, characterized as brown staining confined to the nucleus of intact hepatocytes, were occasionally observed (Figure 3B) but were less numerous than apoptotic bodies.
Positive control slides of duodenum demonstrated dark brown BrdU-labeled cells that covered the entire surface of the intestinal villi or dark brown TUNEL-labeled cells at the tip of the villi. The pattern of intestinal BrdU or TUNEL labeling was consistent with that described with this tissue after continuous (>4 days) BrdU labeling (deFazio et al. 1987) or TUNEL labeling (Gavrielli et al. 1992). Liver slides of single labeling of BrdU, TUNEL, or GSTP verified that the double labeling technique did not decrease binding of either antibody because staining intensity for BrdU, TUNEL, or GSTP was similar between single and double labeling procedures. Staining of BrdU and GSTP control slides incubated with no primary antibody was negative. Omission of TdT from the TUNEL mix completely abolished labeling.
Quantitation of BrdU and TUNEL labeling indices of nonfocal and focal (GSTP+) hepatocytes is presented in Table 1. Proliferation and apoptosis within focal hepatocytes were increased approximately fourfold greater than nonfocal hepatocytes, indicating the utility of demonstrating differences in proliferation and apoptosis between focal and nonfocal hepatocytes.
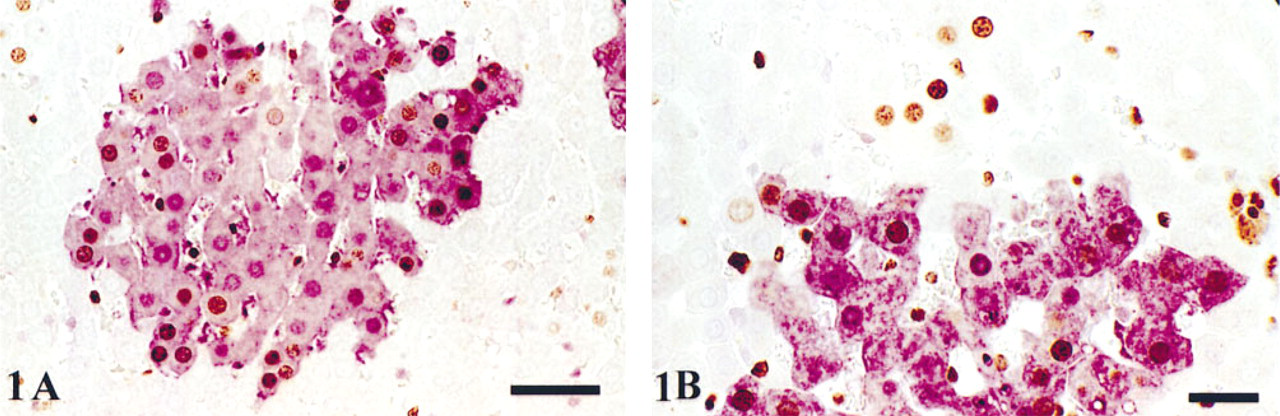
Photomicrograph of BrdU/GSTP double labeling in rat liver. (
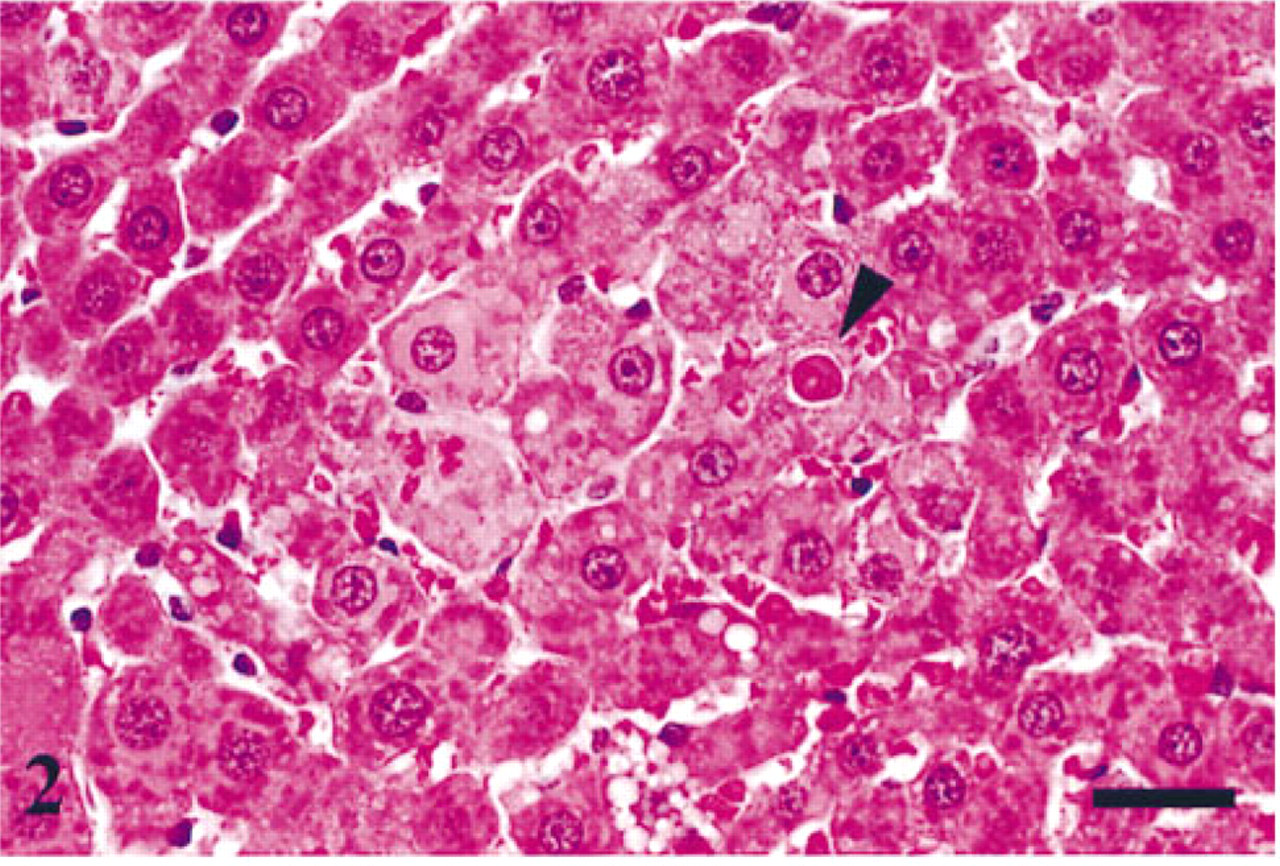
Photomicrograph of H&E-stained liver demonstrating eosinophilic hepatic focus with apoptotic body (arrowhead). Bar = 25 μm.
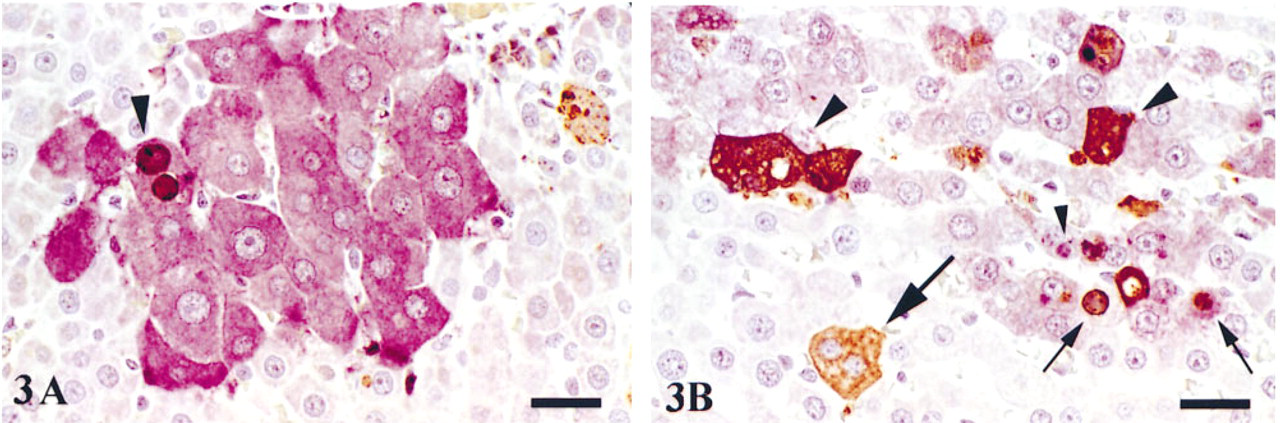
Photomicrographs of TUNEL/GSTP double labeling in rat liver. (
Discussion
Double labeling of BrdU or TUNEL and cytoplasmic proteins in formalin-fixed, paraffin-embedded liver or other tissues from rats by manual methods has been described (Buchmann et al. 1994; Tischler 1995; Ubink et al. 1995), but this is the first report of an automated capillary gap technique for demonstration of BrdU or TUNEL in single or double labeling procedures. Automation of double labeling using the TechMate staining system (Myers et al. 1995) has enabled this typically time-consuming and tedious procedure to be performed with a minimal effort in an accurate, consistent, and expedient manner.
Use of steaming compared to microwave treatment for heat-induced epitope retrieval (HIER) is equally effective for most antigens (Myers et al. 1995). Incorporation of an automated steaming step for BrdU and GSTP on the TechMate (Myers et al. 1995), rather than the manual steaming procedure conducted in this study, would complete the automated procedure. HIER was not conducted with the TUNEL procedure, because we have observed decreased specificity after microwaving and the use of HIER for ISEL is controversial (Lucassen et al. 1995; Negoescu et al. 1996). Several other modifications were made to the automated double labeling techniques described by Myers et al. (1995). Because proteolytic enzymatic digestion is often required to expose crosslinked sites to allow adequate antigen–antibody binding, digestion with proteinase K was conducted in this study before the BrdU or TUNEL procedure. The utility of proteolytic digestion after antigen retrieval is questionable. However, we determined that this step was beneficial to staining tissue fixed in formalin for 3 days but was not necessary for tissue stained for 1 day (data not shown). A 1-hr incubation in 1 N HCl was added for BrdU labeling to intensify staining in tissue fixed in formalin for 3 days (Frank et al. 1993). We also determined that rat liver fixed for 24 hr or less did not require HCl incubation for BrdU staining, and this step was omitted for TUNEL because HCl may increase the number of single- or double-stranded DNA breaks (Wijsman et al. 1993).
BrdU and TUNEL labeling indices of nonfocal and focal hepatocytesa
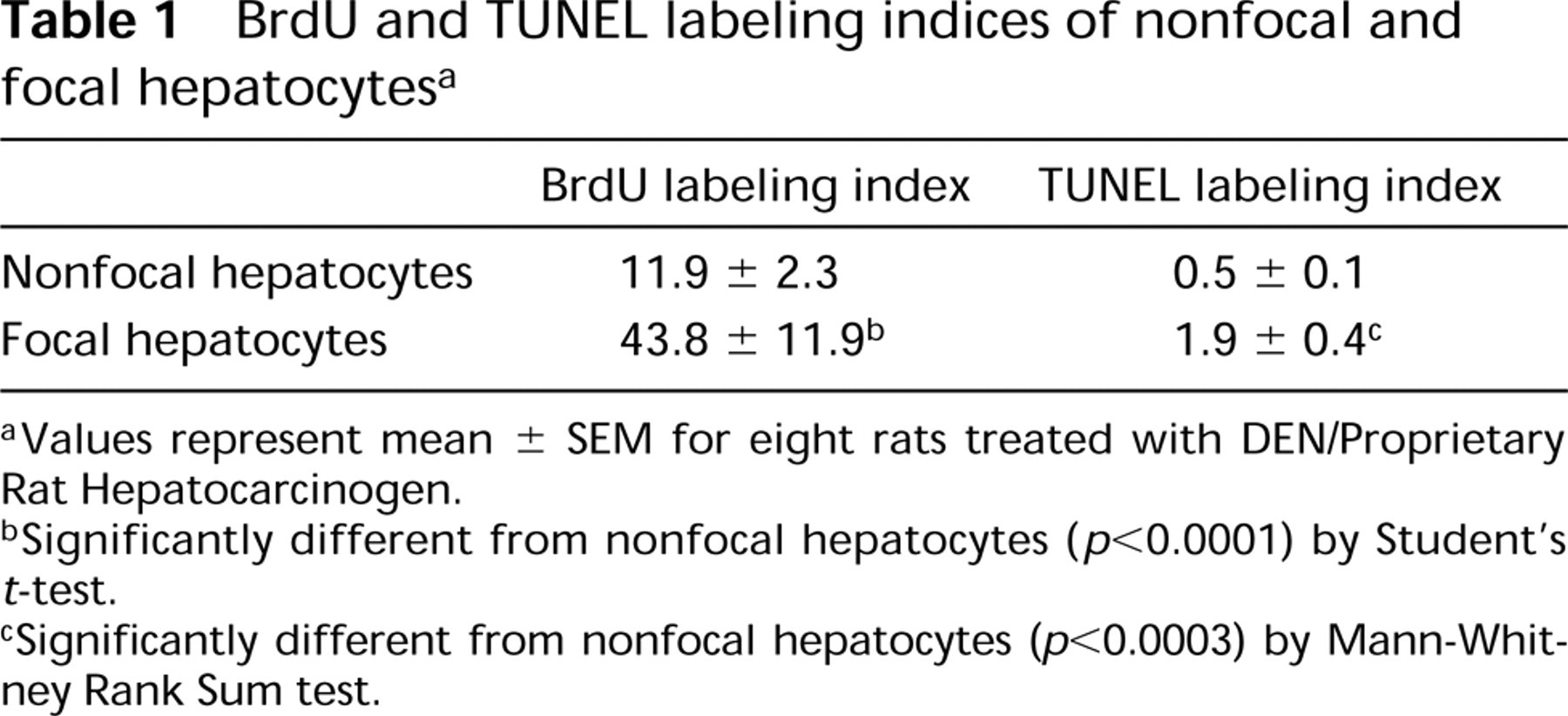
Values represent mean ± SEM for eight rats treated with DEN/Proprietary Rat Hepatocarcinogen.
Significantly different from nonfocal hepatocytes (p<0.0001) by Student's t-test.
Significantly different from nonfocal hepatocytes (p<0.0003) by Mann-Whitney Rank Sum test.
The utility of an avidin–biotin blocking kit in our second label (GSTP) after HIER for 20 min is unknown, because the residual link and label system may have been deleted by HIER (Lan et al. 1995). We suggest that this step may be superfluous and can be omitted without consequence. The product of the DAB chromagen used for BrdU or TUNEL was intensified by addition of nickel chloride (Hsu and Soban 1982). For GSTP staining, we also used aminoethyl-carbazole (AEC), which produced a red stain that was acceptable for distinguishing from brown-stained BrdU-positive nuclei or apoptotic cells or bodies. However, hepatocyte staining with the purple VIP chromagen contrasted better than red AEC in double labeling using modified DAB and had the added advantage of being insoluble in organic solvents (Pastore et al. 1995).
Results of this study demonstrate the utility of BrdU for detection of proliferating hepatocytes within GSTP+ foci. The advantage of BrdU is that it can be administered to animals for prolonged periods, increasing the cumulative total of cells undergoing DNA synthesis compared to that detected with endogenous markers, such as PCNA. Immunostaining for PCNA has the potential to identify cells in the G1, S, G2, or M phases of the cell cycle and can be used in archival tissue sections (Eldridge and Goldsworthy 1996). The choice of which measure of cell proliferation is the most relevant depends on the objective of the study (Goldsworthy et al. 1995). PCNA has been successfully used in double labeling immunohistochemistry studies of rat liver in this laboratory with a similar automated procedure (data not shown).
Apoptosis is best characterized by evidence of DNA fragmentation and morphological features by H&E, but these present serious drawbacks in quantitation (Negoescu et al. 1996). ISEL, by coupling morphology and DNA fragmentation into a unique assay, has become the method of choice for apoptosis investigation. The drawbacks of ISEL are that apoptotic bodies lacking visible chromatin may not stain (false-negative), whereas other cells with DNA strand breaks, including necrotic cells, may be stained (false-positive) (Goldsworthy et al. 1996). ISEL techniques have been compared to H&E, quantitation of apoptosis in the rat liver. Less apoptosis was found by the ISNT technique using DNA polymerase compared to identification by H&E, (Wheeldon et al. 1995), but TUNEL using enzyme predigestion has been shown to be comparable to that with H&E, staining (Kong and Ringer 1995). In other organs, TUNEL has been shown to be more sensitive than ISNT (Gold et al. 1994). Transforming growth factor-β1, another marker for apoptosis in rat liver (Oberhammer and Roberts 1994), has also been demonstrated within GSTP+ foci in rat liver by double labeling immunohistochemistry (Stinchcombe et al. 1995). The utility of ISEL techniques compared to TGF-β1 as markers of apoptosis is currently under investigation (Goldsworthy et al. 1996).
Because DNA fragmentation is associated with various biological and artifactual situations, the ISEL technique is selective (rather than specific) for apoptotic nuclei (Ansari et al. 1993). An optimized TUNEL will find a threshold at which only cells with a high degree of DNA fragmentation (apoptosis) are labeled. The use of pretreatments to optimize TUNEL sensitivity and specificity has been addressed (Gold et al. 1994; Kong and Ringer 1995; Lucassen et al. 1995; Negoescu et al. 1996). Briefly, pretreatments such as proteolytic digestion or HIER methods such as microwave irradiation will increase sensitivity but may also decrease specificity unless used below certain limits of concentration, temperature, and exposure times. The above considerations have led others to recommend standardizing pretreatment and fixation within experiments, accepting apoptotic labeling only if it is strong compared to background and using morphological assessments to support counting of labeled cells or bodies (Negoescu et al. 1996). In addition, use of automated techniques for TUNEL presented here would standardize this procedure and may help to reduce variability between experiments.
Fewer TUNEL-labeled apoptotic cells were observed in this study compared to TUNEL-labeled apoptotic bodies. This was an expected finding, because the appearance of early apoptotic cells is limited to only a few minutes and apoptotic bodies are visible by light microscopy in the liver for a few hours (Goldsworthy et al. 1996). Apoptotic bodies exhibit diverse forms and sizes and can be categorized into chromatin- and nonchromatin-containing bodies (Goldsworthy et al. 1996). Most apoptotic bodies have been engulfed by surrounding hepatocytes and consequently are visualized as intracellular bodies. Some investigations of TUNEL labeling in liver have not made a clear distinction between apoptotic cells vs apoptotic bodies (Hino et al. 1996), since both may contain TUNEL-positive chromatin. Most of the TUNEL-positive nuclei in our tissue have been phagocytized and are therefore classified as apoptotic bodies rather than apoptotic cells, which follows the classification proposed by Goldsworthy et al. (1996).
Diffuse cytoplasmic TUNEL staining observed in this study has been suggested to represent leakage of DNA fragments out of the nucleus of the phagocytized apoptotic body, either as part of the apoptotic process or as an artifact of the subsequent tissue processing and staining procedure (Wijsman et al. 1993; Wheeldon et al. 1995). Occasional bodies did not stain positive, which may indicate that condensed chromatin of some apoptotic bodies is poorly accessible (Wijsman et al. 1993).
The automated BrdU/GSTP and TUNEL/GSTP double labeling methods presented here produce material suitable for image analysis quantitation of proliferation and apoptosis within rat hepatic foci. Automating the quantitation of BrdU- or TUNEL-labeled cells in normal hepatocytes from rat and mouse liver is described, as well as image analysis of hepatocytes expressing GSTP (Carter et al. 1992; Soames et al. 1994; Kong and Ringer 1995,1996), but automated quantitation of these phenomena within rat hepatic foci has not. Use of automated double labeling techniques described here will permit rapid and accurate assessments of proliferation and apoptosis and may aid efforts aimed at automating the quantitation of the net growth potential of preneoplastic and neoplastic lesions.
Footnotes
Acknowledgements
We thank Thomas Covatta for assistance in preparation of the photomicrographs.
