Abstract
The human arylamine N-acetyltransferases (NATs) NAT1 and NAT2 are enzymes responsible for the acetylation of many arylamines and hydrazines, thereby playing an important role in both detoxification and activation of many drugs and carcinogens. Both enzymes show polymorphisms but exhibit key differences in substrate selectivity and tissue expression. In the present study, reverse transcriptase-PCR, Western blotting, and immunohistochemistry were used to investigate the expression of the NATs in human skeletal muscle. Despite the presence of its mRNA, NAT2 enzyme level was below the limit of detection. In contrast, both NAT1 mRNA and enzyme were readily detected in fetal, newborn, and adult muscles. In addition, punctate cytoplasmic and perinuclear NAT1 immunostaining was observed in all tissue sections, the staining being more intense in the fetal tissue. High expression of NAT1 enzyme in fetal muscle was also suggested by Western blotting. Because skeletal muscle accounts for a large proportion of body mass, muscle NAT1 expression may contribute significantly to the total activity in the body. These results further support the involvement of skeletal muscle in the metabolism of xenobiotics.
M
The XMEs comprise oxidative (phase I) and conjugating (phase II) enzymes involved in biotransformation of many chemical stressors. The acetyl-CoA:arylamine N-acetyltransferases (NATs; EC 2.3.1.5) are phase II enzymes that catalyze the transfer of an acetyl group from Ac-CoA to the nitrogen or oxygen atom of primary arylamines, hydrazines, and their N-hydroxylated metabolites. Therefore, the NATs play an important role in the detoxification and potential metabolic activation of many xenobiotics (Upton et al. 2001). The two functional isoforms of human NATs, NAT1 and NAT2, are encoded by two distinct loci located on chromosome 8 (Matas et al. 1997). Genetic interindividual variations in NAT2 protein levels and functional activity lead to the well-known isoniazid acetylation polymorphism. Susceptibility to certain malignancies and drug response phenotypes has been associated with both slow and rapid acetylator phenotypes. NAT1 expression also displays marked interindividual variations that may be a source of pharmacological susceptibility (Hein et al. 2000). NAT1 and NAT2 exhibit marked differences in substrate specificity (Blum et al. 1990) and tissue expression pattern. NAT2 is mainly expressed in liver (Evans and White 1964) and gut (Hickman et al. 1998), whereas NAT1 is found in a wide range of tissues, including liver (Ohsako and Deguchi 1990), colon (Ilett et al. 1994), blood (Irshaid et al. 1993), bladder (Stanley et al. 1996), placenta (Derewlany et al. 1994), skin (Kawakubo et al. 1990), and gingiva (Meisel et al. 2002).
Although the liver is the organ with the primary detoxification functions, skeletal muscle is also involved in the detoxification process. Skeletal muscle cells have been shown to express different types of XMEs, including cytochrome P450 (CYPs) (Smith et al. 2000) and glutathione S-transferases (GSTs) (Hussey et al. 1991). Despite the low level of observed cytochrome P450-dependent metabolism of skeletal muscle microsomes (Crosbie et al. 1997), ethanol consumption has been shown to lead to rhabdomyolysis in humans by inducing skeletal muscle cytochrome P450 expression (Riggs 1998). The relative amount of GSTs in skeletal muscle is only approximately 4% that of the liver. However, because skeletal muscle accounts for 40% of body mass, the GST activity of the liver is estimated to be similar to that of skeletal muscle (Meyer and Ketterer 1995). Therefore, skeletal muscle appears to be a potentially important component of extra-hepatic XME activity.
These observations prompted us to analyze whether NAT enzymes are expressed in human skeletal muscle. We carried out RT-PCR and immunoblotting on muscle homogenates. Immunohistochemical (IHC) analyses of NAT expression was also carried out on fetal, newborn, and adult human skeletal muscle biopsy specimens.
Materials and Methods
Antibody Preparation
Polyclonal antibodies that distinguish between human NAT1 and NAT2 were obtained in rabbits using an antipeptide strategy in which antisera were raised against the divergent C-termini of human NAT1 and NAT2. These antibodies were previously described and used for analysis of NATs expression (Stanley et al. 1996). IgGs were purified from sera using protein A–Sepharose affinity chromatography (Sigma; Saint-Quentin Fallavier, France) and stored in PBS buffer containing 20% glycerol (v/v) and protease inhibitors (Roche; Meylan, France).
Muscle Biopsies
Biopsies from the quadriceps muscle were obtained during diagnostic or surgical procedures in accordance with French legislation on ethical rules. All samples were derived from individuals with no history of muscle disease. Fetal tissues were obtained immediately after abortion induced by mifepristone (RU486). A postmortem biopsy of a 37-week-old infant and biopsies from a 57- and a 77-year-old were obtained during surgical procedures.
RNA Isolation and Reverse Transcription-polymerase Chain Reaction (RT-PCR)
Total RNA was isolated from skeletal muscle using TRIzol (Gibco BRL; Cergy–Pontoise, France) according to the manufacturer's instructions. For cDNA synthesis, 6 μg of total RNA was used at 70C in a final volume of 20 μl using the C. therm Polymerase RT-PCR kit (Roche). For PCR analysis, 10 μl of cDNA was used as a template in a 50-μl amplification mixture containing 200 μM of each deoxynucleotide triphosphate, 10 mM Tris-HCl, pH 8.3, 1.5 mM MgCl2, 50 mM KCl, 0.3 μM of each primer, and 2.5 U of Expand High Fidelity DNA polymerase (Roche). The NAT primers used in the PCR were as follows: for human NAT1 (specific 387-bp fragment), sense 5'-tagaagacagcaaatacc-3', antisense 5'-agttgataactggtgagc-3'; for human NAT2 (specific 526-bp fragment), sense 5'-tgccaaagaagaaacacc-3', antisense 5'-tgaaaatgtgtccttatg-3'. The PCR conditions were as follows: 35 cycles each consisting of 1-min annealing at 47C, 1-min extension at 72C, and 1-min denaturation at 94C. To assess whether the RNA samples were contaminated by genomic DNA, an additional PCR reaction was carried out under the same conditions with primers for human β-crystallin (CRYAB); sense 5'-aaggagctgaccagccagct-3', antisense 5'-actggtggggaaactttcttg-3'. According to the intron/exon organization of the human αB-crystallin gene, a specific 623-bp fragment is generated from cDNA, whereas a 3055-bp fragment would be amplified from genomic DNA. Absence of genomic DNA contamination was also confirmed using a negative control with no reverse transcriptase. The PCR products were separated by gel electrophoresis in ethidium bromide-stained agarose (1.8%).
Protein Sample Preparation, SDS-PAGE, and Immunoblot Analysis
Muscle proteins were extracted by crushing muscle biopsy specimens (20–30 mg) in 500 μl Tris-HCl 25 mM, pH 7.5, 1 mM DTT, 1 mM EDTA, 0.2% SDS, 0.5% Triton X-100, and protease inhibitors. To selectively separate proteins in the muscle extracts from contaminating species, such as nucleic acids and lipids, that would otherwise interfere with electrophoresis, trichloroacetic acid (TCA) precipitation was used. In addition, TCA precipitation allows more proteins to be loaded in acrylamide gels. Briefly, cold TCA was added (to a final concentration of 25%) to 250 μl of muscle extracts (protein concentration adjusted to 3 mg/ml) and the proteins were allowed to precipitate on ice for 30 min. The mixture was then centrifuged for 30 min at 15,000 × g and the protein pellet washed with cold acetone to remove residual TCA. After centrifugation (30 min, 15,000 × g), proteins were resuspended in a small volume of 4 × SDS sample buffer (30–40 μl) and heated at 100C before SDS-PAGE (10% acrylamide) under reducing conditions. After electrophoresis, proteins were transferred to a nitrocellulose membrane. Immunodetection of NAT1 or NAT2 isoform was carried out using the polyclonal antibodies described above at a dilution of 1:1500. Visualization of immunoreactivity was performed using a secondary goat anti-rabbit IgG conjugated with horseradish peroxidase. Recombinant human GST-NAT1 and GST-NAT2 proteins were used as controls.
Immunohistochemical Analysis
All experiments were performed on 8- μm frozen transverse or longitudinal cryostat sections. Double immunofluorescence was performed using the purified polyclonal anti-NAT1 antibody (1:100) with either a monoclonal antibody against the slow myosin heavy chain (1:5) (NCL-MHCs; Novacastra, Newcastle upon Tyne, UK) or a monoclonal antibody against the intermediate filament protein, desmin (1:200) (D33; Dako, Trappes, France). Briefly, on unfixed sections, primary antibodies were simultaneously incubated overnight at 4C. The anti-NAT1 antibody was visualized with an AlexaFluor 488 (Molecular Probes; Montluçon, France) directly coupled to an anti-rabbit secondary antibody, and the anti-slow and anti-desmin antibodies were visualized with an AlexaFluor 594 directly coupled to an anti-mouse secondary antibody. Secondary antibodies in the absence of the primary antibodies were used as negative controls. Finally, to visualize nuclei, the sections were mounted in medium (Mowiol; Calbiochem–Novabiochem, San Diego, CA) containing bis-benzimide (0.0001% w/v, Hoechst no. 33258; Sigma, St Louis, MO). All images were digitalized using the MetaView image analysis system.
Results
RT-PCR Analysis
We initially investigated the expression of NAT1 and NAT2 in human skeletal muscle using RT-PCR. The primer pairs were designed to yield PCR products of different sizes (387 bp for NAT1 and 526 bp for NAT2). Both NAT1 and NAT2 PCR-derived bands were detected in two adult muscle samples (Figure 1, Lanes 1 and 4 for NAT1; Lanes 2 and 5 for NAT2). Identical results were obtained from mRNA extracted from a third adult muscle biopsy specimen (data not shown). These results strongly suggest that NAT1 and NAT2 mRNA are present in adult skeletal muscle. The co-amplification of αB-crystallin (CRYAB) mRNA revealed no genomic DNA contamination (Figure 1, Lanes 3 and 6). Absence of genomic DNA contamination was also confirmed using a negative control with no reverse transcriptase in the RT experiment (Figure 1, Lane 7; data not shown for other subjects).
Detection of NAT Expression by Western Blot Analysis
To study NAT expression at the protein level, we used specific NAT antibodies. We first investigated whether NAT1 and NAT2 proteins were present in human skeletal muscle extracts. To do this, fetal (n=2) and adult (n=2) muscles were used to prepare total protein extracts at a concentration of 3 mg/ml. Equal amounts of extracts were then concentrated by TCA precipitation to selectively separate the proteins from the contaminating species (e.g., nucleic acids, lipids) and to increase the protein fraction of the samples. Western blot analysis with the specific antibody against human NAT1 revealed the presence of NAT1 protein in all of the skeletal muscle protein extracts tested (Figure 2). In addition, the results suggested that NAT1 protein was expressed at a high level in fetal tissue (Figure 2, Lanes 1 and 3). The expression of NAT2 protein in these samples was also studied by Western blot analysis. However, the NAT2 protein was not detectable at the level of sensitivity attained in these experiments (data not shown).
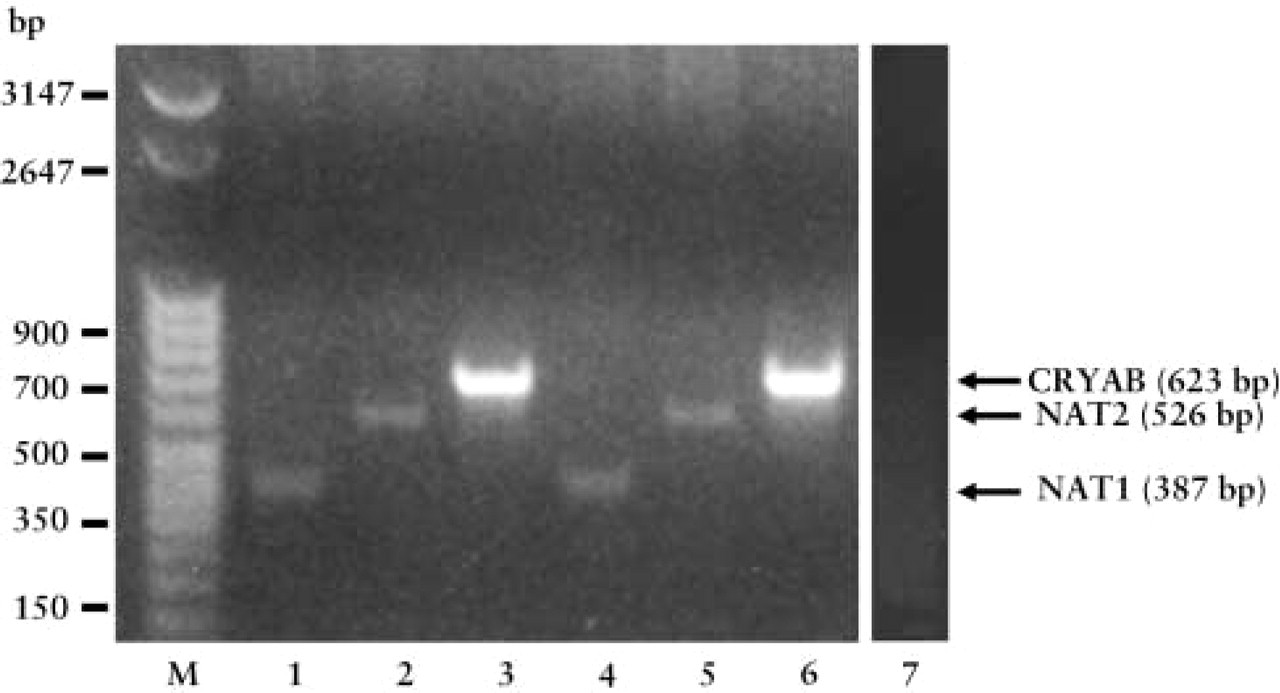
Sequence-specific primers for human NAT1 and NAT2 were used for RT-PCR of total RNA from human adult skeletal muscle (two subjects). Specific primers for the human CRYAB gene were used as controls for genomic DNA contamination (see Materials and Methods). RT-PCR products were revealed on ethidium bromide-stained agarose gels (1.8%) after electrophoresis. A characteristic 387-bp fragment was obtained for NAT1 (subject 1, Lane 1; subject 2, Lane 4), and a 526-bp fragment was obtained for NAT2 (subject 1, Lane 2; subject 2, Lane 5). A specific 623-bp fragment for CRYAB (subject 1, Lane 3; subject 2, Lane 6) was obtained as a positive control. Absence of genomic DNA contamination was also confirmed using a negative control with no reverse transcriptase in the RT experiment (subject 1, Lane 7).
Immunohistochemistry
To further analyze the expression of NAT1 protein in human skeletal muscle, IHC experiments were performed on fetal, newborn, and adult muscle sections. In transverse sections, the most intense NAT1 immunostaining was observed in fetal muscle (Figure 3A) whereas weaker immunostaining was obtained in newborn and adult muscles (Figures 3B and 3C, respectively). To determine whether the localization of NAT1 was fiber type-specific, double immunofluorescence staining with the slow myosin heavy chain antibody was carried out (Figures 3D–3F). The results obtained demonstrate that NAT1 is evenly distributed within slow and fast fibers, with no preferential localization in either type of muscle fibers. The localization of NAT1 immunostaining in fetal muscle was similar to that observed in newborn and adult muscles, with punctate cytoplasmic staining but also with a punctate perinuclear arrangement (Figures 4A–4F). When immunostaining was performed in the absence of the primary antibody, no staining of the muscle sections was visible (data not shown). These results were consistent with the Western blot data that suggest a strong expression of NAT1 protein in fetal tissues (Figure 2). To further characterize the immunostaining of NAT1, we used an antibody against desmin, a Z-line-associated intermediate filament that appears as collars surrounding Z-discs of the myofibril (Ip 1999). The subcellular localization of NAT1, in relation to desmin, was investigated by carrying out double immunofluorescence staining on longitudinal sections of adult muscle. The localization of NAT1 appeared punctate and striated (Figure 5A). NAT1 (Figure 5A) and desmin (Figure 5B) immunostainings are clearly distinct (Figure 5D), although both appeared in a striated pattern. This indicated that NAT1 is unlikely to be localized at the Z-lines.
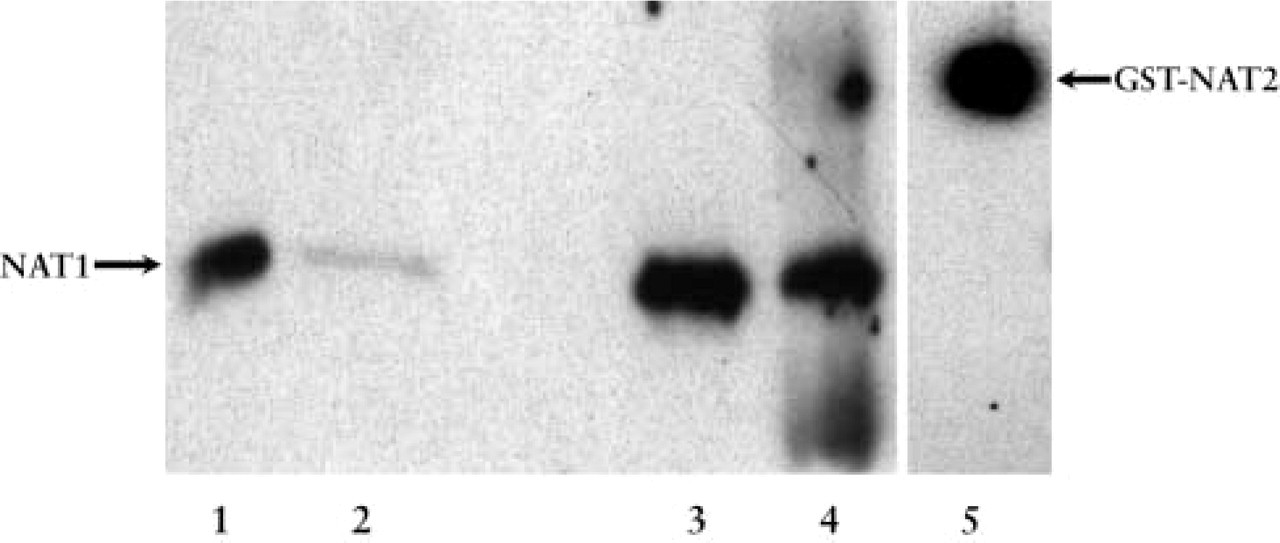
Total protein extracts from human fetal and adult skeletal muscle were used for Western blotting with specific polyclonal antibodies against human NAT1 and NAT2. Equal amounts of protein extracts were precipitated by TCA and then subjected to SDS-PAGE under reducing conditions. After transfer to a nitrocellulose membrane, the primary antibodies were used at 1:1500 dilution. Immunoreactivity was detected by the SuperSignal Kit (Pierce; Rockford, IL). Expression of NAT1 protein was detected in two fetal muscle extracts (subject 1, Lane 1; subject 2, Lane 3) and in two adult muscle extracts (subject 1, Lane 2; subject 2, Lane 4). No protein was detected using the specific anti-NAT2 antibody (data not shown). A positive control of the anti-NAT2 antibody was made using recombinant GST-NAT2 (Lane 5).
Discussion
We have demonstrated by RT-PCR, Western blotting, and IHC, that human skeletal muscle synthesizes the arylamine N-acetyltransferase NAT1. Although RT-PCR revealed that the NAT2 mRNA is also present in human skeletal muscles, its expression product was not detected by immunoblotting. Such a difference between the expression profiles of NAT1 and NAT2 has been reported in other cell types. Both NAT1 and NAT2 mRNA have been detected in liver, gastrointestinal tract tissues, ureter, bladder, lung (Windmill et al. 2000), and mammary gland (Sadrieh et al. 1996). Nevertheless, the expression of NAT2 enzyme has been demonstrated only in the liver and colon epithelium, whereas the NAT1 enzyme is widely distributed throughout the human body (Sim et al. 2000). RT-PCR is the most sensitive method for detection of low-abundance mRNA (Wang et al. 1989). Therefore, it can be suggested that the NAT2 gene is expressed in human muscle at a very low level, which implies that NAT1 is the predominant isoform found in human skeletal muscle.
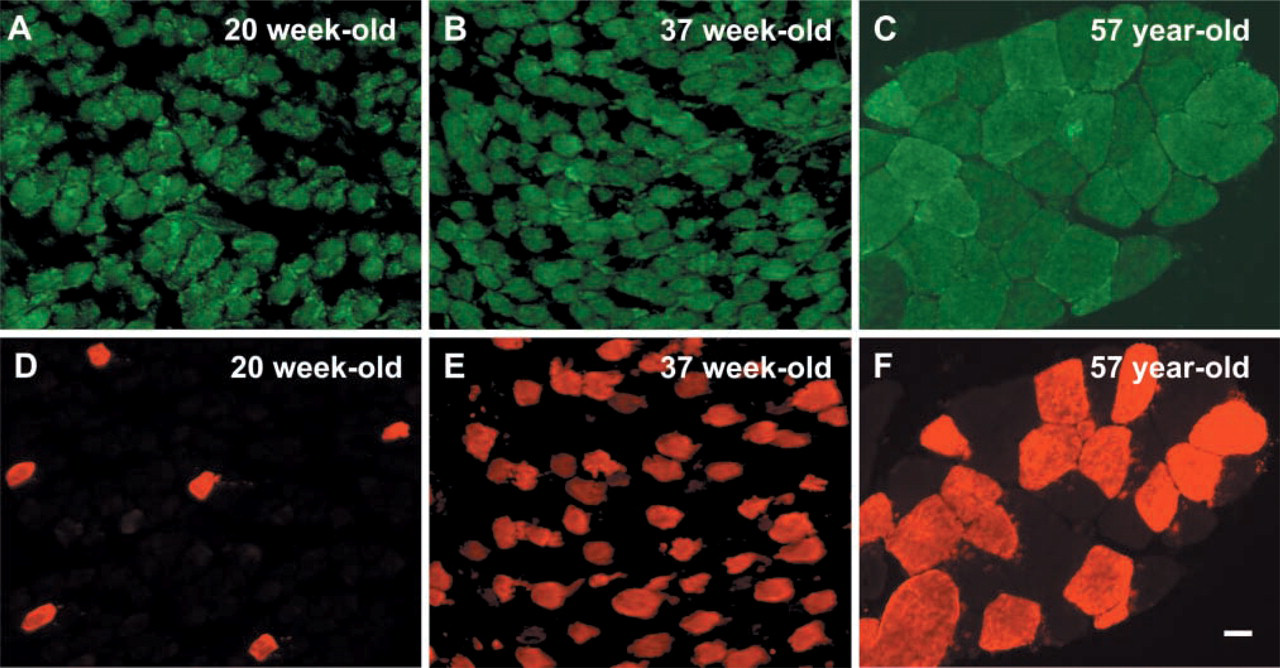
Double immunofluorescence staining of NAT1 (
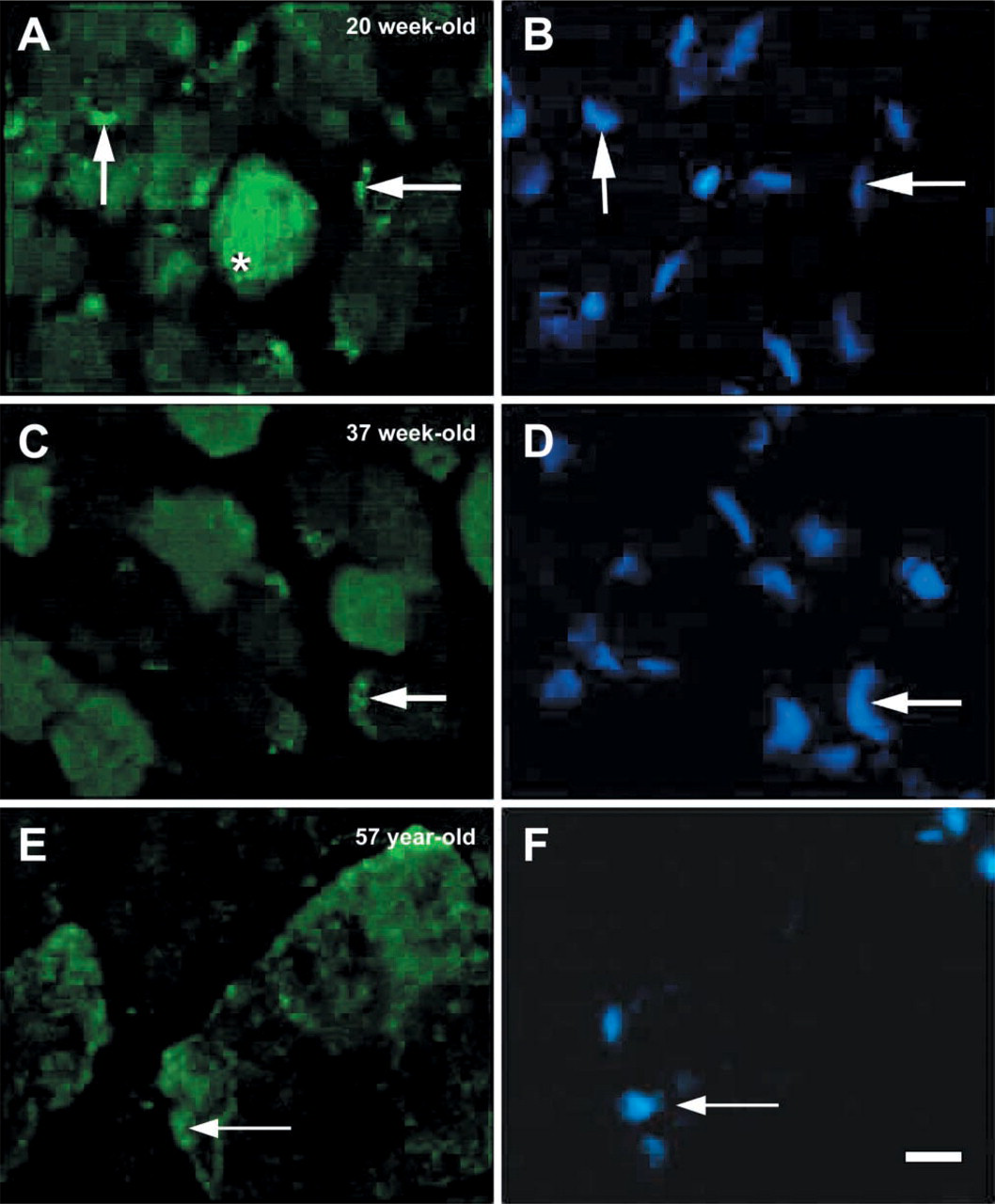
Immunofluorescence staining of NAT1 in transverse cryostat sections of human fetal (
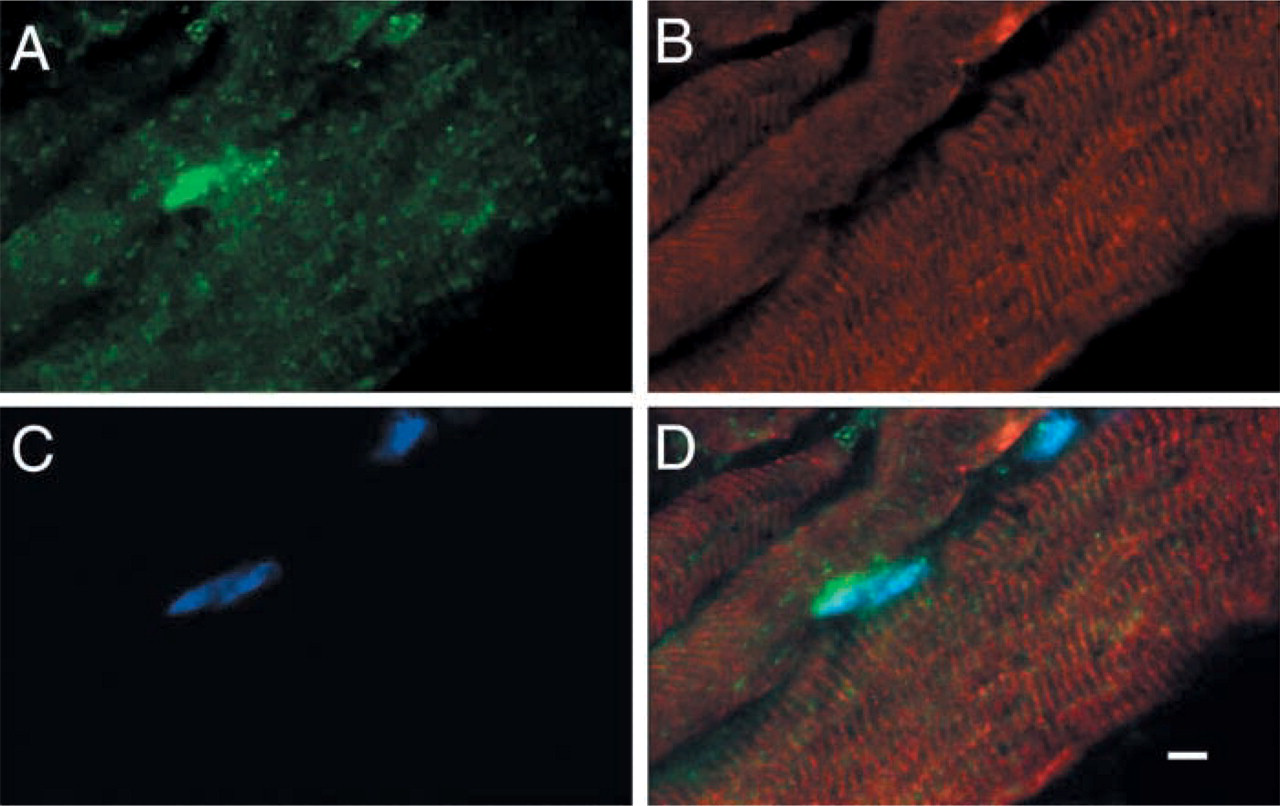
Double immunofluorescence staining of NAT1 (
NATs are polymorphic enzymes and the differences in immunoreactive recombinant protein variants have been reported in yeast expression systems. Thus, NAT1 variants encoded by NAT1∗14B, NAT1∗15, NAT1∗17, NAT1∗19, and NAT1∗22 have been shown to exhibit NAT1 protein expression levels below the limit of detection as measured by Western blotting (Fretland et al. 2001a). Nevertheless, these alleles account for less than 3% of NAT1 alleles found in whites (Hein et al. 2000). Similarly, reductions in the expression of NAT2-immunoreactive protein were observed for NAT2 variants possessing T341C (13fold reduction), A434C (50-fold reduction), or G590A (threefold reduction) variations (Fretland et al. 2001b). Given the relative frequency of T341C and G590A-containing alleles in whites (45% and 28%, respectively) (Hein et al. 2000), the lack of detection of NAT2 in skeletal muscle may be due to samples carrying genotypes combining such alleles. Overall, we cannot rule out that the expression of NAT1 and NAT2 enzymes in skeletal muscle is underestimated in our study.
Our results suggest that NAT1 is expressed at a high level in fetal compared to newborn and adult skeletal muscle (Figures 2 and 3). In contrast, the level of expression of several metabolic enzymes is developmentally regulated, with increased levels being present in the adult (Davidson et al. 1983). NAT1 activity has been reported in other fetal tissues, including the lungs, kidneys, and adrenal glands (Pacifici et al. 1986). Such a fetal extrahepatic activity has also been shown in rodents (Sonawane and Lucier 1975). More recently, using RT-PCR, Smelt and colleagues (2000) have demonstrated NAT1 expression in pre-implantation embryos at the blastocyst stage, as well as in human placenta as early as 5.5 weeks of gestation. Earlier placental expression cannot be ruled out (Smelt et al. 2000). Overall, prenatal NAT1 expression appears to be among the earliest xenobiotic-metabolizing enzyme genes whose expression has been detected (Smelt et al. 2000). Altogether, our data suggest a decrease in NAT1 expression during skeletal muscle differentiation.
Skeletal muscle enzymes, such as β-enolase (Fougerousse et al. 2002), and amorphin (Chowrashi et al. 2002) are present at the Z-line or M-band of skeletal muscle, resulting in a striated immunoreactivity in longitudinal sections. Our double immunolabeling results (Figure 5) on longitudinal adult muscle sections showed a distinct striated pattern for both NAT1 and desmin, a Z-line-associated filament (Ip 1999). This absence of co-localization indicates that NAT1 is unlikely to be present in the Z-line of mature muscle. Further experiments using immunoelectron microscopy on ultrathin sections are needed to clarify whether the striated pattern of NAT1 is associated with M-bands.
The early expression of NAT1 and its persistence in adult skeletal muscle have potential pharmacological consequences. Because skeletal muscle constitutes a large proportion of total body mass, it is likely that the total activity of NAT1 in muscle contributes significantly to the total activity in the body, as suggested for other xenobiotic-metabolizing enzymes (Meyer and Ketterer 1995).
It has been proposed that human NAT1 plays a role in folate catabolism through the acetylation of the folate para-aminobenzoylglutamate (Minchin 1995). Given the ability of maternal folate supplementation to protect against neural tube defects (NTDs), NAT1 expression may be implicated in the correct developmental patterning of the neural tube (Smelt et al. 2000). Arylamine N-acetyltransferases are also involved in the metabolic activation of a variety of carcinogenic compounds, such as benzidine and β-naphthylamine (Guengerich 1992). Epidemiological studies have found associations between NAT1 polymorphisms and certain malignancies (Hein et al. 2000). Nevertheless, these findings await further association studies, together with other susceptibility genes and quantitative estimates of carcinogen exposure (Hein 2002). The association between arylamine N-acetyltransferases and drug response has been extensively investigated (Meisel 2002), and it has been hypothesized that NAT1 could affect the efficacy and side effects of 5-aminosalicylic acid, a drug commonly used to treat inflammatory bowel diseases (Hickman et al. 1998). Nevertheless, no association with NAT1 genotype has been reported thus far (Ricart et al. 2002). Considering the potential impact of drug toxicity in development, the prenatal pattern of neuromuscular NAT1 activity should be examined.
Variations in muscle cell activity related to aging (Pansarasa et al. 1999), chronic immobilization (Kondo et al. 1993), or exercise (Atalay et al. 1996) modulate the activities of antioxidant enzymes such as cytosolic and mitochondrial superoxide dismutases, catalase, glutathione S-transferase, glutathione reductase, and glucose-6-phosphate dehydrogenase. Ethanol consumption has also been shown to induce CYP1A1/2 in rat skeletal muscle (Smith et al. 2000), which supports the potential role of ethanol in the pathogenesis of alcohol-associated rhabdomyolysis in humans (Riggs 1998). In contrast, NATs are generally believed to be constitutively expressed enzymes (Meisel 2002). However, Butcher and colleagues (2000b) have provided evidence for a substrate-dependent downregulation of NAT1 in cultures of human peripheral blood mononuclear cells, which suggests an effect on drug toxicity and cancer susceptibility. These investigators have also shown that the hydroxylamine of the NAT1-selective substrate p-aminobenzoic acid irreversibly inactivates NAT1 both in cultured cells and cell cytosols, possibly via the formation of covalent modifications at the co-factor-binding site (Butcher et al. 2000a). Although the exact mechanisms remain to be elucidated, these findings show that genetic controls account for only part of the observed variability in NAT1 activity (Hughes et al. 1998). The exact relationship between NAT1 activity in muscle cells and clinical or toxicological parameters needs to be further investigated.
In conclusion, the expression of NAT1 in muscle cells may be an important factor in the detoxification/activation processes because of the potential involvement of the muscle in the pharmacokinetics of many xenobiotics (Khazaeinia et al. 2000). In a broader sense, the involvement of other muscular xenobiotic-metabolizing enzymes to pharmacological insults appears as a potential target for future pharmacogenetic studies.
Footnotes
Acknowledgements
Supported by grants from the Association Française contre les Myopathies, Alliance program, and the Association pour la Recherche contre le Cancer.
