Abstract
Polymorphonuclear white blood cells from patients with low serum vitamin B12 (cobalamin) have ultrascopic appendages that project from their nuclear membranes. These appendages are most often found in the shape of blebs and stalks. Cytoplasmic rings that may be separated from the nucleus have also been seen. There is no known function for these appendages. Blood from 11 patients with low serum B12 was processed for electron microscopic examination. In situ end-labeling of DNA and subsequent electron microscopic examination were performed. DNA was localized in all of the visualized appendages and rings. Treatment with DNases I and II decreased the labeling of these appendages by 63%. These DNA-laden appendages are a unique ultrastructural finding and may function to transfer fragmented DNA, which occurs in vitamin B12 deficiency, from the nucleus into the cytoplasm.
Keywords
U
There has been no speculation about the possible function of these appendages. All of the conditions in which appendages have been identified are also characterized by disorders of DNA. It is possible that the role of these appendages is to remove or isolate abnormal DNA from the nuclear body. Eleven patients with a wide range of abnormal serum B12 values were selected for study from a bank of such patients. Although there is no single definitive test for the diagnosis of vitamin B12 deficiency (Stabler et al. 1990), all patients had low serum B12 levels and most had other positive tests used to confirm the diagnosis, including serum methylmalonic acid, serum homocysteine, serum intrinsic factor antibody, the Schilling test, and nuclear hypersegmentation on routine blood smear. Blood from these patients was prepared by buffy coat for electron microscopic examination and the thin sections were labeled with an in situ DNA end-labeling method (Migheli et al. 1995) to determine if DNA is associated with appendages.
Materials and Methods
Clinical Parameters
Eleven patients with low serum B12 values were selected for study from a bank of such patients kept in the Pathology Department of the Massachusetts General Hospital, Boston. The patients were selected to include a wide range of abnormal serum B12 values and no other selection criteria were used. The hospital Human Studies Committee approved this study. Age and gender were recorded. Routine hematological parameters were determined by standard methods. Serum intrinsic factory antibody (IFAB), serum methylmalonic acid, and homocysteine were drawn before the administration of vitamin B12 in the Schilling test.
Smears of peripheral blood were blinded and lobe counts made using the method of Nath and Lindenbaum (1979). The number of band forms was recorded although not used in calculating the mean lobe count (MLC). The segmentation index (SI) of Edwin (1967) was also calculated.
Sample Preparation
Blood was drawn in EDTA tubes before the administration of the Schilling test and immediately processed for electron microscopic studies. Samples were centrifuged at 1000 rpm for 10 min at room temperature (RT), pipeting off the plasma and gently adding Karnovsky's II fixative to the top of the buffy coat. After 1.5 hr the buffy coat was removed intact from the tube and fixed for another 1.5 hr in a petri dish. It was then cut into full-thickness segments and stored temporarily in cacodylate buffer. Segments were then post-fixed with osmium tetroxide, stained en bloc with uranyl acetate, dehydrated in graded ethanol solutions, infiltrated with propylene oxide/epon mixtures, and embedded in epon. One-μm sections were cut, stained with toluidine blue, and examined using the light microscope. The number of apoptotic cells/5000 granulocytes was counted on these preparations. After representative areas were chosen, thin sections were cut and mounted on nickel grids.
Immunoelectron Microscopy to Detect DNA
Grids were labeled using the method of Migheli et al. (1995). The grids were rinsed in terminal deoxynucleotidyl transferase (TdT) buffer (25 mM Tris-HCl, 200 mM sodium cacodylate, 5 mM cobalt chloride) and then incubated with 2 U of TdT (Boehringer Mannheim; Mannheim, Germany) and 0.5 nM digoxigenin-11-dUTP (Boehringer Mannheim) in 100 μl TdT buffer for 60 min at 37C. The reaction was terminated by rinsing with 300 mM sodium chloride-30 mM sodium citrate for 30 min at RT. The grids were briefly rinsed in Tris-buffered saline (TBS) and then incubated at RT for 15 min with bovine serum albumin (BSA) in TBS (10 mg BSA/1 ml TBS). The grids were then incubated for 60 min at RT with anti-digoxigenin antibody from sheep conjugated to ultrasmall (1-nm) colloidal gold (Boehringer Mannheim), diluted 1:30 in TBS-BSA. The grids were then washed in TBS, followed by rinsing in double-distilled water. The gold particles were silver-enhanced using a kit (Boehringer Mannheim) by incubation for 7–10 min at RT. The grids were then washed in double-distilled water and stained with 2% uranyl acetate for 20 min, washed in double-distilled water, and stained with lead citrate for 10 min, followed by washing in double-distilled water. The grids were examined in a Phillips 301 electron microscope.
Controls were made by omitting the TdT, digoxigenin-11-dUTP, or both.
DNase Treatment
A DNase I reaction mixture of 10 ml 0.1 M sodium acetate buffer, pH 5.0, and 0.5 ml 0.1 M magnesium sulfate was prepared. Five mg DNase I (deoxyribonucleate 5′-oligonucleotidohydrolase from bovine pancreas; Roche Diagnostics, Chicago, IL) was dissolved in 0.5 ml double-distilled water and added to 9.5 ml of the reaction mixture. Grids from two patients were incubated on this solution at RT for 30 min and then rinsed in double-distilled water. The grids were then DNA-labeled as described above.
A DNase II reaction mixture of 0.75 ml 1 M sodium acetate buffer, pH 4.6, 0.375 ml 20 mM magnesium sulfate, 6.375 ml double-distilled water was prepared. To 2.5 ml of this mixture was added 0.4 ml 150 mM NaCl in water. To this latter solution was added either 0.1 ml DNase II (deoxyribonucleate 3′-oligonucleotidohydrolase from porcine spleen; Worthington Biochemicals, Lakewood, NJ) solution, (2 mg dissolved in 2 ml water) or 0.1 ml water as a control. Grids from two patients were incubated on these solutions at RT for 100 min and then rinsed in double-distilled water. The grids were then DNA-labeled as described above.
Grids that had been treated with DNase II and control solution as described above were then treated with DNase I and control solution as described above and then rinsed in double-distilled water. The grids were then DNA-labeled as described above.
The mean number of colloid labels per whole polymorphonuclear white cell was determined on grids that had not been treated with any DNase, grids that had been treated only with DNase I, grids that had been treated only with DNase II, and grids that had been treated with both DNases I and II. In addition, the mean number of colloid labels per appendage was counted on 22 appendages from grids that had not been Dnase-treated and 16 from Dnase I and II-treated grids.
Results
There were eight male and three female patients with a mean age of 77.4 years, range 58–88 years. The mean serum B12 was 138.6 ± 57.1 pg/ml, range 13–202 pg/ml (normal 220–900 pg/ml). All had normal serum folate greater than 3.0 ng/ml, range 4.3–12.1 ng/ml. All but three of the patients were anemic (hemoglobin less than 14.0 g/dl for men and 12.0 g/dl for women) with a mean hgb of 11.95 ± 1.5 g/dl, range 10.1–14.8 g/dl. One patient had a low white blood cell count (less than 5000/mm3) and platelet count (less than 150,000/mm3); one had an elevated platelet count (greater than 400,000/mm3). Four patients had elevated mean corpuscular volume (normal 94 fl or less for men and 99 fl or less for women). One patient had a positive IFAB and another was indeterminate. Three of nine patients had elevated methylmalonic acid levels (greater than 270 nmol/liter), range 112–93,242 nmol/liter, and seven of nine patients had elevated homocysteine levels (greater than 16 micromoles/liter), range 8.8–197 micromoles/liter. Five patients had abnormal Schilling tests, part 1 (less than 10% excretion). These results did not correct to normal with intrinsic factor in part II of the test in the two patients who were tested. Nuclear hypersegmentation was found in four patients when measured by the MLC (greater than 3.5 lobes per cell) and in the same patients and one other when measured by the SI (hypersegmentation defined as greater than 31.5). Ultrascopic nuclear appendages on polymorphonuclear white blood cells were found in all of the patients and the prevalence varied from 2.0 to 21.2% of the cells examined. Abnormal mitochondria shaped like “water wings” (Bunting et al. 1996) were present in five patients, and spontaneously occurring apoptotic cells (Bunting et al. 1997) were found in peripheral blood of 10 patients. No appendages were seen in lymphocytes.
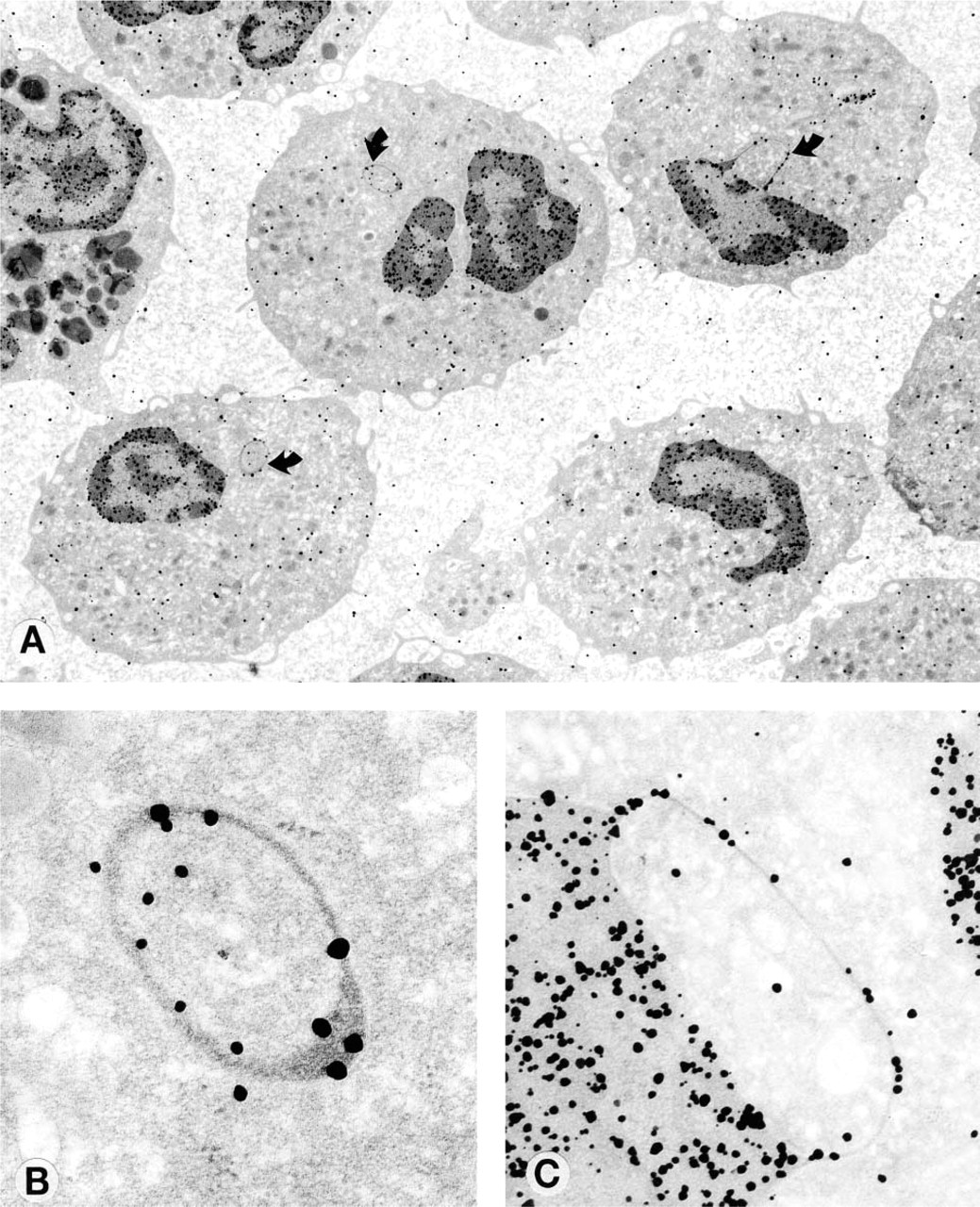
White cell nuclear appendages from patients with low serum B12 with DNA labeling. (
Electron microscopic evaluation of the labeled cells showed that DNA was associated with all of the appendages examined. The number of colloid labels varied from a minimum of two to a maximum of 30 per appendage. The morphology of the appendages was varied. All blebs showed DNA labeling on the bleb (Figure 1C). Some blebs showed accumulations of labeled DNA at one end (Figure 2A, arrow), and the morphology of these blebs often suggested that they were about to rupture at this end of the bleb and form a filament (Figure 2A, arrow). Filaments that were clearly separated from the nuclear membrane (Figure 2B) were also visualized; these forms always showed DNA labeling at the end of the filament. Other blebs appeared as rings that were about to be extruded into the cytoplasm (Figure 2C). DNA was also found in one appendage in the shape of a balloon (Figure 2D). Many rings appeared to be isolated from the main body of the nucleus (Figure 1A), but serial sectioning of one bleb (Figures 3C and 3D) showed the bleb as a ring. No labeling of cells or appendages was seen in controls.
DNase Treatment
Figure 4 shows the results of DNase treatment of the grids before DNA labeling, shown as percent of the value from untreated grids. Labeling decreased by 22.6% (p<0.05) with DNase I treatment alone, 37.8% (p<0.01) with DNase II treatment alone, and 59.3% (p<0.01) when grids were pretreated with DNases I and II before labeling. Figure 5 shows the mean number of colloid labels in appendages from grids that had been treated with DNases I and II and untreated. There were four blebs, 12 rings, and six filaments in the no-treatment group and four blebs, six rings, and six filaments in the DNase I and II-treated group (no significant difference). The number of colloid labels per appendage decreased from 12.7 ± 6.7 in the no-treatment group to 4.0 ± 2.9 in the DNase I and II-treated group (p<0.01). Figure 3 shows the results of the same cell and ring appendage DNA-labeled without treatment with DNase (Figure 3A) and also pretreated with DNases I and II (Figure 3B) before labeling.
Discussion
Nuclear appendages that isolate genetic material have previously been described. Barr and Bertram (1949) reported finding a bleb on the large nucleolus that occurs in neurons of the mature cat. They noted that this appendage was found only in females, suggesting that it contains an × chromosome. These observations were later extended to human skin (Barr et al. 1950). Similar findings in polymorphonuclear leukocytes from human females were reported by Davidson and Smith (1954), who described appendages in the shape of drumsticks that extend from the nuclear membrane of these cells. These appendages are visible with the light microscope. The number of drumsticks in a cell is always one less than the total number of × chromosomes in that cell, and the hypothesis that the drumsticks contain an × chromosome was later proved using an × chromosome-specific nucleic acid probe and an in situ hybridization technique (Hochstenbach et al. 1986). It has been shown that the inactive × chromosome is peripherally located (Comings 1980), and it is possible that the × chromosome isolated in the drumsticks may be inactive. There is currently no understanding of the chemical or physical mechanisms that produce this isolation of the × chromosome by appendages.
The removal of DNA from the nucleus by the formation of nuclear blebs has recently been reported by Shimizu et al. (1998) using a human colon cancer cell line in tissue culture. Double-minute chromosomes are circular, autonomously replicating DNA fragments that occur in colon and other cancer cells and mediate oncogene amplification. These extrachromosomal structures do not contain a centromere or telomere and are localized at the nuclear periphery and replicate there. These DNA structures are removed from the nucleus when they are entrapped in a pouch or bud that forms in the nuclear membrane during S phase. These buds are later separated from the nuclear membrane, forming a micronucleus in the cytoplasm. It was later shown that the double minutes could also be extruded from the cytoplasm into the extracellular fluid (Shimizu et al. 2000). The mechanism by which this DNA extrusion occurs is unknown, and it has not been studied using ultrastructural techniques.
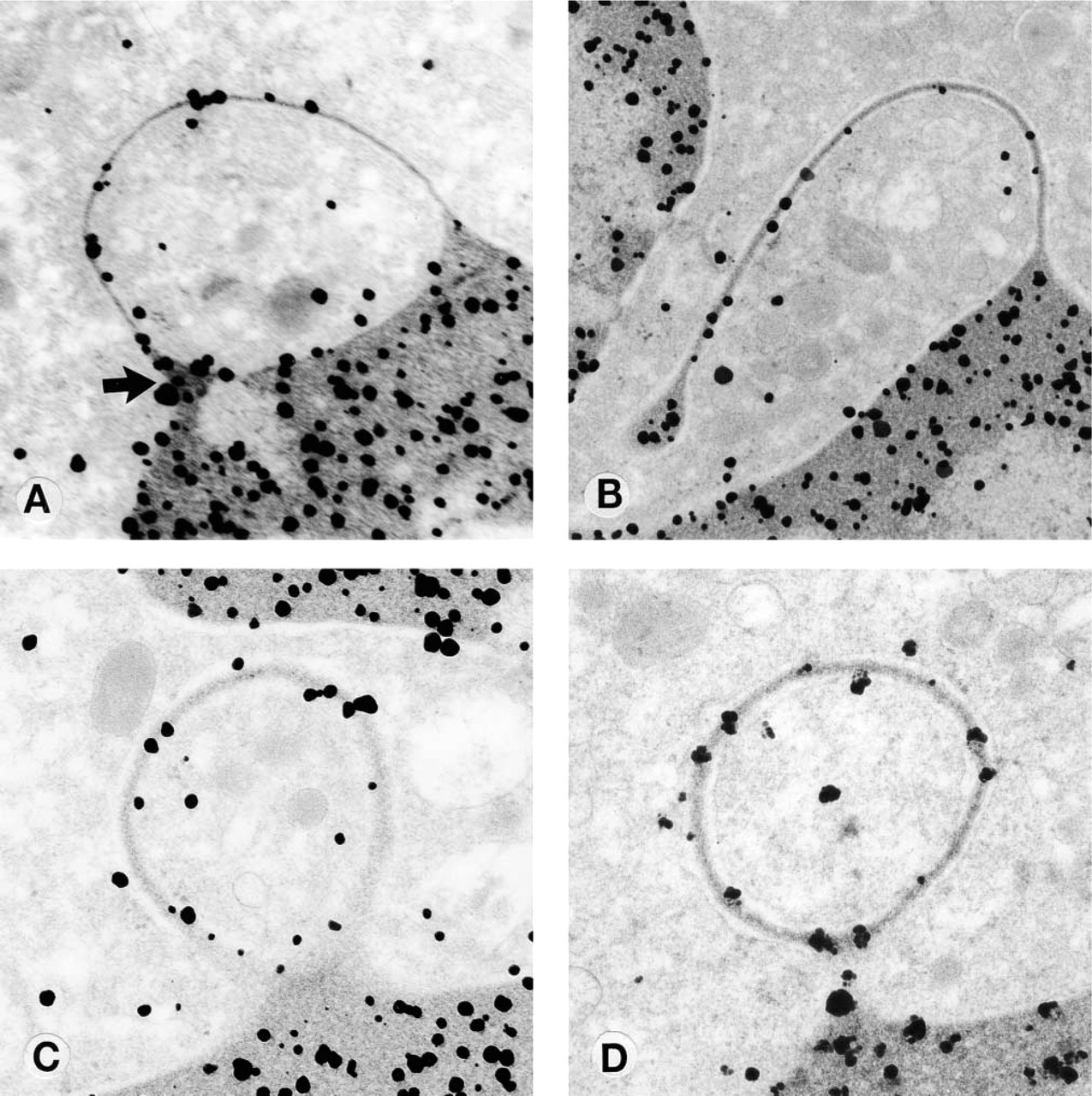
White cell nuclear appendages from patients with low serum B12 with DNA labeling. (
Extranuclear DNA has also been previously studied in folate- and vitamin B12-deficient populations (Fenech et al. 1997, 1998). Micronuclei are thought to form during mitosis when lagging chromosome fragments are excluded from the nucleus as the nuclear membranes reform at the end of mitosis, and the prevalence of cytoplasmic micronuclei has been used as a measure of chromosome breakage (Heddle and Carrano 1977). Fenech et al. (1997) determined the folate, vitamin B12, and homocysteine status of 64 healthy men between the ages of 50 and 70 and correlated these results with the prevalence of chromosome breakage, as measured by the prevalence of cytoplasmic micronuclei in lymphocytes. They found three subjects who were deficient in vitamin B12 and, although their micronucleus results were not reported specifically, it was reported that the percentage of micronuclei significantly increased as serum levels of B12 decreased, even in those subjects who were not B12 deficient. In a later study, similar results were obtained in a study population of 106 young men and women aged 18–32 years (Fenech et al. 1998).
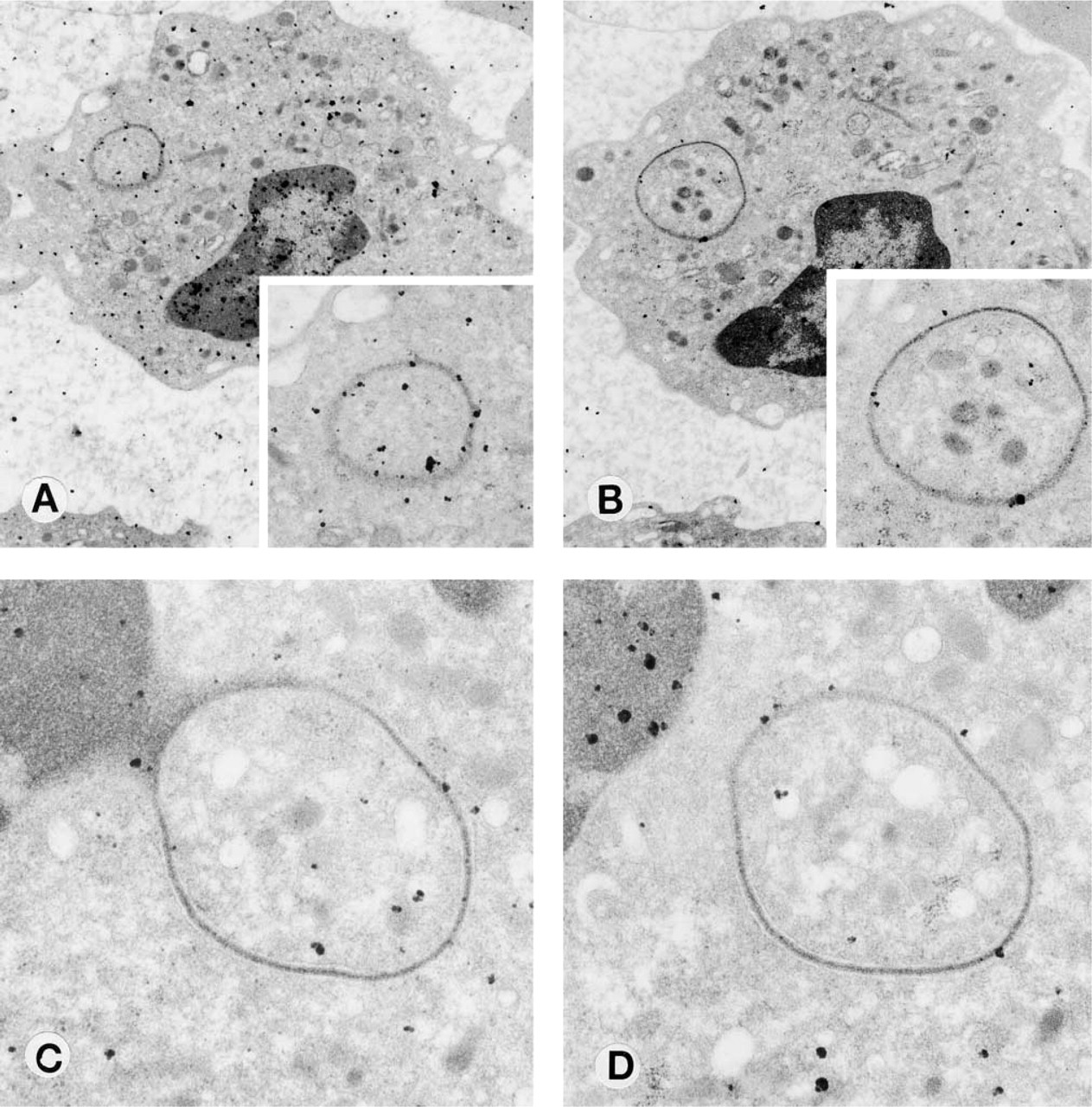
DNase treatment of polymorphonuclear granulocytes before DNA labeling. (
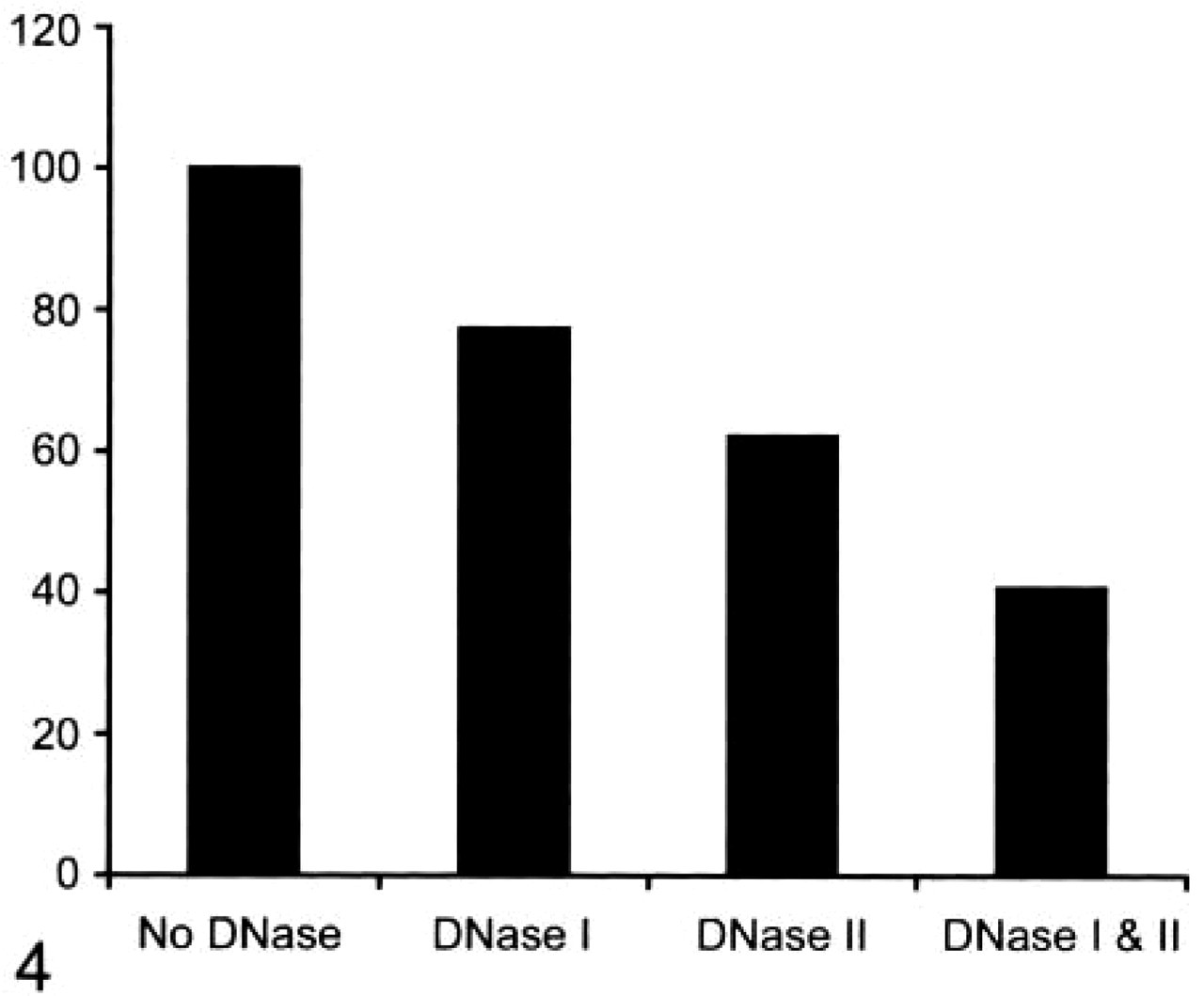
Whole polymorphonuclear white cells were treated with DNase I alone, DNase II alone or DNases I and II before DNA labeling. The figure shows the percent of labeling per cell compared with cells with no DNase treatment (100%).
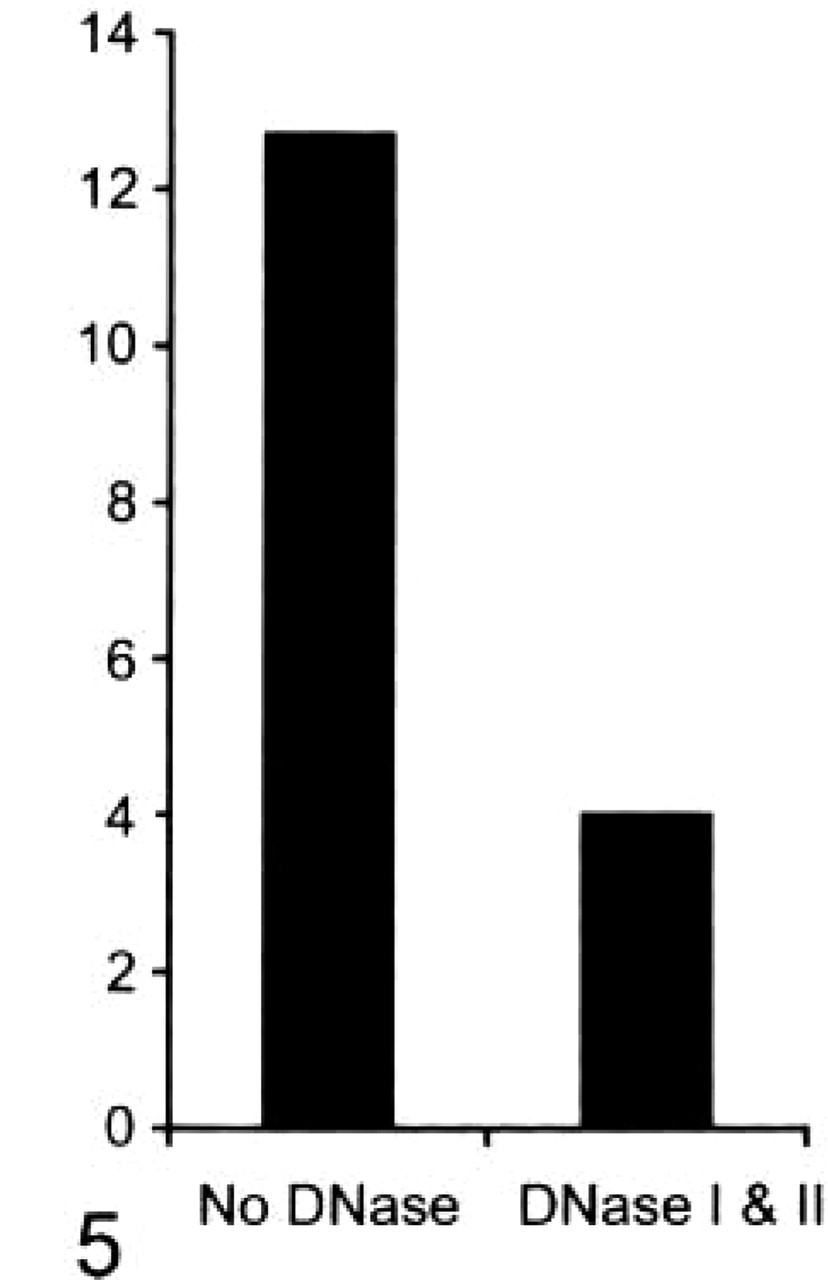
Mean number of colloid labels per appendage from cells that were treated with DNases I and II before DNA labeling compared to mean number of colloid labels per appendage from cells that had not been treated with DNase.
The isolation of DNA from the nucleus by submicroscopic appendages and blebs has not previously been described. This mechanism may be biologically advantageous to vitamin B12-deficient cells. DNA fragmentation occurs in vitamin B12-deficient cells and may be harmful to the cell (Fenech et al. 1997, 1998). Although the exact mechanism is uncertain, fragmentation may occur because uracil is misincorporated into DNA (Luzzatto et al. 1981; Wickramasinghe and Fida 1994) and subsequent excision of uracils by uracil-DNA glycolase may cause DNA breaks (Grafston et al. 1978; Blount et al. 1997). Although the removal of fragmented DNA from the nucleus is a hypothetical function for these appendages, it certainly cannot be concluded that all of the DNA labeled in these experiments was fragmented. Although the DNA labeling method using terminal deoxynucleotidyl transferase favors 3′-OH ends, these ends may be produced artifactually from section cutting and amplified by the favorable labeling conditions that were chosen here to visualize the DNA in the delicate appendages (Migheli et al. 1995). The finding that DNases I and II both decrease the labeling also suggests that not only 3′-OH ends are being labeled but that some end tailing or labeling of all types of DNA ends may occur with the high concentrations of cobalt that are used in this method. That treatment with both enzymes does not completely eliminate DNA labeling is probably explained by the familiar problem of steric hindrance (Migheli et al. 1995), which occurs with many chemical reactions that are attempted on the hard plastic epon.
It seems unlikely that the DNA isolation demonstrated here is the same process as that of micronucleus formation described by Fenech et al. (1997, 1998). Nuclear appendages were not found in lymphocytes, as are micronuclei. Furthermore, nuclear appendages were found in cells that are clearly not undergoing mitosis and they are rarely found in normal controls, in contrast to micronuclei, which are found in both young and old subjects who do not have vitamin B12 deficiency. Although micronuclei are homogeneous and stain densely (Fenech and Morley 1985), it is possible that the ultrascopic cytoplasmic rings are, in fact, balloons and if seen in three dimensions might appear the same as micronuclei.
The transfer of DNA into the cytoplasm by appendages appears more similar to the process described by Shimizu et al. in that both use blebs to effect the transfer. There are also substantial differences. The nuclear double-minute chromosomes are described as being encased by the bleb and not attached to the membrane, as is the case with the DNA in this study. There is also no mention of stalk formation in the Shimizu studies, nor the clusters of DNA that appear bound to the membrane in apparently forming stalks and which may possibly trigger appendage formation. It is, of course, difficult to compare these electron microscopic results with those of the confocal microscopic results of Shimizu.
Polymorphonuclear white blood cells from patients with low serum B12 isolate DNA from the body of the nucleus using a highly organized mechanism of nuclear appendages. The production of these appendages may function as an active mechanism for the removal of fragmented DNA from the nucleus and may be unique to cells with segmented and connected nuclei.
Footnotes
Acknowledgments
We would like to thank the Aid for Cancer Research (Boston, MA) for their assistance in funding this project.
We also thank Dr Pamela Roman (Cold Spring Harbor, NY) for reviewing the manuscript.
