Abstract
A
The AIRE gene encodes a 58-kD protein containing a nuclear location signal and several structural domains characteristic of transcription regulators (Finnish-German APECED Consortium 1997; Nagamine et al. 1997). APECED protein contains two plant homeodomaintype (PHD) zinc fingers found in several proteins with a role in the chromatin-mediated regulation of transcription (Aasland et al. 1995). Recently, a domain designated SAND has been identified in APECED protein and in several nuclear proteins such as SP100 and SP140/LYSP100 (Gibson et al. 1998). The N-terminal region of the APECED protein has also been shown to share a high degree of homology with the domain called HSR (homogeneously staining region) (Gibson et al. 1998). The HSR domain has also been found in SP100 and SP140/LYSP100 and has recently been shown to be involved in protein dimerization (Sternsdorf et al. 1997a, 1999; Pitkänen et al. 2000). The HSR domain has also been named as an ASS domain (Mittaz et al. 1999). In addition, the APECED protein contains four LXXLL motifs that have been shown to mediate the interactions of steroid receptor co-activators with nuclear hormone receptors (Heery et al. 1997).
In transient expression studies using mammalian tissue culture cell lines, dual subcellular localization of the APECED protein, nuclear and cytoplasmic, has been observed (Björses et al. 1999, 2000; Heino et al. 1999a; Rinderle et al. 1999). In the nuclei, APECED protein distribution is restricted to discrete punctate structures resembling nuclear bodies, although the relation to any known specific forms of nuclear domains remains to be determined. Cytoplasmic distribution of APECED protein has been shown to completely overlap with that of vimentin. The correct nuclear targeting of the APECED protein appears to depend on the intact structure of the PHD zinc finger domains. The recombinant APECED proteins mimicking disease mutations disrupting and/or deleting PHD fingers were shown to be generally unable to enter the nucleus and were retained in the cytoplasm (Björses et al. 2000). Similar results have been obtained with deletional mutants of the APECED protein (Rinderle et al. 1999).
On the basis of structural similarity to other proteins with an established role in the regulation of transcription and its subcellular location, APECED protein has been suggested to be a transcriptional regulator. Our recent results show that the human APECED protein is indeed able to promote transcription from the minimal promoter in mammalian one-hybrid assays (Björses et al. 2000).
Although the biological function(s) of the APECED protein have just began to emerge, the molecular mechanisms triggered by the mutations in the AIRE gene that finally result in the development of APECED, characterized by such a broad spectrum of clinical symptoms, remain elusive. Defining the biological functions of the AIRE and eventually understanding the pathogenesis of APECED is intimately bound to knowledge about where and when the AIRE gene is expressed.
In human, the expression of the APECED protein has been characterized in immunologically relevant tissues and was shown to be restricted to limited cell populations (Björses et al. 1999; Heino et al. 1999a). Our previous studies have shown that, in the thymus, APECED protein is expressed in epithelial cells of thymic corpusculi, epithelial reticular cells, and medullary thymocytes. In the spleen, APECED was expressed in the red pulp lymphocytes and neutrophilic granulocytes, whereas proliferating lymphocytes in the white pulp did not show any expression. In the lymph nodes, an intense staining was observed in lymphocytes and neutrophilic granulocytes (Björses et al. 1999). In addition, the lymphocytes, neutrophilic granulocytes, and monocytes of peripheral blood leukocytes exhibited APECED protein expression (Björses et al. 1999). In a similar immunohistochemical study, APECED protein was found in the thymic medullary epithelial cells and in cells that were defined as a probable subset of dendritic cells belonging to the monocyte lineage in the thymus and lymph nodes (Heino et al. 1999a). AIRE mRNA has been detected in human thymus, lymph nodes, appendix, and peripheral blood lymphocytes by Northern blotting hybridization (Nagamine et al. 1997). In situ hybridization analyses have revealed AIRE transcripts in the thymus, spleen, lymph nodes, and fetal liver, but no expression was detected outside the immune system (Heino et al. 1999a). The expression of mouse Aire mRNA has been mostly studied by the RT-PCR technique. Aire mRNA expression was detected in adult thymus, ovary, lung, heart, spleen, adrenal gland, thyroid gland, skeletal muscle, kidney, testes, and 11 dpc embryos (Blechschmidt et al. 1999; Ruan et al. 1999). mRNA hybridization in situ showed Aire mRNA expression in a few cells in the embryonic thymus at 14.5 dpc (Blechschmidt et al. 1999). No data have yet been published on the distribution of either Aire mRNA or protein expression at the cellular level in adult mouse tissues.
To extend our knowledge of the biological function of Aire gene, we explored its expression in transfected cell lines and in different mouse tissues using several different experimental approaches, i.e., RT-PCR, in situ mRNA hybridization, and immunohistochemical staining.
Materials and Methods
cDNA Library Screening
An oligo(dT)-primed λZAPII 17d mouse kidney library (kindly provided by Dr. V. Olkkonen; National Public Health Institute, Helsinki, Finland) was screened using human AIRE full-length cDNA as a probe. The positive clones were plaque-purified through three subsequent screening cycles, subcloned into the plasmid vector, and characterized by ABI Big Dye terminator sequencing (Perkin Elmer; Foster City, CA). The nucleotide data reported will appear in DDBJ, EMBL, and GenBank nucleotide sequence databanks under the accession number AJ243821.
RNA Preparation and Reverse Transcriptase-assisted PCR
RNAs were prepared from mouse tissues by the CsCl method (Chirgwin et al. 1979). cDNA first strand was synthesized by polydT priming using mouse thymus total RNA as a template with Superscript II reverse transcriptase (Life Technologies; Rockville, MD). Amplifications were carried out in a 50-μl volume using 5 μl of the first-strand reaction product as a template. The oligonucleotide primers used for amplification were chosen as follows: GGACTAGTACCATGGCAGGTGGGGATGGAAT (nucleotides 2970–2989; GenBank entry AJ007715) and GCTCTAGAGTGGCAGCGCCAGTGGAAG (nucleotides 1356–1375; GenBank entry AJ243821). The recognition sites for restriction enzymes SpeI and XbaI, respectively, were introduced for cloning. The Kozak consensus sequence was introduced in front of the initiator Met codon (Kozak 1991). Oligonucleotides were purchased from Genset. Amplification products were digested with appropriate restriction enzymes and sub-cloned into the pBluescript SK (Stratagene; La Jolla, CA) plasmid vector.
The oligonucleotide primers used for detection of Aire transcripts were CAGCAACTCTGGCCTCAAAG and CTTCGAACTTGTTGGGTGTATAA (GenBank entry AJ243821; nucleotides 431–450 and 693–715, respectively).
mRNA In Situ Hybridization
Adult NMRI mice were anesthetized with pentobarbital and perfused transcardially with physiological saline, followed by 4% paraformaldehyde in PBS for 4 min. Then the tissues were excised and immersed in the same fixative for an additional 60 min. Thereafter, the tissues were processed for paraffin embedding and 5-μm sections were mounted on Polysine glasses (Menzel; Braunschweig, Germany). The sections were deparaffinized, rehydrated, dehydrated, and dried. The sections were hybridized in a hybridization cocktail containing a mixture of three different 33P-labeled (Life Science Products; Boston, MA) cRNA probes (1 × 107 cpm/ml) covering mouse Aire cDNA sequence nucleotides 1–421, 542–879, and 1287–1906 (accession number AJ243821) for 18 hr at 55C. Then the slides were washed twice (15 min) in 2 × SSC at room temperature, treated with 100 μg/ml RNase A (Ambion; Austin, TX) for 30 min at 37C, and washed with 0.5 × SSC and 0.1 × SSC at 60C for 15 min each. The slides were dipped in water, dehydrated through graded ethanol, and either covered with Kodak MR5 (Eastman Kodak; Rochester, NY) autoradiography film or dipped in Kodak NTB2 emulsion. The films were exposed for 2–60 days, developed with Kodak LX 24 developer, and fixed with AL4 (Eastman Kodak) fixative. The dipped sections were exposed for 60 days and develop with D19 (Eastman Kodak) developer, fixed with G333 (Agfa Gevaert; Cologne, Germany) fixative, counterstained with cresyl violet, and coverslipped. The mixture of the sense probes was used as a negative control. Additional control experiments were carried out by treatment of the tissue sections with RNase A (100 μg/ml) for 30 min at 37C before hybridization with antisense probes.
Cell Culture and Transfections
The pAP42 construct containing the full-length mouse Aire cDNA in pcDNA3.1+ (Invitrogen; Carlsbad, CA) was transfected into cultured cell lines by using the FuGene 6 transfection reagent according to the manufacturer's instructions (Roche Diagnostics; Indianapolis, IN). Altogether, five different cell lines (human 293, monkey CV-1 and COS-1, mouse NIH3T3, and hamster BHK cells) were used in this study. For indirect immunofluorescence detection, cells were fixed with 3.5% paraformaldehyde 48 hr after transfection. The fixed cells were permeabilized with 0.1% Triton X-100 and blocked with 0.2% BSA. Tetramethylrhodamine isothiocyanate (TRITC)-conjugated anti-rabbit IgG (Jackson ImmunoResearch Laboratories; West Grove, PA) were used as secondary antibodies.
Production of the Mouse APECED Protein-specific Polyclonal Antisera
The peptide sequence KTKPPKKPDGNLESQHL, corresponding to amino acids 160–176 of mouse APECED protein, was chosen for production of antisera. Immunizations were performed with custom-synthesized MAP peptides (Research Genetics; Huntsville, AL). The immune sera were affinity-purified using respective synthetic peptides attached to CNBr-activated Sepharose 4B columns according to the manufacturer's instructions (Amersham Pharmacia Biotech; Piscataway, NJ).
Western Blotting
The tissue culture samples for SDS-PAGE electrophoresis were collected 48 hr after transfection. Proteins were separated on 11% SDS-PAGE and analyzed by Western blotting with specific polyclonal antisera raised against a synthetic peptide corresponding to amino acids 160–176 of the mouse APECED protein.
Immunohistochemistry
For immunohistochemistry, adult mice were fixed as described for in situ hybridization. The samples were cryoprotected with 15% sucrose in PBS, frozen with carbon dioxide, and 10-μm sections were cut. Mouse peripheral blood leukocytes (PBLs) were separated by centrifugation, smeared onto Polysine glasses, and fixed by immersion in 4% paraformaldehyde for 5 min. Endogenous peroxidase activity was inhibited by immersing the sections in 0.3% hydrogen peroxide in PBS for 5–20 min. The sections were incubated with rabbit affinity-purified anti-peptide antibody raised against mouse APECED protein (amino acids 160–176; dilution 1:100–200) at 4C overnight in a buffer containing 1% BSA and 0.3% Triton X-100. Then the sections were incubated with biotinylated goat anti-rabbit IgG (diluted 1:300) and ABC complex (Vectastain Elite Kit; Vector Laboratories, Burlingame, CA) for 30 min each. The immunoreaction was visualized with nickel-intensified DAB as a chromogen. The sections were then dehydrated and embedded in Entellan. Controls included preimmune serum and antibody presaturated with 20 μg/ml of the peptide used for immunization. In addition, extracts prepared from COS-1 cells transfected with full-length Aire cDNA and empty expression vector were used for preadsorbtion experiments.
In Vitro Translation
The full-length Aire cDNA in Bluescript SK+ was in vitro translated using the TNT-coupled Reticulocyte Lysate System according to the manufacturer's instructions (Promega; Madison, WI). The protein samples were analyzed on an 11% SDS-PAGE, followed by autoradiography.
Results
Mouse Aire cDNA Cloning
The mouse cDNA library was screened with human AIRE cDNA as a probe. Positive clones were plaque-purified through subsequent screening cycles and characterized by nucleotide sequencing. One of the isolated clones encoded for amino acids 430–552 of mouse APECED protein, as deduced from the mouse Aire genomic DNA sequence (Blechschmidt et al. 1999; Mittaz et al. 1999; Wang et al. 1999). To obtain full-length Aire cDNA, we applied RT-PCR to amplify the sequence encoding for the missing N-terminal amino acids. Total mouse thymus RNA was used as a template for cDNA first-strand synthesis. Amplification was performed with mouse Aire-specific oligonucleotide primers. Amplification products were subcloned into the plasmid vectors and further subjected to nucleotide sequencing. Full-length mouse Aire cDNA was constructed by ligation of amplified DNA fragments to the cDNA library clone C1_2. The ability of the obtained cDNA clone to encode for a full-length APECED protein was confirmed by translation of the cDNA in vitro. Electrophoretic analysis of in vitro translation products yielded a major translated polypeptide of 59 kD (data not shown), which is in a good agreement with the predicted molecular weight of the mouse APECED protein (Blechschmidt et al. 1999; Mittaz et al. 1999; Wang et al. 1999). Human AIRE cDNA was used as a positive control.
Expression of Aire cDNA in Mammalian Tissue Culture Cells
The subcellular localization of mouse APECED protein was studied by transient expression and indirect immunofluorescence experiments using the antiserum raised against a synthetic APECED peptide. The analysis of different cell types (BHK, NIH3T3, CV-1, COS-1, and 293) revealed three different staining patterns: (a) speckled nuclear distribution excluding the nucleoli (Figure 1A); (b) filamentous cytoplasmic pattern resembling the staining characteristic to vimentin (Figure 1B); and (c) both of these two distributions in the same transfected cell (Figure 1C). In BHK cells, over 30% of the transfected cells had a speckled nuclear APECED protein distribution, 30% exhibited filamentous cytoplasmic staining, 30% exhibited both distributions, and the remaining cells had other staining patterns, such as clustering of the protein in the nuclear envelope. In mouse NIH3T3 cells, over 70% of the transfected cells showed cytoplasmic staining only. Speckled nuclear staining was observed in less than 10% of transfected cells. A mixed localization pattern was observed in less than 10% of the cells, and the rest exhibited other staining patterns. In addition, the CV-1, COS-1, and 293 cell lines transiently transfected with mouse Aire cDNA displayed the different staining patterns mentioned above (data not shown). The staining pattern varied among the different cell lines. The CV-1 and COS-1 cells had a predominant filamentous cytoplasmic staining pattern, whereas in the majority of transfected 293 cells APECED protein was localized in nuclear speckles. The subcellular distribution of mouse APECED protein therefore closely resembles that of its human counterpart (Björses et al. 1999; Heino et al. 1999a; Rinderle et al. 1999).
Analysis of Aire mRNA Expression by RT-PCR and mRNA In Situ Hybridization
The expression of mouse Aire was first studied by RT-PCR. Analysis of mouse total RNA revealed the amplification of Aire-specific transcripts in all the tissues studied, i.e., thymus, spleen, lymph node, liver, kidney, testis, brain, and fetal liver (data not shown). In general, our RT-PCR results are in agreement with recently published studies (Blechschmidt et al. 1999; Ruan et al. 1999). It should be noted, however, that contrary to a report by Blechschmidt and colleagues, whose study detected no Aire transcripts in brain tissue, our results show clear PCR amplification product in mouse brain (Blechschmidt et al. 1999). In addition, Ruan and colleagues (1999) did not detect any Aire transcripts in spleen and liver whereas specific amplification products were detected in the spleen in this study and in that published by Blechschmidt et al. (1999). No amplification products were observed in negative control reactions containing RNA with no first-strand synthesis or in reagent blanks (data not shown).
Aire expression in mouse tissues was further studied by mRNA in situ hybridization. Our initial experiments with radioactively labeled oligonucleotides failed to detect Aire transcripts even with rather long exposure times (data not shown). To improve the sensitivity of our assays, we decided to use radioactively labeled mouse Aire cRNA as the hybridization probe. Additional improvement in sensitivity was achieved by using a mixture of three cRNA probes covering different parts of the Aire transcript (see Materials and Methods). In the thymus, a strong signal for Aire mRNA could be seen in a small population of cells in the medulla. These cells were larger than thymocytes, stained lightly with cresyl violet, and obviously represented reticular (epithelial) cells of the thymus. Part of the epithelial cells clearly lacked Aire mRNA. A moderate level of staining was also seen in a small population of medullary thymocytes (Figures 2A–2A).
Our results on Aire transcript expression in mouse thymus are in agreement with recently published results showing scattered cells expressing the AIRE mRNA in the medullary zone of the thymus of 17-day mouse embryo and in human juvenile thymus (Blechschmidt et al. 1999). In spleen, strongly labeled cells were scattered along the border of the white and red pulp (Figure 2F). In the lymph nodes, a few strongly labeled cells were seen in the paracortical region (Figure 2G). In bone marrow, a large population of cells exhibited moderate signal (Figures 2I and 2J). In the ovary, a moderate signal could be observed in the granulosa cells of growing follicles (Figures 2K and 2L).
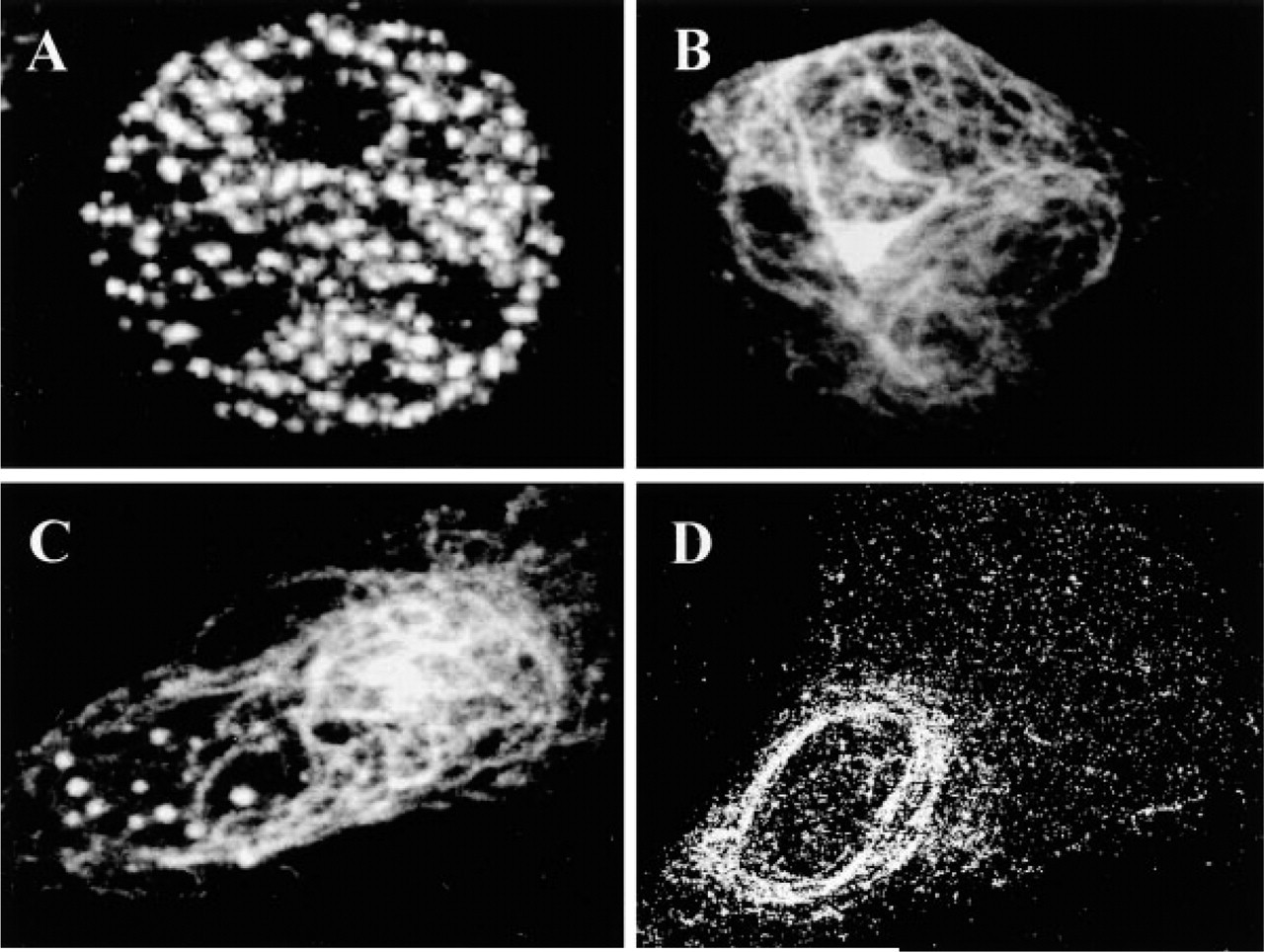
Subcellular location of mouse APECED protein in transiently transfected cell lines. Mouse APECED protein shows three main distributions: (
Immunohistochemistry
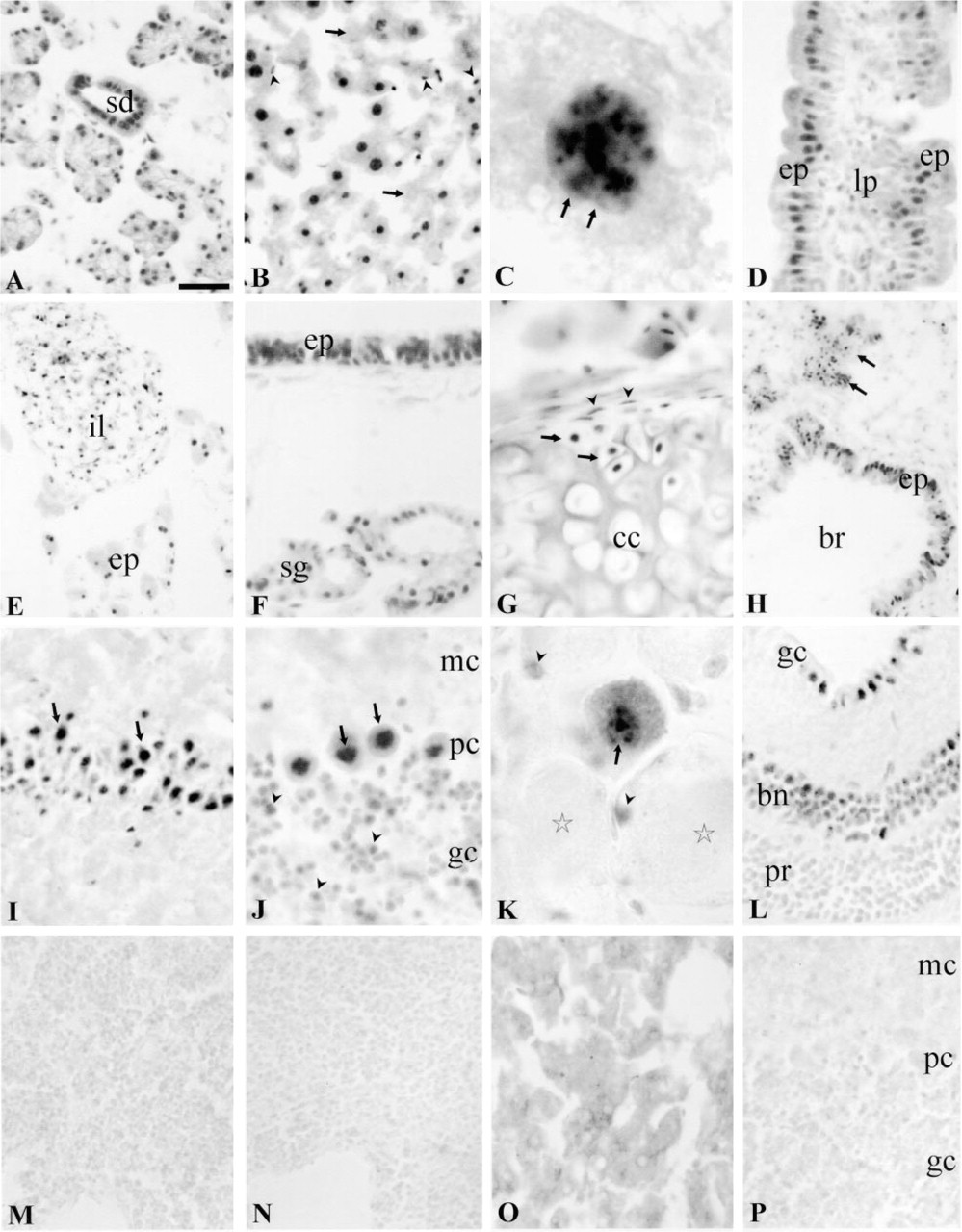
To gain information on the distribution of the APECED protein at the cellular level, we carried out immunohistochemical stainings of mouse tissue sections with antisera raised against a synthetic polypeptide corresponding to mouse APECED protein. The specificity of the immunohistochemical reaction was demonstrated by the lack of immunostaining of tissue sections incubated with antisera preadsorbed either with the synthetic peptide used for immunization (Figures 5M, 5O, and 5P) or extracts prepared from COS-1 cells transfected with full-length Aire cDNA (Figure 5N). The staining was not affected in control experiments performed with antisera preadsorbed with the extracts prepared from COS-1 cells transfected with empty expression vector (data not shown). The specificity of the antiserum was further studied by staining the Western blots of the lysates of NIH-3T3 and COS-1 cell lines transiently transfected with Aire cDNA (Figure 3). A single major protein of the expected molecular weight can be seen in Aire-transfected cell lines (Figure 3, Lane A), whereas cells transfected with empty expression vector remain completely unstained (Figure 3, Lane C). Additional evidence for the specificity of the antiserum was obtained by staining Aire-knockout mouse tissues. The thymus, liver, and cerebellum of the Aire-knockout mouse stained with Aire-specific antiserum did not show any staining (data not shown; Puhakka et al. unpublished observations).
Immune System. A small number of intensely Aire-immunoreactive (Aire-IR) cells was observed in the medulla of the thymus. These cells were larger than thymocytes and had large non-labeled cytoplasm characteristic of reticular (epithelial) cells. The labeling was localized in distinct dots in the nucleus (Figure 4A). Some of the reticular cells were devoid of labeling. In addition, the cells in the thymic corpuscles were intensely stained. A small number of medullary thymocytes exhibited low to moderate staining (Figure 4B). Cortical thymocytes were devoid of immunostaining.
In the red pulp of spleen, large cells possessing prominent nuclei were clearly labeled. These cells most probably represent tissue macrophages (Figure 4C). In addition, a large number of lymphocytes and some neutrophilic granulocytes in the red pulp were Aire-IR (Figure 4C). A few labeled reticular cells could be observed. Light staining was seen in the smooth muscle cells of the trabecules. The white pulp of spleen was devoid of immunostaining.
In the medulla of the lymph nodes, a population of the lymphocytes was clearly stained, but the cortical germinal centers remained non-stained. Some labeled reticular cells could be seen in the medulla. Mast cell nuclei appeared to be unlabeled, although the cytoplasmic granules exhibited strong nonspecific staining which sometimes obscured the nucleus (Figure 4D).
In bone marrow, a subpopulation of large megacaryocytes showed intense staining. The labeling was localized to distinct dots in the nuclei. In addition, scattered smaller cells exhibited clear staining; these cells probably represent lymphoblasts and myeloblasts (Figure 4E).
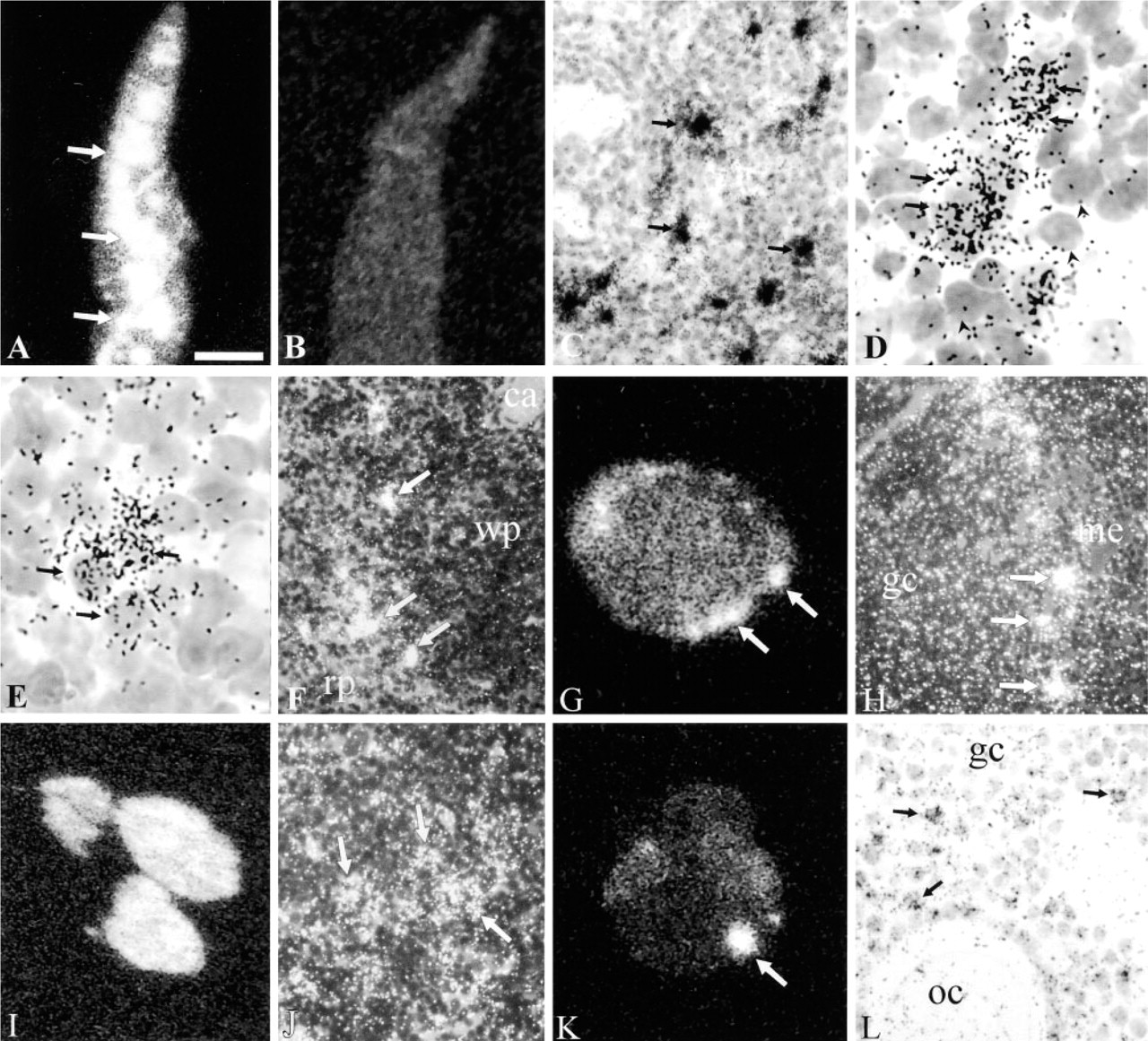
Expression of Aire mRNA in mouse tissues detected by in situ hybridization. (
In the peripheral blood smears, strong nuclear staining could be seen in lymphocytes, polymorphonuclear leukocytes, and monocytes. A small population of lymphocytes appeared to be devoid of staining (Figure 4F).
Urinary Tract. In kidney, the epithelial cells of the proximal and distal convoluted tubules were labeled. The loops of Henle and collecting ducts were mainly unlabeled. The dark cells of the collecting tubules exhibited cytoplasmic staining close to the cell membrane. The podocytes of glomeruli were slightly labeled. Weak staining was also seen in the transitional epithelium in the kidney pelvis (Figure 4G).
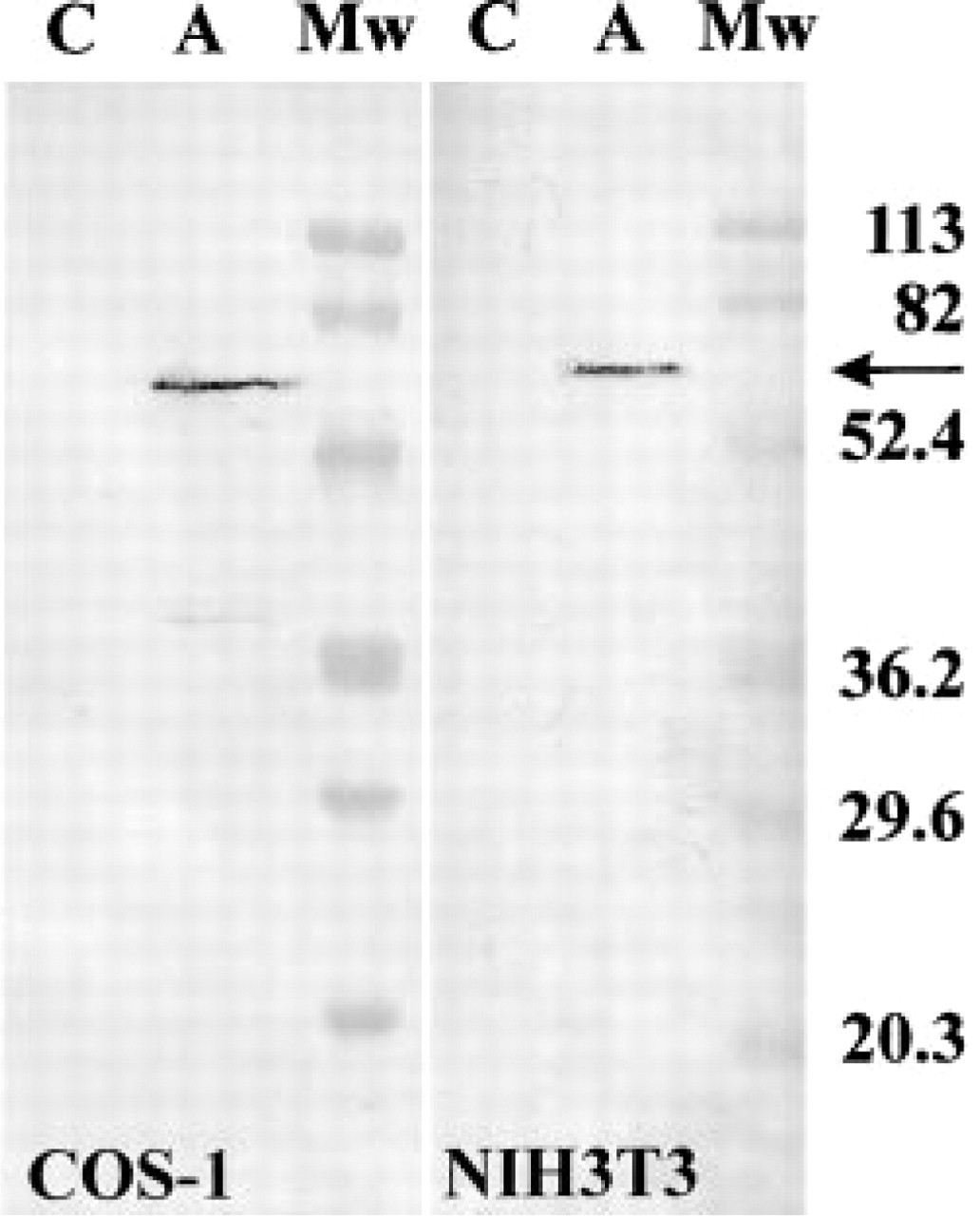
Western blotting analysis of the specificity of antisera. The APECED protein (arrow) is detected as a 59-kD band with polyclonal antibody in the extracts from COS-1 and NIH3T3 cells transiently transfected with the mouse Aire cDNA (Lanes A). Extracts prepared from the cells transiently transfected with the empty pcDNA3.1 + expression vector do not show any staining (Lanes C). Protein molecular mass marker (MW) sizes (in kD) are indicated at right.
In urinary bladder, the transitional epithelium was strongly Aire-IR. The smooth muscle cells of bladder exhibited weak staining (Figure 4H).
Genital Organs. In the testis, immunostaining could be seen in a stage-specific manner in a few seminiferous tubules. Staining was localized in distinct dots in the nuclei of pachytene spermatocytes and round spermatids. In a few tubules, some spermatogonia and Sertoli cells showed Aire-IR (Figure 4I). In the interstitial tissue, most of the cells exhibited nuclear staining. A population of these cells also showed moderate cytoplasmic staining, which was abolished by presaturation (Figure 4J). In epididymis, seminal vesicle (Figure 4K), and prostate the nuclei of epithelial cells stained strongly.
In the ovary, nuclear staining could be observed in the granular cells of different size follicles. The oocytes were also labeled. The cells of corpora lutea and interstitial tissue exhibited strong cytoplasmic staining without clear nuclear labeling. This staining could not be abolished by presaturation of the antibody (Figure 4L). In the uterus, the epithelial cells of the mucosa and secretory glands were clearly stained. The smooth muscle cells of the myometrium were also immunopositive.
Endocrine Organs. Large numbers of cells in anterior and intermediate lobes of pituitary were Aire-IR (Figure 4M). In the adrenal gland, clear nuclear staining could be observed in the cells of zona glomerulosa and in the medullar chromaffin cells (Figure 4N). The cells of the zona fasciculata and reticulata exhibited strong cytoplasmic staining without a clear nuclear signal. This cytoplasmic staining was abolished by presaturation (data not shown). Most of the cells in the adrenal medulla displayed clear staining (Figure 4O). In the thyroid gland, a variable degree of immunostaining could be seen in the epithelial cells of the follicles. Clear staining was observed in parafollicular cells (Figure 4P).
Alimentary Tract. In the salivary glands, most of the secretory cells of the acini and secretory ducts were moderately to strongly Aire-IR (Figure 5A). In the liver, most of the hepatocytes were very intensely Aire-IR, although some were devoid of labeling (Figure 5B). The staining intensity was similar in different regions of the liver lobule. Küppfer cells were also stained (Figure 5B). The staining pattern of the hepatocyte nuclei was uneven and resembled that of transiently transfected cell lines (Figure 5C). The epithelial cells and glandular cells in different parts of the alimentary canal were Aire-IR (Figure 5D). In the pancreas, both the exocrine and endocrine cells were Aire-IR (Figure 5E).
Respiratory System. In the respiratory tract, Aire immunoreactivity could be seen in the respiratory epithelium of large airways from the larynx to the respiratory bronchioles (Figures 5F and 5H). The alveolar Type I and Type II cells displayed weak staining, and alveolar macrophages were clearly labeled. In tracheal cartilage, the undifferentiated cells in the perichondrium and differentiating chondroblasts were stained, but mature chondrocytes were not labeled (Figure 5G).
Nervous System. APECED immunostaining was widely distributed in the central nervous system. The cerebral cortex, hippocampus (Figure 5I), amygdala, thalamic nuclei, hypothalamus, cerebellar cortex, various brainstem nuclei, spinal cord, and dorsal root ganglia (Figure 5K) all revealed strong immunoreactivity. In the cerebellum, intense Aire-IR was observed in the nuclei of the Purkinje cells, while the granular neurons exhibited modest staining (Figure 5J). The most prominent staining was in neurons, but different types of glial cells were also labeled. In the eye, the ganglion cells and bipolar neurons of the retina showed moderate staining, but the rods and cones were negative (Figure 5L).
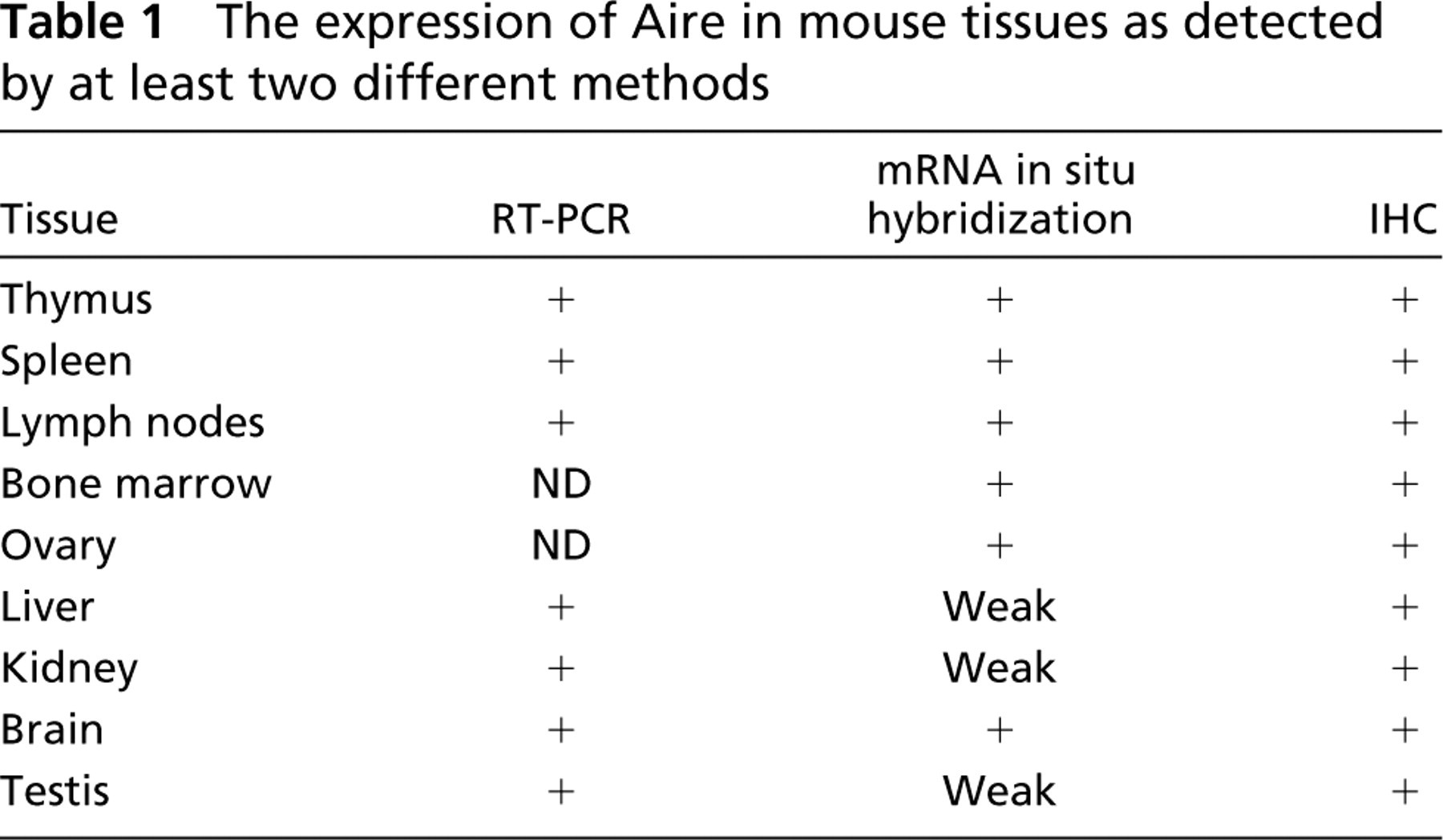
Those tissues in which Aire expression was detected with at least two different experimental techniques are listed in Table 1.
Discussion
To understand the biological function of APECED protein, we studied the spatial expression pattern of the Aire gene in mouse using several different techniques such as RT-PCR, in situ mRNA hybridization, and immunohistochemical staining. Our results revealed expression of Aire in different cell populations in a broad range of rather different types of mouse tissues. Because APECED is an autoimmune disorder, the immunologically relevant organs would be expected to express Aire. In the thymus, spleen, lymph nodes, and bone marrow, Aire expression was detected in different cell populations of lymphoid and myeloid lineages. It should be noted that no double stainings with specific cell-surface markers were performed and therefore the precise identity and stage of differentiation of cells expressing Aire remain to be determined. In addition the reticular epithelial cells and corpuscle cells of the thymus and epithelial cells of the lymph vessels expressed Aire. Our findings in thymus, spleen, and lymph nodes are in a good agreement with earlier studies on human AIRE expression by us and others (Björses et al. 1999; Heino et al. 1999a). Aire was also expressed in the lymphocytes, monocytes, and polymorhphonuclear leukocytes of the peripheral blood.
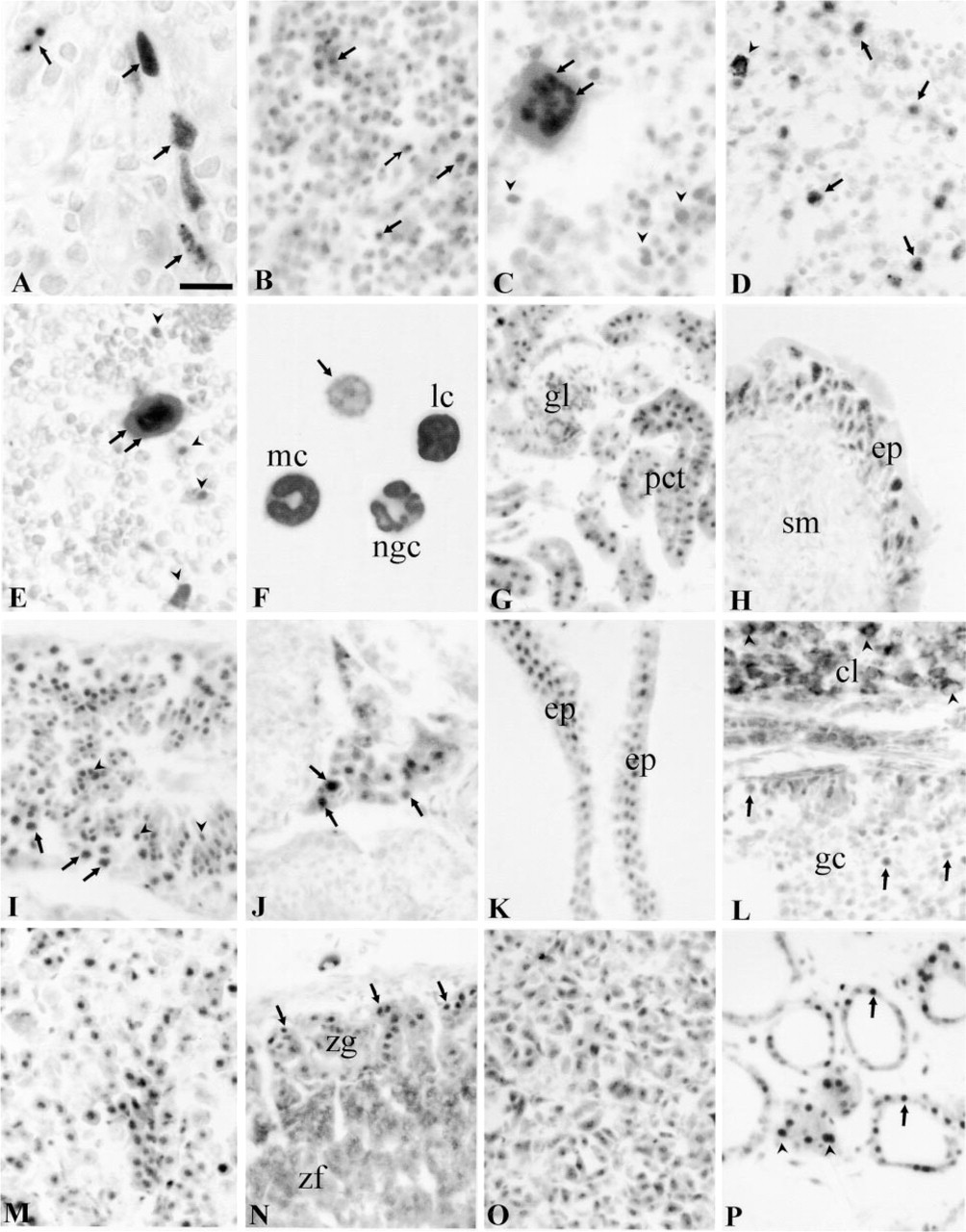
APECED protein detected by immunohistochemical staining I. (
Outside of the immune system, the APECED protein was detected in the brain, liver, kidney, pancreas, intestinal canal, gonads, pituitary, thyroid, and adrenal glands. The cells expressing the APECED protein represent rather diverse cell types, ranging from epithelial cells in several different organs to neurons and glial cells in the central nervous system. Our results showing a broad range of tissues expressing APECED protein are further supported by our studies of the rat. Immunohistochemical staining of rat tissues with antiserum raised against human APECED protein have revealed an almost identical expression pattern as that observed in the mouse (unpublished results).
In all the cell types that showed nuclear labeling in adult mouse tissues, the staining was unevenly distributed and resembled the speckled, nuclear body-like staining that we observed in mammalian cell cultures transfected with mouse Aire cDNA. Similar results have been obtained in experiments with the human counterpart of Aire (Björses et al. 1999, 2000; Heino et al. 1999a,b; Rinderle et al. 1999). The discrete localization pattern of APECED protein inside the nucleus provides an interesting path to follow for dissecting its biological functions. Many different nuclear inclusions have been recognized on the basis of morphological criteria and biochemical constitution (see Matera 1999 for review). Although the association of the APECED protein with any of those entities remains to be investigated, a structure of particular interest is the PML body [also called the nuclear dot (ND), PML oncogenic domain (POD), nuclear domain 10 (ND10) or Kr body], which contains PML, Sp100, and at least 16 other proteins (Matera 1999). PML bodies have been shown to regulate diverse biological processes, such as tumor suppression, transcription regulation, expression of the major histocompatibility complex (MHC), and apoptosis (Mu et al. 1994, Zheng et al. 1998; Torii et al. 1999; Pearson et al. 2000). Interestingly, one of the main components of these bodies, the Sp100 protein, shares HSR and SAND/HNPP domain homology with APECED protein and has been suggested to be derived from a common ancestor (Gibson et al. 1998). Recently, it was suggested by Sternsdorf and colleagues (1999) that the HSR domain may contain a PML body targeting signal, and therefore it is possible that APECED protein could in some cellular context be targeted to PML bodies. However, our preliminary co-localization experiments in transiently transfected cell lines overexpressing APECED protein could not demonstrate co-localization of the protein with PML bodies (Björses et al. 1999). It has been recently shown that PML bodies are dynamic structures with a heterogeneous constitution that depends on a particular phase of the cell cycle (Bloch et al. 1999). Therefore, it would be interesting to further study the possible co-localization of APECED protein with PML bodies and other subnuclear domains in the context of the cell cycle and in cell lines that endogenously express the APECED protein to overcome the problems related to overexpression.
It should be noted that in a few cell types, such as interstitial cells of the testis and few neurons in the trigeminal ganglion, cytoplasmic staining was observed in addition to nuclear labeling. Our results show that, in transiently transfected tissue culture cells, mouse APECED protein also has a cytoplasmic vimentin-like distribution. The proportion of the different distribution types varies depending on the cell line used in the transient expression. It has been speculated by Björses et al. (1999) that the cytoplasmic location of the APECED protein could be caused by the high expression level of the particular cell line which may, in turn, saturate the nuclear transport mechanisms. Alternatively, it can be speculated that APECED protein is imported into the nucleus in complex with a “carrier” protein. In that scenario, the nuclear vs cytoplasmic location of the APECED protein could be dependent on the availability of a protein complex partner needed for efficient nuclear translocation. It is of interest that both the PML and the SP100 protein, the main components of PML bodies, have been reported to be located not only in the nucleus but also in the cytoplasm (Sternsdorf et al. 1997b).
APECED protein detected by immunohistochemical staining II. (
The expression of Aire in mouse tissues as detected by at least two different methods
It has been suggested that APECED protein might be involved in the regulation of self-tolerance either by clonal deletion in the thymus or by induction of peripheral T-cell anergy (Peterson et al. 1998). This assumption is based on the multiplicity of autoimmune diseases caused by mutations in AIRE and on experimental results that have revealed the APECED protein expression pattern to be limited to the immune system (Peterson et al. 1998). Our results show that, in addition to the immune system, Aire is expressed in a broad range of tissues and in rather different cell types. Although the possible involvement of APECED protein in regulating self-tolerance remains to be experimentally explored, it is sound to assume that APECED protein is likely to regulate different biological processes elsewhere in the organism, e.g., in tissues such as the brain or liver. The regulation of completely different developmental pathways by the very same protein, depending on the physiological context provided by a given cell, has been widely recognized. Furthermore, the expression of the same transcription factor in different cellular backgrounds is an elegant mechanism that allows the creation of structural and functional novelties (see Ganfornina and Sanchez 1999 for review).
Conclusions
In this study, we have shown by in situ hybridization and immunohistochemistry that Aire is widely expressed throughout the entire organism. For the first time, we have shown the presence of the APECED protein in multiple tissues outside the immune system in very diverse cell types. We have shown that APECED protein has dual subcellular localization in transiently transfected cell lines. In vivo it was predominantly located in nuclei. However, in a few cell types cytoplasmic localization was also observed. Our results imply that APECED protein most likely has different biological functions in different tissues depending on the tissue- and cell type-specific background. Identification of the protein partners interacting with APECED protein and the cellular pathways regulated by its activity will be of critical importance to understanding the biological role of the APECED protein. It will also provide us with valuable information on the molecular mechanisms behind APECED pathology and new insights into autoimmune diseases in general.
Footnotes
Acknowledgments
Supported by the Academy of Finland, the Ulla Hjelt fond of the Foundation for Pediatric Research, the Medical Research Fund of Tampere University Hospital, and by resources from the Helsinki Biomedical Graduate School MD/PhD program.
The excellent technical assistance of Ulla-Margit Jukarainen, Tuula Airaksinen, Anne Vikman, and Katri Miettinen is greatly appreciated. Donald Smart is acknowledged for language revision.
