Abstract
To extend our recent observation that villin mRNA, encoding an apical microvillous protein, is dichotomously localized in the basal region of human enterocytes, we examined the localization of mRNAs for brush border myosin I (BBMI) and intestinal fimbrin (I-fim). In situ hybridization indicated that BBMI mRNA localized to the basal region of human enterocytes, whereas the mRNA for I-fim distributed diffusely. To facilitate study of potential mechanisms of mRNA targeting, we cloned a full-length cDNA for BBMI including its 5'- and 3'-untranslated regions (UTRs). This cDNA shares 86% sequence identity with bovine BBMI and 85% with rat BBMI. Sequence analysis revealed no obvious similarity between the 3'-UTRs of BBMI and villin. This study provides evidence of novel sorting pathways for intestinal microvillous cytoskeletal proteins.
mRNA
Materials and Methods
Materials
Digoxigenin-11-UTP (DIG-11-UTP) was purchased from Boehringer Mannheim (Indianapolis, IN). [32P]-α-UTP and [32P]-α-dCTP were from DuPont-New England Nuclear (Boston, MA). All restriction enzymes and DNA-dependent RNA polymerase were from Gibco BRL (Grand Island, NY). All other reagents were from Sigma (St Louis, MO). Human jejunal tissue was obtained from adults undergoing elective gastric bypass surgery with a protocol approved by the Human Investigation Review Committee of the New England Medical Center Hospitals.
In Situ Hybridization
A human BBMI (hBBMI) cDNA fragment (603-1057 bp) was subcloned into BlueScript II KS (+) vector at Xho I/Hind III sites. A 510-bp fragment of human I-fim (745-1254 bp) was subcloned into BlueScript II KS (+) vector at BamH I/ Cla I sites. DIG-labeled RNA probes were prepared using DIG-11-UTP according to the manufacturer's instructions. Quantification and dot-blot assay of the RNA probes were performed as previously described (Li et al. 1998). In situ hybridization was carried out as described previously (Li et al. 1998). Controls included in situ hybridization using sense probes and treatment of sections with RNase before hybridization.
Cloning of hBBMI
Human jejunal total RNA was isolated by homogenization of mucosal scrapings in guanidine isothiocyanate, purified through cesium chloride centrifugation as previously described (Li et al. 1998). RT-PCR was carried out as previously described with following modifications (Li et al. 1998), using an oligo T primer (5 '-TTCTAGAATTCAGCGGCCGC(T)30N-1N-3'). With the upstream primer (5'-GGCTAGTAAGCTAGTGATGTC-3') and downstream primer (5' -GGAGAAGAGCTGGTTCAGCTTTG-3') derived from rat BBMI (Balish and Coluccio 1995), a 1.1-kb fragment was generated using the Expand High Fidelity PCR system (Boehringer Mannheim), and directly cloned into pGEM-4 T Easy vector (Promega; Madison, WI). DNA sequencing yielded the sequence from 557 to 1617 bp of the human BBMI (positions according to GenBank accession #AF105424) with 88% and 86% sequence homology to bovine and rat BBMI, respectively.
The 5'-end of the mRNA was cloned by the 5'-Rapid Amplification of cDNA Ends Strategy (5' RACE) (Borson et al. 1992). Briefly, an anchored sequence was added to the 3'-end of the first strand cDNA using terminal deoxynucleotidyl transferase and dGTP. With an anchored primer, poly(C)18, and a gene-specific primer (GSP1: 5'-GGCCTTCAGCAGCTGTTCATCTGC-3'), a 0.8-bp 5' RACE product was amplified and cloned into the pGEM-4 T Easy vector. To obtain the full-length sequence information at the 5'-end of the mRNA, 62 clones containing the BBMI sequence were selected from the more than 200 clones. Among the BBMI-positive clones, four clones containing the longest PCR fragments were sequenced at both ends. All four clones have the same coding sequence at their 3'-ends, with 100% identity to the hBBMI obtained above. All of them also contain the same 5'-end sequence.
The 3'-end of the mRNA was cloned by 3' RACE. Briefly, PCR amplification was accomplished using an anchored primer (5'-TTCTAGAATTCAGCGGCCGCTTT-3'), and a gene-specific primer (GSP2: 5'-GCTCATTGAGCATAATCAGCGAGG-3'), which is located at 1529 bp of cDNA (position according to AF105424). The approximately 2.0-bp 3' RACE product was cloned into the pGEM-4 T Easy vector and sequenced. DNA sequencing was performed by the Protein Core Facility, Department of Physiology, Tufts University, Boston, MA. Sequence was determined on both strands and analyzed using Genetics Computer Group's computer software.
Northern Blotting
Northern blotting was performed as described previously (Li et al. 1998). The full-length hBBMI cDNA was used as a probe and random-labeled with [32P]-dCTP (3000 Ci/mmol) using an oligonucleotide labeling kit (Pharmacia-Biotech; Piscataway, NJ).
Results
mRNA Localization
All jejunal tissue samples were examined and demonstrated normal morphology and cellular architecture as shown previously (Li et al. 1998). The BBMI mRNA signal was found only in the epithelial cell layer, with no detectable signal in the lamina propria, suggesting that hBBMI is expressed only in enterocytes (Figure 1A). Within enterocytes, BBMI mRNA was predominantly localized at the basal region below the nuclei (Figures 1A and 1B). BBMI mRNA expression extended from the crypt-villous junction to the tips of villi (not shown). BBMI mRNA signal was also detected in enterocyte nuclei as dense dots (Figures 1A and 1B), suggesting that they may represent premature, unprocessed mRNAs. Because of the asymmetric distribution of hBBMI mRNA, we further examined the mRNA distribution pattern for I-fim, another microvillous cytoskeletal protein. I-fim mRNA was detected only in enterocytes, but in contrast to BBMI mRNA it was distributed diffusely throughout the cytoplasm (Figures 1D and 1E). No mRNA was visualized when sense probes were used (Figures 1C and 1F).
In control samples, human SI mRNA localized only in the apical domain of the enterocytes, with no mRNA detected in any other region of the cells, similar to our previously reported results (Li et al. 1998) (Figure 2A). Human Apo A-I mRNA was expressed throughout the cytoplasm of the enterocytes, as shown in Figure 2B, and villin transcripts were localized to the basal region of human enterocytes (Figure 2C). Again, no mRNA was visualized when sense probes were used (Figures 2D–2F).
In all of our in situ hybridization studies, we failed to see any signal if the sample was pretreated with ribonuclease A, indicating that the signals observed were due to hybridization to RNA targets (not shown). In addition, the specificity of the mRNA signals was also suggested by the absence of reaction product in the lamina propria.
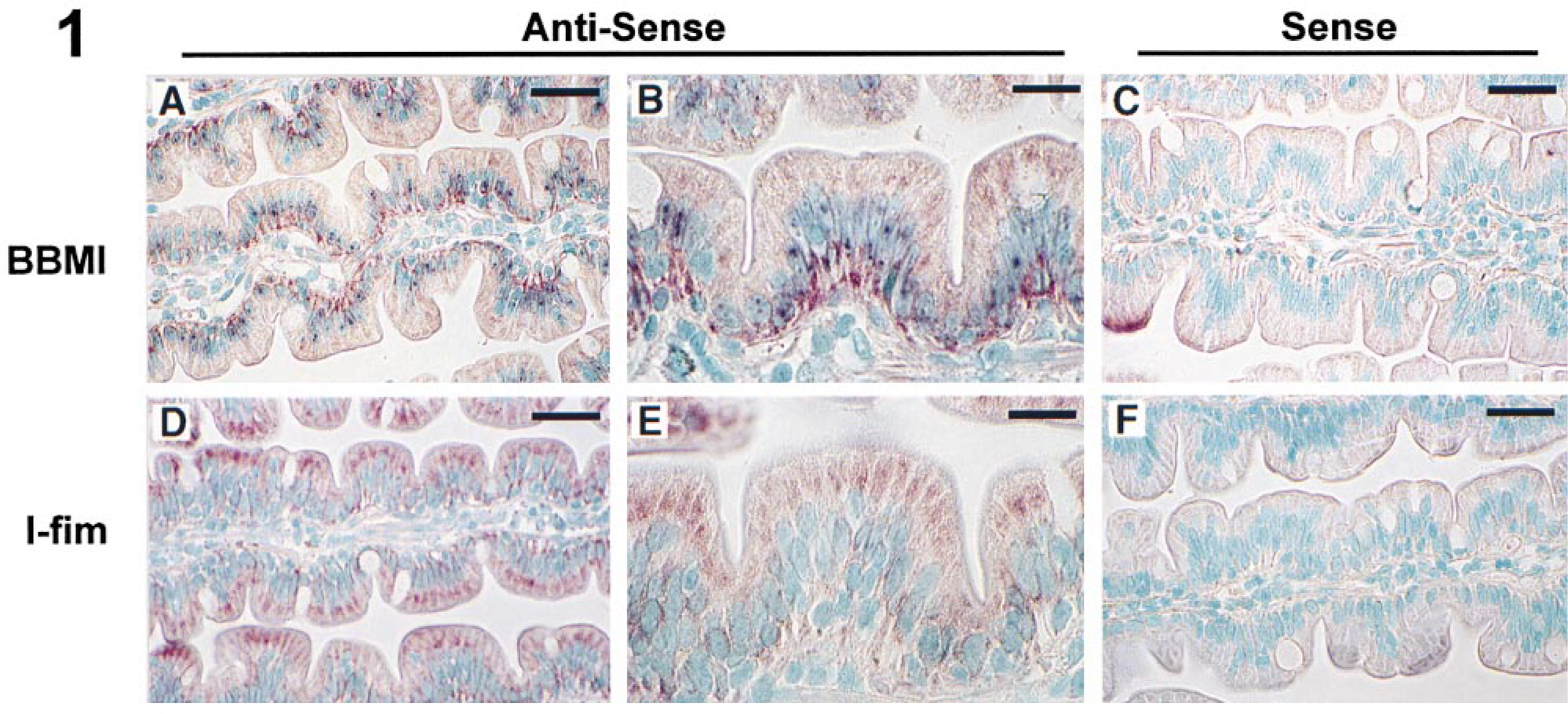
In situ hybridization for hBBMI and I-fim. (
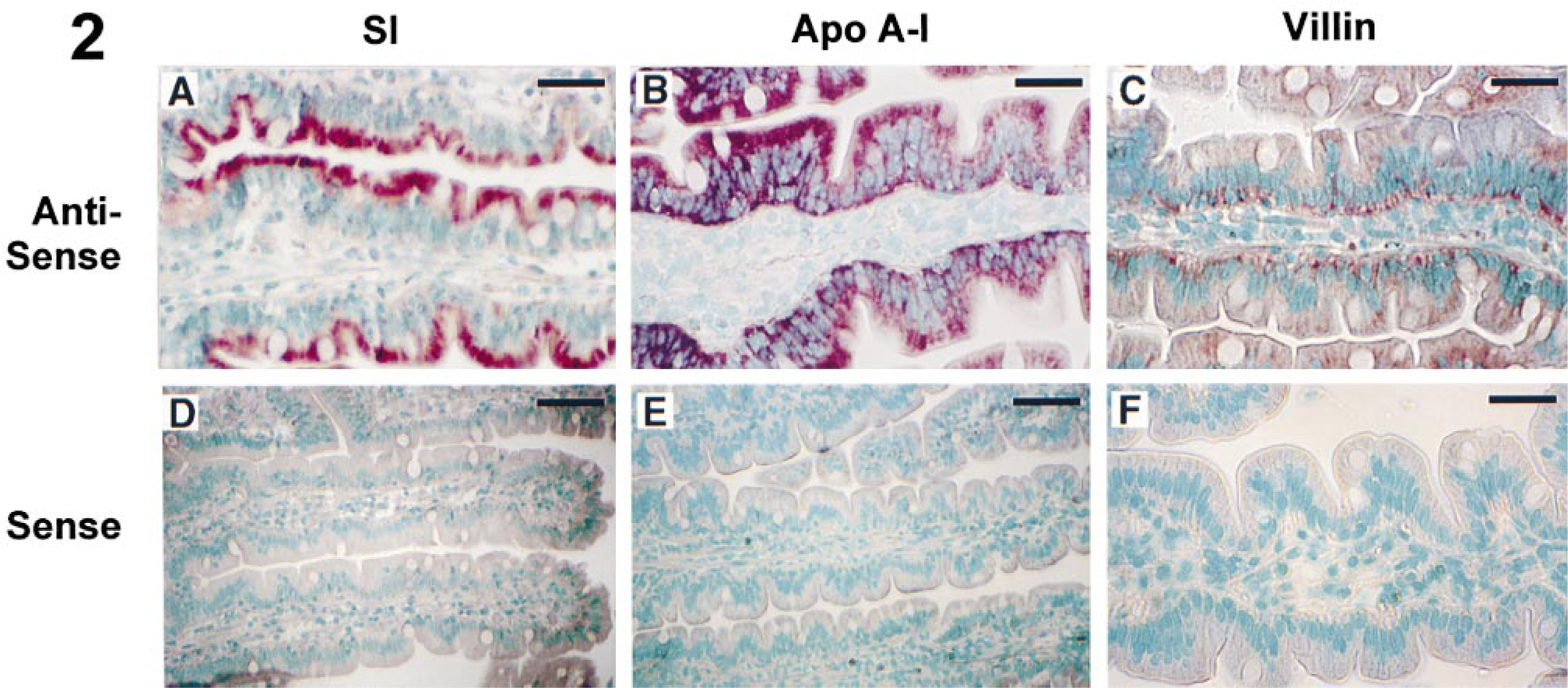
In situ hybridization for human SI, Apo A-I, and villin. (
Human BBMI cDNA
To date, most of the cis-acting elements responsible for mRNA sorting have been found in the 3'-UTR (Hesketh 1996; Oleynikov and Singer 1998). To facilitate the study of potential mechanisms of mRNA targeting in enterocytes, a full-length human cDNA, including the 5'- and 3'-UTRs, was cloned by 5' and 3' RACE as described in Materials and Methods. To obtain full-length sequence information at the 5' end of the mRNA, 62 clones containing the BBMI sequence were selected from more 200 clones containing PCR inserts. Four clones containing the longest PCR fragments were sequenced from both directions to obtain the complete 5'-UTR sequence information. The full-length human BBMI cDNA sequence was available from GenBank (accession #AF105424).
Sequencing of hBBMI revealed a single open reading frame of 3132 bp. The ATG start codon was found 210 bp downstream from the 5'-end of the cDNA. An in-frame termination codon was present 48 bp upstream of the translation start codon. The 3'-UTR was 237 bp before the poly(A) tail. A polyadenylation signal was also present in the 3'-UTR. A search of the GenBank revealed that our cDNA is highly homologous to another recently cloned hBBMI cDNA containing only the full-length coding region (Skowron et al. 1998) (GenBank accession #AF009961). Sequence analysis indicated that the two sequences differ only in six bases in the coding region, which include G at 383 in our clone→C at 174 in AF009961, A1546→C1337, G1547→C1338, A1616→G1407, C2343→G2134, and G2726→A2517. The coding sequence showed 86%, 85%, 86%, and 65% homology with bovine (Hoshimaru and Nakanishi 1987), rat (Balish and Coluccio 1995), mouse (Skowron and Mooseker 1999), and chicken (Collins and Matsudaira 1995) BBMIs, respectively. It also had 99% homology with a 102-bp PCR fragment of previously identified human unconventional myosin heavy chain (Bement et al. 1994). The comparative analysis of available sequences for the 5'-UTR showed approximately 78% and 42% sequence homology with its peers from bovine and chicken BBMIs, respectively. The 3'-UTR showed around 71% and 46% sequence homology with those of bovine and chicken, respectively. In the 3'-UTR of hBBMI, a series of sequence repeats was identified. Sequences CCCCTCTG and TCCTCCAA occurred twice, and ACCCTT occurred three times (Figure 3).
The deduced hBBMI amino acid sequence encodes a 1043-residue polypeptide with a deduced molecular weight of 118,387 D. It differs only in two amino acid residues from AF009961: Q446→P446 and R712→G712. It shares 87% identity and 90% similarity with bovine BBMI, 88% identity and 91% similarity with rat BBMI, and 64% identity and 70% similarity with chicken BBMI. hBBMI has a putative ATP binding sequence at its N-terminus and an actin binding domain in the middle region (Kawakami et al. 1992). A second putative ATP/GTP binding site, which is absent in the bovine BBMI, is also present at amino acid position 669. The functional significance of the additional ATP/GTP binding site in hBBMI is unknown. Three putative calmodulin binding domains are also highly conserved at the IQ region, as previously described (Kawakami et al. 1992).
Northern Blot Analysis
Using the full-length hBBMI cDNA as a probe in Northern blot analyses, we demonstrated that hBBMI was detected as a ~3.6-kb single band species in total RNA extracts from human jejunum and Caco-2 cells (Figure 4). This size is consistent with our cloned cDNA and also suggests that no alternative splicing occurs in the tissue and cells examined, as suggested for rat BBMI (Balish and Coluccio 1995).
Discussion
In the past decade, intensive investigation of protein transport has yielded important insights into the mechanisms involved in protein localization in epithelial cells (Wollner and Nelson 1992; Drubin and Nelson 1996; Caplan 1997). However, the role of mRNA sorting and its asymmetrical localization in the control of protein expression in these cells has not been as fully delineated. Previous studies have demonstrated that mRNA sorting may play an important role in the establishment and maintenance of cellular protein gradients and morphological polarity (Gavis and Lehmann 1992; St Johnston 1995). As elaborated above, the dichotomy between basal localization of villin mRNA and apical distribution of its encoded protein has raised an interesting question about the role of mRNA sorting in the establishment and maintenance of enterocyte polarity. To search for other similar dichotomies and to investigate the role of mRNA sorting in enterocyte polarity, we examined hBBMI and I-fim mRNA distribution patterns in human jejunal enterocytes. Our results revealed a dominant basal localization of BBMI mRNA, similar to that of villin mRNA distribution. This coincidental similarity between the mRNA distributions for two major microvillous core cytoskeletal proteins (villin and BBMI) supports our previous speculation (Li et al. 1998) that these proteins may be initially translated at the basal region and assembled into an increasingly more mature complex during transport to the microvillous core. The speculation is further supported by our previous finding that β-actin mRNA is also localized at both apical and basal regions in human enterocytes (Barth et al. 1998). Translocation of BBMI from the cytoplasm to the microvillous core has also been suggested by the previous detection of vesicle-associated BBMI in the cytoplasm (Drenckhahn and Dermietzel 1988; Hayden et al. 1990).

The 3-UTR of hBBMI cDNA. Polyadenylation signal is bolded. The three repeating elements (E1, E2, and E3) in the 3'-UTR discussed in the text are underlined. The full-length human BBMI cDNA clone has been deposited in the GenBank with accession #AF105424.
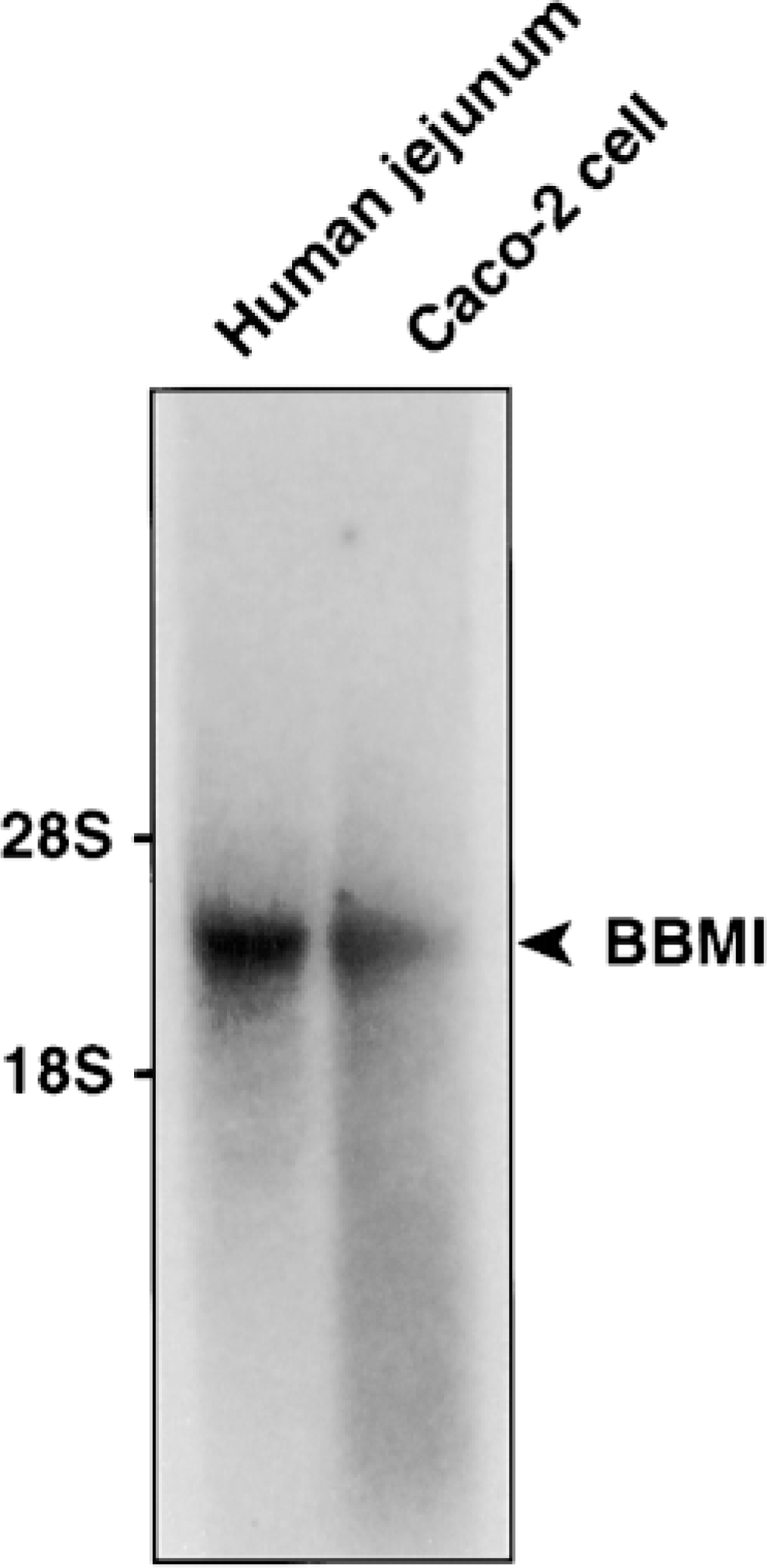
Northern blot. Total RNA (10 μg/lane) from human jejunum and Caco-2 cells were analyzed by Northern blotting using full-length hBBMI cDNA as a probe. Only a single ~3.6-kb mRNA species was present in both samples.
The mRNA distribution of I-fim, another microvillous core cytoskeletal protein, was also examined by nonradioactive in situ hybridization. In contrast to the basal localization of villin and BBMI mRNAs, I-fim mRNA distributes diffusely throughout the cytoplasm in enterocytes. This distinction also indicates that the mRNAs for microvillous core cytoskeletal proteins distribute in enterocytes in different patterns. Because of the previous demonstration of apical localization of I-fim protein in the microvillous core region (Shibayama et al. 1987; Drenckhahn and Dermietzel 1988; Ezzell et al. 1989), I-fim must be translated in different regions of the cytoplasm and then translocated to the apical region.
The mechanism of mRNA sorting remains largely unknown, although some studies have linked sequence repeats in the 3'-UTR to the anchoring site for mRNA binding to cytosolic proteins. For example, Kislauskis et al. (1994) reported that if the localization of β-actin transcripts at the leading lamellae of chicken embryonic fibroblasts was disrupted by antisense oligonucleotides directed against the repeated localization elements (GGACT and AATGC) in the 3'-UTR, β-actin protein was no longer concentrated at the leading edge of the cells, although levels of both protein and mRNA did not change. This alteration eventually resulted in the collapse of the lamellae, and the cells became symmetric. The cytosolic protein responsible for the mRNA binding to the ACACCC repeats from the 3'-UTR of β-actin was also cloned recently (Ross et al. 1997). Similarly, sequence repeats in the 3'-UTR of Vg1 mRNA also bind to a cytosolic protein responsible for the mRNA localization to the vegetal cortex of Xenopus oocytes (Deshler et al. 1997).
Although sequence analysis of the hBBMI cDNA revealed several repeats in its 3'-UTR (ACCCTT, TCCTCCAA and CCCCTCTG) (Figure 3), the significance of these repeats for mRNA asymmetrical distribution is unclear. These repeats are not found in the 3'-UTR of β-actin mRNA nor in the 3'-UTR of human villin mRNA although, as discussed above, different repeated sequences are found in the β-actin 3'-UTR (Ross et al. 1997). Functional assessment of these regions awaits further study.
The cloning of the hBBMI cDNA revealed some unique features, as well as conserved sequences shared with bovine, rat, and chicken BBMIs. The hBBMI cDNA has the highest homology to bovine BBMI, followed by rat and then chicken BBMI; a similar pattern is found in analysis of the deduced amino acid sequence. Similar to bovine, rat, and chicken BBMI, hBBMI has a putative ATP binding domain at its N-terminus followed by an actin binding site and an IQ domain containing three putative calmodulin binding motifs. However, hBBMI possesses an additional putative ATP/GTP binding domain between the actin binding site and calmodulin binding sites. The functional significance of this site is unknown. It will be interesting to examine the functional difference of mutated hBBMI lacking the second ATP/GTP binding site.
The mechanisms controlling mRNA sorting in intestinal epithelial cells and their role in the establishment and maintenance of epithelial polarity remain to be elucidated. The distinct localizations of hBBMI and I-fim mRNAs, as well as the basal localization of villin mRNA in enterocytes and the cloning of the hBBMI cDNA, including its 3'-UTR, offer the opportunity to expand our understanding of the functional significance of mRNA sorting in enterocytes.
Footnotes
Acknowledgements
Supported by National Institutes of Health Research Grant R01-DK-32658, Digestive Disease Core Center Grant P30-DK-34928, and Pediatric Gastroenterology Research Training Grant T32-DK-07471.
We wish to thank Drs Robert Montgomery, Stephen Krasinski, Menno Verhave, and Diana Bianchi for scientific discussion, Dr Scott Shikora for human specimens, and Dr Anne Kane (Director of the Microbiology Core, GRASP Center) for plasmid preparations.
