Abstract
Fibroblasts are the principal mesenchymal cell type found within the connective tissues of all organs. Once thought to play a passive role in tissue remodeling, fibroblasts have now emerged as a key player in regulating structural immunity and modulating the reparative injury response. A recent surge in single-cell RNA sequencing studies has advanced our understanding of the biology of fibroblasts, highlighting their cellular diversity and organization across health and diseased conditions at an unprecedented resolution. In this review, we discuss up-to-date literature on fibroblast subpopulations identified from 2 distinct barrier tissues: oral mucosa and skin. We focus on the transcriptomic signatures that distinguish subsets of fibroblasts in homeostasis and perturbed conditions (i.e., wound healing or chronic inflammatory diseases), and we link them to mechanistic studies that provide functional insights. A deeper understanding of fibroblast diversity and its functional significance may uncover tissue-specific roles in regeneration and immunomodulation, which will be crucial for the development of precision therapy that directly targets fibroblast subsets.
Keywords
Introduction
Fibroblasts were first described in the 19th century as spindle-shaped cells of the connective tissue. Perhaps owing to this designation based on morphology, there had been ambiguity to whether fibroblasts should be truly classified as a single-cell type. We now know that morphology alone is inadequate and does not convey the functional significance of fibroblasts. Fibroblasts are commonly identified by the positive cell surface protein expression of pan-fibroblast markers such as CD140A and/or CD90 and the lack of leukocytic, endothelial, epithelial, and erythrocyte lineage markers (CD45-, CD31-, CD326-, Ter119-, respectively). However, the aforementioned criterion is not an absolute definition of fibroblasts, since it also identifies multipotent skeletal/mesenchymal stromal cell subsets in the long bones (Ambrosi et al. 2019). Thus, single-cell RNA sequencing (scRNA-seq) studies on fibroblasts typically select for additional extracellular matrix genes, such as
The oral cavity is subject to repeated trauma from mastication and microbial insult. Despite this perpetual injury, oral mucosae heal exceptionally fast and with minimal scar formation (Iglesias-Bartolome et al. 2018). Moreover, the damage induced by periodontal pathogens in periodontitis is typically gradual and rarely results in sepsis. The resilient nature of the oral mucosa had been attributed to the neural crest developmental origin of oral fibroblasts (Rinkevich et al. 2015). In contrast, skin fibroblasts have multiple developmental origins: paraxial mesoderm gives rise to dorsal skin fibroblasts and lateral plate mesoderm to ventral trunk and limb skin, and, uniquely, facial skin fibroblasts are derived from the neural crest. Facial fibroblasts exhibit higher proliferative capacity and contribute to an improved wound-healing profile as compared with other skin regions (Usansky et al. 2021), which is linked to the continued expression of tissue-specific homeobox genes throughout adulthood (Pfeiferová et al. 2025). These studies link the developmental origin to the niche-specific function of the fibroblasts, but the distinguishing features between oral and skin fibroblasts at a single-cell level had been underexplored. Emerging scRNA-seq studies have addressed this challenge, revealing that multiple fibroblast subpopulations are found in the oral mucosa and skin and providing novel insights into understanding their functional significance in health and disease.
Oral and cutaneous barriers closely resemble each other at the histologic level. Yet, fibroblast composition and function in these anatomic sites are expected to differ. In this review, we provide an up-to-date review on research development in the single-cell biology of fibroblasts in the oral mucosa and skin. Specifically, we discuss the gene markers that define fibroblast diversity in these barrier tissues from mouse and human models. We further discuss the scRNA-seq and mechanistic studies that investigate the role of these fibroblast subpopulations in the context of wound healing and chronic inflammatory disease conditions.
Overview of the Structural Similarities and Differences in Oral Mucosa and Skin
Oral mucosa and skin share structural similarities and differences (Fig. 1). In humans, keratinized oral mucosa, also known as gingiva, and skin possess stratified squamous epithelial structures. The underlying connective tissue compartments are divided into papillary and reticular layers based on location and extracellular matrix features. The papillary layer is located beneath the epithelium, in which thin collagen and elastin fibers form tightly packed projections into the epithelium. The reticular layer is deeper and possesses thick and interwoven collagen and elastic fibers, creating a meshwork that provides mechanical support. In palatal gingiva, there is a submucosal adipose layer that contains minor salivary glands, similar to a human subcutaneous adipose layer surrounding eccrine and apocrine glands. In contrast, skin has other specialized apparatuses, such as hair follicle, sweat glands, and arrector pili muscles that are not present on oral mucosal layers. In the human oral cavity, buccal mucosal lining consists of nonkeratinized epithelium and is highly elastic and rich in glandular tissues, contrasting gingiva that is immovable and keratinized. The dorsum of the tongue has a thick keratinized layer and specialized papillar structures, whereas the ventrum is thin, nonkeratinized, and highly vascular. The oral mucosae in mice share these features, although all regions of the murine oral mucosa are keratinized and the thickness of each histologic layer differs between species. This review focuses on fibroblasts that are found in the connective tissue layers of oral mucosa and skin.
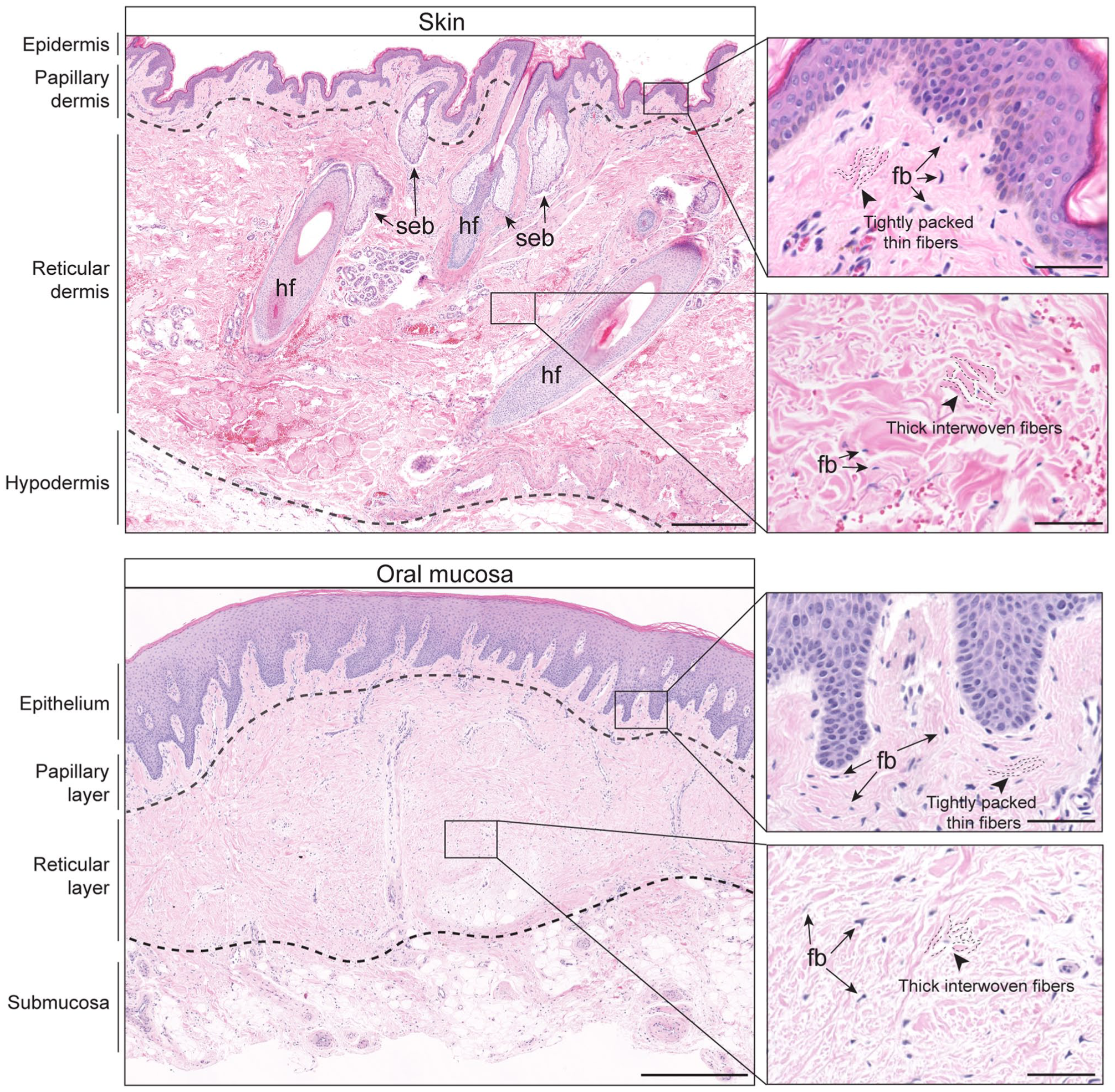
Histologic features of human skin and oral mucosa. Top: hematoxylin and eosin–stained image of human forearm skin shows different histologic compartments of epidermal, dermal, and hypodermal layers demarcated by dashed lines. Specialized appendages such as hair follicles (hf) and sebaceous gland (seb) are present in normal skin. Scale bar, 500 µm. Bottom: hematoxylin and eosin–stained image of human palatal gingiva with similarly layered architecture as compared with skin composed of keratinized stratified squamous epithelium, underlying lamina propria and submucosa. Scale bar, 1 mm. Inset images show high-magnification view of a papillary connective tissue layer that has thin and tightly packed collagen fibers and a reticular layer with thick and interwoven collagen fibers as indicated by dotted lines. Arrows point to spindle-shaped fibroblastic cells (fb). Scale bar, 50 µm.
Fibroblast Heterogeneity in the Oral Mucosa and Skin during Homeostasis
Dermal Fibroblasts
Since the 1970s, it was speculated that dermal fibroblasts from the papillary versus reticular layers were distinct by their differential proliferative capacity in vitro (Harper and Grove 1979). This was later demonstrated in murine models, in which fibroblasts within the papillary layer of neonatal skin were CD26+ (also known as DPP4) and the reticular layer and hypodermis consisted of DLK1+ or SCA1+ fibroblasts, respectively (Driskell et al. 2013). These early findings have been validated and utilized by multiple scRNA-seq studies (Thompson et al. 2022; Almet et al. 2025). Joost et al. (2020) identified 4 distinct fibroblast subpopulations from a total of 340 cells, each marked by the differential expression of
In an scRNA-seq study, Tabib et al. (2018) found 2 major dermal fibroblast populations in healthy human forearm skin, in which
Single-Cell RNA Sequencing Studies of Fibroblasts in Homeostatic Skin and Oral Mucosa.

NA, not available.
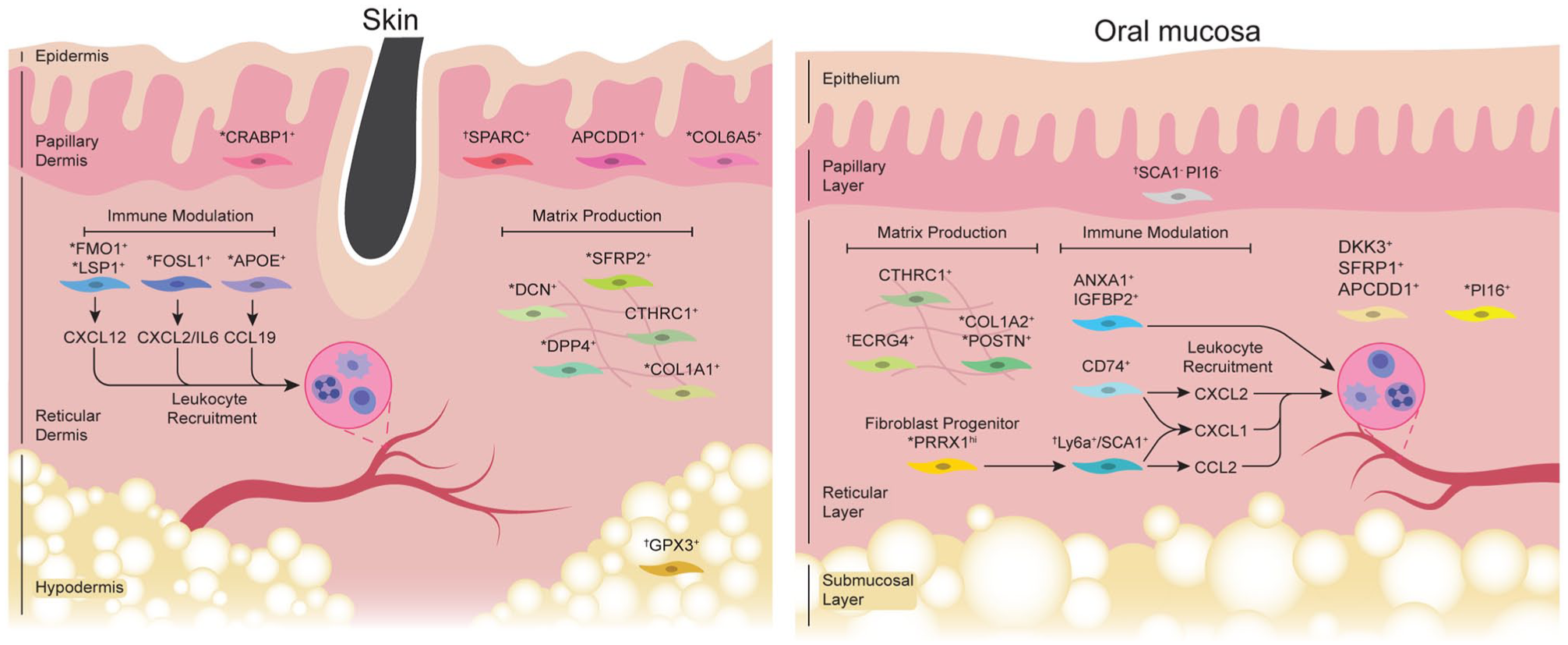
Fibroblast subpopulations in skin and oral mucosa during homeostasis. Left: fibroblast subpopulations identified from normal skin. The papillary layer is composed of
Oral Fibroblasts
Our group reported that murine palatal gingiva harbors 6 fibroblast subtypes as identified from the scRNA-seq (Ko et al. 2023). The top 4 fibroblast clusters had enriched expression of
There is an abundance of human scRNA-seq studies that examine healthy oral tissues. Anterior hard palate contains
Fibroblast Subpopulations in Wound Healing
Wound healing is a complex biological process characterized by hemostasis, inflammation, proliferation, and remodeling. Fibroblasts play a critical role in every phase, with distinct subpopulations emerging over time and across spatial niches in dermis (Fig. 3A). In a multi-omics study, Foster et al. (2021) examined murine dorsal skin wound healing across 2-, 7-, and 14-d time points and found an “activated-responder” fibroblast subset that expressed inflammation-related genes such as
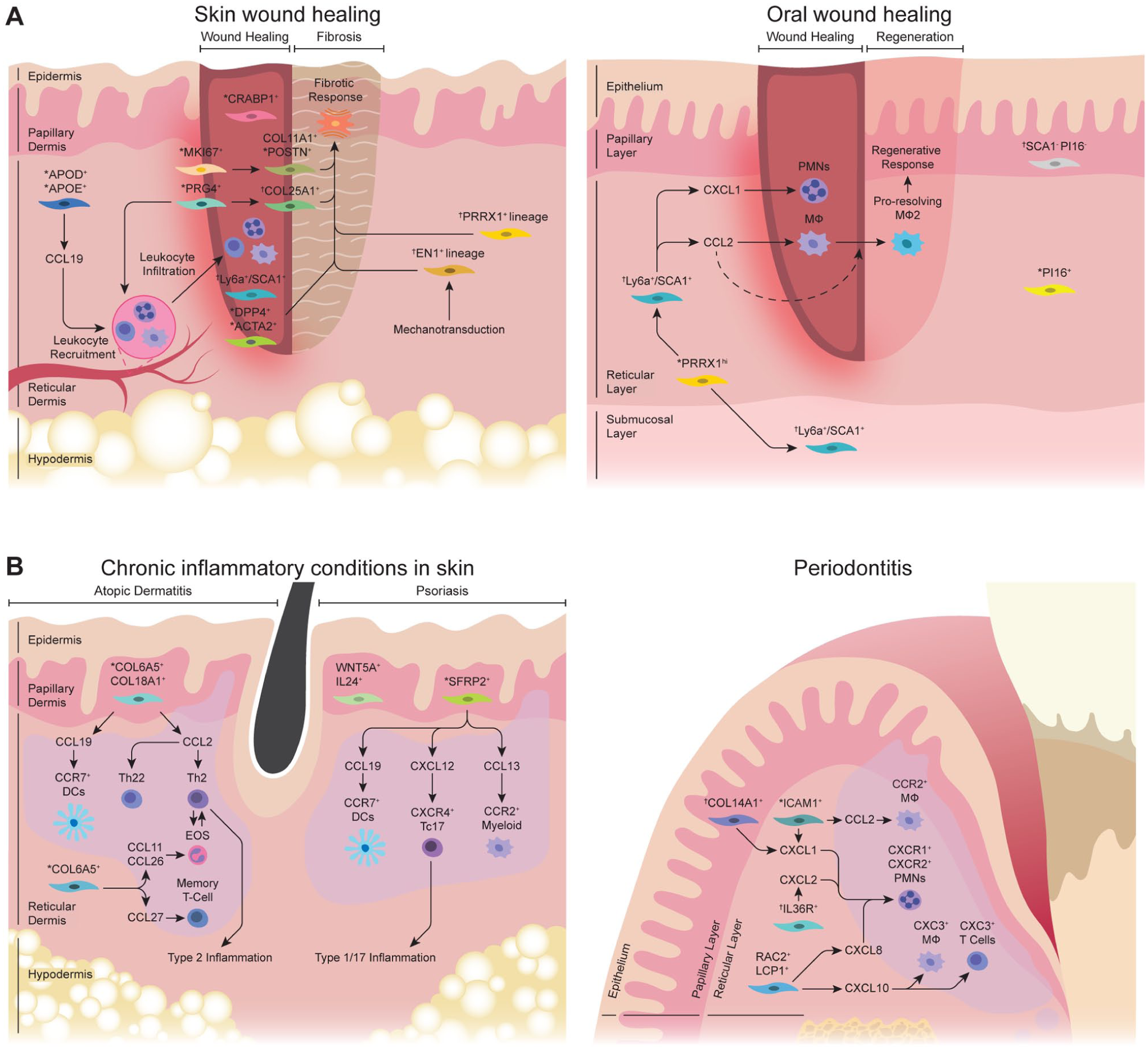
Fibroblast subpopulations under perturbed conditions in skin and oral mucosa. (
In a recent study, Liu et al. (2025) employed scRNA-seq and spatial transcriptomic approaches to examine human skin wound repair over 1-, 7, and 30-d time points. The study identified 4 major fibroblast clusters, each designated as a mesenchymal (
Functional In Vivo Oral and Skin Wound-Healing Studies
scRNA-seq studies have identified distinct fibroblast subsets during wound healing, but their biological relevance is underexplored. Our group reported that postnatal fibroblasts with active
Fibroblast Subpopulations in Chronic Inflammatory Conditions
Periodontal Disease
Periodontitis is a prevalent chronic inflammatory disease characterized by persistent connective tissue inflammation and progressive destruction of tooth-supporting structures. There exists abundant scRNA-seq studies that investigate the cellular and transcriptional dynamics in human periodontitis (Caetano et al. 2021; Williams et al. 2021; Caetano et al. 2023). These studies collectively point to an inflammatory gingival fibroblast subset expressing CCL- or CXCL- cytokines as a prominent feature when comparing periodontitis with healthy groups (Fig. 3B). Williams et al. (2021) demonstrated that the fibroblast subset marked by
Murine Studies on Fibroblasts in Periodontitis
Several studies have investigated the role of fibroblasts using murine models of periodontitis. Kondo et al. (2022) demonstrated that
Chronic Inflammatory Diseases of the Skin
Similar to oral mucosa, the skin barrier is susceptible to chronic inflammation (Fig. 3B). Atopic dermatitis (AD) is a prevalent chronic skin disease characterized by overt type 2 inflammation and allergic pruritus. scRNA-seq studies on human atopic skin have largely validated dysregulated epithelial barriers and inflammatory milieu but also revealed unexpected changes in dermal fibroblasts. Reynolds et al. (2021) reported that the number of fibroblast subsets that expresses
Psoriasis is another common chronic skin disease characterized by thick scaly plaques and is biased toward type 1 and 17 inflammation (Greb et al. 2016). scRNA-seq studies on psoriasis identified a proinflammatory fibroblast population marked by upregulated expression of
Functional Studies on Fibroblasts in Skin Inflammatory Diseases
In a murine AD study using epicutaneous ovalbumin sensitization, fibroblast subsets exhibited an inflammatory phenotype marked by increased expression of
Single-Cell RNA Sequencing Studies of Fibroblasts for Perturbed Conditions in Skin and Oral Mucosa.

FB, fibroblast; NA, not available; POD, postoperative day.
Concluding Remarks and Future Perspectives
Fibroblasts constitute heterogeneous subpopulations, each equipped with a specialized gene expression profile and spatial distribution in skin and oral mucosa. Other than extracellular matrix–associated genes, these fibroblast subsets differentially express transcripts that are implicated in inflammation and distinct molecular pathways. A major limitation in interpreting individual scRNA-seq studies is the inconsistent cell nomenclature and varying number of cell identities. This likely stems from a combination of technical variations, such as cell dissociation protocols and/or computational workflow, which can vary widely among studies. To address this, an integrative approach to categorize fibroblast subsets has been performed for skin fibroblasts (Ascensión et al. 2021; Almet et al. 2025), which consistently identified
An integrated scRNA-seq atlas of human oral and craniofacial tissues is available (Fernandes Matuck et al. 2024). However, a detailed fibroblast-centric analysis across oral mucosal sites is lacking and remains a promising future direction. In the oral cavity, different anatomic sites are under various functional demands, and site-specific disease susceptibility is a well-known phenomenon. Similar fibroblast features implicated in chemotaxis and complement activation are found in healthy gingiva and buccal mucosa (Williams et al. 2021). This may be explained by the shared neural crest developmental origin and/or exposure to oral microbiota. Yet, it is worth noting that fibroblasts in these anatomic sites have not been compared under perturbed conditions (i.e., wounding), which may reveal divergent molecular pathways. An integrative oral fibroblast atlas may also identify stable and intrinsic markers for distinct oral fibroblast subset that must precede development of new mouse models for mechanistic studies in the future.
While informative, scRNA-seq analysis alone cannot resolve the spatial context of cells in a given niche. Spatial transcriptomics is a rapidly evolving methodology that can unravel gene expression patterns within complex tissues, and its application for examining oral structures is continually expanding (Caetano and Sharpe 2024; Haller et al. 2024). In the skin biology field, this technology has been employed for human wound healing and chronic inflammatory diseases conditions (Ganier et al. 2024; Steele et al. 2024; Liu et al. 2025). As the detection voxel size improves down to 2 µm (Visium HD) and the single-cell resolution becomes capable of detecting up to 5,000 transcripts (Xenium), we anticipate that future studies will provide a spatial context to oral fibroblast identity and function at an unprecedented resolution. A comprehensive analysis of various oral tissues in health and disease will be the crucial next step toward understanding remarkable fibroblast heterogeneity in the oral cavity, identifying functionally relevant fibroblast subsets, and developing precision therapy targeting distinct subpopulations.
Author Contributions
K. Prasongyuenyong, K.I. Ko, contributed to conception and design, data acquisition, analysis, and interpretation, drafted and critically revised manuscript; W.S. Kim, contributed to conception and design, data acquisition, analysis, and interpretation, drafted and critically revised manuscript; Z. Chen, contributed to conception and design, drafted the manuscript. All authors gave final approval and agreed to be accountable for all aspects of the work.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by funding from the National Institutes of Health (to K.I.K.; R01-DE030415 and R01-AR082951).
