Abstract
More than 1 billion individuals worldwide have experienced dental trauma, particularly children aged 7 to 12 y, predominantly affecting the anterior teeth, which has a significant impact on oral health and esthetics. Rapid emergency restorations using composite resin are followed by medium-term lab-fabricated mock-ups. Recent advancements in artificial intelligence (AI) assist dental restorations, and the objective of this study was to compare the performances of different AI approaches for the learning and reconstruction of central incisors. The study was approved by ethical committees and followed AI in dentistry recommendations. STL files of mature permanent maxillary incisors without severe wear were collected from 3 universities. Principal component analysis (PCA) and Deep Learning of Signed Distance Functions (DeepSDF) models were trained using these files. The learning of PCA and DeepSDF approaches were 3-fold cross-validated, and their performances were assessed using the following metrics to measure the reconstruction accuracy: the difference of surfaces, volumes, lengths, average Euclidian distance, Hausdorff distance, and crown–root angulations. Explainability was assessed using feature contribution analysis for PCA and Stochastic Neighbor Embedding (t-SNE) for DeepSDF. DeepSDF showed significantly better precision in surface, volume, and Hausdorff distance metrics compared with PCA. For reconstructions, the lower size of the latent code of the DeepSDF model demonstrated lower performances compared with higher sizes. In addition, DeepSDF raised concerns about explainability. This study demonstrates the potential of PCA and DeepSDF approaches, particularly DeepSDF, for the learning and reconstruction of the anatomy of upper central incisors. To foster trust and acceptance, future research should, however, focus on improving the explainability of DeepSDF models and considering a broader range of factors that influence smile design. These high performances suggest potential clinical applications, such as assisting practitioners in future smile designs and oral rehabilitation using AI approaches.
Keywords
Introduction
More than 1 billion individuals worldwide have experienced a dental trauma, with a notably higher prevalence in children between 7 and 12 y old, and predominantly affecting the anterior teeth (Petti et al. 2018). Such traumas significantly impair oral health, masticatory function, esthetics, and psychological well-being. In emergencies, practitioners must rapidly restore the original shape of the teeth, typically using direct chairside layering of composite resin. However, achieving the ideal tooth shape and ensuring patient satisfaction with the rehabilitation plan remains challenging due to the subjective nature of esthetic perceptions (Tosun and Kaya 2020) and because technical expertise is acquired through many years of practice (Machado et al. 2013; Pham and Nguyen 2022).
Artificial intelligence (AI) has emerged as a transformative tool, enabling dental practitioners to design, plan, and restore patients’ smiles with enhanced customization and predictability (Jafri et al. 2020; Touati et al. 2022). However, the development and implementation of these AI-based systems are predominantly driven by manufacturers, leading to a reliance that may limit practitioners’ understanding of the underlying processes (Rokhshad et al. 2024). Moreover, these systems currently lack transparency and often fail to provide sufficient explainability, making the decision-making process opaque and raising concerns regarding potential errors and accountability.
Traditional approaches such as principal component analysis (PCA) have been extensively used in dental shape reconstruction (Mehl et al. 2005; Litzenburger et al. 2013). However, PCA is primarily a statistical dimensionality reduction tool that relies on a mesh. This dependence on predefined landmarks limits its ability to capture fine shape variations but also can introduce inconsistencies due to misalignment or variability in landmark placement (Mehl et al. 2005; Litzenburger et al. 2013; Iyer et al. 2023). More recently, deep learning approaches, such as generative adversarial networks (GANs), have achieved high performances in prosthesis design (Chau et al. 2024). However, these methods remain constrained by voxel-based representations, requiring extremely fine resolutions to capture anatomical details, which significantly increases computational complexity. Moreover, these approaches are often perceived as black-box systems, raising concerns about their explainability (Chau et al. 2024).
Deep Learning of Signed Distance Functions (DeepSDF) has recently emerged as an elegant alternative, as it does not learn a discrete mesh but instead represents a surface continuously, making it less prone to discretization artifacts (Park et al. 2019). As opposed to GANs, which implicitly learn data distributions without explicit geometric representation, DeepSDF explicitly encodes the distance of any 3-dimensional (3D) point to the nearest surface, enabling precise and accurate reconstructions even from partial data. This method has been successfully adapted to reconstruct fractured incisors from a dataset of 80 anatomies acquired by cone-beam computed tomography (CBCT), demonstrating its potential for dental applications (Chen et al. 2024). The hypothesis is that DeepSDF could manage a broader dataset of incisors scanned with high-quality intraoral scanners, potentially surpassing PCA techniques in performance, while still maintaining a high explainability. Therefore, the present study aimed to evaluate the performances of DeepSDF approaches in comparison with PCA, with a primary focus on their effectiveness in learning the anatomy of upper central incisors. Secondarily, the objectives aimed to evaluate their ability to explain shape variability in dental morphology and to determine their potential to achieve clinically relevant reconstructions with high anatomical fidelity.
Materials and Methods
Dataset
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the institutional review board of the Hospices civils de Lyon (ethical approval No. HCL-23_5271). It was also conducted and reported following the recommendations on AI in dentistry (Schwendicke et al. 2021) (Appendix Table 1). A set of STL files was retrospectively collected from the Indiana University School of Dentistry (Indianapolis, IN, USA; ethical approval No. NS0911-07), Lyon Dental University (Lyon, France; ethical approval No. HCL-23_5271), and the Faculty of Medicine (Leuven, Belgium; ethical approval No. B322201525552). Only mature permanent maxillary central incisor teeth without severe tooth wear were included (tooth wear index ≤ 2); all teeth presenting decay, enamel defects, restorations, or cervical lesions were excluded. For each tooth, the STL file was classified by 2 dental experts in 3 basic groups, according to previous studies: square, ovoid, and triangular shapes (Hussain et al. 2016; Mahn et al. 2018). Intraexaminer reliability was tested through determination of the Cohen’s kappa coefficient. In case of discrepancies, a third expert evaluated the tooth to allow a consensus to be reached. To ensure the robustness of the statistical analyses, the recommendations for events per variable (EPV) in logistic regression analysis were followed (Austin and Steyerberg 2017). EPV was applied to evaluate the relationship between 5 anatomical parameters and eigenvalues derived from PCA, requiring a minimum of 50 samples. In addition, PCA typically requires 5 to 10 samples per dimension, suggesting that retaining 10 eigenvectors would necessitate at least 100 samples (Jolliffe 2002).
Preprocessing
Each STL file underwent a preprocessing step to ensure consistency in alignment and mesh quality. The first tooth was arbitrarily used to establish a coordinate reference position, scale, and rotation, to automatically align all shapes, using an iterative closest-point algorithm (Billings et al. 2015). The long axis of each tooth was therefore aligned on the same axis and checked for low distorted mesh by an expert with more than 5 y of experience using the computer-aided design software Rhino 7 (Rhinoceros 3D, Robert McNeel & Associates).
Remeshing and Application of PCA
For the PCA approach, the surfaces of each tooth were remeshed to construct a similarly structured homologous and isotopological mesh following the MEG-IsoQuad method (Maquart et al. 2021). Based on the STL files isotopologic to a surface, the surface was partitioned into cuboids using a geometry-feature-aware algorithm. This involved applying a global parameterization to align the principal curvature directions and sharp features on the surface. Node connection constraints were used to restrict each arc to lie on a common isoparametric curve. The global parameterization was optimized using a gradient-based algorithm by iteratively repositioning the nodes based on the gradient of the objective function until reaching a satisfactory global embedding quality. The coordinates of the landmarks of all the remeshed files were then used as input parameters for PCA, to extract the main shape variations. The influence of each mode M was then evaluated using the compactness, that is, the proportion of the first M modes on the total variance of all modes, and only the most influent modes were retained to reproduce the shape (Mehl et al. 2005).
Modeling Continuous Signed Distance Functions
For the DeepSDF approach, a dataset of pairs X, consisting of 3D point samples (500,000 points) and their corresponding Signed Distance Field (SDF) values, was generated to learn the shape of each STL file. To ensure high-density sampling near the tooth surface, 94% of the points were placed within 0.25 mm from the surface, randomly perturbed along their normal vectors using Gaussian noise (σ = 0.25 mm). The remaining 6% of the points were sampled randomly within a bounding box extending 2 mm beyond the tooth’s maximum extents. For each point, the SDF value was calculated, which indicated the proximity to the nearest surface and whether the point was inside (negative) or outside (positive) the surface. A neural network was then trained to predict the SDF value for any spatial query point related to each specific STL file. The network, functioning as an auto decoder, comprises several fully connected (FC) layers that generate the SDF values (Fig. 1). In the DeepSDF learning configuration, both the network parameters and the latent code z were optimized concomitantly. The loss function was calculated following methodologies from previous studies (Park et al. 2019). Once trained, the network provides a continuous function that predicts the SDF value for any 3D point in space. Using this function, the tooth’s surface can be reconstructed by extracting the points for which the SDF value is zero: representing the interface between the inside and outside of the surface. These zero-crossing points are then processed by the Marching Cubes algorithm.
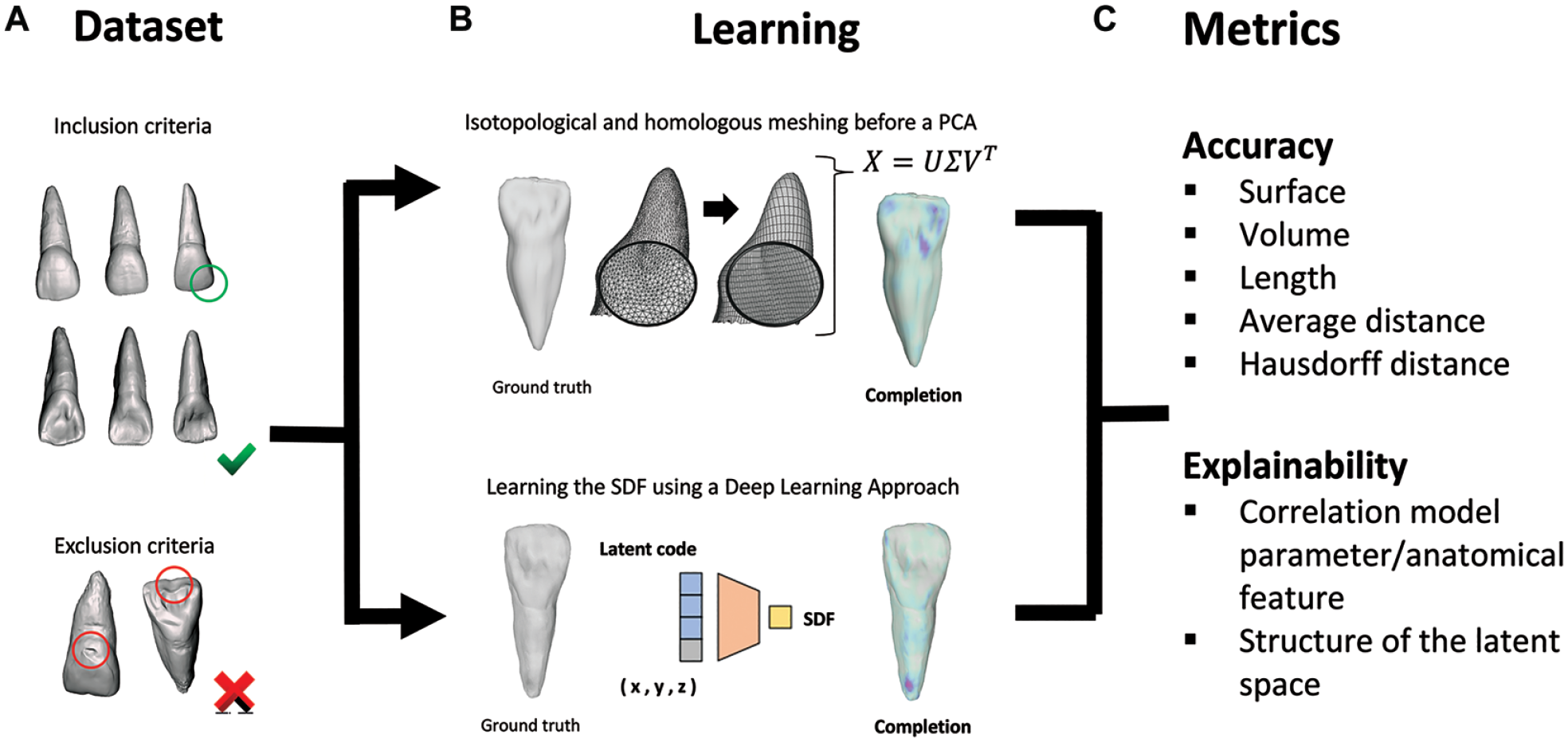
Workflow for evaluating the learning with (
Validation and Performance Metrics
The learning of PCA and DeepSDF approaches were 3-fold cross-validated, performed with random splits across all STL files, and their performances were evaluated using the following metrics to measure the reconstruction accuracy: the difference of surfaces, volumes, lengths; the average Euclidean distance defined as the mean positional error between each point and its nearest triangle in the other shape; the Hausdorff distance, defined as the maximum distance from any point in 1 point cloud to the nearest triangle in the other; and crown–root angulations (Binvignat et al. 2024). For the DeepSDF approach, performances of different sizes of the latent code (8, 16, 64, 128, and 256) were compared. To further evaluate the clinical applicability of the DeepSDF models, an external test dataset was used, consisting of 900 intraoral optical scans of upper dental arches from the publicly available 3DTeethSeg’22: 3D Teeth Scan Segmentation and Labeling Challenge dataset (Appendix Table 2) (Ben-Hamadou et al. 2023).
Explainability
For the PCA approach, explainability was assessed using the Feature Contribution Analysis to understand how each principal component influenced the shape variation (Binvignat et al. 2024). For the DeepSDF approach, explainability was assessed using shape interpolation between 2 latent codes. In addition, the latent space was analyzed using Stochastic Neighbor Embedding (t-SNE) (van der Maaten and Hinton 2008; Luo et al. 2024) and the performance of a random forest algorithm to classify tooth shape categories (triangular, rectangular, ovoid) based on the values of the latent code (Binvignat et al. 2024).
Five anatomical features were extracted from each tooth: volume of the tooth, external surface, tooth inclination, crown–root angulation, and tooth length. After normality testing using Kolmogorov–Smirnov, descriptive statistics were expressed as the mean and standard deviation. Thereafter, the Pearson correlation coefficient was calculated between the coefficients of the anatomical features and the principal modes and similarly between the coefficients of the anatomical features and the values of the latent code.
Virtual Simulation of Tooth Reconstruction
Fractures were simulated by defining a cutting plane 1 mm below the highest point of the principal axis of each tooth, inclined randomly between −50° and +50° using a script developed in Python 3.8. The section below this plane was retained as the fractured model. Each original model underwent 5 different simulated fractures to create a dataset of 1,425 fractured incisors. Subsequently, shapes were reconstructed using an auto decoder based on the values of the latent code. The performance of reconstructions was evaluated by comparing differences in the 5 anatomical features and the distances between nodes on the intact and repaired surfaces.
Implementation Details and Network Training
The network was implemented using PyTorch, and the Adam optimizer was used to train the network (Kingma and Ba 2015), in which the learning rate was set at 10−4. Overall, the network was trained for 2,000 epochs during approximately 12 h using a Nvidia 1080Ti GPU, and the optimal weights were taken at 1,000 epochs. The auto-decoder network comprises 8 FC layers, each with 512 neurons. In addition, the training involved setting a variance parameter of 0.000001, to control the noise added during training, and a near-surface sampling ratio of 1, to balance the distribution of the 500,000 points. All surfaces from ground truth and inference were displayed on the Desk web interface (Jacinto et al. 2012). The code used for this study is available in the following GitHub repository: https://github.com/valette/DeepSDF.
Results
Training
Among the 345 STL files collected, 285 were included. For both PCA and DeepSDF approaches, the shapes were learned with a Hausdorff distance lower than 0.3 mm. For all metrics, the DeepSDF approach demonstrated significantly lower values for the test dataset (P < 0.05; Table 1; Appendix Fig. 1).
Performances of Principal Component Analysis (PCA) and Deep Signed Function (DeepSDF) for Learning the Anatomy on the Test Dataset.
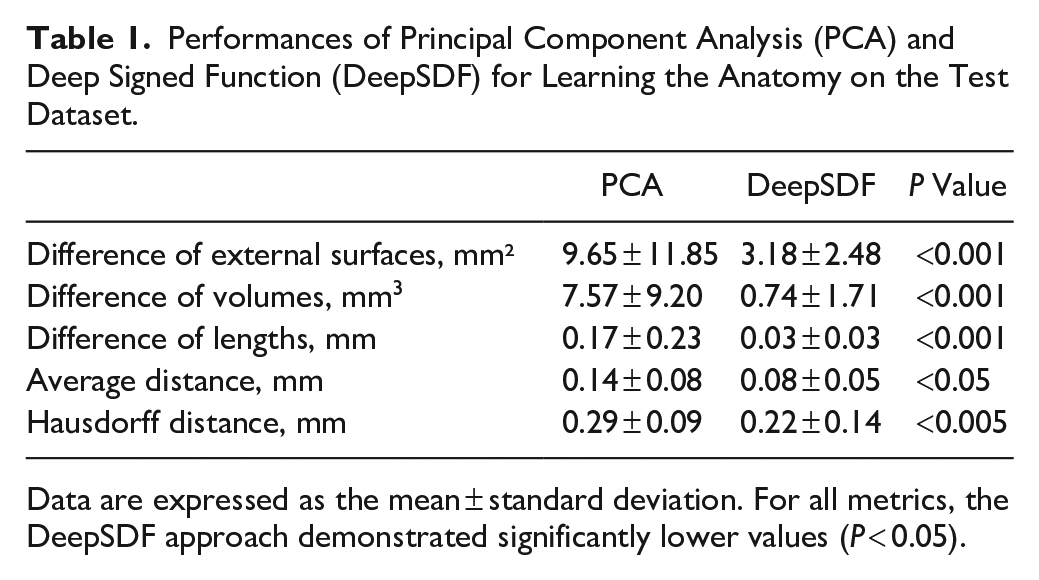
Data are expressed as the mean ± standard deviation. For all metrics, the DeepSDF approach demonstrated significantly lower values (P < 0.05).
Model Analysis and Performances
For the PCA approach, the first 5 modes of shape variations accounted for 70.9% of the overall shape variability in the population; the eigenvalues of the higher modes were negligible compared with the first modes (Appendix Fig. 2). For the DeepSDF approach, convergence was observed after 1,000 epochs (Appendix Fig. 3). The average Euclidian distance between ground truth and inferences was significantly smaller for high sizes of the latent code, compared to the lower sizes (p < 0.01; Appendix Table 3, Fig. 2A). On the dental arches of the clinical dataset 3DTeethSeg’22, the DeepSDF models demonstrated stable reconstruction performances, with significantly lower accuracy observed for models with smaller latent code size (Appendix Table 3, Fig. 2B, C).
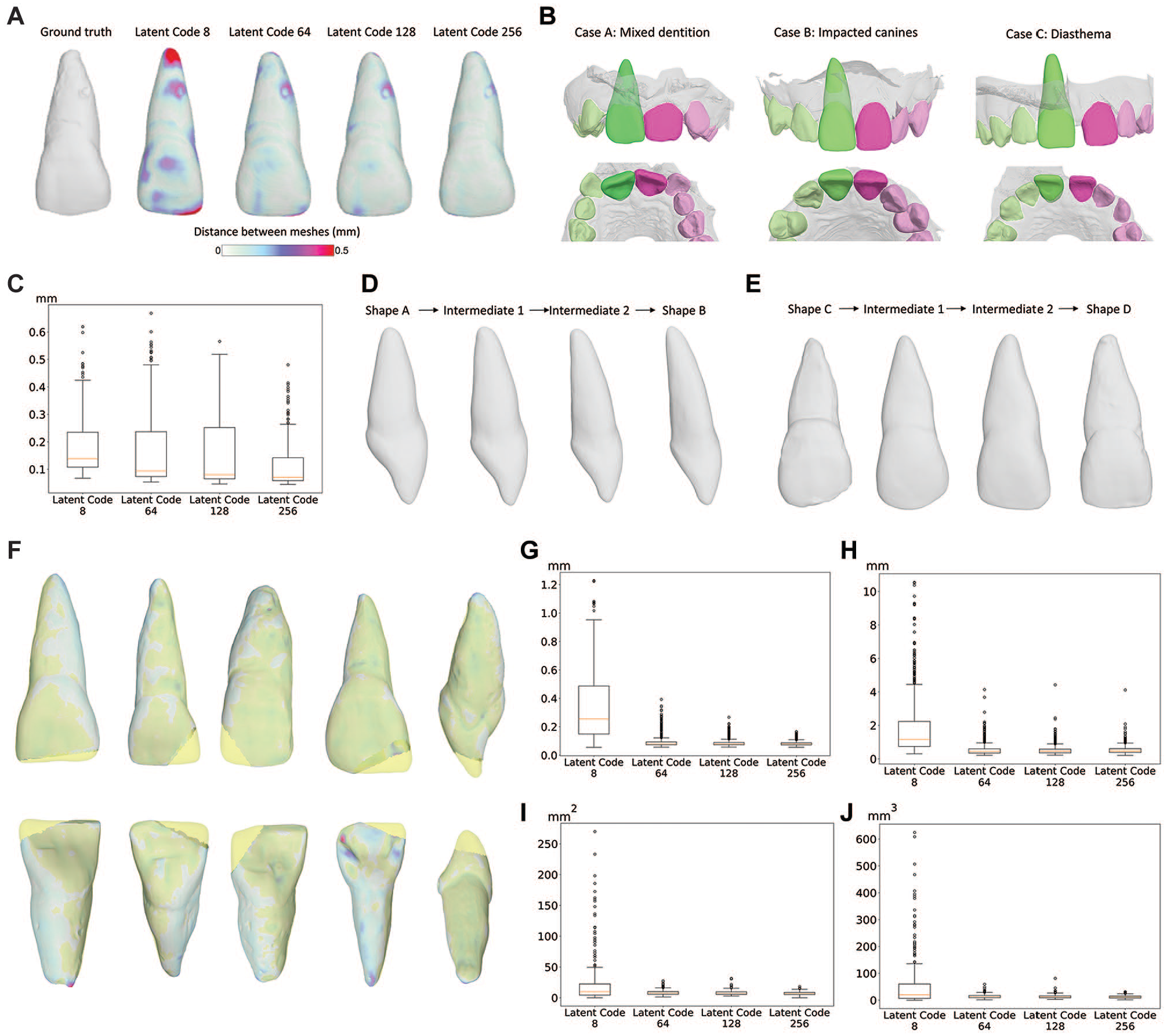
Influence of the size of the latent code on learning and reconstruction of central incisors using DeepSDF models of different sizes with (
Explainability
For the PCA approach, the surface and volume of the tooth could mostly be explained by the variations in the mode 1 (63.2% and 50.1%, respectively, both P < 0.01; Appendix Table 4). For the DeepSDF approach, the average Euclidian distance between ground truth and inferences was significantly smaller for high sizes of the latent code, compared with the lower sizes for the test dataset. The interpolated shapes between 2 latent codes demonstrated smooth transitions from the first to the second form while preserving continuity in the signed distance function (Fig. 2D, E). The t-SNE visualization, combined with an agglomerative clustering approach, identified 3 distinct clusters for latent size ≥64 (Appendix Fig. 4). In addition, the 3 clusters were not related to the tooth shape, and the performances of the random forest to classify the tooth shape, according to the latent code, were low (Appendix Table 5). The Cohen’s kappa unweighted estimate of agreement between both experts was 0.74 (95% confidence interval [CI] [0.65, 0.82]).
Virtual Simulation of Tooth Reconstruction
For all metrics, the DeepSDF model for the latent code of size 8 demonstrated significantly higher values compared with higher sizes of the latent code (P < 0.05). For the Hausdorff distance, there was a trend toward higher values for the latent code of size 64 compared with 128 but without reaching statistical significance (P = 0.113). For the volume distance, there was a trend toward higher values for the latent code of size 64 compared with 256 (0.059; Fig. 2F–J; Table 2).
Performances of Deep Signed Function (DeepSDF) Models of Different Sizes of Latent Code for Reconstructing the Anatomy on the Test Dataset.
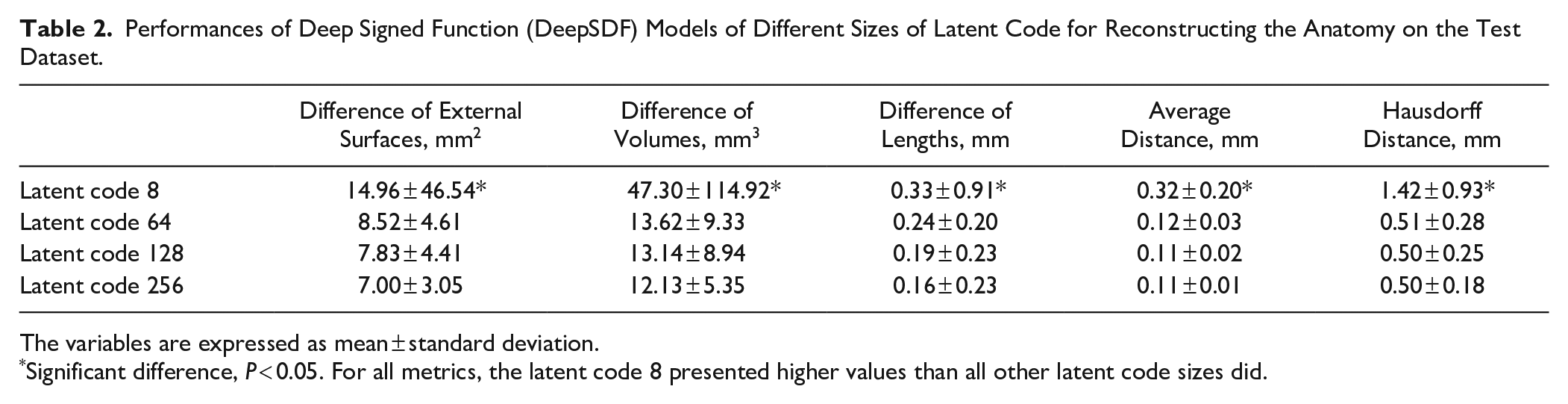
The variables are expressed as mean ± standard deviation.
Significant difference, P < 0.05. For all metrics, the latent code 8 presented higher values than all other latent code sizes did.
Discussion
The present study found that DeepSDF outperformed PCA in learning the shape of central incisors, particularly in capturing fine anatomical details while maintaining a level of explainability through its ability to construct interpolated shapes. More precisely, DeepSDF models with higher latent code sizes demonstrated greater performance in reconstructing the initial shape of the maxillary central incisors compared with smaller latent sizes. However, the deeper understanding of the latent space structure remains limited, highlighting the need for further investigation regarding how shape variations are encoded.
One of the key methodological aspects of the present study was the precise learning of DeepSDF, consistent with previously reported performances on in vitro datasets (Chen et al. 2024). In this regard, it is of particular importance to underscore that PCA does not inherently learn shape structures but rather performs statistical morphing based on predefined meshes, while DeepSDF learns a continuous signed distance function. These differences likely explain the significantly greater ability of DeepSDF to capture fine anatomical details compared with PCA. The ability to capture fine anatomical variations was further facilitated by the high precision of intraoral scan surfaces. In comparison, many training datasets rely on CBCT data, and associated learning approaches, such as CNNs and GANs, are based on fixed-resolution grids (Broll et al. 2024). However, representing anatomical variations on a grid requires a substantial increase in resolution, which significantly raises computational costs. In addition, the nature of voxels introduces a stair-step effect, limiting shape accuracy, whereas microtexture precision is essential for personalized smile design (Machado et al. 2013; Pham and Nguyen 2022).
These high learning abilities also enable accurate shape reconstruction, even in cases of incomplete or partially captured anatomies. While practitioners reconstruct dental shapes by inferring the original form based on prior anatomical knowledge, DeepSDF completes missing structures by leveraging a learned implicit representation of shape variations derived from the training dataset. In the present study, the teeth were reconstructed and presented a difference in average distances of approximately 0.1 mm, which was close to those of incisors’ full crowns obtained using the CEREC software (Wang et al. 2020) or of fractured incisors using DeepSDF (Chen et al. 2024). Recently, GANs also demonstrated high performances to automatically design dental crowns for molars, confirming the potential of AI approaches for large defects (Chau et al. 2024). Clinically, these AI-assisted tools could support practitioners chairside in future smile design workflows, as they have been shown to improve the reproducibility of crown designs and reduce design time by up to 90% compared with traditional wax-up methods (Liu et al. 2024). Following this design phase, 3D printing of the generated model could serve as a mock-up guide for restorations (Gürel et al. 2021). Beyond coronal reconstruction, a key aspect of the present study is that DeepSDF was trained on complete tooth anatomies, encompassing both coronal and root regions. When applied to the 3DTeethSeg’22 dataset, DeepSDF successfully predicted full tooth morphology, including the root structure, highlighting promising clinical applications in prosthodontics and endodontic (Frese et al. 2012; Wu et al. 2016; Choi et al. 2023; Mohammad-Rahimi et al. 2024).
A critical factor before clinical implementation, and essential for maintaining the trust of both practitioners and patients, is the ability to explain model predictions and ensure their generalizability (Chau et al. 2024; Rokhshad et al. 2024). Regarding PCA, explainability is straightforward, as shape variations are explicitly described by eigenvectors; herein, more than 10 modes were required to capture the large-scale shape variations, which is consistent with previous findings related to the occlusal surface of premolars and molars (Mehl et al. 2005; Binvignat et al. 2024). However, in case of missing structures, PCA can only project the remaining shape into its learned space and approximate reconstruction by combining principal modes. Since the generated shape is necessarily constrained by the statistical variations present in the training dataset, it is less adaptable to anatomical cases outside the learned distribution. In comparison, DeepSDF learns a continuous surface representation, making it more flexible to reconstruct missing anatomies. In addition, the fact that DeepSDF maintained high reconstruction accuracy on an independent dataset suggests a strong ability to generalize unseen anatomies. Second, unlike GANs, which implicitly learn data distributions, DeepSDF explicitly models a signed distance function, making it inherently more interpretable. This explicit representation allows interpolation between learned shapes, generating intermediate forms that blend features from different anatomies. Such interpolations could assist practitioners when performing individualized smile designs and addressing complex esthetic considerations. Addressing these complex demands requires a highly detailed learning of fine anatomical structures, which in the present study was achieved only with models using the highest latent code size. Besides, it is interesting to note that clustering analysis revealed distinct patterns between small and large latent codes, suggesting different learning patterns. However, the latent space was not fully optimized for classification tasks, indicating that future research should focus on developing disentangled latent representations to enhance our understanding of the learning process and improve the adaptation of restoration designs to specific anatomical features (Rotem et al. 2024).
Smile design relies on multiple factors, and the current dataset has limitations including the absence of adjacent teeth or lack of facial scan data, despite the reported influence of these factors (Pereira Cenci et al. 2023). Surprisingly, no method that uses the entire dentition for the reconstruction process of a single tooth has been reported (Broll et al. 2024). However, a promising approach could involve training multiple DeepSDF models to leverage the full contextual information of the data (Yang et al. 2022). Furthermore, the learning of the shapes mostly involved usual and intact shapes from 3 different populations, and therefore, very rare anatomies such as peg-shaped incisors might be wrongly reconstructed, due to their absence in the training dataset. In future studies, it might be worth evaluating whether increasing the number of these rare anatomies in the training set proves to be more effective. Moreover, the models were primarily tested on simple fracture lines; further investigations are therefore needed to evaluate their applicability for complex fractures involving multiple lines as well as lower-quality STL files acquired under clinical conditions. All of these observations highlight that both practitioners and patients should rely on the predicted shapes with caution. Despite the high performance of these AI-based reconstruction models, the outcomes represent only statistically probabilistic shapes and may not always accurately reflect future occlusal alterations.
This study demonstrates the potential of AI approaches, particularly DeepSDF, for the learning and reconstruction of the anatomy of upper central incisors. To foster trust and acceptance, future research should, however, focus on improving the explainability of DL models and considering a broader range of factors that influence smile design.
Author Contributions
P. Binvignat, contributed to conception, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; S. Valette, contributed to conception, design, data analysis and interpretation, critically revised the manuscript; A. Hara, P. Lahoud, contributed to conception, data acquisition, drafted and critically revised the manuscript; R. Jacobs, contributed to conception, data interpretation, drafted and critically revised the manuscript; A. Chaurasia, contributed to conception, data analysis and interpretation, critically revised the manuscript; M. Ducret, contributed to conception, drafted and critically revised the manuscript; R. Richert, contributed to conception, design, data analysis and interpretation, drafted and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345251344548 – Supplemental material for AI in Learning Anatomy and Restoring Central Incisors: A Comparative Study
Supplemental material, sj-docx-1-jdr-10.1177_00220345251344548 for AI in Learning Anatomy and Restoring Central Incisors: A Comparative Study by P. Binvignat, S. Valette, A. T. Hara, P. Lahoud, R. Jacobs, A. Chaurasia, M. Ducret and R. Richert in Journal of Dental Research
Footnotes
Acknowledgements
The authors would like to express their gratitude to Tristan Macquart (PhD; LaMCoS) for his contribution in explaining, developing, and presenting the MEG-IsoQuad method on Rhino 7 for dental applications. The authors would like to thank Shanez Haouari (PhD; Hospices Civils de Lyon, France) for help in manuscript preparation.
A supplemental appendix to this article is available online.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
