Abstract
Background
Thiopurine S-methyltransferase (TPMT) is crucial for metabolizing thiopurine drugs. This study aimed to establish the cutoff values for TPMT activity in a cohort of healthy individuals. We defined normal TPMT activity ranges and identified clinically applicable thresholds to distinguish individuals with normal TPMT function from those with reduced or deficient activity.
Methods
A total of 457 participants, including 207 children and 250 healthy adults without prior thiopurine drug exposure, were enrolled. TPMT activity was measured and common defective genetic variants (TPMT*3A, TPMT*3B, and TPMT*3C) were detected. To determine TPMT activity cutoff values and maximize sensitivity and specificity, receiver operating characteristic curve analysis was employed.
Results
The cutoff values for TPMT activity in children were ≥52.9 nmol 6-MMP/g Hb/h for persons of the wild type and <52.9 nmol 6-MMP/g Hb/h for individuals who were heterozygous. In adults, the cutoff values were ≥44.6, 31.58–44.5, and <31.58 nmol 6-MMP/g Hb/h for wild-type, heterozygous, and compound heterozygous individuals, respectively. The sensitivity and specificity were 79.29% and 100% in children, whereas, in adults, they were 61.86% and 78.57%, 38.46% and 64.73%, and 100% and 95.98% in the wild-type, heterozygous, and compound heterozygous, respectively.
Conclusions
Identifying TPMT activity cutoff values is crucial for managing patients receiving thiopurine therapy, especially in Thailand. This approach allows for personalized treatment plans and minimizes the risk of adverse drug reactions. Since TPMT activity cutoff values can differ by population and testing methods, it is important to establish specific cutoff values locally.
Keywords
Introduction
Thiopurine S-methyltransferase (TPMT) catalyzes the methylation of several purine analogues, including thiopurine medications like azathioprine (AZA), 6-mercaptopurine (6-MP), and 6-thioguanine (6-TG). In addition to treating rheumatic disorders, inflammatory bowel disease, and children’s acute lymphoblastic leukaemia (ALL), these medications also act as immunosuppressants to stop allograft rejection.1,2 The active metabolite 6-thioguanine nucleotide (6-TGN), which is closely linked to mercaptopurine toxicity, can be produced from 6-MP, which is quickly produced once thiopurine medications are consumed. 3 6-MP is changed into its inactive metabolite, 6-methylmercaptopurine (6-MMP), by the enzyme thiopurine S-methyltransferase (TPMT). Variants in the TPMT gene can reduce enzyme activity, leading to elevated levels of 6-TGN. The accumulation of this active metabolite, 6-TGN, results in myelosuppression.4,5 Additionally, patients with extremely high TPMT activity are at risk of hepatotoxicity.6,7.
The US Food and Drug Administration states that before receiving 6-MP treatments, people should have their TPMT genotype and phenotype (activity) evaluated in accordance with the Clinical Pharmacogenetics Implementation Consortium Guidelines. TPMT phenotype and genotype screening are used to find patients who are TPMT-deficient and at risk of experiencing severe myelosuppression. 8 There are currently 46 allelic variations of the TPMT gene known to exist, according to the *allele nomenclature (https://liu.se/en/research/tpmt-nomenclature-committee). Reduced TPMT activity and alterations in amino acids are the outcomes of several of these. To find the most prevalent TPMT alleles, such as the *2, *3A, *3B, and *3C alleles, specific genetic variants are found. This allows for accurate risk classification. While 9% of Thai people have heterozygous variants, 90% of them have wild-type TPMT. 5 Common variant alleles in the Thai population include the *2, *3A, *3B, and *3C alleles; however, this approach may not be suitable to detect rare or novel variants.
TPMT enzyme activity testing aids in directly quantifying enzymatic activity and identifying the functional phenotype. It accounts for the variability in TPMT enzyme activity resulting from non-genetic factors, such as liver disease and drug interactions. TPMT enzyme activity testing can overcome the rare variants in TPMT genotyping. Lennard (2014) noted that identification of TPMT-deficient individuals cannot be 100% accurate based on either the TPMT genotype or phenotype alone. 9 Since TPMT variations are uncommon in the Thai population, not all people with rare variants are included in TPMT genotyping. Although TPMT enzyme activity testing is advised, there is currently no recognized reference interval for TPMT activity. This study evaluated TPMT activity cutoff levels to determine who had physiological, moderate, and low activity. The goal was to offer clinically relevant reference activity values for the regular evaluation of thiopurine toxicity susceptibility. We gathered discordant samples between TPMT genotype and TPMT activity for sequencing to find novel SNPs, and we employed multiple linear regression to determine the association between TPMT activity and demographic parameters (age and gender).
Subjects and methods
Study protocol and participants
Four hundred fifty-seven participants had their blood drawn at the transfusion medicine department of Ramathibodi Hospital, Mahidol University, and Chulalongkorn University’s Faculty of Medicine, both in Bangkok, Thailand. The sample included 207 children (≤18 years old) and 250 adults (>18 years old). All samples were collected from fasting subjects who were not consuming prescribed drugs, without apparent disease, not receiving thiopurine therapy, and without blood transfusion within 3 months of the study. The Institutional Review Board, Faculty of Medicine, Chulalongkorn University (COA No 1030/2016, IRB No 512/59), and the Ethics Committee of the Faculty of Medicine, Ramathibodi Hospital, Mahidol University (protocol code COA.MURA.2018/241), approved this study.
TPMT enzyme activity
High-performance liquid chromatography-tandem mass spectrometry (LC-MS/MS) (Agilent Technologies Foster City, CA, USA) was employed to assess the TPMT enzyme activity in whole blood using a previously reported methodology. 10 In summary, the amount of 6-methylmercaptopurine (6-MMP) that was produced following 1 hour of incubation at 37°C was used to measure TPMT activity. This was done by converting 6-MP (Sigma-Aldrich Co. LLC, Singapore) with S(5ʹ-denosyl)-L-methionine iodide (SAM) (Sigma-Aldrich Co. LLC, Singapore) as the methyl donor. The amounts of 6-MMP generated in this assay were found to be directly correlated with the enzyme levels in the samples. Nanomoles of 6-MMP generated per g of haemoglobin per hour (nmol 6-MMP/g Hb/h) for one unit of enzyme is the unit used to express TPMT activity. The percentage coefficients of variation (%CV) for the intra- and inter-assay were 3.7% and 2%, respectively.
TPMT genotyping
The most common defective genetic variants, TPMT*3A (rs1800460, c.460G>A and rs1142345, c.719 A>G), TPMT*3B (rs1800460, c.460G>A), and TPMT*3C (rs1142345, c.719 A>G), were detected using the TaqMan method (Applied Biosystems® ViiATM7 Real-time PCR system). One millilitre of whole blood was treated with the MagNa Pure automated extraction system (Roche Applied Science, Penzberg, Germany) to extract DNA. The NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) was used to evaluate the purity of DNA. It can detect spectra in a dynamic range of roughly 220–750 nm. 5 ng/mL of genomic DNA, 12.5 µL of TaqMan™ Genotyping Master Mix (Thermo Fisher Scientific Inc., Washington, USA), and 1.25 µL of TaqMan™ Drug Metabolism Genotyping Assay (Thermo Fisher Scientific Inc., Washington, USA) were used in the 11.25 µL reaction volume used for genotyping. The PCR conditions were as follows: 50 cycles of 15 s at 92°C and 90 s at 60°C were performed after a 10-min initial denaturation at 95°C. TaqMan Genotyper Software was used to interpret the genotypes following the manufacturer’s instructions.
The linear regression model
Putative determinants of TPMT activity were classified into age, gender, and TPMT genotype (wild-type, heterozygous, and compound heterozygous) categories. A univariate linear regression analysis was used to identify the factors correlating with TPMT activity. Factors associated with TPMT activity with a P-value of <0.1 were entered in a forward selection multiple regression model. The significance level was set at a two-sided P-value <.05.
TPMT cDNA sequencing
To identify novel SNPs from the wild-type TPMT genotype and the discordantly low TPMT enzyme activity, 13 DNA samples were sequenced. Following the manufacturer’s instructions, whole blood was drawn into EDTA tubes, and DNA was extracted using the DNA Extraction/Genomic DNA Purification kit (Geneaid, New Taipei City, Taiwan). The isolated DNA was kept at −20°C until analysis, and the concentration of genomic DNA was measured using the NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). To prevent amplification of the TPMT pseudogene on chromosome 18, the TPMT cDNA sequence was amplified using 3 µL of DNA in a 50 µL reaction volume that contained HotStarTaq PCR reagents (Qiagen) and primers for cDNA TPMT exons 2–9 or 5–10. A 40 µL reaction mixture including 50–100 ng of DNA, 0.25 µM primers, 500 µM dNTPs, 2.5 U Taq DNA polymerase (Invitrogen, USA), and 2.75 mM MgCl2 was used to conduct PCR. A T100 Thermal Cycler (Bio-Rad, USA) was used for amplification with a program consisting of an initial denaturation at 95°C for 3 min, 35 cycles with the following conditions: 95°C for 30 s, 59°C for 40 s (64°C for exon 2 primers), 72°C for 1 min, and a final extension at 72°C for 10 min. Primers were designed using Primer 3 Plus (https://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi) based on the NCBI RefSeqGene for Homo sapiens thiopurine S-methyltransferase (TPMT) on chromosome 6 using the NCBI Reference Sequence NG_012137.3.
The resulting PCR products, a positive control (TPMT*1/*1), and a 100-bp DNA Ladder (New England BioLabs, USA) were separated using agarose gel electrophoresis using 2% agarose gels. PCR products of appropriate length were purified using the Gel/PCR DNA fragment kit (Geneaid, Taiwan) and sequenced using the ABI 3730XL DNA Analyzer.
Statistics
STATA 10 software (Stata Corporation, College Station, Texas) was used to conduct statistical analyses. The participants’ clinical features were described using descriptive statistics, which were reported as the mean ± SD. A student t test was employed for statistical comparisons. Receiver operating curve (ROC) analysis was used to assess cutoff values. An ideal cutoff value can be found by locating the point on the ROC curve that maximizes Youden’s index. 11 Graphs were generated using GraphPad Prism 8 (GraphPad Software, San Diego, US). Simple and multiple linear regressions were used to determine the relationship between two or more continuous variables to predict TPMT enzyme activity. The regression model included age, gender, and TPMT genotype. Statistical significance was defined as a P-value of less than 0.05.
Results
Demographic data of the participants
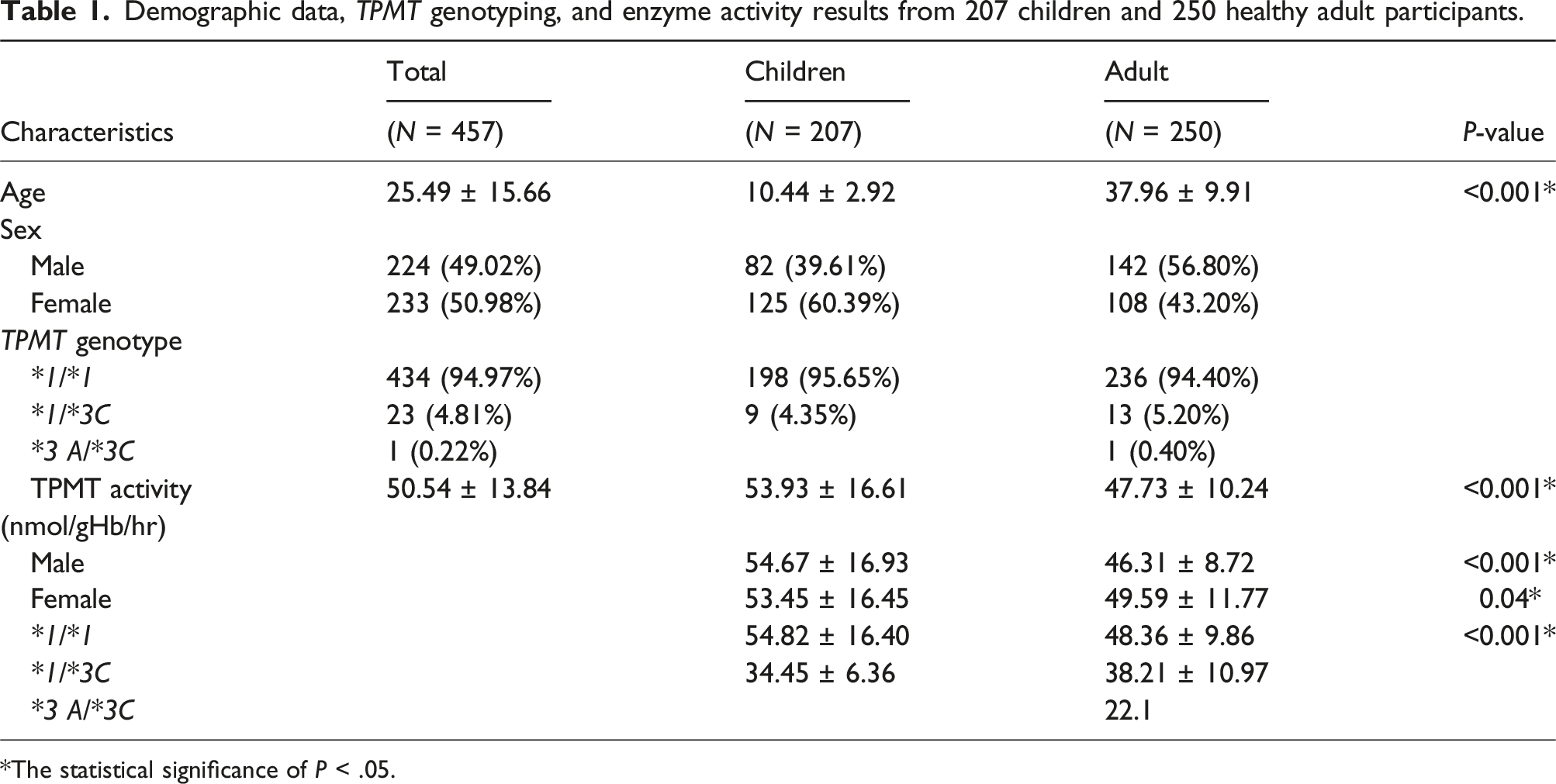
In this study, 457 healthy participants were recruited, consisting of 207 children and 250 adult participants. Of these, 39.61% (82 of 207) were male children and 60.39% (125 of 207) were female children. In the adult group, 56.80% (142 of 250) were males, whereas 43.20% (108 of 250) were females. The average age was 10.44 ± 2.92 years among children and 37.96 ± 9.91 years among adults. TPMT genotyping results indicated that 94.97% (434 of 457) of the participants had the wild-type genotype, 4.81% (23 of 457) were heterozygous, and 0.22% (1 of 457) were compound heterozygous. A TPMT activity distribution graph for the entire study group is illustrated in Figure 1. The values for the TPMT enzyme activity assays were 53.93 ± 16.61 nmol 6-MMP/g Hb/h for children and 47.73 ± 10.26 nmol 6-MMP/g Hb/h for adults, indicating a significant difference between the two age groups. The male and female children exhibited significantly higher enzyme activity compared with that observed in the adults, with median values of 54.67 ± 1.87 nmol 6-MMP/g Hb/h (P < .001) and 53.45 ± 1.47 nmol 6-MMP/g Hb/h (P = .04), respectively. The demographic data are summarized in Table 1, and the observed genotype frequencies are consistent with Hardy–Weinberg equilibrium. A graph displaying the distribution of TPMT activity among 207 children and 250 adult participants. Demographic data, TPMT genotyping, and enzyme activity results from 207 children and 250 healthy adult participants. *The statistical significance of P < .05.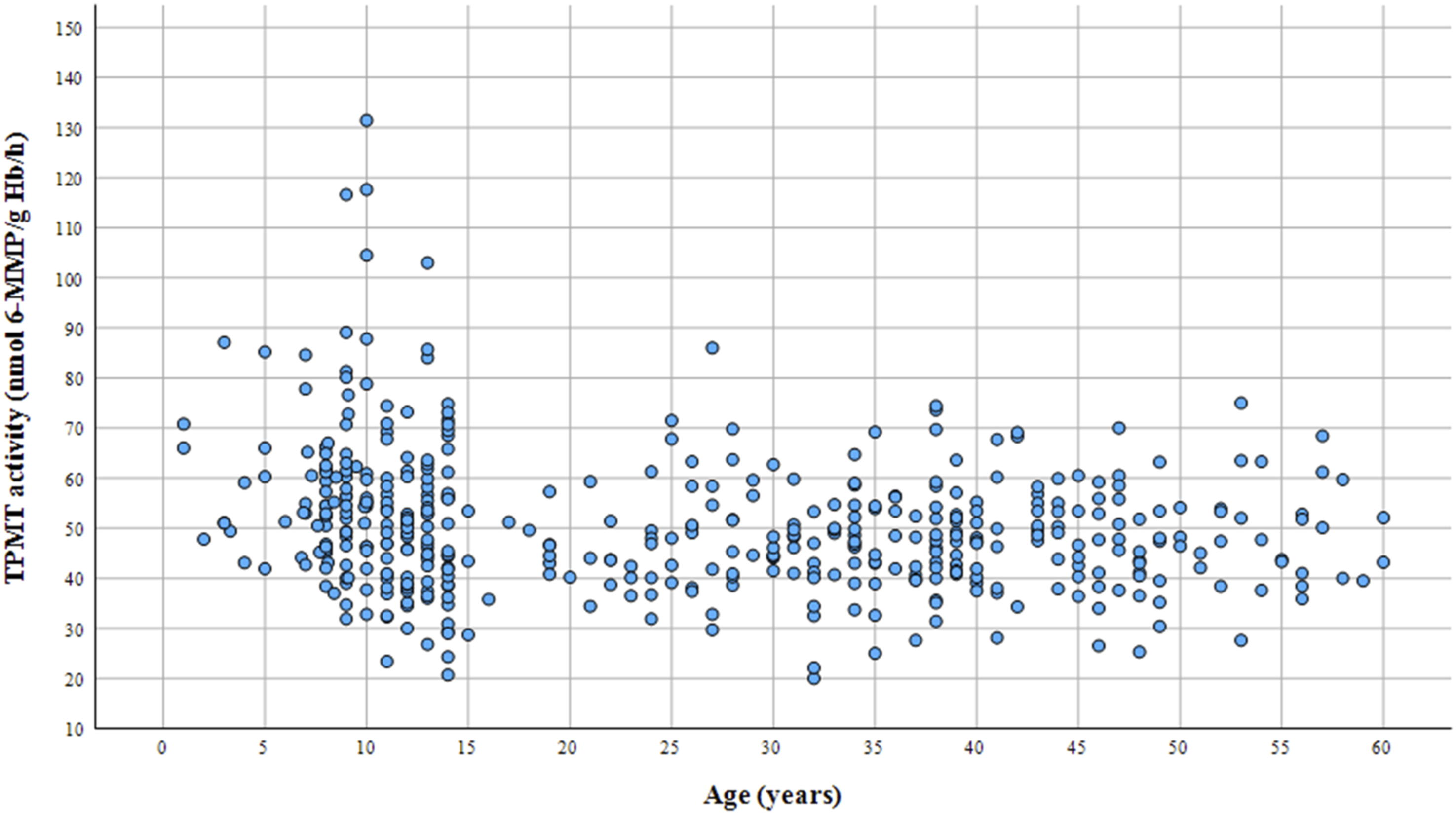
TPMT enzyme activity cutoff values
The cutoff for distinguishing thiopurine S-methyltransferase (TPMT) activity was evaluated using ROC curve analysis based on the phenotype–genotype concordance (Figure 2). Children with normal enzyme activity had TPMT activity cutoff values of ≥52.9 nmol 6-MMP/g Hb/h, whereas those with low enzyme activity had cutoff values of <52.9 nmol 6-MMP/g Hb/h. Evaluation of the cutoff distinguishing thiopurine S-methyltransferase (TPMT) activity using ROC curve analysis based on phenotype–genotype concordance in children (A) and adults (B).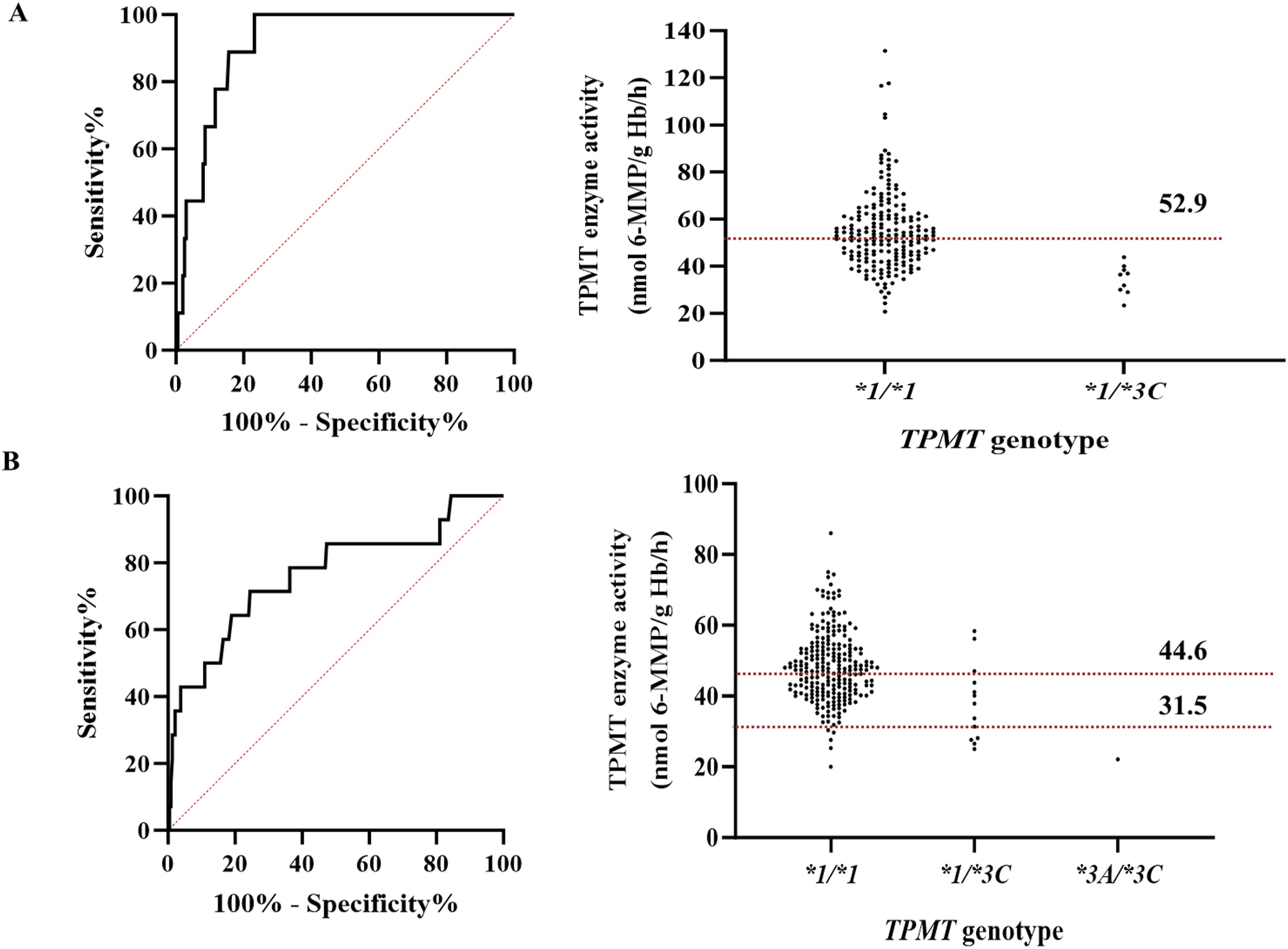
The sensitivity and specificity were 79.29% and 100%, respectively (Figure 2(a)). In adults, the values were ≥44.6 nmol 6-MMP/g Hb/h for individuals with physiological enzyme activity, 31.58–44.5 nmol 6-MMP/g Hb/h for those with intermediate enzyme activity, and <31.58 nmol 6-MMP/g Hb/h for individuals with low enzyme activity (Figure 2(b)). The sensitivity and specificity were 61.86% and 78.57%, 38.46% and 64.73%, and 100% and 95.98% for the wild-type, heterozygous, and compound heterozygous variants, respectively.
Linear regression model
Simple and multiple linear regression analyses of the TPMT enzyme activity in adults and children.
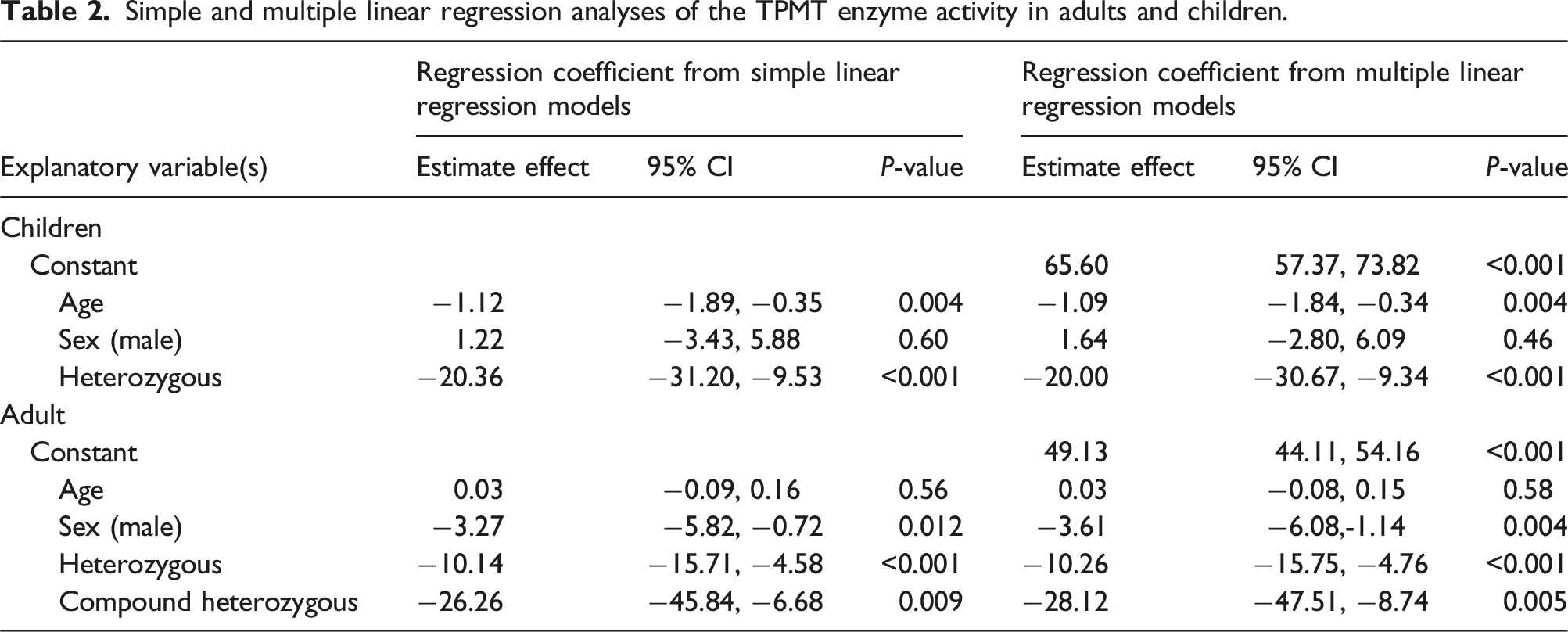
The multiple linear regression model for TPMT activity in children is as follows:
TPMT activity = 65.60 - (1.09 × Age) - (20 × TPMT Heterozygous Genotype).
In adults, gender and TPMT genotype affected TPMT activity. The estimated effect of the TPMT heterozygous genotype based on multiple linear regression analysis revealed a decrease in TPMT activity by 10.26%. In contrast, the estimated effect of the TPMT compound heterozygous genotype resulted in a decrease in TPMT activity by 28.12%, which indicates that the TPMT compound heterozygous genotype has a significantly negative influence on TPMT activity.
The equations for determining TPMT activity in adults are as follows:
TPMT activityHET = 49.13 - (3.61 × Male) - (10.26 × TPMT Heterozygous Genotype).
TPMT activityComHET = 49.13 - (3.61 × Male) - (28.12 × TPMT Compound Heterozygous).
Discrepancies in TPMT genotype and TPMT activity
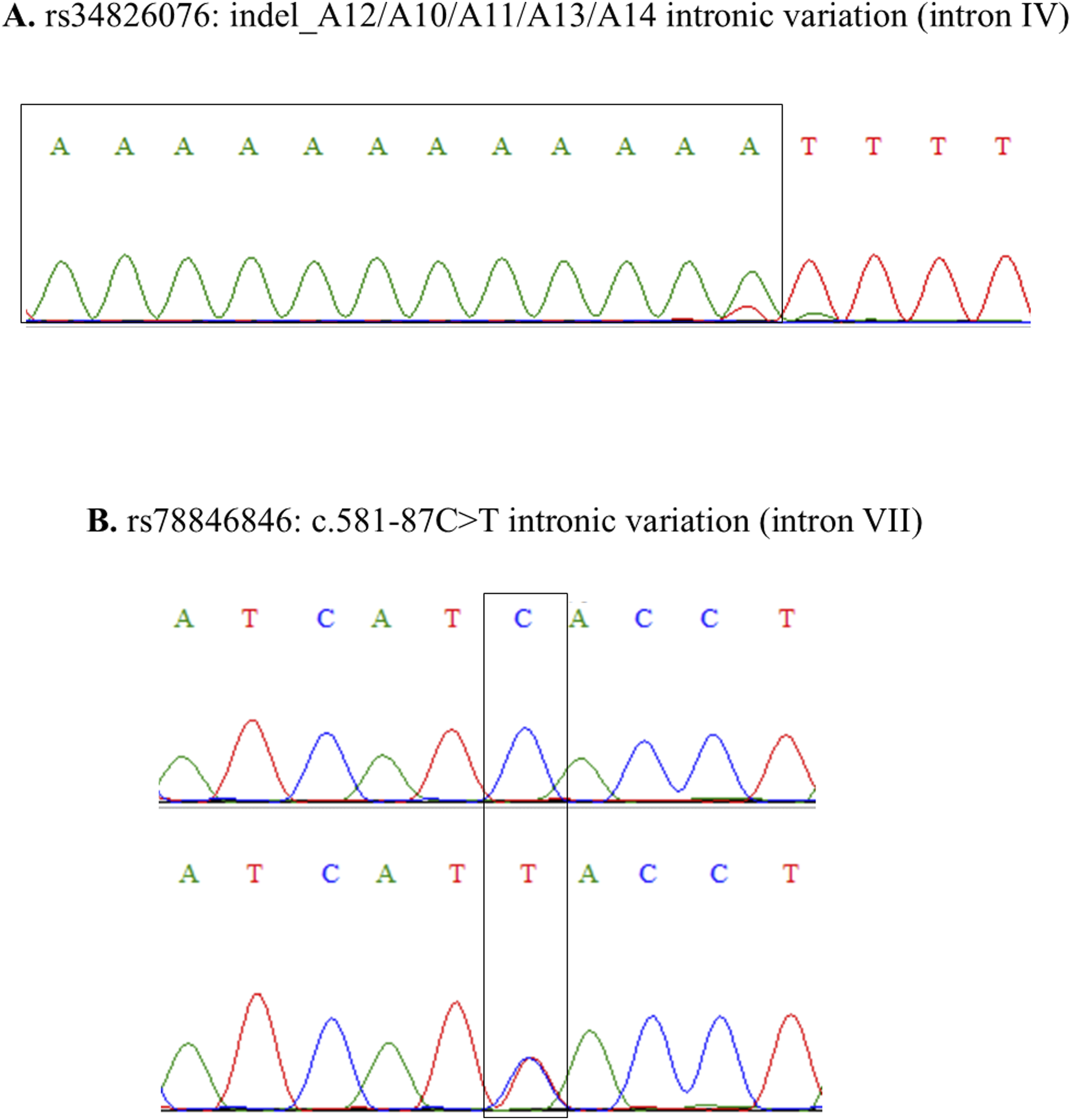
We identified 13 subjects exhibiting discordance between the wild-type TPMT genotype and low TPMT activity, and their DNA was sequenced to identify TPMT polymorphisms (Figure 3). All the variants detected were located in the introns. One sample carried the heterozygous genotype for rs3931660 (c.140+114T>A) (Figure 3(a)), one carried the homozygous genotype for rs12529220 (c.141-101A>T) (Figure 3(b)), one carried the homozygous genotype for rs2518463 (c.366+58T>C), and one carried the homozygous genotype for rs747392577 (ins_T5/T6) (Figure 3(c)). In addition, three samples carrying the homozygous genotype for rs12529220 and the heterozygous genotype for rs2518463 were identified (Figure 3(d)). The remaining samples did not show any distinct pattern. We sequenced three samples with heterogeneous and exhibited higher TPMT activity. The results indicated that all three samples carried the variant rs34826076 (A12/A10/A11/A13/A14) (Figure 4(a)), and two of them also carried rs78846846 (c.581-81C>T) (Figure 4(b)). TPMT sequencing variation with discordant wildtype TPMT genotype and low TPMT enzyme activity: (A) rs3931660:c.140+114T>A, (B) rs12529220:c.141-101A>T, (C) rs747392577:ins_T5/T6, and (D) rs2518463:c.366+58TC. TPMT sequencing variation with discordant heterozygous TPMT genotype and high TPMT enzyme activity: (A) rs34826076: indel_A12/A10/A11/A13/A14 and (B) rs78846846:c.581-87C>T.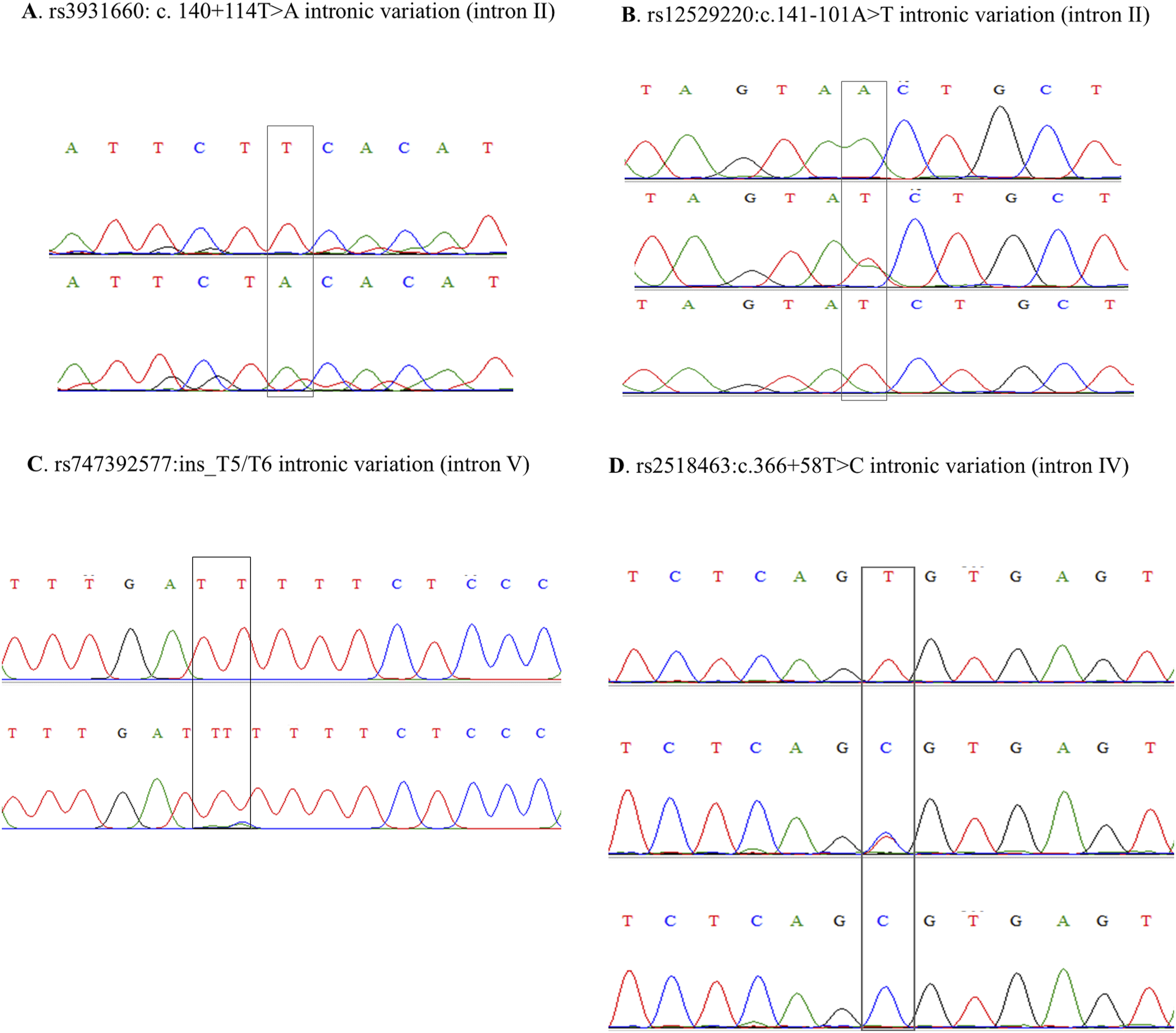
Discussion
In this study, a cohort of healthy Thai individuals was examined with respect to TPMT genotype and TPMT activity. The objective of TPMT screening is to detect TPMT-deficient patients who are at risk of developing severe bone marrow suppression. 8 The results indicated that children have increased TPMT activity compared to adults. This is consistent with recent observations showing that neonates had greater TPMT activity than adults.12,13
The cutoff TPMT activity values used to distinguish physiological and intermediate TPMT activity in healthy children were ≥52.9 nmol 6-MMP/g Hb/h for normal individuals and <52.9 nmol 6-MMP/g Hb/h for individuals with low TPMT activity. In adults, the cutoff values were defined as ≥44.6 nmol 6-MMP/g Hb/h (physiological), 31.5–44.5 nmol 6-MMP/g Hb/h (intermediate), and <31.5 nmol 6-MMP/g Hb/h (low) activities. TPMT activity levels in whole blood were higher than in packed erythrocytes, with threshold values of ≥9 U/ml pRBC (physiological), 2.5–8.9 U/ml pRBC (intermediate), and <2.5 U/ml pRBC (low). 13 The erythrocyte isolation process and the impact of various sample matrices on analytical performance are the causes of the variations in cutoff values between isolated erythrocytes and whole blood. Because it is simpler, quicker, and less prone to error, TPMT activity was assessed directly in whole blood lysates in this investigation.
Regression analysis was conducted to predict TPMT activity based on TPMT genotype and age for children as well as TPMT genotype and gender (male) for adults. The effect of gender on enzyme activity has also been studied previously14–16; however, the differences in TPMT activity between genders may not be consistently observed across all studies or populations.14,15 Factors such as study design, sample size, and the genetic diversity of the population can all influence the results. It would be cautious about using these equations in clinical practice as a substitute for direct TPMT activity measurement. While the equations may provide an estimate based on genotype, age, and gender, TPMT activity can be influenced by several non-genetic factors, including red blood cell turnover, underlying disease states, and potentially unidentified genetic variants. Furthermore, the relatively small sample sizes in the variant genotype cohorts limit the accuracy and generalizability of these equations. Until validated in larger, independent cohorts, and across diverse clinical settings, these models should be used with caution and not replace direct enzymatic testing where available.
The present study had a concordance rate of 80.19% (95% CI: 74.10–85.39) in children and 60% (95% CI: 53.63–66.12) in adults between the TPMT genotypes and phenotypes. We found that the TPMT wild-type alleles in 20.70% (41/198) of children and 38.13% (90/236) of adults would have been misclassified based on genotype, as they empirically exhibited intermediate or low TPMT activity. Because patients with wild-type TPMT are less likely to experience bone marrow toxicity 17 and therapy termination, it is intriguing from a clinical standpoint that overall, 28.66% (131/457) of the wild-type people exhibited lower enzyme activity than would be predicted from genotype alone18,19 if administered a normal dose (2–3 mg/kg/day) of thiopurine. Moreover, 3/13 (23.07%) of the TPMT heterozygote adults in this cohort would have been misclassified by genotyping, as their enzyme activity assays yielded values above the cutoff for intermediate TPMT activity. If TPMT function was only evaluated by genotyping, people who had higher TPMT activity than anticipated concerning the genotype results could be in danger of obtaining a subtherapeutic thiopurine medication.
Such genotype–phenotype discordance has been observed in other studies. The percentage of wild-type individuals with low activity, for instance, ranged from 0 to 23%, according to a meta-analysis by Booth et al. (2011). This was lower than our findings (131/434, 30.18%), while the percentage of heterozygotes with physiological activity ranged from 0 to 8%, which was lower than what we found in our study (3/22, 13.63%). 20 The discrepancy in our results may be attributed to the TPMT activity measurements being influenced by physiological or environmental factors. We assayed TPMT activity in whole blood, which represents the metabolic processes throughout the body. We observed a high potential for a discordant TPMT genotype–phenotype, which may occur from the lower TPMT variant allele frequency in the Asian population.
In discordant samples, we observed that wild-type participants had low TPMT activity who harbour an intronic polymorphism (rs3931660, 140+114 T>A), (rs12529220, 141-101 A>T), (rs2518463, 366+58 T>C), which is part of the TPMT*3E allele. In a previous study, logistic regression analysis in patients with myasthenia gravis revealed mutations in the intronic T140+114A (rs3931660) variant to be associated with azathioprine intolerance (P < .01) after Bonferroni’s correction for multiple testing. 21 Moreover, previous reports of rs2518463 indicated no support for an adverse association with thiopurine. First, a study of inflammatory bowel disease patients receiving combined AZA and 5-aminosalicylic acid (5-ASA) therapy indicated that rs2518463 carrier status was not associated with increased 6-thioguanine nucleotide (6-TGN). 22 Second, rs2518463 did not influence either the risk of childhood ALL or the survival rate of patients. 23 Studies in larger cohorts are necessary to determine whether other SNPs associated with discordance between the TPMT genotype and TPMT activity are associated with response to thiopurine treatment. Moreover, we sequenced three samples with heterogeneous characteristics that exhibited higher TPMT activity. The results indicated that all three samples carried the variant rs34826076, and two of them also carried rs78846846. The variant rs34826076 is an indel (A12/A10/A11/A13/A14) located in intron IV, while rs78846846 (c.581-81C>T) is found in intron VII. However, there is no evidence to support that these SNPs are associated with elevated TPMT activity.
Several factors explain the typical concordance of the TPMT genotype and TPMT activity in a healthy cohort. First, uncommon genetic variations found in promoters, non-coding regions, or epigenetic changes may cause decreased TPMT activity. The genotype of the highly polymorphic TPMT gene promoter is substantially correlated with TPMT activity. It has a variable number of tandem repeats (VNTR), 24 ranging from 3 to 9 (VNTR*3 to VNTR*9).25,26 The highest level of TPMT activity was specifically linked to genotypes where at least one allele had more than five repetitions. Zukiv (2010) added that the upstream regulatory region of the TPMT promoter and the quantity and kind of VNTRs influenced the total amounts of TPMT gene transcription. 27 In the present study, we did not examine the effect of promoter characteristics on TPMT gene expression and hence TPMT activity. Second, in this study, we focus on individuals visiting the Transfusion Medicine department who have been screened for Hepatitis B and C, HIV, and syphilis. However, it’s important to consider that other diseases or treatments may affect thiopurine methyltransferase (TPMT) activity. For example, individuals with impaired renal function or those who have undergone specific treatments may experience a decrease in TPMT activity. 28 Third SAM level influences TPMT activity, particularly in TPMT heterozygous individuals. 29 According to Tai et al. (1997), SAM makes the TPMT protein more stable, and its impact is more noticeable for the mutant protein than for the wild-type protein. 30 This may explain why some individuals with a heterozygous TPMT genotype exhibit physiological TPMT activities. Accordingly, determination of the SAM levels may be important as an additional biomarker for selecting thiopurine treatment. More specifically, assessing SAM levels in TPMT-mutant patients can improve treatment from both diagnostic and economic perspectives. Standard doses of 6-MP can be given to patients with high SAM levels to increase the effectiveness of treatment, whereas SAM supplementation during 6-MP therapy can help lower the risk of adverse effects 29 for patients with low SAM levels; however, further research is needed in this area.
In summary, we established cutoff values for TPMT activity in healthy Thai individuals and identified SNPs in discordant subjects through extended sequencing. Further assessment of these SNPs will be necessary in future studies. It’s important to note that TPMT activity cutoffs can vary based on the population and the methods used. This study focuses specifically on a Thai population, so the cutoff values should be determined locally.
Moreover, the frequency of variant alleles in the Asian population is lower than in the Caucasian population, which suggests that measuring TPMT activity could be beneficial for identifying deficiencies. However, this study has a limited sample size, particularly within the compound heterozygote group. As such, any proposed cutoff should be interpreted with caution. Larger studies are required to accurately define a cutoff between heterozygous and compound heterozygous samples. Therefore, the suggested cutoff is based on our current data and serves as a preliminary reference rather than a definitive threshold.
Footnotes
Acknowledgements
The authors would like to thank and acknowledge Assoc. Prof. Fabienne Thomas for valuable assistance with logistic regression analysis. We also acknowledge the Pharmacogenomics Laboratory, Ramathibodi Hospital, for providing the necessary facilities and resources to support this study.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Materials and services were supplied by the Thailand Center of Excellence for Life Sciences (Public Organization), Thailand, and the Ratchadapiseksompotch Fund, Faculty of Medicine, Chulalongkorn University, grant number RA60/48.
Ethical approval
The study was approved by Mahidol University Institutional Review Board (approval no. COA.MURA.2018/241) on June 01, 2018 and Institutional Review Board, Faculty of Medicine, Chulalongkorn University (approval no. COA No 1030/2016, IRB No 512/59) on December 22, 2016.
Consent to participate
Before participating in this study, each subject gave written informed consent.
Guarantor
Assoc.Prof. Dr. Apichaya Puangpetch, Ph.D.
Contributorship
Pimonpan Jinda managed activity to annotate, maintained research data, and wrote the initial draft. Pimonpan Jinda, Santirhat Prommas, Rattanaporn Sukprasong, and Jiratha Rachanakul analyzed the TPMT genotype and TPMT activity. Pimpun Kitpoka, Wimol Thienphopirak, Suwat Chiawchan, Orapa Suteerojntrakool, Chansuda Bongsebandhu-phubhakdi, and Therdpong Tempark recruited patients, collected samples, and acquired clinical data. Sittiphong Hunthai and Supaporn Wiwattanakul Apichaya Puangpetch conceptualized the research, and conducted research and investigation process. The final version of the text was approved after revisions by all authors.
Generative AI
This content did not use generative AI.
Data Availability Statement
The corresponding author can provide the datasets used in this study upon reasonable request.
