Abstract
Background
Healthcare laboratory systems produce and capture a vast array of information, yet do not always report all of this to the national infrastructure within the United Kingdom. The global COVID-19 pandemic brought about a much greater need for detailed healthcare data, one such instance being laboratory testing data. The reporting of qualitative laboratory test results (e.g. positive, negative or indeterminate) provides a basic understanding of levels of seropositivity. However, to better understand and interpret seropositivity, how it is determined and other factors that affect its calculation (i.e. levels of antibodies), quantitative laboratory test data are needed.
Method
36 data attributes were collected from 3 NHS laboratories and 29 CO-CONNECT project partner organisations. These were assessed against the need for a minimum dataset to determine data attribute importance. An NHS laboratory feasibility study was undertaken to assess the minimum data standard, together with a literature review of national and international data standards and healthcare reports.
Results
A COVID serology minimum data standard (CSMDS) comprising 12 data attributes was created and verified by 3 NHS laboratories to allow national granular reporting of COVID serology results. To support this, a standardised set of vocabulary terms was developed to represent laboratory analyser systems and laboratory information management systems.
Conclusions
This paper puts forward a minimum viable standard for COVID-19 serology data attributes to enhance its granularity and augment the national reporting of COVID-19 serology laboratory results, with implications for future pandemics.
Keywords
Introduction
With the emergence of the coronavirus severe acute respiratory syndrome-coronavirus-2 (SARS-CoV-2) in late 2019 and the subsequent pandemic, the need to understand and study the virus became paramount.1–4 The ability to diagnose and understand a health threat and develop processes, behaviours, vaccines and therapeutics to fight it comes in part from the ability to amass, assess and analyse timely and meaningful data as quickly as possible. 5 At the start of the pandemic, there was no comprehension of whether people who had contracted COVID-19 would be immune to later infection and, if so, how long immunity would last. Understanding serology and seroprevalence was key to determining how effective vaccines can be for both current and future variants.6,7 Vendors were developing new assays but could not calibrate the results as there was insufficient data on how to equate antibody binding units with immunity. Laboratories that were developing new assay techniques were not able to share or compare data between the differing techniques, compounding the problem.8–11
Within England, National Health Service (NHS) laboratories can send data by way of Labgnostic (The National Pathology Exchange, NPEx). The data that is sent varies lab to lab and health board to health board. There is no standard approach, nor is there a core minimum defined set of data attributes that are reported. This is in part due to the multitude of systems that exist for the creation and reporting of the data. There is no onward automated feed from Labgnostic to National Health Service (NHS) Digital (now NHS England), meaning that linkage to other relevant nationally collected health datasets does not take place routinely.
When the pandemic started, SARS-CoV-2 serology data was added to the data feeds sent from Test and Trace laboratories via Labgnostic to NHS Digital. However, data from NHS laboratories were not sent. High-level qualitative results were reported, that is, SARS-CoV-2 test results stating outcomes, those being either positive, negative or indeterminate. Although many laboratories produced and recorded quantitative data and results, very few reported this granular level data to Labgnostic, moreover, to be able to report this would have necessitated the development of new data pipelines to transfer the augmented SARS-CoV-2 test results to organisations such as NHS Digital.
The consistent reporting of qualitative data has proven a useful approach to understanding levels of seropositivity with the population; however, it is binary in nature (positive or negative) and reactive. To gain a greater understanding of the range of antibody levels within a seropositive population, and potentially gauge the consequences with respect to new SARS-CoV-2 variants, there was a need for more granular data to be reported from laboratories nationally.12–14 A good example of the minimum data variables (attributes) was described by the Centre for Controlled Disease, 15 which stated 20 data variables as a core minimum to the recording and reporting of SARS-CoV-2 testing, together with associated guidelines. 16 Additionally, the World Health Organisation (WHO) has put forward an International Standard (IS) for SARS-CoV-2, for which it states seven data variables assessed across a range of different SARS-CoV-2 S protein assays. 17 They identified the limitation of sample sizes and recognised that more data was needed to further develop the standardisation approach and harmonisation efforts. The application of the WHO standard has shown that it promotes the ability to better analyse and compare immunology data across different datasets. 18 However, emphasis was placed upon the need for ‘a standardised quantification of anti-SARS-CoV-2 antibodies’, being of ‘the utmost importance’. 18 A call for action by the scientific community during 2022 called for more immunology data to help facilitate the comparison of quantitative assays results data to help better understand immune responses. 19 Additonally, others also recognised this needed to be done to further develop SARS-CoV-2 data representation and standardisation to enable comparability across different testing methods and for population based studies too.20–24
The precise, unambiguous and standardised representation of data, its quality, subsequent harmonisation and interoperability is important as it constitutes the foundation for the accurate and meaningful analysis of multiple sources of data from different domains, institutions and countries.25,26 However, being able to collate, harmonise and represent data from different types of laboratory systems can be problematic.27–33 There are multiple variations that apply to laboratory testing procedures. Firstly, there are different assays that can be used to test for SARS-CoV-2 which, dependent upon the test being performed, produce different numerical outcomes.3,34 Such results are not easily comparable due to the difference in ranges and values between differing assay types, due to different target antigens and different detection technologies.35–39 Secondly, test kit batch variability within a given test type can introduce variance into the measurement process within laboratories.40–42 Thirdly, laboratory analyser systems themselves are subject to variance, this can in part be due to how machines are setup and calibrated thus producing bias and could include analytical imprecision.43,44 Lastly, there are a variety of different Laboratory Information Management Systems (LIMS), which record and represent data generated from the laboratory analyser systems. Moreover, these can be individually configured for the purpose at hand; thus, variation can exist between different instances of the same LIMS software, that is, laboratory A’s LIMS software configuration could be different from that of laboratory B, even though they both use the same piece of LIMS software.
When studied from a holistic viewpoint, this equates to multiple sources of variance throughout the laboratory testing process.43,45 To be able to fully understand and grasp how such variance exists and occurs throughout the testing process, the data must be accurate in the first place; moreover, the representation of the data itself must be formal and standardised.46–49 It is crucial that the description of each data attribute is unambiguous and explicit, so that it precisely describes what it represents, thereby removing the ability to misinterpret its meaning.50–52 Thus, the use and application of naming and naming conventions are paramount to present a standardised approach and, where possible, foster and enable interoperability.
Many laboratories name data attributes according to their own specifications and needs, thus producing local encoded messages. Whilst this is perfectly fine within the context of a singular or group of laboratories that utilise this code, when trying to share or interpret such encoding outside of that localised context, for example, nationally, it can be incredibly difficult to decipher such local naming conventions. Thus, trying to glean what testing kits and platforms are being used and how the actual results have been arrived at from reported laboratory data presents a serious problem.
Definitions and standards for data attributes and specification for many laboratory assays and test results are available within the United Kingdom, including specifications for interoperability via Labgnostic (e.g. Health Level 7 messaging).53–56 Additionally, there are standardised naming conventions for assays and test kits. However, the thorough and correct application and usage of such standards varies between laboratories.
The authors of the paper were involved in a range of initiatives across the country responding to the pandemic for example the National Core Studies Programme and found that granular COVID-19 data was not captured nor was it available for the Scientific Advisory Group for Emergencies (SAGE) to help answer pandemic questions. 4 Sir Michael Fergusson chair of UK Health Security Agency (UKHSA) Scientific Advisory Group for antibody testing which, in its final lessons-learned recommended that pandemic preparedness should include ‘The ability to safely and quickly link serological/immune surveillance data to clinical and genomic data to answer research questions that can inform public health policy’. 57
The CO-CONNECT project was a 2-year research programme funded by the Medical Research Council and the Department of Health and Social Care to build a federated platform streamlining the process for researchers to find and access COVID-19 related datasets from around the United Kingdom.58,59 Serology data was key to informing the COVID-19 response, with researchers requiring access to granular level serology data linked to relevant longitudinal healthcare records. This paper presents a defined minimum set of standardised data attributes for the reporting of SARS-CoV-2 laboratory results developed by the CO-CONNECT project, together with a set of formalised names for laboratory analyser systems, LIMS software providers and SARS-CoV-2 serology measurements as part of the CO-CONNECT standardised vocabulary.
Method
The method applied for the COVID-19 serology minimum data standard (CSMDS) is illustrated in Figure 1. An inductive mixed methods approach was adopted utilising Delphi and teach-back qualitative methods.60–64 Co-Connect COVID-19 serology data standard development methodology.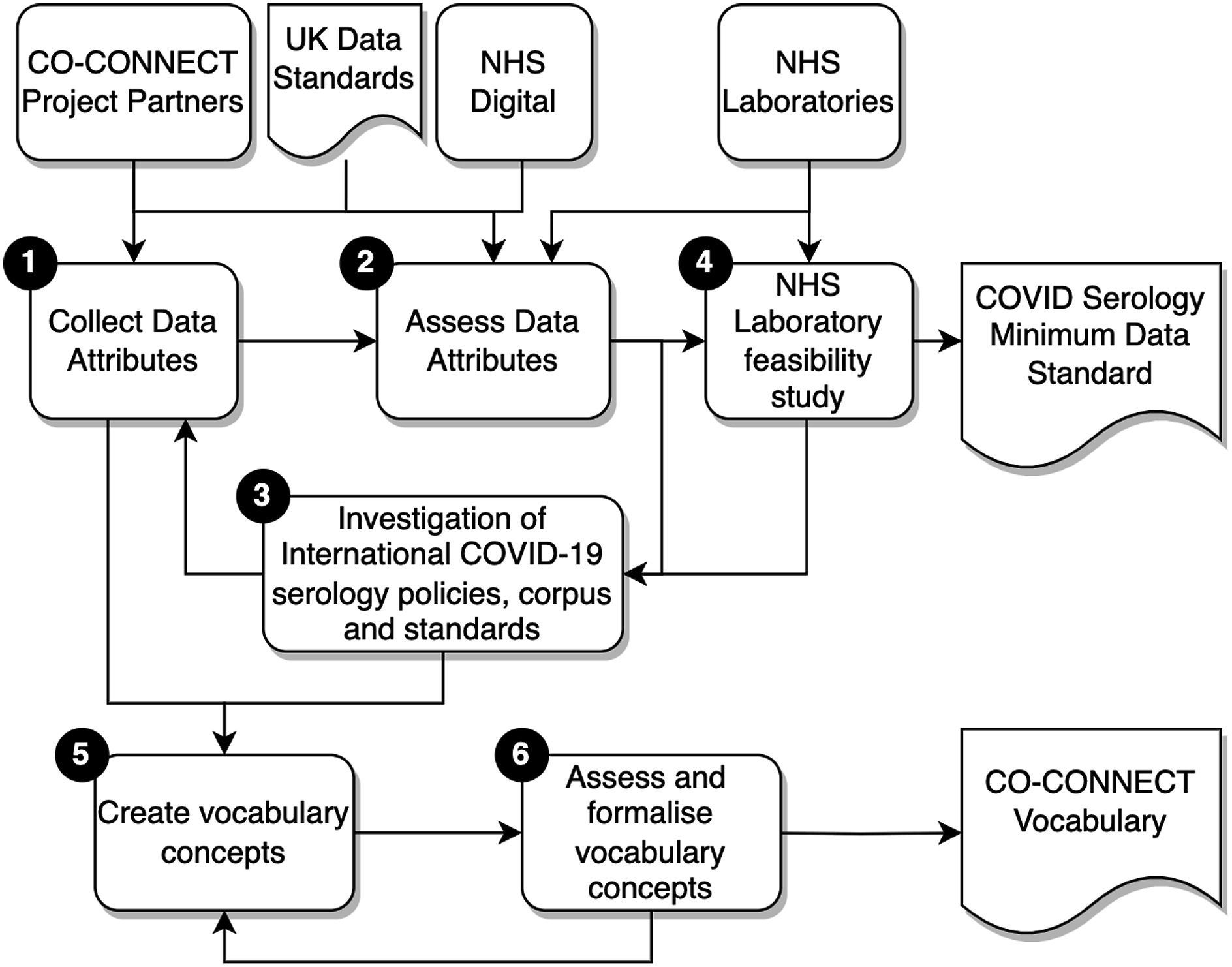
The first three stages formed an iterative feedback process loop, to enable the development of a representative set of data attributes for a minimum data standard. The fourth stage was a laboratory feasibility study to assess the CSMDS. Stages 5 and 6 were used to develop the CO-CONNECT vocabulary. The six stages employed in the method were thus: • Stage 1: Collection of data attributes: The data came from the CO-CONNECT partners organisations and three NHS laboratories. Each of the CO-CONNECT partners were collecting relevant COVID-19 serological data, from research cohorts and unconsented longitudinal datasets. After the relevant access permissions were gained from each organisation, metadata was extracted from anonymised data exports utilising the ODSHI White Rabbit statistical profiling tool
65
and shared with the CO-CONNECT team for analysis. Three NHS laboratories at Barts Health NHS Trust (Barts), NHS Great Ormond Street Hospital for Children (GOSH) and NHS Tayside acted as exemplar sources of laboratory data, providing excerpts of data being reported. These different data feeds provided an initial set of 36 individual data attributes. • Stage 2: Assessment of data attributes. Figure 2 depicts the approach taken for Stage 2, to which a Delphi method was used to help reach a consensus.61,66,67 Weekly workpackage meetings enabled debate and feedback against progress utilising expert knowledge from the project group, together with findings from stage three and UK data standards. Individual expert assessments took place independently of weekly meetings. From these processes ranked data attribute lists were produced together with collected expert feedback. The teach-back method was used, combining views and Delphi ranking votes, to reach a consensus. The iterative approach enabled the group to focus on the importance of data attributes within the standard highlighting those that were considered critical for reporting granular data. Those that were not were removed from the standard. Questions were collected to be answered in Stage 3. • Stage 3: Investigation of International COVID-19 serology policies, corpus and standards: To further support the development of the CSMDS, other sources of information were studied, such as national standards, international standards, work from other organisations, other nations and the corpus of literature upon the subject.15,16,49,54,55,68–70 These fed into Stages 1 and 2 to inform the debate and further refine the approach. • Stage 4: NHS laboratory feasibility study: Once the CSMDS had reached a level of maturity, the next stage was to understand whether it represented the COVID-19 serology test result data being collected within the CO-CONNECT partner organisations and pilot NHS laboratories and if it was fit for purpose. To accomplish this, a 4-step process was applied to each of the pilot laboratories at NHS Barts, NHS GOSH: Step 1: Assessment of laboratory HL7 messages – Anonymised excerpts of HL7 COVID-19 serology messages from the three laboratories were studied to understand how they represented the test data within the HL7 message syntax. Step 2: Comparison of laboratory HL7 messages against the CSMDS - the CSMDS was compared against each laboratory HL7 COVID-19 serology message to determine which CSMDS data attributes were represented and which were not, in effect a gap analysis was performed. The missing data attributes were highlighted. Step 3: Definition of HL7 changes – Utilising the highlighted missing CSMDS data attributes, changes were defined for each of the laboratories' HL7 messages so that they could be represented within them. In effect, where to put each data attribute within the message and accordingly the data type to use. Step 4: Laboratory assessment of the proposed HL7 changes – each of the three NHS laboratories assessed the proposed changes to their existing HL7 COVID-19 serology messages to move towards CSMDS standardisation. Specifically: (a) was it possible to make the necessary changes to their LIMS software and output the data attributes needed for the CSMDS; and (b) how easy was it to accomplish these changes and were there any complications that might hinder the move towards CSMDS standardisation? • Stage 5: Creation of vocabulary concepts: Data attributes and exemplar HL7 laboratory messages were collected throughout the process of developing the CSMDS, these were examined to understand the types of laboratory analyser systems and LIMS software that were being used to capture the data. To support this, a web-based evaluation of current analysers and LIMS software was undertaken, together with a study of numerous hospital trust ISO 15189 accreditation certificates.
71
From these, a list of laboratory analyser systems and LIMS software was compiled with the associated manufacturers. • Stage 6: Assessment and formalisation of vocabulary concepts: The CO-CONNECT project data team reviewed the entries within the list of analysers and LIMS software and assessed whether they were suitable and fit for purpose. Outputs from these assessments formed a feedback loop to Stage 5 and changes were made to the compiled list. Stage 2 - assess data attributes and information processes.
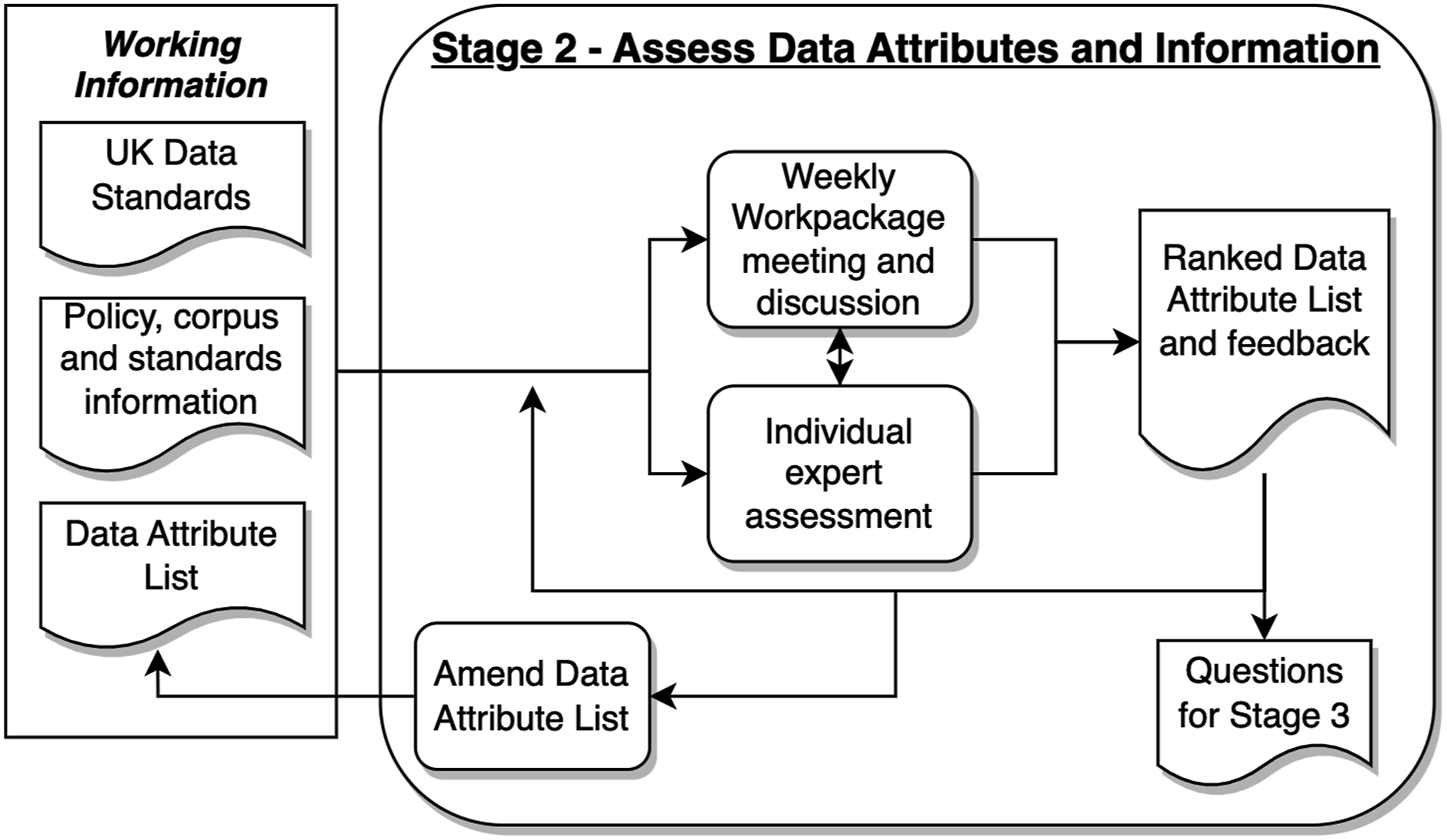
Results
The final outputs were (i) a COVID-19 serology minimum data standard comprised of 12 data attributes and (ii) a formalised CO-CONNECT vocabulary for laboratory analyser systems, LIMS software providers and SARS-CoV-2 serology measurements:
COVID-19 serology minimum data standard
COVID-19 serology minimum data standard data attributes.
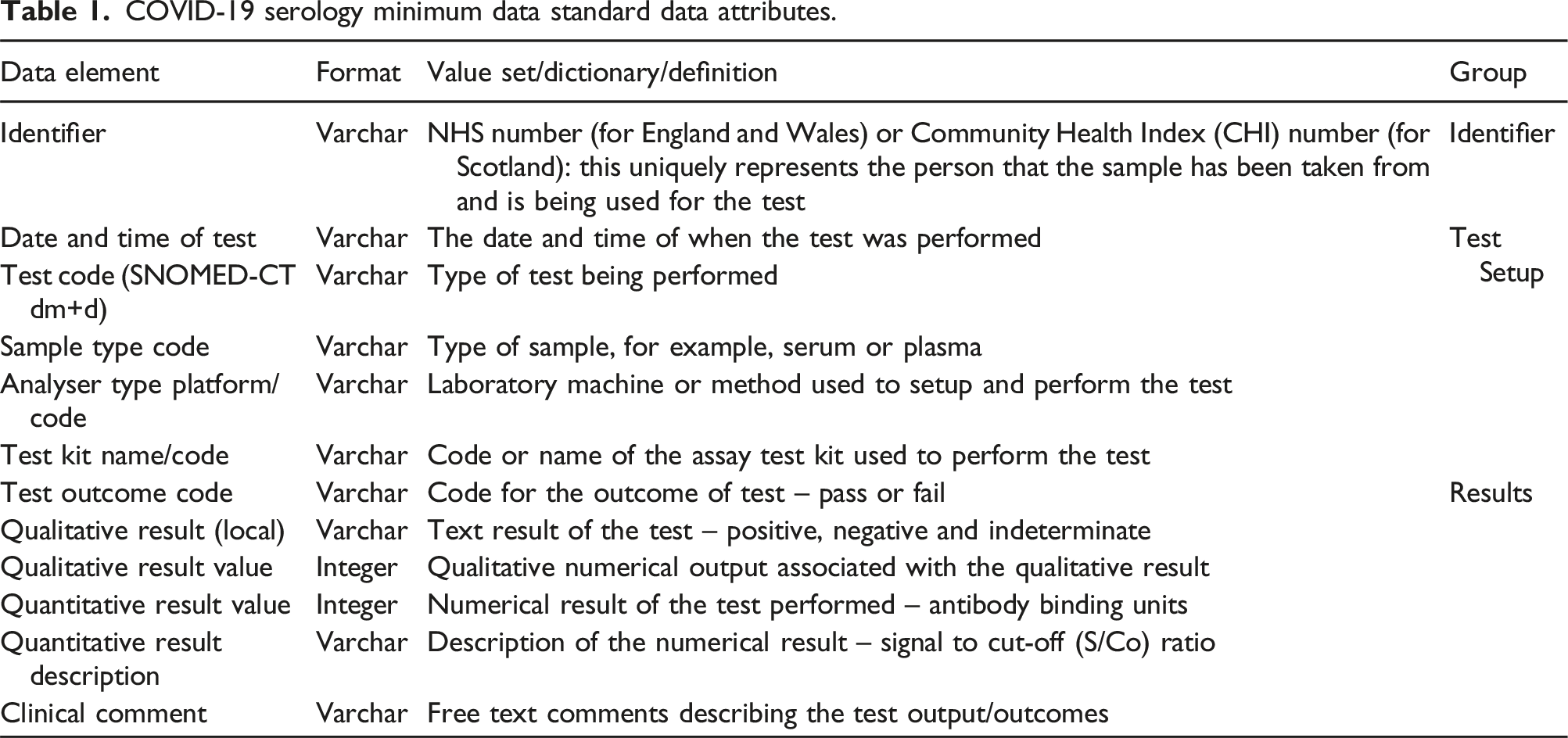

The 12 data attributes can be grouped into 3 main areas: (a) identifier, (b) test setup and (c) results as shown in Table 1.
Standardised vocabulary
Laboratories often used local vocabulary within the HL7 test messages. This brought about difficulties when sharing data between different laboratories or on a much grander scale, that is, nationally. Two such instances of this were the naming of laboratory analyser systems and LIMS software. No standardised naming convention (vocabulary) was observed across a randomly selected number of different laboratory result reports within England.
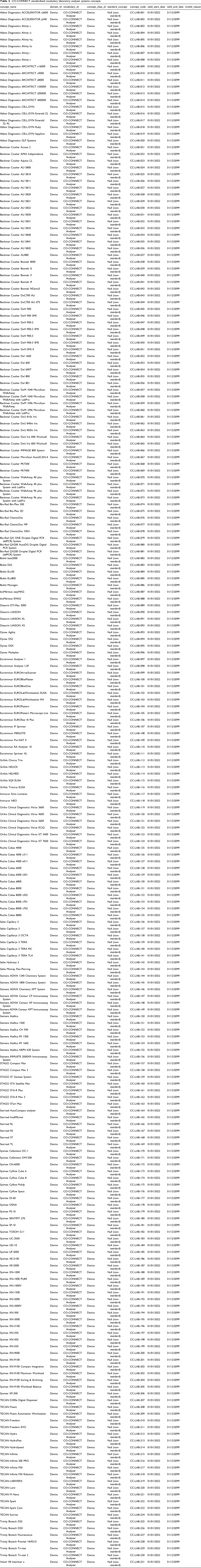
CO-CONNECT standardised vocabulary laboratory analyser systems concepts.
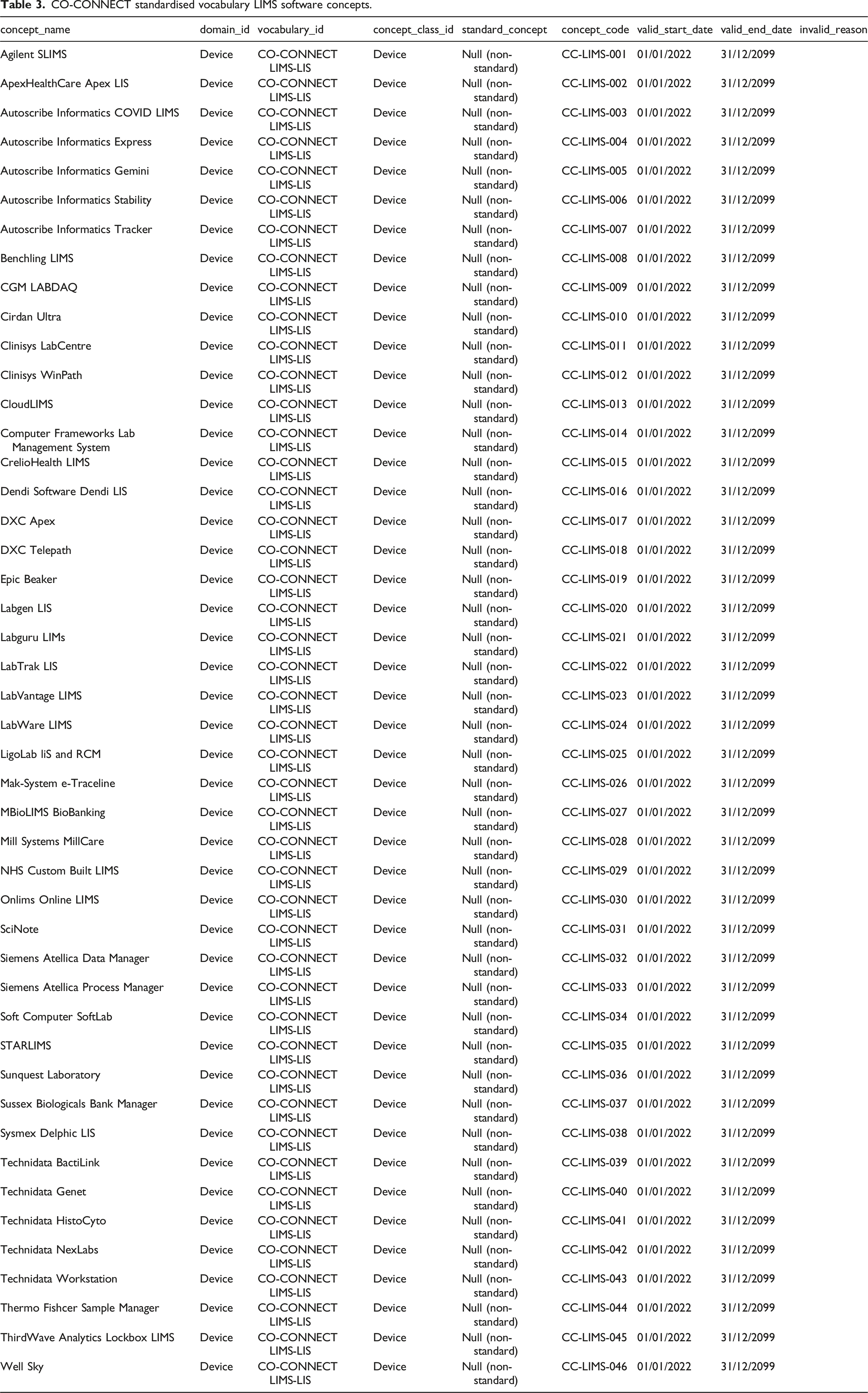
CO-CONNECT standardised vocabulary LIMS software concepts.
CO-CONNECT standardised vocabulary SARS-COV-2 serology measurement concepts.
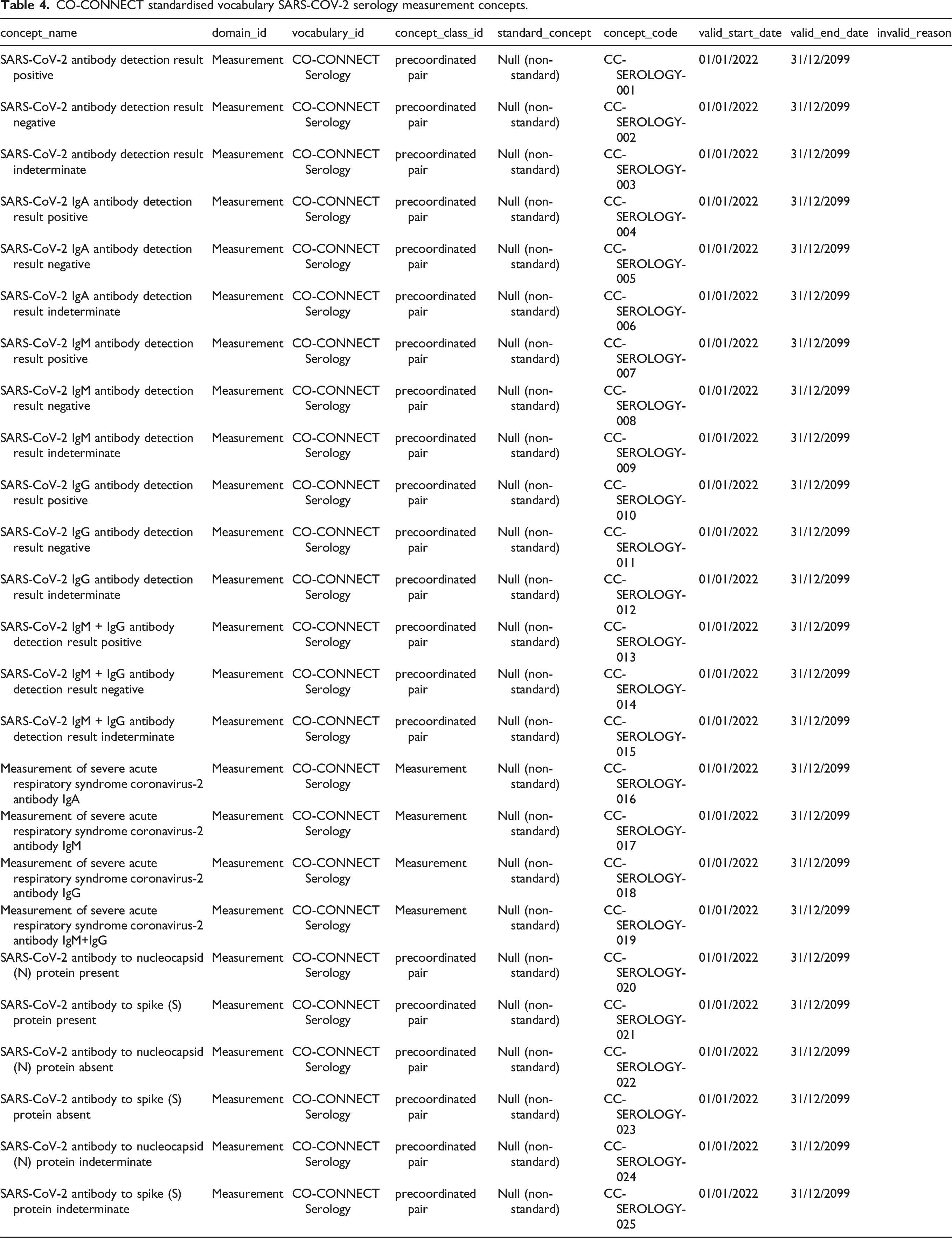
The Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM) approach to the description of concepts was utilised to structure the expression of the CO-CONNECT vocabulary concepts.72,73 Employing the OMOP CDM schema to describe concepts in a standardised manner supports interoperability and reuse.74–77 Tables 2–4 illustrate how the concepts were formalised by IQVIA (a company who provide OHDSI and OMOP services) and form part of the larger standardised CO-CONNECT vocabulary that has been published for universal use.
Each concept of the CO-CONNECT vocabulary has a ‘concept_name’ and a ‘domain_id’ (a unique domain identification). The laboratory analyser systems and LIMS software have a domain_id of device (see Tables 2 and 3), whilst the SARS-CoV-2 serology measurement domain_id is a measurement (see Table 4). The ‘vocabulary_id’ is a code that represents each vocabulary. For the laboratory analyser systems it is CO-CONNECT Analyser, for LIMS software it is CO-CONNECT LIMS-LIS and for the SARS-CoV-2 serology measurements it is CO-CONNECT Serology. The ‘concept_class_id’ is a semantic tag which is a unique identifier for the class a specific concept belongs to.
74
The concept_class_id for laboratory analyser systems (see Table 2
As part of the process, the CO-CONNECT concepts were, where possible, mapped to the representative standard SNOMED codes. These concepts can now be used to represent the respective devices and measurements within the OMOP CDM. Each concept has a unique ‘concept_code’ (source code) for each of the three CO-CONNECT vocabulary sections.
53
These source identifiers follow the nomenclature of: • CC-LAB-xxx for laboratory analyser systems; • CC-LIMS-xxx for LIMS software; • CC-SEROLOGY-xxx for SARS-CoV-2 serology measurements.
The CC stands for CO-CONNECT, whilst the xxx signifies an assigned integer, for example, 001. The fields of ‘valid_start_date’, ‘valid_end_date’ and ‘invalid_reason’ signify the lifecycle of a vocabulary. The ‘valid_start_date’ for all concepts was set as the first of December 2022 and the ‘valid_end_date’ was set as the thirty first of December 2099. There were no values for ‘invalid_reason’ as the vocabulary is currently valid. These concepts form part of the larger CO-CONNECT formalised vocabulary, which augments the currently available international standardised vocabularies, such as SNOMED. 68
As data completion rates were low, it was necessary to specifically ask for the additional data points and national laboratories had to invest time and effort into providing these as it was not normal practice to share more than the result. This points to fundamental differences in the data requirements of the clinical and research use cases. Most NHS labs are configured to support clinical use cases and the additional data items required to provide research-quality data mean additional configuration, testing and work that cannot be justified within the clinical context. The paucity of implementable standards in this domain only makes it harder. Data points such as device type/id and the specific methodology used are not seen as critical/high value items in the clinical domain.
Feasibility of adoption of the CSMDS
During the laboratory feasibility studies at Barts, GOSH and Tayside, it was found that it would be straight forward to change their laboratory systems to report the CSMDS granular data. Ten of the major LIMS software providers were contacted, introduced to the proposed CSMDS and asked to complete a survey on whether their software could accommodate the reporting of the CSMDS data attributes. Three LIMS software providers replied to the survey, with two stating that there would be no foreseen problems in implementing such a CSMDS, they could capture and report all of the data attributes and in fact were currently doing so for most assays. The third responded that instances of their software where setup differently with potentially different interfaces and therefore they were unsure if they could support the CSMDS. Further analysis is needed to fully understand at a national level what the barriers to implementation might be.
Discussion and conclusions
The CO-CONNECT project investigated how COVID-19 serology laboratory results are captured and reported within the UK, focussing on how to improve these processes and how to capture more granular data. High-level qualitative results (positive, negative, etc.) were found to be routinely captured, yet many testing laboratories produce an array of quantitative data which could also be relatively easily reported. How data is reported via national systems such as Labgnostic (in England) and Scottish Care Information Store (equivalent system in Scotland) varies across different laboratories, ranging from full reporting to almost no reporting of quantitative data. This is in part due to how individual laboratories are configured, the analyser machines they utilise and the LIMS software they employ to capture, structure and then report said data. There was no nationally specified minimum set of data attributes for the reporting of COVID-19 serology results that laboratories could apply and adhere to, to support the reporting of granular results across the United Kingdom. The COVID-19 Serology Minimum Data Standard (CSMDS) fills this gap and sets out a clear and structured approach to what should be captured and reported, stipulating 12 distinct data attributes. The approach and method applied could be further refined and applied on a larger scale.
We found locally encoded naming conventions are often applied to data attributes, for example, the name given to the laboratory analyser system conducting the test. Utilising controlled vocabularies such as LOINC and SNOMED-CT resolves this variance supporting comparisons at a national level across assays, test kits, laboratory analyser systems and LIMS software. Our work has produced a set of formalised concepts as part of the CO-CONNECT standardised vocabulary for the representation of laboratory analyser systems, LIMS software names and SARS-CoV-2 serology measurements to support such an approach, with the view to understanding and potentially reducing variability of reporting testing data. These new concepts form the first step towards a standardised representation of laboratory analyser systems and LIMS software, not only for serology, but potentially for numerous other laboratory tests. Such an approach, if adopted nationally, can enable clear, unambiguous identification and reporting of the laboratory analyser systems, the LIMS software being used and the results for testing at a given point in time.
Historically, the granular level of detail has not been routinely captured as the calibrations and research questions generally have already been undertaken over many years prior to use in clinical care. COVID-19 was a new disease, and a rapid testing program was delivered within a clinical setting without prior research and calibration. To be able to respond in a timelier fashion to future pandemics and to support research requiring national level laboratory data linked to other relevant healthcare records, we highly recommend that standardised, granular level, laboratory data is captured nationally and shared using automated pipelines with organisations collecting other national health datasets. Although in England some high-level data is captured from several health trusts, granular level data is not fully captured and none of the information is currently shared and linked to other national health datasets via automated pipelines. Within Scotland, data is captured via the Sci-Store system but is also not standardised and automatically linked. When we investigated if granular level data could be captured at a Scottish national level, we were informed that the systems for capturing the data were antiquated and cannot be modified to capture more fields without risking stability.
Looking forward, there are three potential areas to address for this work to progress: 1. The work set out herein has utilised three NHS laboratories to gain a snapshot of the current situation, together with exemplar data from other CO-CONNECT partner NHS laboratories, thus it is not wholly representative of all organisations across the United Kingdom. This is therefore a limitation. Across England, there are multiple laboratories, using many different types of analysers and LIMS software, ranging from brand new state-of-the art systems to antiquated legacy systems. To facilitate adoption of the CDMDS, a thorough assessment of a wider range of organisations, technologies, software and reported laboratory messages must be performed to appreciate who does what, with what and how. 2. There is a need to apply a standardised vocabulary for the reporting of laboratory results. The use of more detailed standardised naming conventions (for laboratory analyser systems, LIMS software and SARS-CoV-2 serology measurements) will enable the reporting of higher quality detailed data. This will potentially allow for greater understanding of variability in laboratories which could support further work to address and decrease this. 3. To support future pandemics and research using test data, a new data pipeline should be built and configured, so that results can be reported to organisations such as NHS England.
Footnotes
Acknowledgements
The authors wish to thank all the staff members involved from NHS Tayside, NHS GOSH and NHS BARTS for their efforts with data collection and assessment work as part of this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We acknowledge the support from CO-CONNECT funded by UKRI (MRC) and DHSC (NIHR) (MR/V03488X/1) and supported by a consortium of 29 partner organisations, NHS Tayside, NHS GOSH and NHS BARTS whose infrastructure made this research possible.
Ethical approval
No ethical approval was required for this research work, and the basis for the research was infrastructure.
Guarantor
ERJ.
Contributorship
ENU contributed to writing – original draft, writing – review and editing, methodology, data collection, investigation, formal analysis and supervision; JM contributed to writing – review and editing, methodology, investigation and formal analysis; NJS contributed to writing – review and editing, methodology, investigation and formal analysis; AH contributed to writing – review and editing, data collection, investigation and formal analysis; JJ contributed to writing – review and editing, methodology, investigation, formal analysis and supervision; EM contributed to writing – review and editing, data collection, investigation and formal analysis; GM contributed to writing – review and editing, investigation and supervision; LM contributed to data collection and investigation; AC contributed to writing – review and editing and investigation; MF contributed to writing – review and editing, methodology, data analysis, investigation and formal analysis; PQ contributed to funding acquisition, writing – review and editing and supervision; ERJ contributed to funding acquisition, writing – review and editing, methodology, investigation, formal analysis and supervision.
