Abstract
Objective
To provide theoretical support for clinical diagnosis of lung cancer through an overview of systematic reviews (SRs) of the diagnostic value of miRNA.
Methods
We searched PubMed, Embase, and the Cochrane Library to collect SRs of the diagnostic value of microRNA for lung cancer until April 2021. A comprehensive database search was carried out, screened, and extracted information independently by two researchers, to compare and analyze the sensitivity and specificity of relevant literatures. The ROBIS tool was applied to assess the risk of bias of included SRs and meta-analysis.
Results
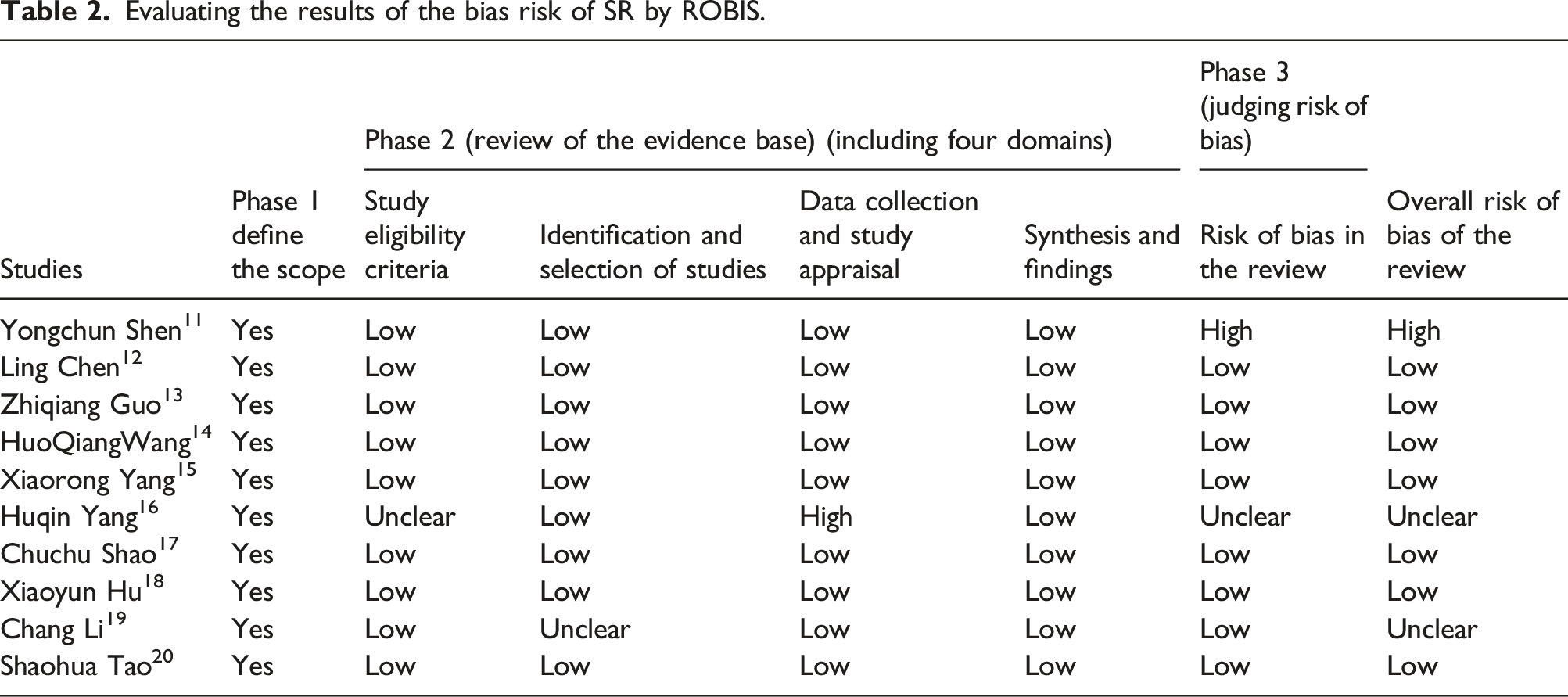
A total of 10 SRs were included the results of risk of bias assessment by ROBIS tool showed: 10 SRs completely matched the four questions of phase one. In phase two, nine SRs were low risk of bias in the including criteria field; one study was uncertain; nine SRs were low risk of bias, one study was uncertain in the literature search and screening field; nine SRs were low risk of bias in the data abstraction and quality assessment field, one SR was high; and 10 SRs were low risk of bias in the data synthesis field. In the phase three of comprehensive risk of bias results, 8 studies were low risk, one was high risk, and one study was uncertain. MiRNA had a pooled sensitivity of 0.77 (95% CI: 0.73–0.81) and specificity of 0.81 (95% CI: 0.79–0.84). The summarized area under the SROC curve was 0.86 (95% CI: 0.83–0.89), and combined diagnostic odds ratio was 14.68 (10.87–19.81). The sensitivity and specificity of multiple MicroRNA was 0.80 (0.73–0.85) and 0.80 (0.76–0.83); the sensitivity and specificity of single MicroRNA was 0.74 (0.69–0.79) and 0.83 (0.80–0.88).
Conclusion
MiRNA is a promising biomarker for the diagnosis of lung cancer in Chinese population, with advanced sensitivity and specificity. It provides a faster and less invasive assessment of lung cancer than other markers that require histopathological analysis. We should improve the reliability of the conclusion. The construction of diagnosis for lung cancer provides reliable evidence support.
Introduction
Lung cancer (LC) is a malignant tumor with extremely high morbidity and mortality. 1 Epidemiological research has indicated that lung cancer will remain the main public health issue in major countries before the 2050s.2,3 Lung cancer is classified according to the type of disease into non-small cell lung cancer (NSCLC) (about 85%) and small cell lung cancer (SCLC) (about 15%). Most patients are found to have advanced lung cancer, 4 and previous studies showed that NSCLC patients have 80% chance to survive 5 years if they are diagnosed at early stage. However, early diagnosis for lung cancer is difficult by the lack of effective methods. 5 Hence, the determination of effective biomarkers related to diagnosis is of great help for clinical application.
MicoRNAs (miRNAs) which are a group of non-coding small RNAs and participate in regulating gene expression; aberrant miRNA expression has been shown to play an important role in incidence of human malignancies, has opened up new perspectives for tumor diagnosis and provides a novel approach for diagnosis of tumors, including lung cancer. 6 Furthermore, the dysregulation of miRNAs has been reported in many diseases, particularly, cancers. 7 Abnormal miRNAs expression has been shown to play an important role in the development of tumors; it may provide a more accurate prediction for patients with cancers, especially NSCLC in their clinical utility as biomarkers. 8 The miRNAs could be stably detected in serum or plasma and have been used as biomarkers for diagnosis, so they are an emerging potential tool for the diagnosis of lung cancer. 9
In recent years, systematic reviews of primary diagnostic studies for all diagnoses and tests for the diagnostic value of miRNAs in lung cancer, but due to the different quality of the studies, the results of different studies may also present biases or contradictions. So, evidence for lung cancer has not been synthesized in one clinically useful format. In this report, to provide more reliable evidence-based medicine evidence to clinicians, the diagnostic value of miRNAs in lung cancer is given by systematically reviewing all the SRs and meta-analysis (MA). Given the heterogeneity in reported miRNA profiles between ethnicities, the studies of review were from different countries and regions, for example, America, Asia, and Africa.
Methods
Retrieval strategy
PubMed, Embase, and Cochrane Library were searched to evaluate diagnostic the value of miRNA in lung cancer. The retrieval time is from the establishment to 15 April 2021. The retrieval strategy is the combination of subject words and free words. The search terms were lung neoplasms OR adenocarcinoma of lung OR bronchioloalveolar adenocarcinoma OR bronchial neoplasms OR bronchogenic carcinoma OR multiple pulmonary nodules OR pancoast syndrome OR pulmonary blastoma OR pulmonary sclerosing hemangioma OR lung neoplasm* OR lung tumor*OR lung carcinoma*OR lung cancer*OR lung tumour* OR pulmonary neoplasm*OR pulmonary tumor* OR pulmonary carcinoma*OR pulmonary cancer*OR pulmonary tumour*OR bronchial neoplasm OR bronchogenic carcinoma*OR bronchial carcinoma* OR lung adenocarcinoma*OR alveolar adenocarcinoma*OR alveolar carcinoma* OR alveolar cell carcinoma*OR bronchiolar carcinoma*OR bronchioloalveolar carcinoma*OR pancoast syndrome OR pulmonary sclerosing hemangioma* OR lung sclerosing hemangioma*OR multiple pulmonary nodule OR non-small cell lung carcinoma*OR non-small cell lung cancer*OR non-small cell lung carcinoma* OR non-small cell lung cancer* OR small cell lung cancer* OR oat cell lung cancer*and meta-analysis and systematic review.
Inclusion criteria and exclusion criteria
The inclusion criteria were as follows: (1) concerned the diagnostic potential of miRNAs for LC, (2) the relationship between miRNA and diagnostic accuracy was studied, and (3) studies must provide sufficient data to assess diagnosis value of miRNA in lung cancer detection. Exclusion criteria were as follows: (1) not related to the diagnostic values of miRNA for LC, (2) reported duplicate data from other studies, (3) took the form of letters, editorials, case reports, or reviews, and (4) the literature studied the relationship between the gene expression of miRNA and diagnostic accuracy, with a receiver operating characteristic curve (ROC), but without report of the specific values of sensitivity and specificity.
Literature screening and data extraction
Each article, title, and abstract were screened by two independent reviewers to determine eligibility. In case of disagreement, consult a third party to assist in judgment, lack of information, and contact the author for more information. When selecting the literature, read the title first and abstract, after exclusion of apparently irrelevant literature, for further reading full text to determine final inclusion. Data extraction of the selected SR/MA included: first author, year, country of corresponding author, inclusion criteria, evidence quality assessment tool, pooled sensitivity, pooled specificity, pooled diagnostic odds ratio (DOR), pooled positive likelihood ratio (PLR), negative likelihood ratio (NLR), area under the curve (AUC), the risk of bias SR, and quality of evidence
Risk assessment of bias included SR
The risk of bias included SR was assessed by two evaluators by the ROBIS (Risk of Bias in Systematic Review) tool, including three stages: (1) assess relevance (optional), (2) identify concerns with the review process, and (3) judge risk of bias. 10 Phase 2 covers four domains through which bias may be introduced into a systematic review: study eligibility criteria; identification and selection of studies; data collection and study appraisal; and synthesis and findings. Phase 3 assesses the overall risk of bias in the interpretation of review findings and whether this considered limitations identified in any of the Phase 2 domains.
Results
Data selection and study characteristics
The results of the literature search were shown in Figure 1(a). A total of 1365 relevant articles were identified in the initial search. One hundred sixty-seven were removed for duplication. After carefully reviewing the titles and abstracts, a further 1065 were excluded, of which 165 were reviews and 1032 were irrelevant articles for our topics. After full-text reading by two reviewers, 120 failed to meet the inclusion criteria. Finally, 10 SRs were included in this overview. Flow diagram of study selection process.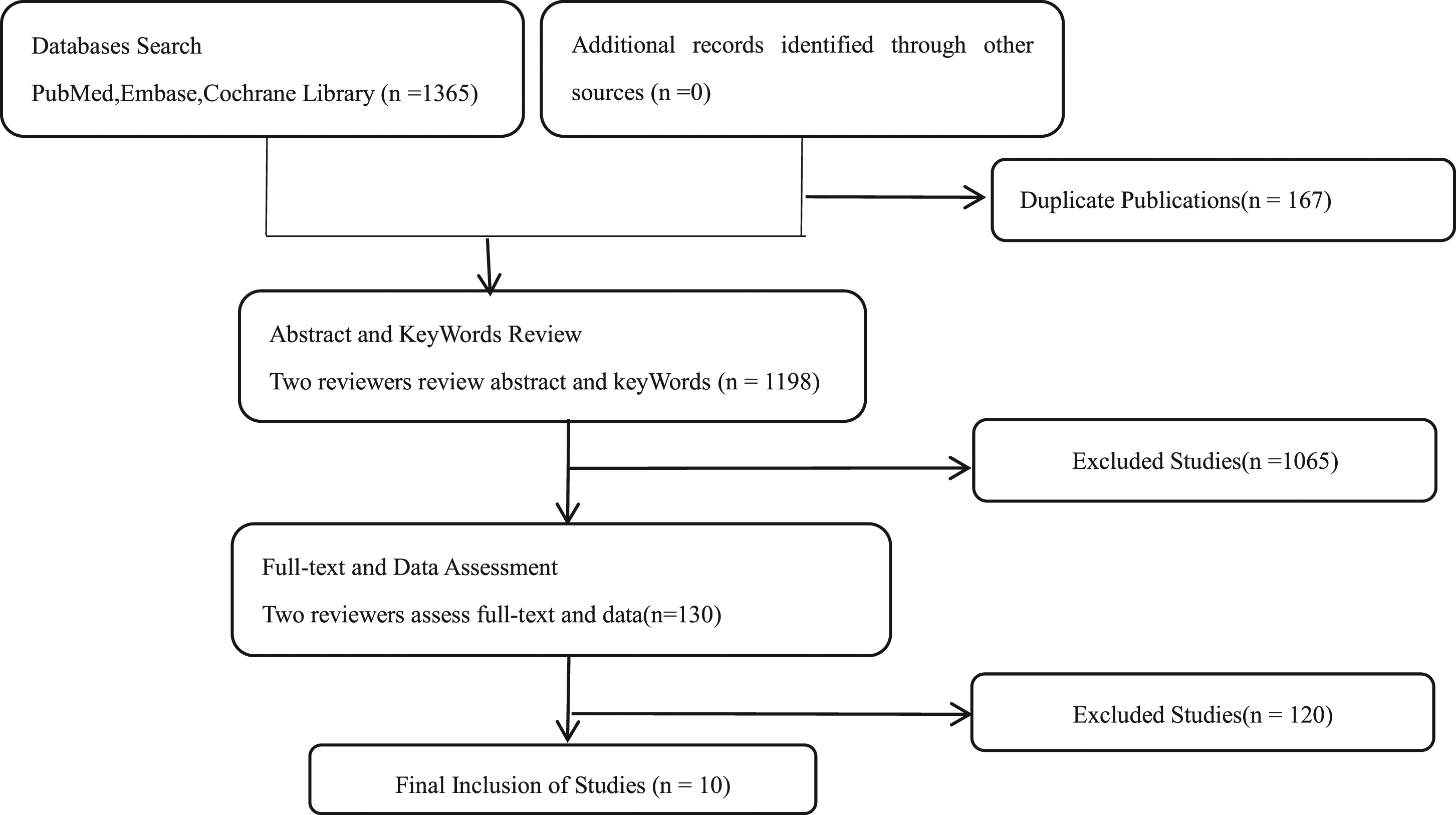
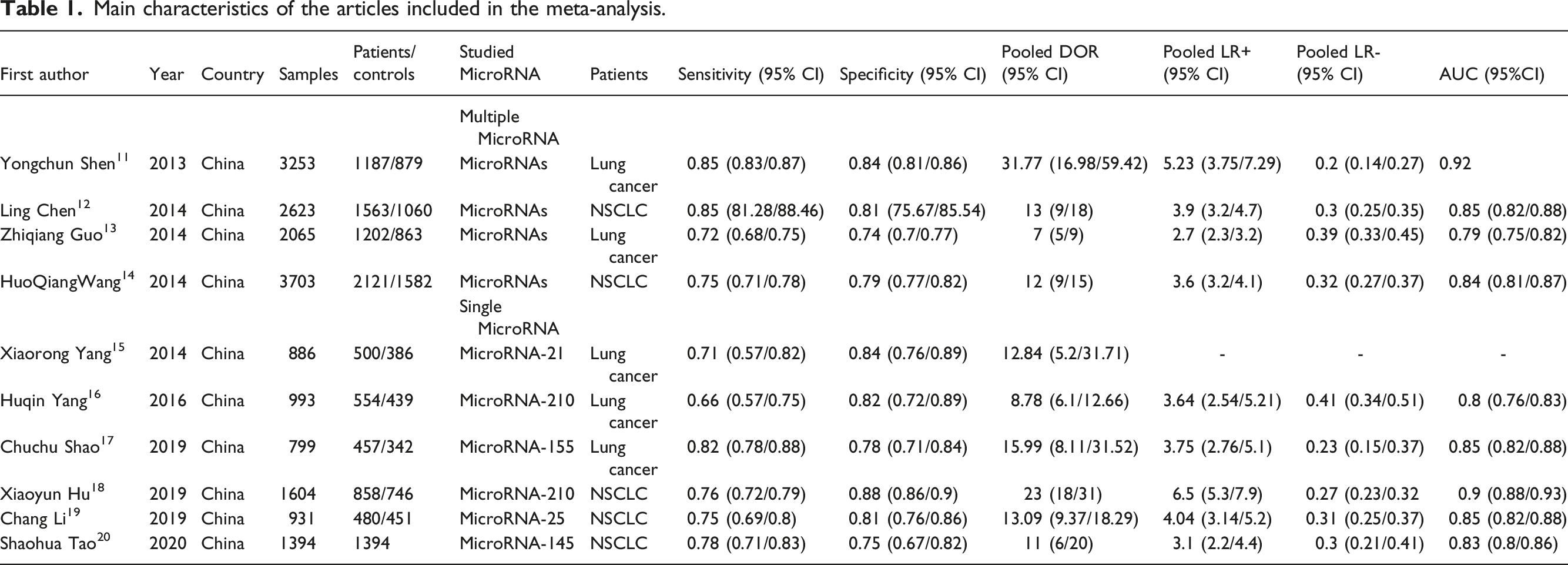
Main characteristics of the articles included in the meta-analysis.
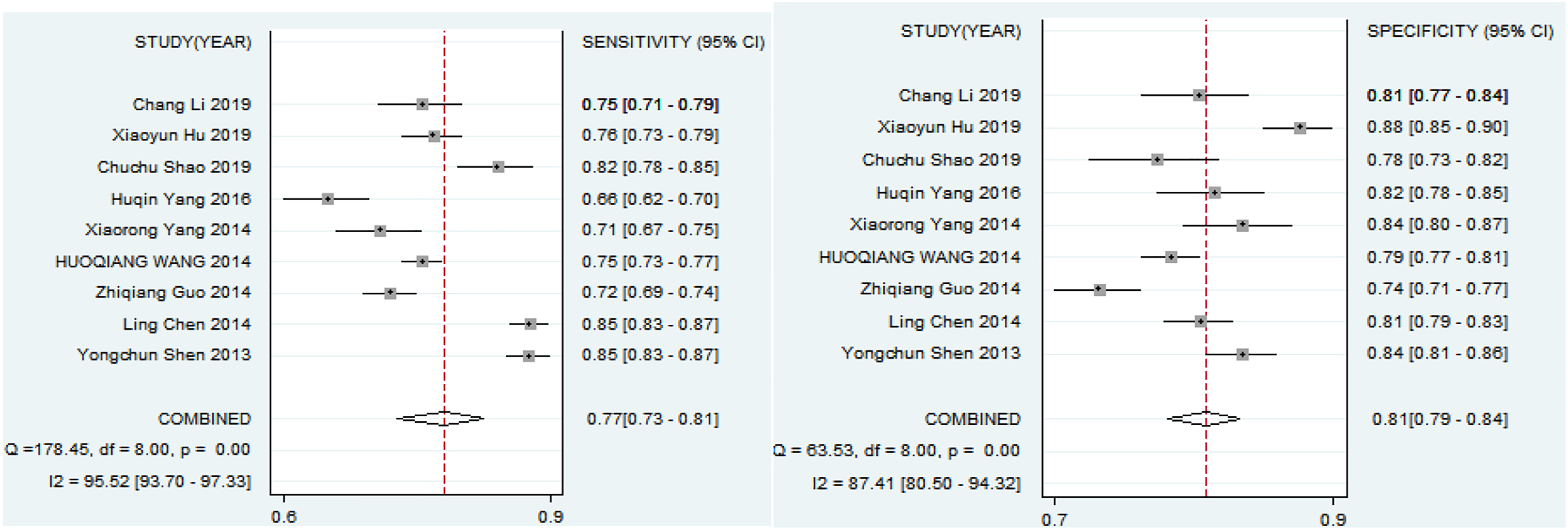

Combined sensitivity and specificity of miRNA in diagnosis of lung cancer assessed by forest plots.
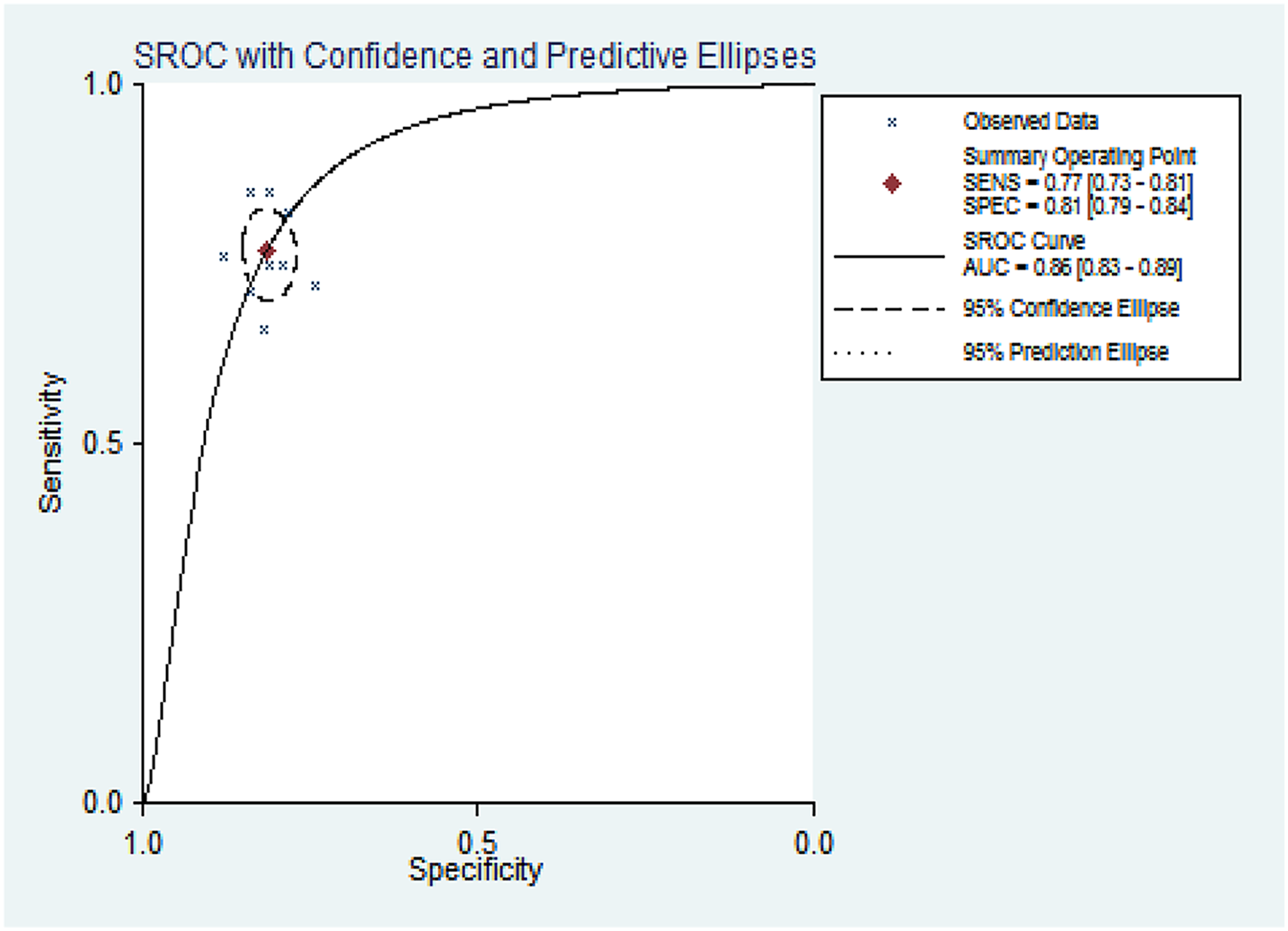
Summary receiver operating characteristic (sROC) curve of miRNA in lung cancer diagnosis.
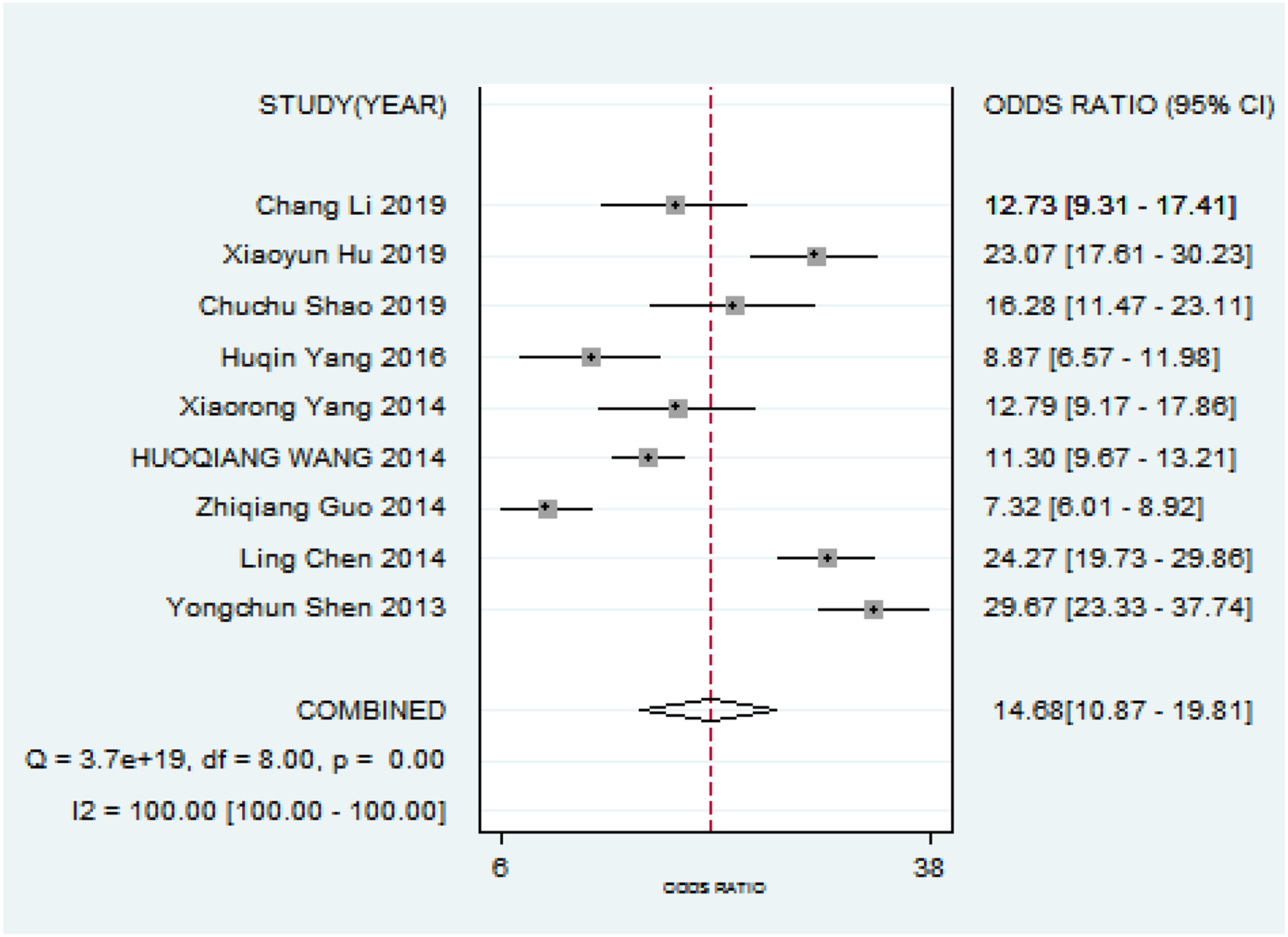
Diagnostic odds ratio of miRNA in lung cancer diagnosis.
Risk-of-bias assessment of the included SRs by ROBIS
Evaluating the results of the bias risk of SR by ROBIS.
Subgroup analysis
Subgroup analysis based on the typle of miRNAs.
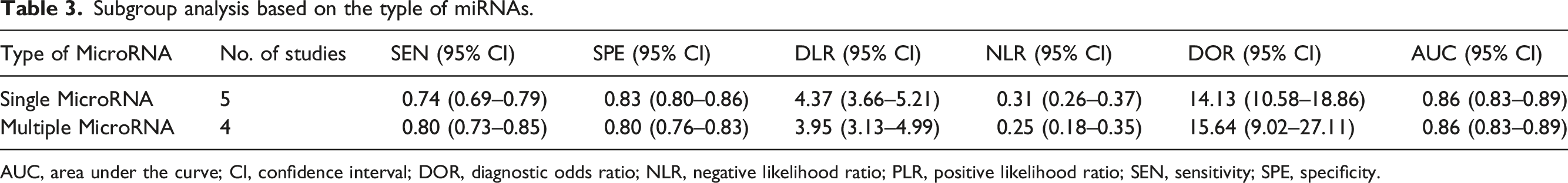
AUC, area under the curve; CI, confidence interval; DOR, diagnostic odds ratio; NLR, negative likelihood ratio; PLR, positive likelihood ratio; SEN, sensitivity; SPE, specificity.
Discussion
Diagnosis of lung cancer by evidence-based medicine of methods and concepts provides efficacy and safety of diagnosis, scientific basis, and actively evaluate evidence-based screening which will help improve the level of lung cancer diagnosis. 21 According to the statistics, the preventive and control situations of NSCLC in most countries remain pessimistic in the future. 22 Duing to controversial results have been reported in different biomarkers, populations, and studies on the predictive role of the mutations identified in disease, whereas there is still a lack of widely used diagnostic biomarkers for lung cancer with satisfactory sensitivity and specificity. 19 Therefore, seeking biomarkers for early diagnosis of lung cancer is the key to improve the survival rate of patients with lung cancer and reduce the financial burden of patients. Emerging evidence has reported that miRNAs are frequently dysregulated in cancer and may be involved in tumorigenesis and several critical cellular processes, such as cellular proliferation, differentiation, and apoptosis.23,24
Studies have shown that 50% of miRNA coding genes identified are located in tumor-related gene regions or fragilesites, and there is a significant difference in expression of these miRNAs between tumor cells and corresponding normal cells, which indicates that miRNA may play a critical role in occurrence and development of human tumors. 19 MiRNA has provided a potential new way for early diagnosis of cancer. Studies show that miRNA can abnormal expression in lung cancer tissues and stably exist in serum. 25 Some miRNAs were found to be dysregulated in NSCLC which may indicate disease status or disease prognosis. 26 Most of studies have investigated the potential correlation between miRNA profile expression in the circulation and its possible clinical use to diagnose NSCLC as the experimental results varied from different studies may influence diagnostic capabilities. 20 On the one hand, this research conducted an overview of systematic reviews to ascertain the overall accuracy of miRNA in detecting lung cancer.
In this study, all cases in enrolled studies received blood tests as minimally invasive (venous puncture). Based on the 10 included articles, miRNA can be detected in the plasma of lung cancer patients with marked differences when compared with controls, suggesting it can serve as a serum-based biomarker for lung cancer detection individually. Our studies demonstrate the potential of miRNAs to be a minimally invasive diagnostic test. MiRNAs to diagnose lung cancer were following advantages: A miRNA-based test would present comprehensive analysis of tumors without invasive, such as lung biopsy or surgical exploration. Sample collection and processing would be relatively inexpensive and simple. Such miRNA could offer a greater sensitivity and specificity than other diagnostic biomarkers, consistent with previous studies. 13 A forest plot depicting the diagnostic accuracy of miRNA for lung cancer indicated that miRNAs from different SRs sensitivity and specificity were different. The combined sensitivity was 0.77 (0.73–0.81) and specificity was 0.81 (0.79–0.84), and AUC was 0.86 (0.83–0.89). The combined specificity and sensitivity are relatively high. There is not much difference for specificity, indicated that miRNA has high specificity in the diagnosis of lung cancer. Nevertheless, sensitivity of miRNA was differences. It may be caused by the types of patients included and the number of studies. It should be noted, however, that the reported miRNA markers in this review were mostly derived from Asia, America, and Africa to take into account heterogeneity of miRNA profiles between ethnicities. Generalization of these miRNA markers thus should be carried out to need more data support. The overall risk of bias in the systematic evaluation of the diagnostic value of miRNA for lung cancer is low.
In this meta-analysis, the diagnostic performance of miRNA assays was further assessed by subgroup analyses as shown that multiple MicroRNA had slightly high sensitivity, the specificity, DOR, NLR, and PLR which were not much difference. From the AUC, diagnostic value of lung cancer all had high diagnostic accuracy. There was no significant difference between single MicroRNA and multiple MicroRNA in the diagnostic value of lung cancer from this study. Therefore, clinicians can select a single miRNA for the diagnosis of patients according to the patient’s situation, which can reduce the economic burden of patients from this study. However, it is necessary for more research to get more accurate conclusions.
However, several limitations still existed in this overview of systematic reviews as follows: (1) We only included articles published in English, but did not cover articles in other languages. (2) The sample size was still relatively small, including only 10 studies. Therefore, larger samples and sufficient data are required to verify the diagnostic and prognostic value of miRNA in lung cancer. (3) There are few studies on lung cancer stage including SRs, and no relevant studies have been conducted in this paper. Our studies were without enough data for the adjustment by other dates such as TNM stage, histological type, and so on. Therefore, further research and studies with high-quality and large sample capacity were needed to complete current findings in subsequent years.
Conclusions
In conclusion, miRNA is a promising biomarker for the diagnosis of lung cancer with potential sensitivity and specificity. It provides a faster and less invasive assessment of lung cancer than other markers that require histopathological analysis, and the construction of diagnosis for lung cancer provides reliable evidence support.
Footnotes
Acknowledgements
We gratefully acknowledge the technical support of Jinyong Hua, Jing Liu, Mingxia Ma, Linsen Xie, and Jinhui Tian for reviewing the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Guarantor
Jinyong Hua.
Contributorship
Jinyong Hua and Jing Liu, CA developed the idea of the study, participated in its design and coordination and helped to draft the manuscript. Mingxia Ma and Linsen Xie contributed to the acquisition and interpretation of data. Jinhui Tian provided critical review and substantially revised the manuscript. All authors read and approved the final manuscript.
