Abstract
Although prostate-specific antigen-based prostate cancer screening had a positive impact in reducing prostate cancer mortality, it also led to overdiagnosis, overtreatment and a significant number of unnecessary biopsies. In the post-prostate-specific antigen era, new biomarkers have emerged that can complement the information given by prostate-specific antigen, towards a better cancer diagnostic specificity, and also allowing a better estimate of the aggressiveness of the disease and its clinical outcome. That means those markers have the potential to assist the clinician in the decision-making processes, such as whether or not to perform a biopsy, and to make the best treatment choice among the new therapeutic options available, including active surveillance in lower risk disease. In this article, we will review several of those more recent diagnostic markers (4Kscore®, [-2]proPSA and Prostate Health Index, SelectMDx®, ConfirmMDx®, Progensa® Prostate Cancer Antigen 3, Mi-Prostate Score, ExoDx™ Prostate Test, the Stockholm3 test and ERSPC risk calculators) and prognostic markers (OncotypeDX® Genomic Prostate Score, Prolaris®, Decipher® and ProMark®). We will also address some new liquid biopsy approaches – circulating tumour cells and cell-free DNA – with a potential role in metastatic castration-resistant prostate cancer and will briefly give some future perspectives, mostly outlooking epigenetic markers.
Keywords
Introduction
Prostate cancer (PCa) is the second most common cancer and the fifth cause of cancer death in men worldwide. 1 This high incidence is largely due to the generalised use of prostate-specific antigen (PSA) in PCa detection over the last 30 years. 2 Although the European Randomized Study of Screening for Prostate Cancer (ERSPC) showed a 21% reduction in PCa mortality in men invited to screening with PSA after 13 years of follow-up, 3 there is still the risk of overdiagnosis and overtreatment.4–7 This controversy has been especially highlighted after the results of the Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial in the US have been published, although a considerable number of contamination was reported in the PSA screening control group. 8 The issues about PSA-based screening are mostly due to its low specificity for cancer, 9 and to its inability of discriminating between indolent versus significant PCa. In fact, there are a significant number of men who undergo unnecessary prostate biopsy, mostly when PSA concentration is below 10.0 ng/mL, where PCa is present in only 25% of men. 10 In order to avoid overdiagnosis and overtreatment, the Public Health England (PHE) does not recommend a national screening programme for PCa. Instead there is an informed choice programme, which is called Prostate Cancer Risk Management Programme (PCRMP). According to this programme, asymptomatic men aged 50 years or over who ask their general practitioner about PSA testing should receive information on the pros and cons of a PSA test and, if they decide to do it, the test is available free. 11 On the other hand, symptomatic men and those aged under 50 who are considered to be at higher risk for PCa, should be managed according to the National Institute for Health and Care Excellence (NICE) guidelines. These guidelines do not recommend a prostate biopsy on the basis of a serum PSA level alone. Rather, it is proposed that the biopsy decision should also include other factors, such as digital rectal examination (DRE) findings, any comorbidities (along with their risk factors, including increasing age and ethnicity) and any history of a previous negative biopsy. If necessary, a multiparametric magnetic resonance imaging (mpMRI) of the prostate can also be performed to help with the decision to perform a biopsy. 12
Although PSA is a key parameter for PCa management, it has important limitations, and 60–80% of all biopsies elicited by an increased PSA value do not show PCa and are, therefore, unnecessary. This low specificity of PSA for cancer comes from the fact that several benign conditions can increase PSA levels, such as inflammation, trauma or benign prostatic hyperplasia. 13 In order to increase PSA specificity, different strategies have been developed in the beginning of the 1990s, including PSA density (ratio of PSA to prostate volume) and PSA velocity (change of PSA over a time period). 10 Also some modifications to the total PSA (tPSA) assay have been used, such as the percent free PSA (%fPSA) and the complexed PSA (cPSA). While the %fPSA has provided some clinical value in distinguishing PCa from benign conditions, mostly in the diagnostic grey zone, the cPSA added very little additional value to the clinical utility of tPSA in PCa detection. 14
In the post-PSA era, new biomarkers have emerged that can help clinicians in the decision making processes, such as whether or not to perform a biopsy and which is the best treatment option to follow. 15 In this review, besides diagnostic and prognostic markers, we will address some biomarkers with a potential role in metastatic castration-resistant prostate cancer (mCRPC) and will also briefly summarise some future perspectives, concerning mostly epigenetic markers.
Diagnostic markers
Although PSA remains the most used biomarker in PCa detection, 16 in the last few years, new biomarkers have been developed that can help clinicians decide whether to perform an initial biopsy, or to carry out a repetition biopsy. 17 We will now discuss some of those biomarkers, which are already available for clinical use.
4Kscore® (ARUP Laboratories, Salt Lake City, USA)
Vickers et al. 18 developed a statistical prediction model for PCa detection, based on a panel of four serum kallikreins: tPSA, free PSA (fPSA), intact PSA (iPSA) and the kallikrein-related peptidase 2 (hK2). The authors performed a first evaluation of this panel in the Göteborg cohort of the ERSPC, in previously unscreened men with a tPSA of 3 ng/mL or higher. 19 Later, they made a second evaluation in previously screened individuals from the Rotterdam arm of the ERSPC. 18 They concluded that the application of this statistical model – which included the four kallikreins, age and the DRE – could accurately predict PCa with a total Gleason score (GS) ≥7 both in men submitted to a first or a repetition biopsy. 20 The model had a higher predictive accuracy for high-grade cancer than only tPSA and age, with an area under the curve (AUC) of 0.798 vs. 0.699. 18 Using a cutoff of 20% for biopsy, the model avoided 513 biopsies in 1000 men not previously screened, missing 66 cancers, of which 12 had a GS ≥7, whereas only 269 biopsies would have been avoided with tPSA alone. 21 In 1000 men with a previously negative biopsy, the statistical model recommended against biopsy in 362 individuals missing 47 cancers, most of which were low grade. 18 Stattin et al. 22 performed a study with these four markers in cryopreserved serum samples from a large representative cohort from Sweden (40,379 men), of which 12,542 were followed for more than 15 years. Their conclusions supported the use of 4Kscore® as a reflex test regarding biopsy decisions, in men with a tPSA >2 ng/mL. According to the authors of this study, the four kallikrein markers allow risk stratification and enhance the prediction of metastasis, compared with tPSA alone. In men with moderated PSA elevations, at the age of 50 and 60, the four kallikreins model yielded C-indexes from 0.82 to 0.88 in predicting documented distant metastasis. 22 In a prospective study undertaken in the US, several cutoffs of 4Kscore® were compared – 6, 9, 12 and 15% – and the results showed that with a 9% cutoff 43% of biopsies would be avoided, while the diagnosis of 2.4% of GS ≥7 PCa would be delayed. However, the authors of this study argue that the cutoff to be used should be personalised. For instance, for an older man with several comorbidities, a higher cutoff of 15% could be more appropriate, than for a younger and healthier patient, to whom a 6% cutoff could be more appropriate. 23
The European Association of Urology (EAU) guidelines on PCa mention the 4Kscore® as a test intended to reduce the number of unnecessary biopsies in PSA-tested men, with an improved prediction of clinically significant PCa when tPSA is between 2 and 10 ng/mL. 24 According to the National Comprehensive Cancer Network (NCCN) guidelines on PCa early detection, this test can be offered to patients before an initial biopsy or for those with a prior negative biopsy, when the risk for clinically significant PCa is thought to be higher. The NCCN panel also underlines that no optimal cutoff has yet been established. 25
This test is currently available only in the US. 26
[-2]proPSA and Prostate Health Index (Beckman Coulter, Inc., Brea, USA)
Comparison of the area under the curve between tPSA, %fPSA, %[-2]proPSA and Prostate Health Index obtained in several studies.
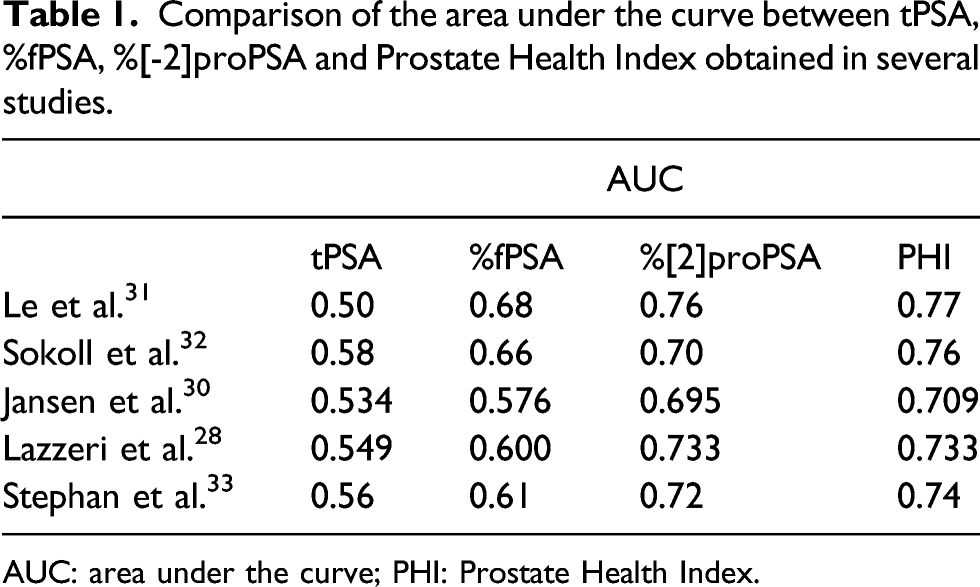
AUC: area under the curve; PHI: Prostate Health Index.
Cutoffs found in different studies for %[-2]proPSA and Prostate Health Index with the same diagnostic sensitivity (90%) and specificity (30%).
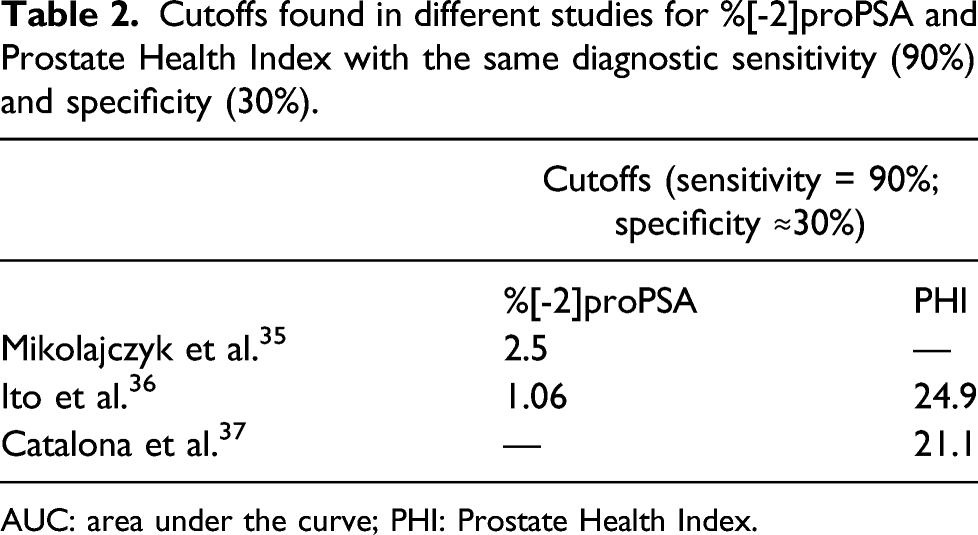
AUC: area under the curve; PHI: Prostate Health Index.
Several published studies also support that %[-2]proPSA and PHI are related to the aggressiveness of the tumour, with higher levels being obtained in patients with a GS ≥7.32,33,37–40 Guazzoni et al. evaluated these [-2]proPSA derivatives in 350 patients submitted to radical prostatectomy, and verified that both %[-2]proPSA and PHI, on univariate analyses, are accurate predictors of several pathologic disease characteristics, such as the presence of pT3 disease, GS ≥7, Gleason sum upgrading and tumour volume <0.5 mL. Moreover, both %[-2]proPSA and PHI were independent predictors when all these four pathologic endpoints were considered (P < 0.001). 39 Thus, these markers could not only reduce the number of unnecessary biopsies, but also allow the selection of patients for active surveillance (AS) or for more aggressive therapy.
Prostate Health Index density (PHID) has also been recently studied, as well as the combination of PHID and mpMRI of the prostate.41,42 Density values were calculated as the biomarker level (PHI) divided by prostate volume in mL. 42 Tosoian et al. 41 found that PHID, which accounts for total prostate volume, had a greater diagnostic accuracy to detect clinically significant PCa (AUC = 0.84) than PHI alone (AUC = 0.76). Moreover, Druskin et al. found that mpMRI could be complementary to PHID in detecting significant PCa. In a study on 104 subjects submitted to mpMRI, these authors reported 100% of diagnostic sensitivity for significant disease when the Prostate Imaging Reporting and Data System (PI-RADS) score was ≥3, or when the PI-RADS score was ≤2 and the PHID was ≥0.44. For PHID alone, using 0.44 as a cutoff, which corresponds to the 25th percentile of PHID, the sensitivity was 92.3% with a specificity of 35.4%, for clinically significant PCa. However, the sensitivity raised to 100% when PI-RADS score was ≥3 or, if PI-RADS score ≤2, the PHID was ≥0.44. 42
According to the EAU guidelines on PCa, PHI can offer further risk assessment to men with a PSA level between 2–10 ng/mL, in order to reduce unnecessary biopsies, outperforming %fPSA and improving prediction of clinically significant PCa. The EAU also highlights that PHI is already FDA approved. 24 On the other hand, the American Urological Association) states that [-2]proPSA can be a secondary screening tool, allowing the reduction of unnecessary biopsies, although there is a need for continuing research to confirm these advantages. 43 The NCCN guidelines mention several studies that have shown the potential utility of PHI in reducing the number of biopsies and in identifying the most aggressive forms of PCa. 25
SelectMDx® (MDxHealth, Irvine, USA)
This test was developed with the purpose of identifying clinically significant PCa. It measures the mRNA levels of two genes – homeobox C6 (HOXC6) and distal-less homeobox 1 (DLX1) – in a post-DRE urine sample. These two genes were selected out of a panel of several other genes studied in post-DRE urine samples by Leyten et al. and Van Neste et al.44,45 In the SelectMDx®, it is also measured the mRNA of KLK3, which encodes for PSA, as a reference for the relative quantification of mRNA from the other measured genes. The results from the two genes panel are combined with other risk factors for PCa – age, PSA density (PSAD), family history and DRE – in an algorithm to calculate the risk of PCa with a GS ≥7. When applied to patients prior to an initial prostate biopsy, this test was shown to reduce the number of unnecessary prostate biopsies, allowing a reduction in overdiagnosis and overtreatment of insignificant PCa, and hence was more cost-effective. 46 In a recently published multicenter validation study of this two-gene molecular test, using urine samples from 1955 men taken prior to an initial prostate biopsy, the AUC was found to be 0.85 with 93% sensitivity, 47% specificity and a 95% negative predictive value (NPV) to detect clinically significant PCa. 47 These data support the use of this test to guide initial prostate biopsy decisions. The NCCN guidelines mention the potential clinical advantages of SelectMDx® in identifying men with clinically significant PCa prior to biopsy, allowing the reduction of unnecessary biopsies. However, the NCCN panel considers this test to be still under investigation at the present time. 25
ConfirmMDx® (MDxHealth, Irvine, USA)
It is known that some epigenetic changes, such as DNA hypermethylation, can occur early in the oncogenic process. In PCa, these cancer-specific DNA methylation can be present in areas of prostate biopsy tissue which are histologically negative, due to a cancer associated halo effect. The ConfirmMDx® is a multiplex epigenetic assay that measures DNA methylation status of three genes – GSTP1 (glutathion-S-transferase P1), APC (adenomatous polyposis coli) and RASSF1 (Ras association domain-containing protein 1) – in prostate biopsy tissue without cancer. This assay was designed to overcome the potential sampling error that may happen during prostate biopsy, aiming to detect the presence of cancer, even when the biopsy is negative, thereby improving accuracy for predicting repeat biopsy outcome.48,49 In a large multicentre study carried out by Van Neste et al., 50 with 7899 prostate core biopsies from 803 patients, this epigenetic assay showed a NPV of 89.2% for all cancers and a NPV of 95.9% for high-grade cancer (GS ≥7), when low levels of DNA methylation were found. This high NPV suggests that ConfirmMDx® may prevent the repetition of a significant number of biopsies. According to the NCCN guidelines, ConfirmMDx® can be considered as an option for men contemplating repeat biopsy, although it is not FDA approved. 25 The EAU states that this test can provide additional information whether to repeat or not the biopsy, after a previously negative biopsy. 24
Progensa® prostate cancer antigen 3 (Hologic, Inc., Marlborough, USA)
Prostate cancer antigen 3 (PCA3) is a large non-coding mRNA chain which is highly expressed in PCa tissue. It can be detected in urine samples after a DRE and isn’t related to prostate size or tPSA level. It has been FDA approved in 2012 as an aid to decide whether or not to repeat a prostate biopsy in men with prior negative biopsies. 51 In a prospective multicentre European study carried out on 463 men with one or two previous negative prostate biopsies, men with a PCA3 score ≥35 had a 39% probability of a positive repeat biopsy, whereas subjects with a score <35 had a probability of 22%. Moreover, the mean PCA3 score was significantly higher in men with a positive biopsy (63.8 vs. 35.5; P < 0.001). In this study, PCA3 had also a higher diagnostic accuracy than %fPSA. 52 Other studies have also evaluated the role of PCA3 in guiding initial biopsy decisions. In a multicentre prospective study with 519 men scheduled for initial biopsy, with a tPSA between 2.5 and 10 ng/mL, PCA3 outperformed tPSA, PSAD and %fPSA. The authors found that men with a PCA3 score ≥35 had a 2.7-fold greater likelihood of having PCa (sensitivity = 64%, specificity = 76%). 53 Wei et al. 51 suggest the use of different cutoffs when PCA3 is used before an initial biopsy or a repeat biopsy. In men submitted to a first prostate biopsy, when PCA3 score was ≥60, the predictive positive value was 80% (sensitivity = 42%, specificity = 91%), whereas in men that underwent a repeat biopsy, a PCA3 score ≤20 provided a NPV of 88% (sensitivity = 75%, specificity = 52%). Several studies have also shown that PCA3 score mean is significantly higher in patients with a GS ≥7,52,53 but there are also studies where a significant difference was not found.54,55 The role of PCA3 in selecting patients for AS has also been investigated. Ploussard et al. 56 verified that a PCA3 score <25 was strongly indicative of low tumour volume and insignificant disease.
According to the EAU guidelines, this test is mainly indicated as an aid to guide the decision of repeat biopsy after a first negative biopsy, although its clinical effectiveness for this goal is still uncertain. 24 According to the NCCN panel, PCA3 is recommended only for men with a previous negative biopsy, in order to avoid an unnecessary repeat biopsy. The panel also states against the use of this marker in the initial biopsy setting. 25
Mi-prostate score (Michigan Medicine Laboratories, Ann Arbor, USA)
The Mi-Prostate Score (MiPS) is a test provided by Michigan Medicine Laboratories (MLabs), from the University of Michigan, that combines tPSA with urinary PCA3 and TMPRSS2:ERG (T2:ERG) expression. 57 T2:ERG derives from the fusion of the transmembrane protease serine 2 gene (TMPRSS2) with the V-Ets erythroblastosis virus E26 oncogene homolog (ERG). 58 This gene fusion is seen in about 50% of PCa in western countries and only in just 11% of PCa in China, due to its wide variation among ethnic groups. 59 Tomlins et al. performed a clinical validation study of MiPS test, using a validation cohort of 1244 men presenting for diagnostic biopsy. The authors used logistic regression models which included T2:ERG, tPSA and PCA3, and verified that the AUC was greater when incorporating all the three markers, rather than only tPSA, tPSA plus T2:ERG or tPSA plus PCA3 score. These risk models, designated MiPS, were thereby validated for predicting PCa and high-grade PCa on biopsy. 60 In a similar study carried out on 443 men scheduled for prostate biopsy, Leyten et al. 58 concluded that T2:ERG added independent additional predictive value to PCA3 for PCa, and that the use of this panel of markers could allow a reduction in the number of prostate biopsies. However, there is no significant association between T2:ERG expression and biochemical recurrence of PCa or lethal disease, although it is associated with stage at diagnosis, in men treated with radical prostatectomy. 61 According to the NCCN panel, MiPS is still under investigation at the moment. 25
ExoDx™ prostate test (Exosome Diagnostics, Inc., Waltham, US)
The ExoDx™ Prostate Test, also referred as ExoDx™ Prostate Intelliscore (EPI), is based on the detection of exosomal RNA from three genes – PCA3, T2:ERG and SPDEF – in a non-DRE, first catch urine sample. Compared to other urinary PCa tumour marker tests, this test does not require a DRE to be made before the urine collection. 15 Exosomes are extracellular vesicles naturally released from cells, that are delimited by a lipid bilayer, containing tumour-derived molecules, such as RNA, and they reflect the intracellular status of their cells of origin. 62 This test is intended to predict high-grade PCa at initial biopsy. 63 It uses an algorithm that combines the weighted expression of this three-gene signature, giving a risk score from 0 to 100. A score higher than 15.6 is related to an increased probability of high-grade PCa, namely, Grade Group (GG) 2 or greater. 15 McKiernan et al. performed two clinical validation studies of EPI, both in men aged ≥50 years. In the first study 1563 men with tPSA between 2–20 ng/mL were enrolled, while in the second one 500 participants with tPSA between 2–10 ng/mL were evaluated.63,64 In the most recent study, with a cutoff of 15.6, 26% of unnecessary prostate biopsies would have been spared, with a NPV of 89%, missing only 7% of PCa GG ≥2. 64 According to the NCCN panel, EPI can be an option for men considering an initial or repeat biopsy. 25
The Stockholm3-test (Karolinska Institutet, Stockholm, Sweden)
The Stockholm3-test (SM3) is a blood-based test, currently available for clinical use in Sweden, Norway, Denmark and Finland. 65 It consists of a prediction model aimed to detect high-grade PCa (GS ≥7). The predictors included in the model encompass clinical variables (age, first-degree family history of PCa and a previous biopsy), blood biomarkers (tPSA, fPSA, hK2, MIC1 and MSMB) and genetic markers (a genetic score based on 254 single-nucleotide polymorphisms [SNPs] and an explicit variable for the HOXB13 SNP). 66 In 2015 the results from a large prospective, population-based study carried out in Sweden – the Stockholm3 study (STHLM3) – were published. It was made using a large random subsample (5426 men submitted to a prostate biopsy and with a tPSA≥1 ng/mL) of the male population aged 50–69 years in Stockholm. The prediction S3M model was compared to tPSA ≥3 ng/mL as a screening tool for PCa with GS ≥7. For the same sensitivity as the tPSA test (cutoff of ≥3 ng/mL) to detect high-risk PCa, use of the S3M could reduce the number of biopsies by 32% (95% CI 24–39) and avoid 44% (95% CI 35–54) of benign biopsies. Moreover, one of the components of the model – the genetic score – only needs to be measured once in a man’s lifetime. Overall, the results showed that SM3 can be applied in men aged 50–69 years with a tPSA ≥1 ng/mL, lowering the number of prostate biopsies and the detection of clinically insignificant disease, but keeping the sensitivity to detect clinically significant PCa, that is, reducing overdiagnosis. 67 The AUC for S3M (AUC = 0.76) is higher when compared to other diagnostic tools, namely tPSA (AUC = 0.58), %fPSA (AUC = 0.64), PSAD (AUC = 0.69), the clinical model included in SM3 alone, which encompasses tPSA, age, DRE and prostate volume (AUC = 0.71), and age plus 4Kscore® (AUC = 0.70). It has been proposed that S3M could be used as a reflex test in PCa screening, for men with tPSA ≥3 ng/mL. 66
One of the limitations of SM3 is that, since the STHLM3 study was done in Stockholm, most of participants were of northern European descent, with the same ethnic origin. 67 However, other SM3 validation studies are being carried out in Germany, Holland and in the UK. Studies validating the test on a non-Caucasian population started in 2020. 65
ERSPC risk calculators (Prostate Cancer Research Foundation, Rotterdam, The Netherlands)
The ERSPC risk calculators (RCs) estimate the individual risk of PCa. They were developed based on the Dutch section of the ERSPC trial 68 and are available online. Both RC1 and RC2 are meant for patient use. RC1 uses family history, age and urinary symptoms, while RC2 is based on tPSA level. The remaining RCs are intended to be used by health care professionals. Combined into one decision tree, RC3 and RC4 rely on tPSA level, DRE, prostate volume, previous negative prostate biopsy and mpMRI. They can also include the PHI result. RC5 estimates the probability of having indolent prostate cancer, whereas RC6 calculates the risk of having PCa in the next 4 years, based on age, tPSA, DRE, family history, prostate volume and previous biopsy status. 69 Several external validation studies, in both European and non-European cohorts, assessed RC3/RC4, showing AUC in the range of 0.71–0.88 for PCa prediction.70–73 In two other cohorts, RC3 and RC4 showed AUC for PCa and significant PCa of 0.66–0.77 and 0.85, respectively.74,75 According to the EAU, the RCs may be helpful to determine the potential risk of cancer, thereby reducing the number of unnecessary biopsies. 24
Prognostic Markers
Four commercially available multigene panels have been shown to provide prognostic information on PCa, helping to differentiate between aggressive cancers requiring treatment, from non-aggressive cancers that could be managed with AS. According to the NCCN, their use may be considered during initial risk stratification to provide prognostic information, such as the likelihood of death with conservative management, biochemical progression after radical prostatectomy or external beam therapy, metastasis development after radical prostatectomy or salvage treatment. 76
OncotypeDX® genomic prostate score (Genomic Health, Inc., Redwood City, USA)
This is a quantitative polymerase chain reaction (qPCR) assay that evaluates the expression of 17 genes, in the form of a score, known as the Genomic Prostate Score (GPS). 77 It is performed on biopsy samples and can predict the risk of adverse pathology at radical prostatectomy. Klein et al. 78 reported that GPS could predict high-grade (odds ratio [OR] = 2.3 per 20 GPS units) and high-stage disease (OR = 1.9 per 20 GPS units) at prostatectomy. OncotypeDX® has been reported as able to help the clinician in adapting treatment choices and reclassifying patients by risk. Albala et al. 79 reported that the incorporation of GPS as part of the decision algorithm for patients with low and very low-risk cancer, led to a substantial increase in AS (21% reduction in interventional treatment) and to a reclassification of 4.3% very low risk and of 35.7% low-risk patients into intermediate-risk. Dall’Era et al. 80 verified an increase of 24% in the use of AS.
Prolaris® (Myriad Genetics, Inc., Salt Lake City, USA)
Prolaris® is a qPCR assay that measures the expression of 46 genes to provide a cell cycle progression (CCP) score, ranging from −1.3 to 4.7. 81 The assay can be used after a positive biopsy or after radical prostatectomy and it is meant to predict disease progression and mortality. Cuzick et al. 82 reported that, after prostatectomy, a high CCP score was predictive of biochemical recurrence, with a hazard ratio of 1.89 for an increase of 1 unit in CCP. Prolaris® has been reported to have good prognostic value, can predict time of death and also can guide changes in treatment directions, mostly towards a reduction in interventional treatment.83,84 Cuzick et al. 85 also reported, in 14% of patients with a low Cancer of Prostate Risk Assessment (CAPRA) score, a risk reclassification to higher risk groups; in 44% of patients with an intermediate CAPRA score, they also found a reclassification to lower risk groups. Moreover, Prolaris® has shown utility in predicting the outcome after primary radiation therapy. 86
Decipher® (Decipher Biosciences, Inc., San Diego, USA)
The Decipher® assay, which is a genomic classifier (GC), evaluates the expression, in tumour tissue, of a panel of 22 genes that are involved in aggressive PCa. Ranging from 0 to 1, the GC score aims to predict the systemic progression of the disease after definitive treatment. It predicts the probability of metastasis, allows risk stratification and assists in decision making regarding adjuvant therapy.87,88 Erho et al. compared this GC to pathology staging parameters, GS, PSA and several clinical models on univariate analysis, and demonstrated that the GC had a higher accuracy for the prediction of distant metastasis, while multivariate analyses consistently demonstrated the GC alone as being statistically significant. They reported that, for the detection of clinical metastasis within 5 years of radical prostatectomy, Decipher® had the highest AUC of 0.75 compared to an AUC of 0.74 for an integrated genomic-clinical classifier and 0.69 for a clinical-only classifier. 88 Ross et al. showed that the GC, after radical prostatectomy, correlated with increased cumulative incidence of biochemical recurrence, metastasis and PCa specific mortality (P < 0.01). After 10 years, cumulative incidence of metastasis was 12% for patients with low and 47% for patients with high GC scores, respectively. 89 In a study with 146 patients with PCa, Michalopoulos et al. 90 reported that, after review of the GC scores, over 60% of high-risk patients were re-classified as low risk and adjuvant treatment recommendations were modified for 30.8% of patients. Den et al. 91 also observed that, from a total of 1586 patients, GC was able to reclassify 52%, 76% and 40% of patients belonging to low, intermediate and high-risk CAPRA Post-Surgical score groups, respectively.
According to the NCCN, Decipher® may be considered as part of counselling for risk stratification in patients with PSA persistence or PCa recurrence after radical prostatectomy. 76
ProMark® (Metamark Genetics, Inc., Waltham, USA)
ProMark® is a multiplex immunofluorescence-based assay that evaluates the expression of eight tissue proteins in a biopsy sample, giving a risk score ranging from 0 to 1, that predicts disease aggressiveness and patient outcome. 92 Blume-Jensen et al. evaluated its performance to predict favourable or nonfavourable pathology at prostatectomy, independently and relative to the NCCN and D’Amico classification systems. The primary goal was to define a model able to distinguish patients who were candidates for AS from those more likely to require prostatectomy. The study reported that, with increasing ProMark® scores, the frequency of nonfavourable pathology increases. Predictive values for favourable pathology in very low-risk and low-risk NCCN and low-risk D'Amico groups were 95%, 81.5% and 87.2%, respectively, at a risk score ≤0.33. These predictive values were higher than those given by these risk classification groups. For nonfavourable pathology, at a risk score of >0.8, the predictive value across all risk groups was 76.9%. 93
Markers with a potential role in metastatic castration-resistant prostate cancer (mCRPC): Liquid biopsy
Circulating tumour cells
Evaluation of circulating tumour cells (CTC) has been shown to have prognostic value in mCRPC. 94 In a study with 231 patients with mCRPC, de Bono et al. verified that patients with higher pretreatment CTC counts (≥5 CTC/7.5 mL) had a shorter overall survival (OS) and posttreatment CTC counts predicted OS better than PSA decrement. 95 Moreover, CTC molecular characterisation could provide a liquid biopsy approach helping to predict the cancer response to androgen receptor (AR) signalling inhibitors used in mCRPC – such as abiraterone and enzalutamide – and to standard chemotherapy. 96
Cell-free DNA
Cell-free DNA (cfDNA) consists of fragmented segments of DNA from tumour cells, as well as from nonmalignant tumour microenvironment cells and non-tumour cells, which are shed into body fluids and the circulatory system. 97 In patients with PCa, cfDNA in blood is detectable in higher levels when compared to controls. 98 In addition, over 50% of blood samples and more than 70% of urine samples from PCa patients show changes on cfDNA that may be used as PCa biomarkers. 99 The inhibition of AR signalling with abiraterone or enzalutamide is an important therapeutic tool for mCRPC. However, some patients will have a poor response, since they develop AR mutations in metastasis or a gain in AR copy number, mostly after hormone therapy.100,101 It has been shown that cfDNA analysis can identify over 90% of somatic mutations present in matched metastatic tissue, 102 as well as detecting the gain in AR copy number, allowing to identify mCRPC patients who will not benefit from AR targeting therapy.100,102,103 Since the collection of circulating cfDNA is minimally invasive and allows to obtain a comprehensive molecular analysis, cfDNA analysis presents as an opportunity to identify novel gene alterations associated with tumour progression. 102
Future outlook
Epigenetic changes, such as histone modifications and variants, DNA methylation, as well as some microRNAs (miRNAs), are common in PCa, and play a role in the pathogenesis of the disease and may be markers of value in diagnosis, prognosis and response to therapy.104–106
Several studies have shown the existence of nucleosomes – basic chromatin units composed of DNA and two copies of the histones H2A, H2B, H3 and H4 – in the serum and plasma of cancer patients. There are, however, several benign situations, such as inflammatory and infectious conditions, autoimmune diseases, ischaemia and trauma, which may lead to increases in circulating nucleosomes. This fact can limit their use in cancer diagnosis. However, several post-translational histone modifications, such as methylation, acetylation and phosphorylation, have been reported in several cancers. 107 In PCa decreased H3K27me3 values were detected in metastatic disease, 107 whereas increased H3K4me values were observed in organ confined PCa. 108 Decreased H3K9me3 and H3K27me3 values were observed after 5-azacitidine therapy (which induces a neuroendocrine differentiation of PCa), 109 In addition, increased H3K4me3,110,111 H3K36me3 112 and H4K20me3 were also observed. 113 Acetylation of lysine 16 from histone 4 (H4K16ac) contributes to androgen-dependent gene activation in PCa cells, playing a role in the pathophysiology of this cancer. 114 It was found that PCa patients with high H3K9ac levels had a longer survival than those expressing lower levels. 115 There is also an increase in H2AX, which is a marker of DNA damage and repair. 116 Histone variant H2A.Z has also been implicated in the development of PCa, with increased expression especially in anti-androgen therapy-resistant PCa, 106 being associated with disease progression. 117 Ellinger et al. studied H3K4 methylation in patients with advanced PCa, and found that this epigenetic modification was a significant predictor of PSA recurrence, following radical prostatectomy. Furthermore, they also found that H3K4me1, H3K4me2 and H3K4me3 levels were significantly increased in hormone-refractory PCa. 118 One of the most studied epigenetic enzymes in PCa is the histone methyltransferase EZH2, responsible for H3K27 trimethylation. Its overexpression is particularly found in mCRPC. 119
Among the DNA binding proteins, the high mobility group box1 (HMGB1) is highly expressed in PCa cells. The release of HMGB1 into the circulation appears to correlate with disease severity and may be a target for therapy.110,120 This nuclear DNA binding protein appears to play a role in the development and progression of PCa. 121 It has been implicated as an androgen receptor co-activator in prostate cells, 122 and it is also an inflammatory mediator affecting different stages of carcinogenesis, such as cell proliferation, angiogenesis, migration and invasion. Some studies point to it as a potential target for PCa therapy. 123 The receptor for advanced glycation end products (RAGE) is a multi-ligand receptor that binds to several molecules, including HMGB1. Activation of RAGE has been implicated in the inflammatory response and in the development of different types of cancer, including PCa. 124 On the other hand side, soluble RAGE, which is shed from the surface of immune cells may act as a decoy receptor for advance glycation end products, such as HMGB1, and prevent further stimulation of immune cells. 125
miRNAs have also been reported as potential biomarkers for PCa risk and disease progression. Those are small non-coding RNAs, about 22-nucleotide long, that regulate gene expression and participate in the regulation of several cellular processes. The most frequently miRNAs studied until now in PCa are miR-1, −21, −106b, −141, −145, −205, −221 and −375. 126
Genetic testing could be also important in PCa since it is well known that family history raises the risk of the disease. According to the NCCN panel, germline genetic testing should be made in PCa patients with any of the following: a positive family history, high-risk PCa, Ashkenazi Jewish ancestry or intraductal histology. Genetic testing should include MLH1, MSH2, MSH6 and PMS2 and the homologous recombination genes BRCA1, BRCA2, ATM, CHEK2 and PALB2. 76
Proinflammatory cytokines could also play a role as biomarkers in PCa, since there have been found higher levels of these markers in PCa patients, highlighting that inflammation can play a part in PCa development.127–129
Also immune-related biomarkers have been identified as potential PCa markers and therapeutic targets. These include the programmed cell death receptor-1 (PD-1) and their ligands (PD-L1 and PD-L2). Cancer cells develop a resistance to proapoptotic signals, through a blockage in the PD-1, PD-L1 and PD-L2 pathway, disrupting signals that allow cancer immune evasion, thus creating an immunosuppressive tumour microenvironment. 130 Increased expression of PD-1, PD-L1 and PD-L2 was found in PCa tissue of invasive prostate tumours in mice. 131 On the other hand, a large prospective study on gene expression data from human prostatectomy samples, showed that PD-L2 was expressed at higher levels than PD-L1. It was associated with a worse prognosis and was a predictor of the response to postoperative radiation therapy. Data from this study suggested that PD-L2 could be a potential therapeutic target in PCa, in combination with radiotherapy. 132 These data, altogether, point that these programmed cell death markers can become very new biomarkers in PCa.
Conclusion
Tumour markers/scores already in use and promising new biomarkers in prostate cancer.
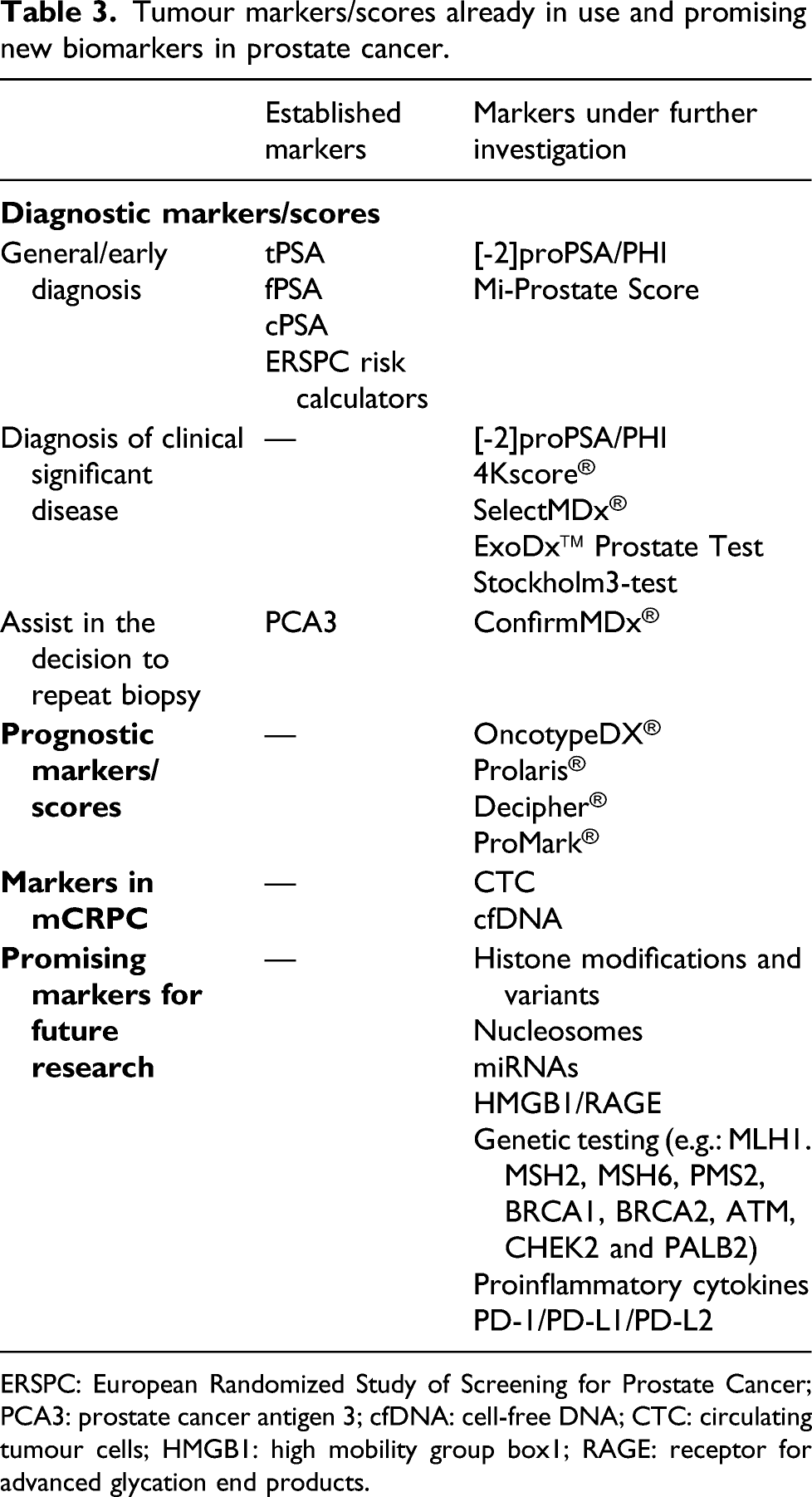

ERSPC: European Randomized Study of Screening for Prostate Cancer; PCA3: prostate cancer antigen 3; cfDNA: cell-free DNA; CTC: circulating tumour cells; HMGB1: high mobility group box1; RAGE: receptor for advanced glycation end products.
Footnotes
Acknowledgements
This article was prepared at the invitation of the Clinical Sciences Reviews Committee of the Association for Clinical Biochemistry and Laboratory Medicine.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Guarantor
The guarantor is the first/correspondent author (Manuel M. Garrido), who is willing to take full responsibility for the article, including for the accuracy and appropriateness of the reference list.
Contributorship
MMG, RMB and JCM researched the literature and participated in writing the article. MMG wrote most of the first draft of the manuscript. SH and JTG reviewed the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
