Abstract
Background
MicroRNA has received considerable attention in the clinical context, and attempts are being made to use microRNA in clinical diagnosis. However, adequate quantities of microRNA required for analysis are challenging to isolate. We tested the effect of various reagents in improving microRNA extraction and compared their efficacy to that of a commercially available extraction kit (HighPure miRNA isolation kit, Roche).
Methods
We used the synthetic oligonucleotide miR-21 and formalin-fixed, paraffin-embedded (FFPE) tissue sections from colon cancer samples (n = 10). We tested increasing volumes (100–600 μL) of 1,4-dioxane, 2-butanol, 2-propanol, acetonitrile, polyethylene glycol (PEG) 600, PEG 1000, PEG 1540, PEG 2000, tetraethylene glycol dimethyl ether (TDE), and tetrahydrofuran, instead of the binding enhancer solution provided in the kit. MiR-21 analysis was performed via stem-loop RT-qPCR using Universal ProbeLibrary probe (Roche).
Results
The optimum amount of each enhancement solution was 200–500 μL. We obtained ΔCp values of optimum additional volume for each solution from 1.04 to 2.50 and compared these with those obtained using the commercially available kit. PEG 1540 and 2000 produced superior reactivity with minimal addition. For FFPE tissue samples, addition of the enhancement solutions PEG 1540 and 2000 resulted in mean crossing point values of 18.15 ± 2.26 and 17.73 ± 3.26, respectively. We obtained a crossing point value of 20.56 ± 4.26 (mean ± SD) using the commercially available kit.
Conclusions
The tested enhancer reagents, which are relatively readily available and easy to use, can improve microRNA extraction efficacy of a commercially available kit.
Keywords
Introduction
MicroRNAs (miRNA) are small (approximately 22 nucleotides) non-coding RNAs regulating numerous essential cell functions.1,2 miRNAs play a critical role in several biological processes, such as cell proliferation and maturation, apoptosis, regulation of chronic inflammation and cancer progression.3,4
miRNA analysis has a wide array of clinical applications, such as in the diagnosis of various diseases including cancer,5,6 in determining patient response to treatment7–9 and in prognosis.10,11 Moreover, therapeutic applications using miRNA are gaining significance,9,12,13 and analysis of miRNA from exosomes14,15 is an attractive option as part of the liquid biopsy.16–18
There are several methods for miRNA extraction using manual or automated extraction protocols. 19 Here we attempted to perform miRNA analysis using clinical samples in our laboratory. However, we encountered challenges because we were unable to obtain the crossing point (Cp) value required for the analysis. Reliable quantification of mRNA and miRNA concentrations for diagnostic and prognostic purposes requires optimal RNA quality and quantity. 20 The principle of this kit is to use an additional reagent solution for enhanced and greater low-size RNA extraction. We examined the ability of various reagents to improve miRNA extraction efficiency and compared their performance to that of a commercially available extraction kit.
Methods
We used the conventional HighPure miRNA isolation kit (Roche Molecular Systems, Pleasanton, CA, USA) for this study. First, we tested the candidate-binding enhancer solutions instead of the binding enhancer (BE) solution (Roche Molecular Systems) included in this kit using synthetic miR-21.
We tested 2-propanol, ethanol, polyethylene glycol (PEG) 600, 1000, 1540, 2000, tetraethylene glycol dimethyl ether (TDE), tetrahydrofuran (THF), 1,4-dioxane, 2-butanol and 2-propanol as binding enhancer solutions instead of the Binding Enhancer (Roche Molecular Systems). Usually, extraction kits use different reagents to enhance the miRNA extraction efficacy, and we used the above solutions instead of the BE solution (Roche Molecular Systems). PEG was used after heating at 60°C. We also used additional volumes ranging from 100 to 600 μL of other binding enhancement solutions.
Subsequently, 20 nM of synthetic miR-21 (Hokkaido System Science, Sapporo, Japan) and 100 ng of K562 total RNA, made up to a total of 150 μL with water, were added to 312 μL to include the binding buffer (Roche Molecular Systems). Specifically, we added 100–600 μL of each candidate-binding enhancer solution or 200 μL of the BE solution (Roche Molecular Systems) and applied a spin column after mixing them well. We prepared five samples from each mix.
We used the HighPure miRNA Isolation Kit (Roche Molecular Systems) following the manufacturer’s instructions using 100 μL of elution buffer for a one-column protocol.
We compared Cp values for miR-21 expression via stem-loop reverse transcription-quantitative PCR (RT-qPCR) using Universal ProbeLibrary (UPL) probe (Roche Molecular Systems).
MiRNA were transcribed into cDNA via an miRNA-specific RT reaction using the designed miRNA specific stem-loop RT primer and Transcriptor First Strand cDNA Synthesis Kit (Roche Molecular Systems). The stem-loop RT primer combines 44 nt of stem-loop sequence 5′-GTCAGAGGAGGTGCAGGGTCCGAGGTATTCGCACCTCCTCTGACTCAACA-3′.
RT reactions were prepared per the manufacturer’s instructions with modifications. We used 2.5 μL of an elute solution containing 50 nM of a stem-loop RT primer, 10 U of transcriptor reverse transcriptase (Roche Molecular Systems), 20 U of protector RNA inhibitor (Roche Molecular Systems) and 1 mM of deoxynucleotide mix. The RT reaction was set at 16°C for 30 min, with 60 cycles of 30°C for 30 s, 42°C for 30 s, 50°C for 1 s, 85°C for 5 min and a hold at 4°C.
We also used the miR-21 Cp values obtained via real-time quantitative PCR (qPCR), which was performed using a LightCycler 96 instrument with LightCycler 1.1 software (Roche Molecular Systems). The following primers were used for amplification: sense 5′-GCCTGCTAGCTTATCAGACTGATG-3′ and antisense 5′-GTGCAGGGTCCGAGGT-3′. We used 2.5 μL of a cDNA solution at five-fold dilution in TE buffer containing 0.2 μM of each primer and 0.2 μM of the UPL probe no. 82 (Roche Molecular Systems). PCR conditions included initial denaturation at 95°C for 10 min, followed by 40 cycles of denaturation (95°C for 10 s, 4.4°C/s) and annealing (60°C for 30 s, 2.2°C/s).
We set an additional optimal volume for the miRNA extraction enhancement solution, which showed a significant difference compared with the Cp value at the additional volume immediately before. In addition, we set an additional effective volume to be added using one-way analysis of variance. We determined the minimum volume to be added to obtain a significant difference at P < 0.01 and the most suitable volume to be added, which was evaluated using Tukey’s method.
Furthermore, we performed miRNA extraction from formalin-fixed paraffin-embedded (FFPE) tissue samples using the most suitable addition volume of the binding enhancement solutions. FFPE tissue samples were obtained via surgical resection from 10 patients with colon cancer who were being treated at the Daiyukai General Hospital. The samples were irreversibly anonymized such that no clinical or patient-related information was available for the study. Ethics approval for the research was obtained from Daiyukai Health System. Tissue samples were serially sectioned (5 µm thick) and the sections were placed onto glass slides. Tissue was scraped from each glass slide with a scalpel. We performed the procedure in accordance with the pretreatment method described in the manufacturer’s instructions. Briefly, we added 800 μL of xylene to one section in a 1.5 mL reaction tube, followed by incubation for 5 min and mixing. Next, we added 400 μL of ethanol, centrifuged for 2 min and discarded the supernatant. Finally, we added 1000 μL of ethanol, centrifuged and discarded the supernatant. The tissue was then dried.
We used the HighPure miRNA Isolation Kit (Roche Molecular Systems, Pleasanton, CA, USA), and the manufacturer’s instructions were followed using 100 μL of elution buffer for a one-column protocol. We analysed the solutions for miR-21 expression level using RT-qPCR. We performed FFPE using the most suitable FFPE additional volume of synthetic miR-21. Additionally, we compared the Cp value to the Roche kit using the candidate-binding enhancer solutions.
Results
The Cp values obtained from RT-qPCR using synthetic miR-21 are shown in Table 1. The changes in Cp values were different in each solution, but all solutions showed an increase in Cp value with additional volume-dependent increases due to the enhancement solutions. All solutions showed superior Cp value compared with the BE solution included in the HighPure miRNA isolation Kit (Roche Molecular Systems). PEG 1000 showed a Cp of 0.032 with 300 µL of additional volume, with no significance at P = 0.01. PEG, particularly at the molecular weight range of 1000–2000, showed a lesser volume requirement compared with the other solutions. These results suggested that PEG 1540 and PEG 2000 exhibit superior effects but small additional volume requirements.
Cp values for the extraction of synthetic miR-21.
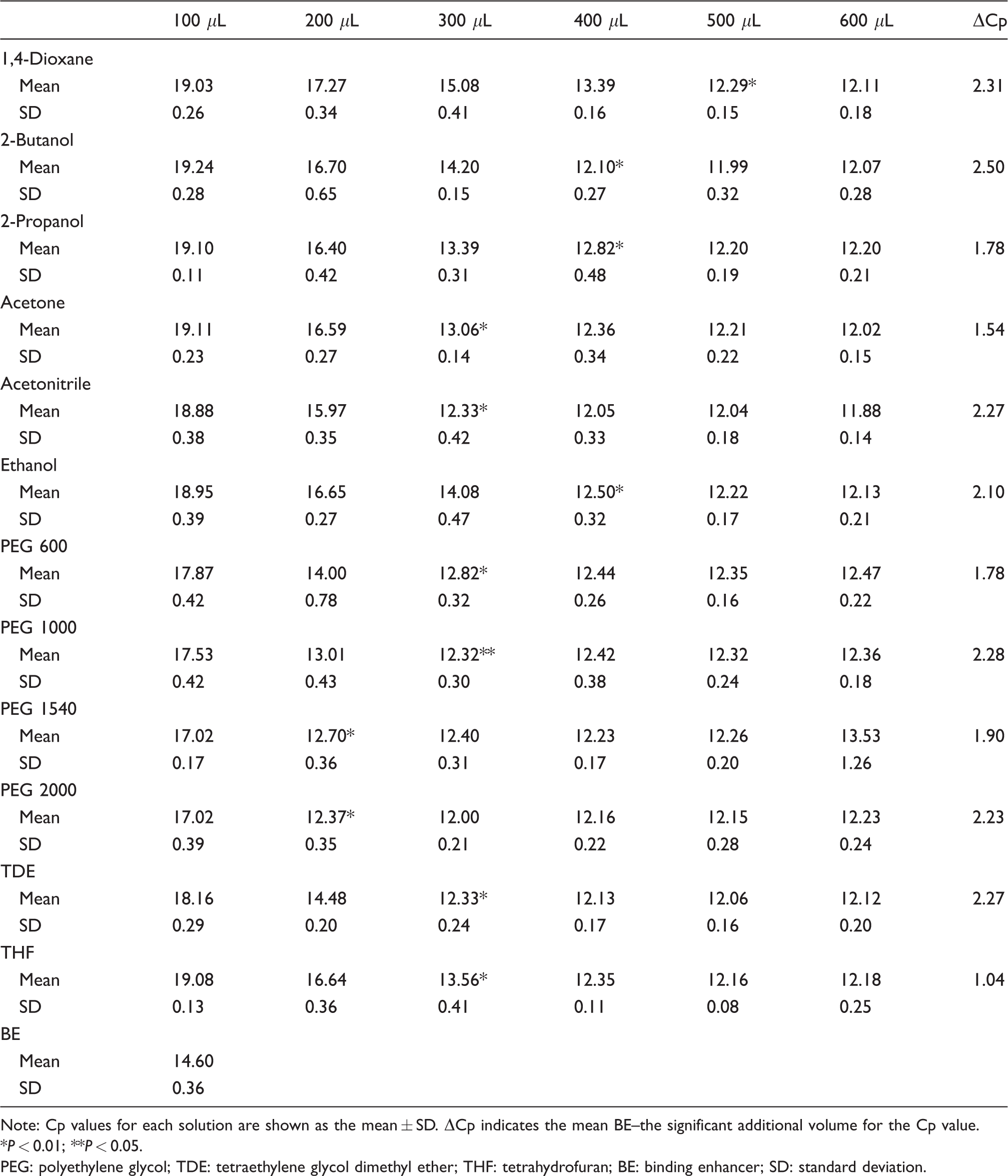
Note: Cp values for each solution are shown as the mean ± SD. ΔCp indicates the mean BE–the significant additional volume for the Cp value.
*P < 0.01; **P < 0.05.
PEG: polyethylene glycol; TDE: tetraethylene glycol dimethyl ether; THF: tetrahydrofuran; BE: binding enhancer; SD: standard deviation.
Next, we performed miRNA extraction from FFPE tissue samples. We used a suitable volume of miRNA enhancement solutions as shown in Table 2. The binding enhancement solutions increased the extraction efficiency from FFPE tissue samples compared with the BE solution (Roche Molecular Systems). The results showed a similar tendency for synthetic miR-21. Particularly, PEG 1540 and PEG 2000 showed a superior tendency, although in small amounts, compared with the other solutions.
Cp values for FFPE tissue samples.
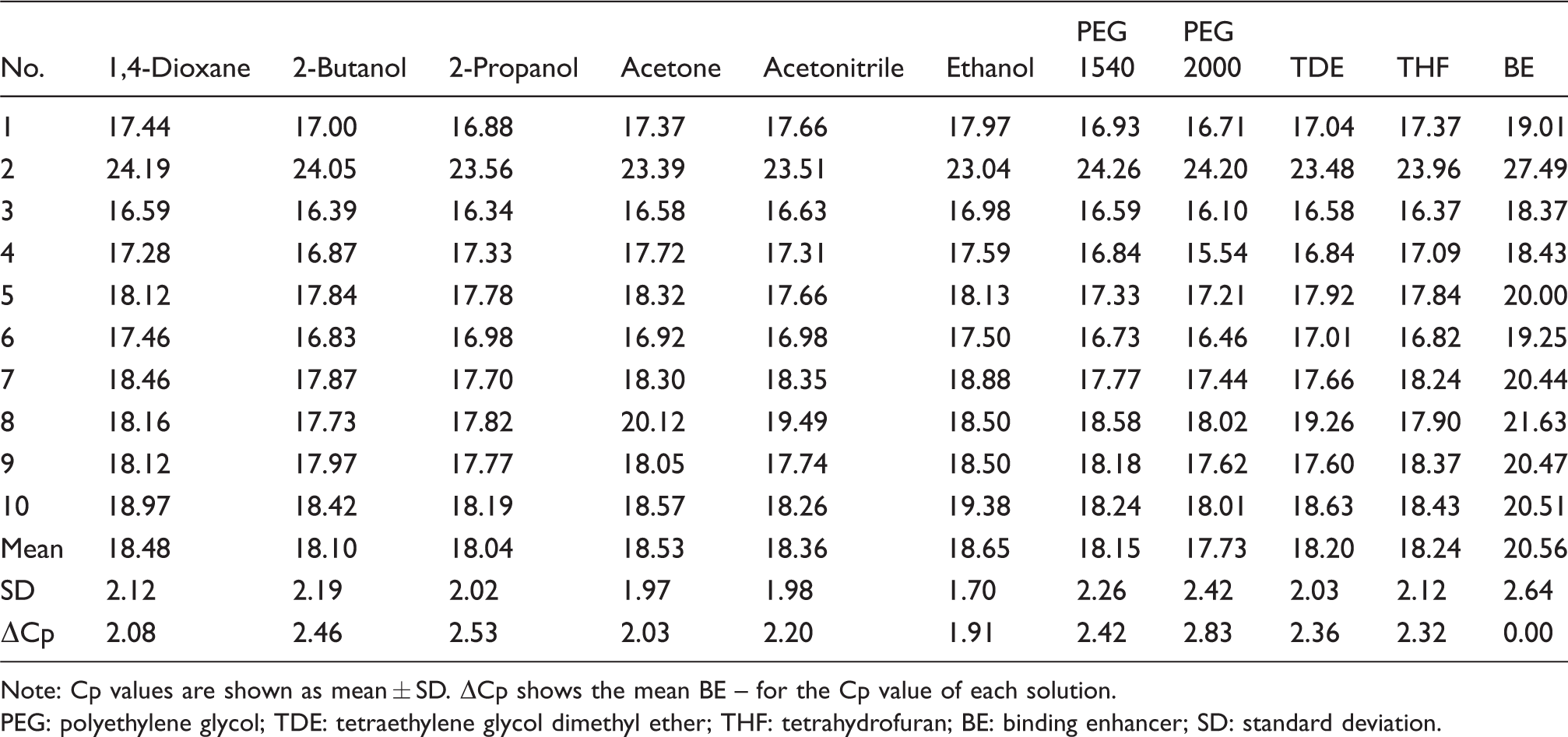
Note: Cp values are shown as mean ± SD. ΔCp shows the mean BE – for the Cp value of each solution.
PEG: polyethylene glycol; TDE: tetraethylene glycol dimethyl ether; THF: tetrahydrofuran; BE: binding enhancer; SD: standard deviation.
Discussion
In clinical laboratories, for miRNA analysis, it is important that the quantity necessary for analysis be determined, in addition to removing PCR inhibitors from clinical samples.
In this study, we used FFPE tissue and peritoneal dialysis fluid samples in our laboratory. However, there were challenges in obtaining sufficient miRNA quantities for analysis. In addition, cross-linking with proteins, enzyme degradation and chemical degradation occurring during fixation reduced the yield, quality and integrity of RNA.21
We used the conventional HighPure miRNA isolation kit (Roche Molecular Systems) and examined different reagents that are effective in improving miRNA extraction efficiency instead of the BE solution (Roche Molecular Systems). We obtained a superior effect using each solution than that obtained using the BE solution (Roche Molecular Systems). Moreover, we observed the same tendency between the synthetic substance and FFPE tissue samples taken from colon cancer patients using PEG 1540 and PEG 2000 at a suitable additional volume.
Using PEG 1540 and PEG 2000 has a few advantages. In general, we considered the low toxicity of PEG as an advantage, particularly in the context of user and workplace safety. Particularly, PEG 1540 and PEG 2000 showed effects even at small amounts compared with other solutions used with the HighPure miRNA extraction kit (Roche Molecular Systems).
Moreover, minimizing the additional volume of enhancement solutions reduced the number of steps and contamination with samples other than nucleic acids. Furthermore, conjugates used as new biomarkers accelerated the use of miRNA, and plasma and urine samples have attracted attention due to their minimal invasiveness. In general, we believe that minimizing the volume of enhancement solutions is useful because, typically, large volumes of such liquid samples are required. When using such liquid samples, minimizing the volume of the binding buffer and enhancement solution is an important factor in nucleic acid binding to the column. However, PEG 1540 and PEG 2000 have high melting points, and these solutions must be warmed to dissolve them before use. Currently, we are working on improving this feature.
Consider the following example: In some situations, miRNA extraction from clinical samples is difficult because of various factors. In these situations, miRNA yield can be improved by using an enhancement solution, such as PEG 1540 or PEG 2000, instead of the BE solution (Roche Molecular Systems).
Moreover, we observed the same tendency of miR-21 using a synthetic product and FFPE of colon cancer in miR-200c (data not shown). We believe that other miRNA species must be considered.
Currently, we used the HighPure miRNA extraction kit (Roche Molecular Systems), which is a conventional miRNA extraction kit. For nucleic acid extraction, we used the Boom method based on the chaotropic effect. 22 We expected our miRNA extraction enhancement solutions to have a dewatering effect and surmised that these solutions might enhance binding to the extraction kit spin column. Although it may be necessary to optimize for additional volume, we believe that we obtained the same effect because other miRNA extraction kits work on the same basic principle.
In this study, we also investigated whether the effectiveness of miRNA extraction using reagents that are readily available in most clinical laboratories can be increased. The findings of this study can help improve miRNA extraction efficiency using available reagents. In situations in which we could not obtain a sufficient miRNA yield for some reason, we replaced the BE solution (Roche Molecular Systems) of the kit with these reagents. We hypothesized that the reagents we tested can improve miRNA extraction efficacy.
The results of this study suggested that it is possible to improve the miRNA extraction efficiency of a commercially available kit that is not specialized for miRNA extraction or an existing total RNA extraction kit.
Conclusion
We identified several solutions that can improve miRNA extraction efficacy of a conventional commercially available miRNA extraction kit.
Particularly, PEG 1540 and PEG 2000 are relatively readily available and easy to use, and can improve miRNA extraction efficacy of a commercially available extraction kit.
Supplemental Material
Supplemental Material1 - Supplemental material for Evaluation of the efficacy of various reagents in improving microRNA extraction
Supplemental material, Supplemental Material1 for Evaluation of the efficacy of various reagents in improving microRNA extraction by Arizumi Kikuchi, Azumi Naruse, Takahiro Sawamura and Kenichi Nonaka in Annals of Clinical Biochemistry
Supplemental Material
Supplemental Material2 - Supplemental material for Evaluation of the efficacy of various reagents in improving microRNA extraction
Supplemental material, Supplemental Material2 for Evaluation of the efficacy of various reagents in improving microRNA extraction by Arizumi Kikuchi, Azumi Naruse, Takahiro Sawamura and Kenichi Nonaka in Annals of Clinical Biochemistry
Footnotes
Acknowledgements
We thank Dr. Noriyuki Okita (Roche Molecular Systems) and Mr. Kazutoshi Sasaki (Roche Molecular Systems; present affiliation, NIPPON Genetics Co., Ltd) for their valuable advice and support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
This study protocol was approved by ethics committee of Daiyukai Health System (Permission number: 2016–023).
Guarantor
AK.
Contributorship
AK, AN, TS and KN designed the study and collected samples. KN was involved in gaining ethical approval, patient recruitment and obtaining informed consent. AK, AN and TS carried the experiments. AK and AN wrote the first draft of the manuscript. All authors analysed, reviewed, edited the manuscript, and approved the final version of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
