Abstract
Background
The distributions of within-subject biological variation are usually described as coefficients of variation, as are analytical performance specifications for bias, imprecision and other characteristics. Estimation of specifications required for reference change values is traditionally done using relationship between the batch-related changes during routine performance, described as Δbias, and the coefficients of variation for analytical imprecision (CVA): the original theory is based on standard deviations or coefficients of variation calculated as if distributions were Gaussian.
Methods
The distribution of between-subject biological variation can generally be described as log-Gaussian. Moreover, recent analyses of within-subject biological variation suggest that many measurands have log-Gaussian distributions. In consequence, we generated a model for the estimation of analytical performance specifications for reference change value, with combination of Δbias and CVA based on log-Gaussian distributions of CVI as natural logarithms. The model was tested using plasma prolactin and glucose as examples.
Results
Analytical performance specifications for reference change value generated using the new model based on log-Gaussian distributions were practically identical with the traditional model based on Gaussian distributions.
Conclusion
The traditional and simple to apply model used to generate analytical performance specifications for reference change value, based on the use of coefficients of variation and assuming Gaussian distributions for both CVI and CVA, is generally useful.
Keywords
Introduction
Numerical estimates of the components of biological variation in healthy humans are mainly described as between-subject and within-subject biological variations and are documented either as standard deviations, sI and sG, or coefficients of variation, CVI and CVG, respectively, as described previously. 1 The general assumption is that CVI is random when the subject is in steady-state and, for calculation of the pooled value usually detailed, homogeneity of the variances of the individuals studied is needed. Estimation of the components of biological variation is best performed by use of nested analysis of variance, ANOVA, from which the estimates are obtained as standard deviations. Most of the subsequent applications of such data are only really correct when standard deviations (s) are applied, but coefficients of variation (CV) are often used. The resulting values for applications become therefore, inexact, especially for large CV, but are generally used instead of s because these are simple to handle and the formulae become more understandable. Moreover, the use of CV makes it easy to compare the effects of variation between the components of analytical variation.
In one of the first publications on generation and application of numerical data on the components of biological variation, Cotlove et al. 2 used s in their primary calculations. However, the data were subsequently applied as CV and the analytical imprecision, CVA, in their hallmark proposal for maximum acceptable CVA was defined as CVA ≤ 0.5 × CVI. If this specification was attained, the total combined, CVI and CVA, CVA+I = (CVI2 + CVA2)½, would not exceed CVI by more than 12%.
Harris and Yasaka
3
introduced the concept of the reference change value (RCV) as a means to assess whether significant changes had occurred between serial results during monitoring of patients. The theory was elaborated using s with the formula RCV = z × 2½ × sA+I but was later generalized using CV but without rigorous mathematical support for this approach. In general, the z score selected was 1.96, appropriate for the probability,
Larsen et al.
4
and, later, Petersen et al.
5
created limits for the acceptable analytical performance (analytical performance specifications) needed for the satisfactory use of RCV using both s and CV interchangeably. The assay imprecision (CVA) and bias between the assays at the time of the two measurements (Δbias) being assessed for a significant change can both affect the RCV. With larger CVA, there needs to be smaller Δbias, and
However, the problem of deciding which of s or CV was most correct and how to apply both variables was still not solved. However, in a discussion on whether log-Gaussian distributions based on natural logarithms (ln-Gaussian) were more correct for the distribution of the CVI of brain natriuretic peptides, Fokkema et al. provided detailed documentation of formulae for RCV generated with ln-Gaussian estimates. 8 The use of this ln-Gaussian approach might be useful in elimination of certain problems with the use of traditional CV – and corresponding s – such as the observation of negative concentration values when CV more than 30 % are (incorrectly) documented and the asymmetry of RCV and solving the intermixing of s and CV. Further, many population-based reference intervals, the dispersions of which are mostly dominated by CVG, can be described by log-Gaussian distributions, and therefore it might be biologically sound to consider that CVI may also be logarithmically distributed. In addition, Lund et al. have based their extension of RCV to involve changes in more than two serial samples from an individual in monitoring on logarithmic distributions.9,10
The purpose of this study was to examine approaches to derivation of RCV with ln-distributions of CVI by establishing the needed formulae for the calculations and to create analytical performance specifications to define the allowable combination of Δbias and CVA for use of the ln-transformed CVI.
Materials and methods
Assumptions for generating analytical performance specifications for RCV using ln-Gaussian CVI distributions are that the data are random, the variances are homogeneous and there is no auto-regression or correlation. Relations between the total logarithmic standard deviation, σA+I, and CVA+I of concentration data: CVA+I = (exp{#x003C3;A+I2} − 1)½8 and σA+I = {ln[CVA+I2 + 1]}½,9,10 where σI is the logarithmic within-subject biological variation, σA is the logarithmic analytical imprecision and σA+I = [σI2+σA2]½.
The basic model for RCV according to Fokkema et al. 8 is that the RCV is described as a ratio relative to the first result, x2/x1. This can then be used to define the performance specifications needed to meet the criteria of total variation being increased by not more than 12%.
Ratio between two consecutive samplings with measurements

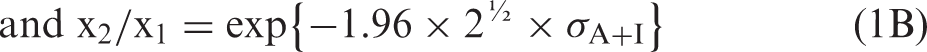
Maximum analytical performance specifications for analytical imprecision are similar to proposal of Cotlove et al.
2
that σA ≤ 0.5 × σI. When σA = 0.0, then the minimum x2/x1, is the simple (x2/x1)MIN = exp{#x0002B;1.96 × 2½ × σI}. For the maximum allowable σAMAX = 0.5 × σI, the maximum x2/x1 is
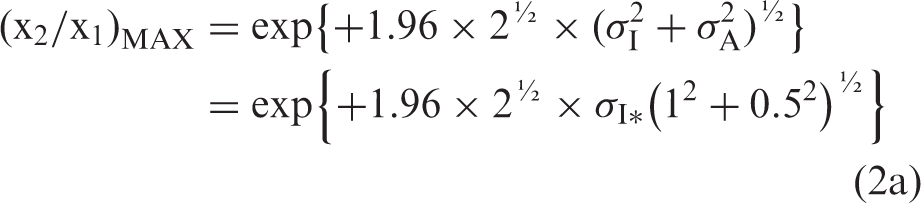
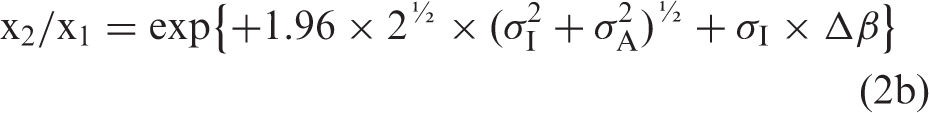
If the formulae using Gaussian distributions4,5,7 are relevant also for ln-Gaussian distributions, then the relation between Δβ and σA is
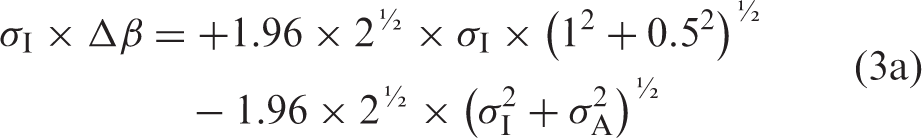
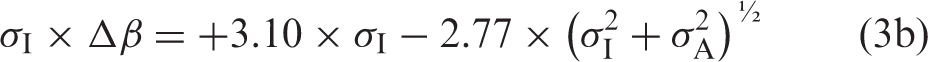
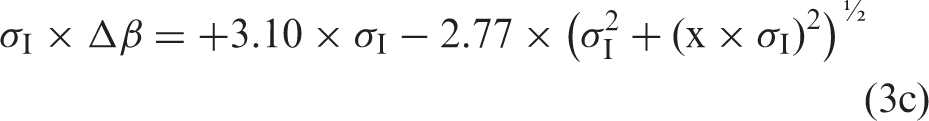
Back transformation from logarithms is performed by a BIASFACTOR
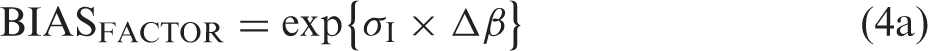
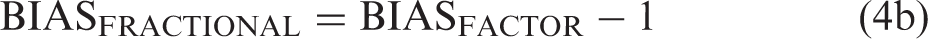
Results
The test for our model will be that the ratio between the variable x2/x1, (x2/x1)VAR and (x2/x1)MAX is equal to 1.0, which is the same as
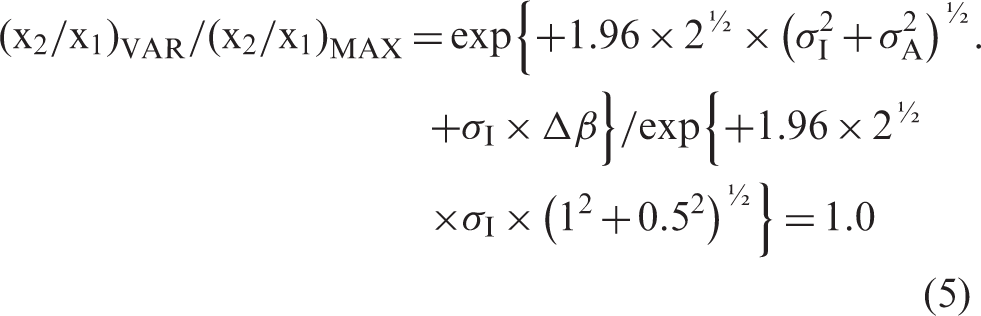
The bias factor, BIASFACTOR, then is BIASFACTOR = exp{#x003C3;I × Δβ} and the fractional bias, BIASFRACTIONAL, becomes
For the lower limit, the same formulae is based on z = −1.96
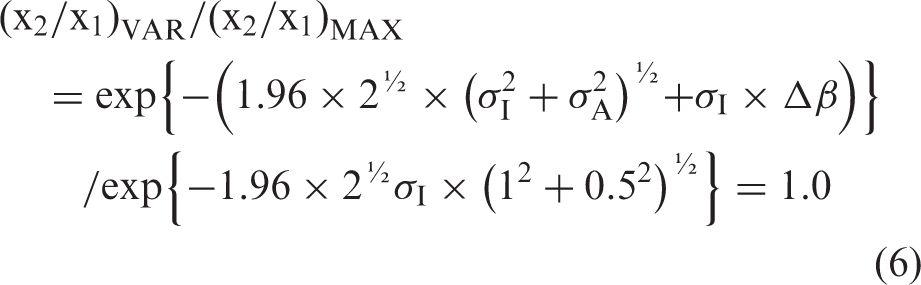
An example of logarithmic bias fraction, σI × Δβ = +3.10 × σI−2.77 × (σI2+σA2)½, as a function of σA = x × σI values from 0.00 to 0.15, for a σI-value = 0.3, is shown in Figure 1.
An example of logarithmic bias fraction, σI × Δβ = + 3.10 × σI − 2.77×(σI2 + σA2)½, as a function of σA = x × σI values from 0.00 to 0.15, for a σI-value = 0.3.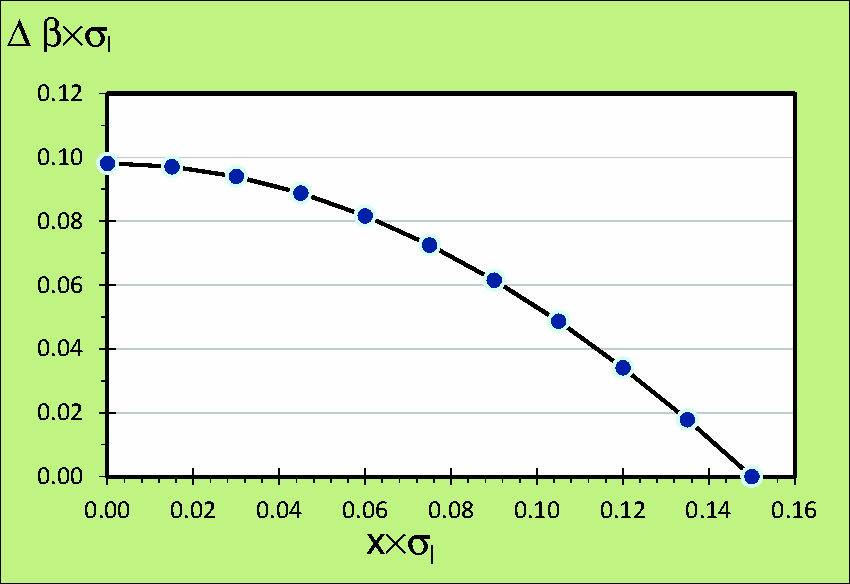
The Δβ is transformed to the BIASFACTOR = exp{#x003C3;I × Δβ} = exp{#x0002B;1.96 × 2½ × σI × (12 + 0.52)½ − 1.96 × 2½ × (σI2 + σA2)½} = exp{#x0002B;3.10 × σI − 2.77 × (σI2 + σA2)½}. The relation between σI and maximum BIASFACTOR for maximum Δβ (i.e. σA = 0) is shown as a straight line in Figure 2. BIASFACTOR = exp{0.33 × σI} and the formula is maximum BIASFACTOR − 1, which is the same as the maximum fractional bias, BIASFRACTIONAL and the maximum fractional bias for σI = 0.1 is BIASFRACTIONAL = exp{0.33 × 0.1}−1 = 1.0335–1 = 0.0335 (≈ 3.35%).
The relation between σI and the maximum bias factor. The relationship is described by the formula σI = exp{#x00394;βMAX}.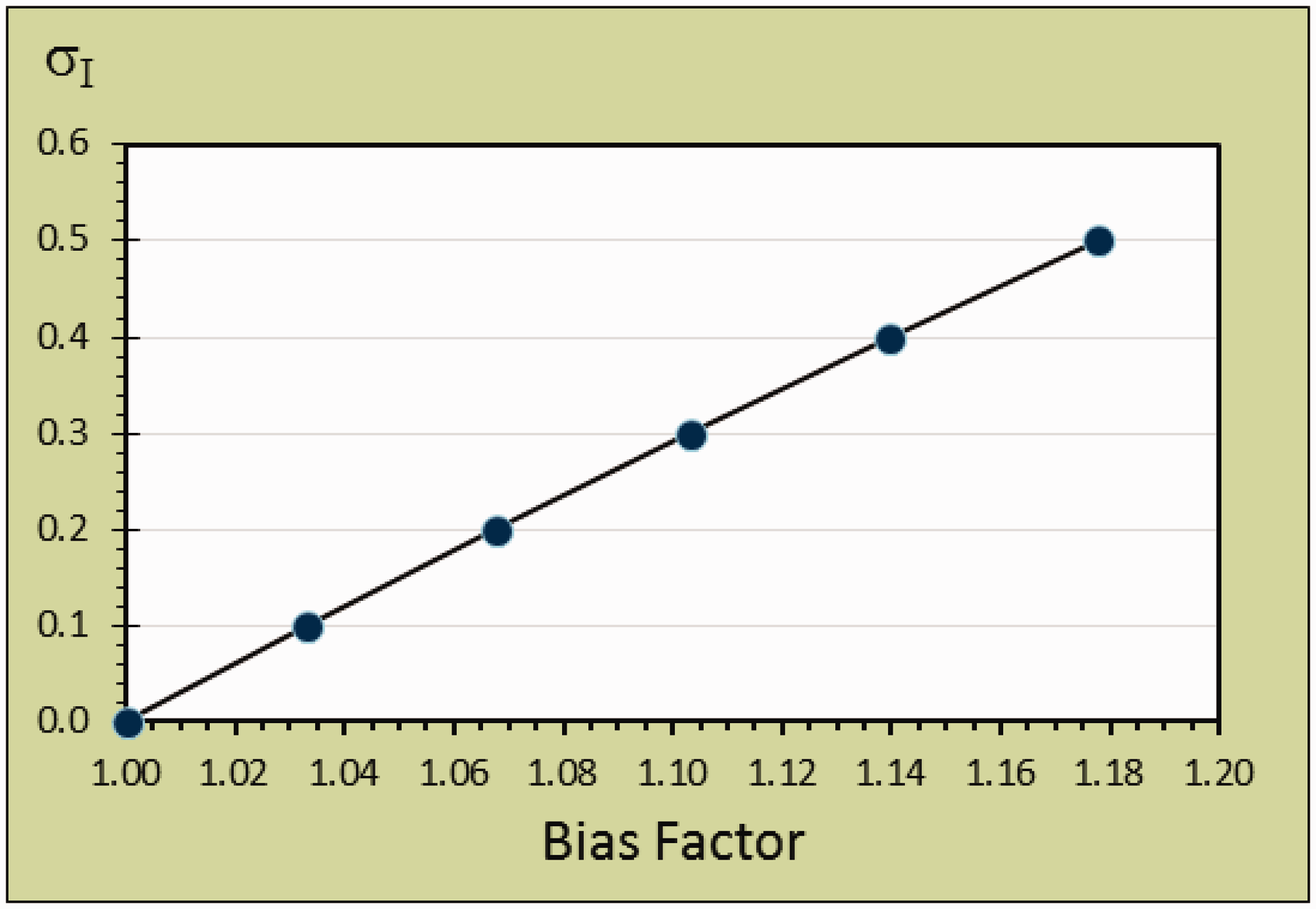
Example 1
Plasma prolactin with CVI = 39.2% = 0.392. 11
According to Lund et al., 9 CVI is transformed to σI using the formula σI = {ln[CVI2 + 1]}½, σI = 0.378.
If a CVA = 5.0% = 0.05 is assumed, the σA becomes 0.05, due to the small value where σI ∼ CVI.
The allowable Δβ will become 0.378 × Δβ = +3.10 × σI − 2.77 × (σI2 + σA2)½ = 1.1718–1.056 = 0.1156, and exp{0.1156} = 1.123, and BIASFRACTIONAL = 0.123, so the result is ∼ 12.3 %.
Figure 3 illustrates the relationships between the allowable bias fraction as function of σA between 0.00 and 0.05.
Fractional bias as function of imprecision between 0.00 and 0.05 for plasma prolactin with CVI = 39.2 % = 0.392 and transformed to σI using the formula σI = {ln[CVI2 + 1]}½, σI = 0.378.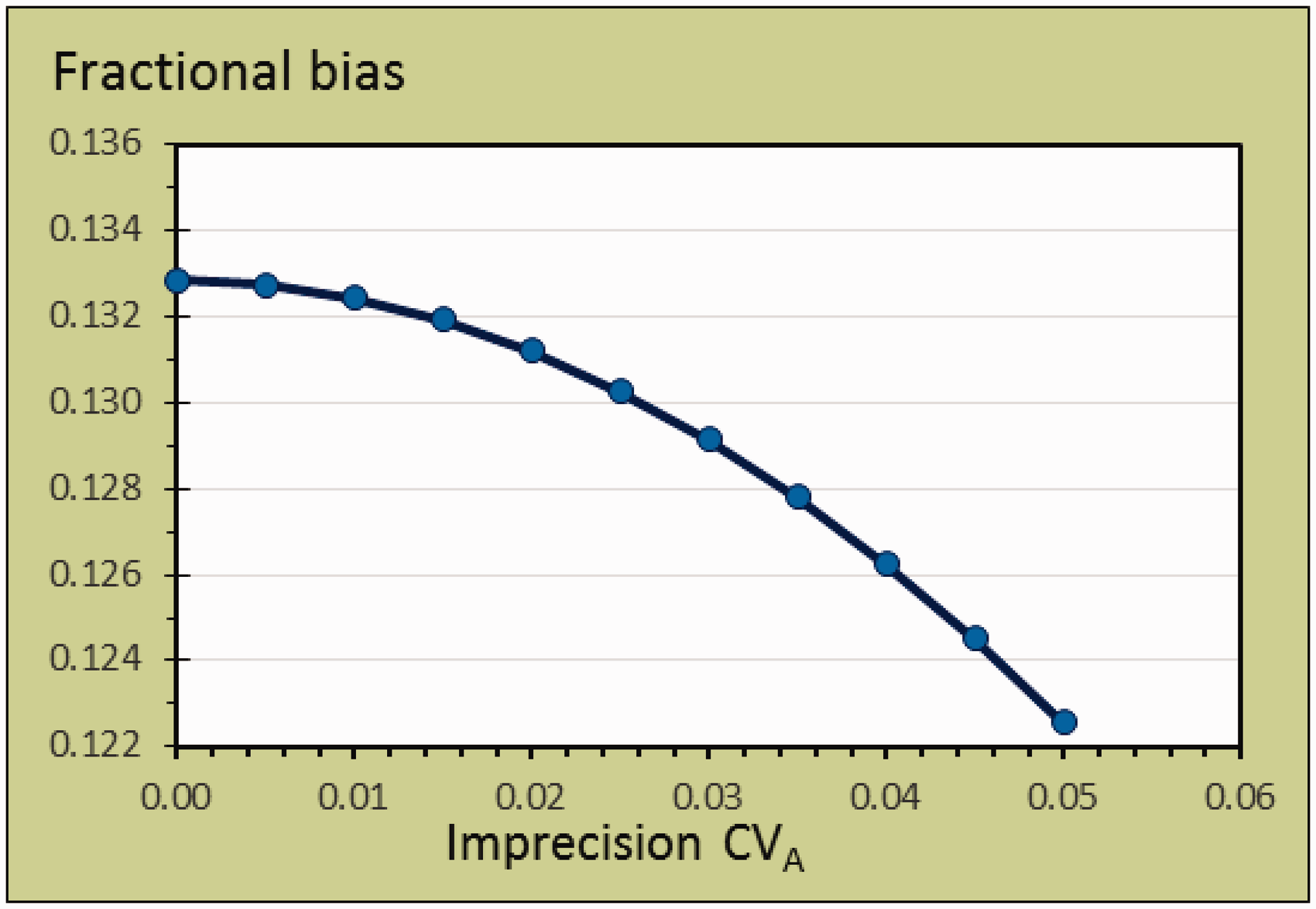
The traditional method4,5 has no transformation of CVI, since the assumption is for a Gaussian distribution, so the formula is
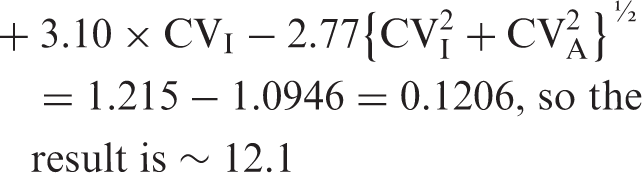
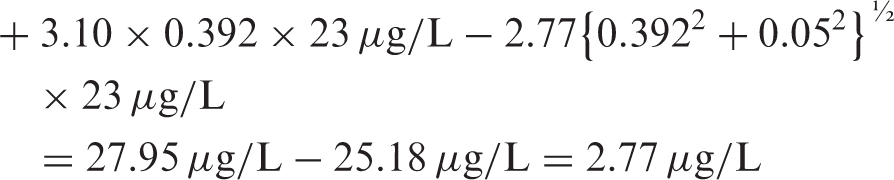
Example 2
Plasma glucose with CVI = 4.5 % = 0.045. 11
According to Lund et al., 9 CVI is transformed to σI using the formula σI = {ln[CVI2 + 1]}½ = 0.045 but, due to the small value, the result becomes indistinguishable from the CVI.
If we assume a CVA = 1.5% = 0.015, the σA becomes 0.015.
The allowable Δβ will become 0.045 × Δβ = 3.10 × 0.045 − 2.77 × (0.0452 + 0.0152)½ = 0.1395–0.1314 = 0.0081, and exp{0.0081} = 1.0081, and BIASFRACTIONAL = 0.0081∼0.8 %.
The traditional method,4,5 both as using s and CV, gives the same result: 0.8%.
Discussion
There are several indications that the distributions of both CVI and CVG are not Gaussian but skewed, as is the total variation of traditional population-based reference values for healthy individuals for many common measurands in laboratory medicine. Moreover, the use of logarithms is the basic assumption for the theoretical investigations both by Fokkema et al., 8 now widely cited and applied, and by Lund et al.9,10 Even if the skewed distributions are better described by other possible models, we consider that logarithmic distributions are the best to describe numerical biological phenomena. In addition, the traditional description has drawbacks for large CV, e.g. the problems with negative concentrations being found when derived RCV are applied if inappropriately generated high CV are used. One of the reasons for the lack, to date, of clear and comprehensive documentation of the type of distribution may be the need for a large number of samples from healthy individuals in steady-state to reveal the exact type of distribution, whether Gaussian, ln-Gaussian or other.
Nonetheless, it is interesting to see that the performance specifications based on our new logarithmic and on the traditional 4 models are practical identical for small CVI values such as for plasma glucose as documented in Example 2, and hardly dissimilar for plasma prolactin with large CVI. This means that both models can be used for generating analytical performance specifications for CV – and corresponding s – except from the fact that only logarithmic models can avoid against negative concentrations being found when RCV are applied if large (but inappropriate) estimates of CV are used. The reason for the nearly identical results may be the transformation of CVI to σI by the formula σI = {ln[CVI2 + 1]}½. This does seem to be very effective since the allowable Δbias for the plasma prolactin example (Example 1) would be 12.1 % instead of the 12.3 %, if not transformed logarithmically.
Both models for generation of analytical performance specifications can be useful for manufacturers of instruments and reagents including calibrators, where each lot has a specific Δbias from the previous lot. This is also vital for medical laboratories, where every change of lot poses a challenge to the desired analytical stability, and therefore needs special attention with necessity of intensive control, especially for measurands such as serum sodium, calcium and chloride which have well-documented small CVI. In spite of the enormous endeavours on standardization of measurements with traceability to a high level of trueness, actual performance in practice may demonstrate large deviations, e.g. for serum sodium measured over eight years in two large Belgian hospital laboratories, where deviations of up to 4 mmol/L were seen. 12 It is important to keep in mind that both Δβ and σA are variables in the formula and that their influences are fundamentally different: in consequence, solving problems and errors in ongoing analytical performance must be addressed separately for each variable.
Another model for RCV related to the change in standard deviation has been described by Jones 13 that RCV% = z × [CVtotal2 + ((1 + RCV%) × CVtotal)2]½. This is complicated to calculate the limits for the RCV, but it might be interesting to estimate the performance specifications derived from this model and compare to our models in future work.
The estimation of analytical performance specifications with our new model seems to be rather more complicated than the traditional theory,4,5 but, even for large CVI, the difference between models is negligible; thus, only for very high CVI is the ln-model of significant advantage.
Conclusions
Our model for analytical quality specifications based on logarithmic distributions of CVA is conceptually more correct than the traditional model based on data derived using assumptions of Gaussian distributions, but results are similar up even with large CVI.
Footnotes
Acknowledgements
We would like to thank Merete Frejstrup Pedersen for her assistance in generating the figures.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Guarantor
FL.
Contributorship
All authors were involved in the performance of the study and wrote the paper.
