Abstract
Raman spectroscopy (RS) is a label-free, non-destructive optical modality that provides a detailed profile of the molecular composition of a sample. There is growing interest in the clinical application of RS to characterize biomolecular signatures associated with radiotherapy response in tumor cells and tissues. A critical step before analyzing Raman data consists of performing spectral pre-processing to increase the quality of the measurements. Spectral pre-processing comprises baseline subtraction, signal smoothing, cosmic ray (CR) correction, and removal of poor-quality measurements. Herein, we present a convolutional autoencoder (AE) for single-step, automated pre-processing of Raman spectra obtained from tumor cells and tumor tissue. We trained two separate models using the same proposed architecture, one for eliminating spectral artifacts from preclinical single-cell line and xenografted tissue spectra exposed to single-fraction radiation, and the other for correcting clinical prostate tumor biopsy spectra collected from patients receiving high-dose-rate brachytherapy (HDR-BT). The autoencoder demonstrated fast, excellent performance in removing baseline, noise, and CRs. For the preclinical data, the model obtained a root mean squared error (RMSE), and a percentage root mean squared difference (PRD) of 7.1 × 10−5 and 3.1%, respectively, between the AE-corrected spectra and their corresponding target data (pre-processed by our current baseline-removal algorithm). Also, the autoencoder successfully removed 94.0% of CRs from the spectra. For the clinical biopsy data, the AE achieved an RMSE and a PRD of 8.1 × 10−5 and 3.7%, respectively, and a CR removal rate of 90.2%. Overall, the AE corrected approximately 11 000 spectra within 2.4 s without the need of a GPU. Furthermore, comparative supervised learning-based post-processing data analyses were performed separately on the spectra pre-processed by the autoencoder versus the target data, and we show consistency in the biochemical radiation response profiles extracted. Finally, the AE architecture was leveraged to train a reconstruction AE to facilitate semi-automated identification of poor-quality prostate biopsy spectra, and we demonstrate 96.4% agreement between AE and manually removed outliers. These results support the development of a deep learning framework for efficient, automated pre-processing of tumor cell and tissue Raman spectra collected for radiation response monitoring studies.
This is a visual representation of the abstract.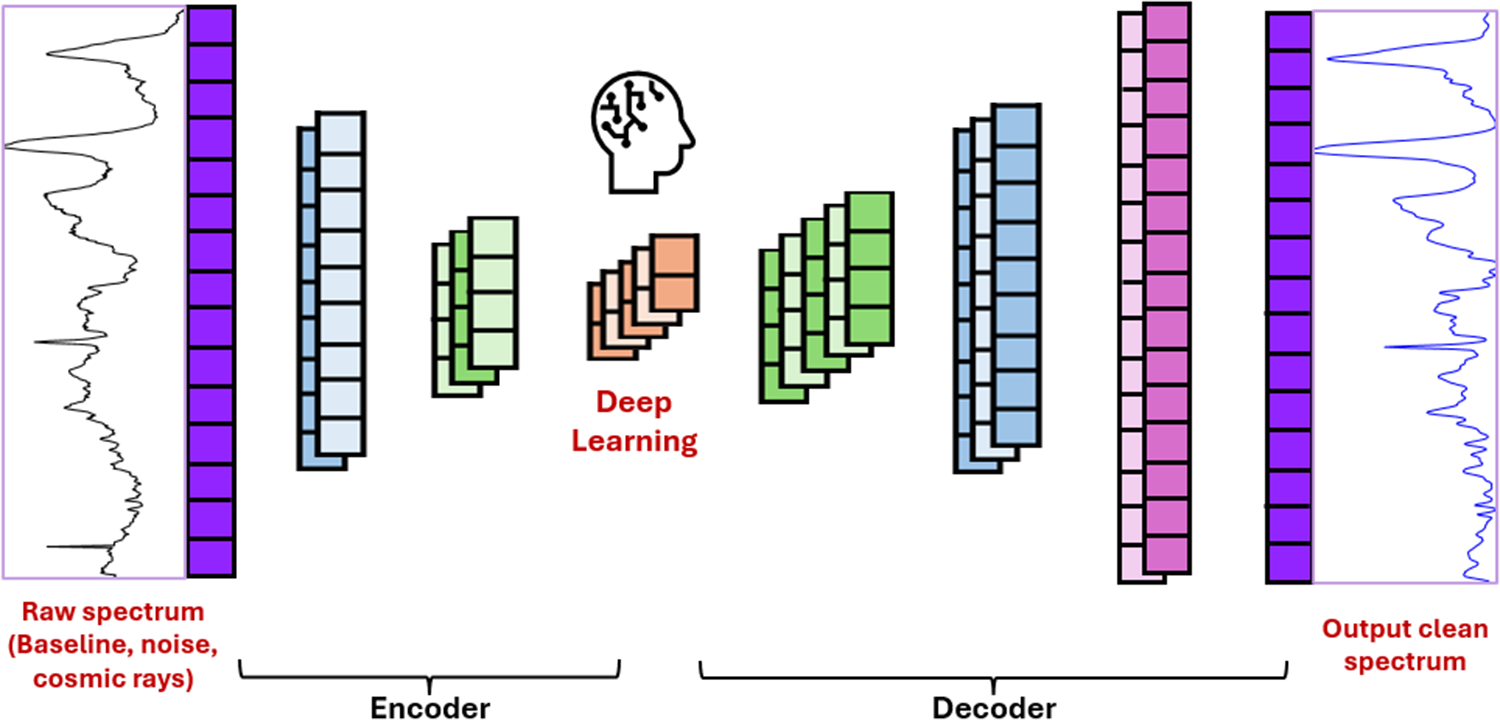
Keywords
Get full access to this article
View all access options for this article.
