Abstract
Mitochondrial dysfunction is a hypothesized component in the multifactorial pathogenesis of migraine without aura (MoA, ‘common migraine') and the related condition of cyclic vomiting syndrome (CVS). In this study, the entire mitochondrial genome was sequenced in 20 haplogroup-H CVS patients, a subject group studied because of greater genotypic and phenotypic homogeneity. Sequences were compared against haplogroup-H controls. Polymorphisms of interest were tested in 10 additional CVS subjects and in 112 haplogroup-H adults with MoA. The 16519C→T polymorphism was found to be highly disease associated: 21/30 CVS subjects [70%, odds ratio (OR) 6.2] and 58/112 migraineurs (52%, OR 3.6) vs. 63/231 controls (27%). A second polymorphism, 3010G→A, was found to be highly disease associated in those subjects with 16519T: 6/21 CVS subjects (29%, OR 17) and 15/58 migraineurs (26%, OR 15) vs. 1/63 controls (1.6%). Our data suggest that these polymorphisms constitute a substantial proportion of the genetic factor in migraine pathogenesis, and strengthen the hypothesis that there is a component of mitochondrial dysfunction in migraine.
Introduction
Migraine is a very common condition, affecting approximately 15% of adults (1, 2), with high economic costs, especially in terms of lost time from employment and as a frequent cause of healthcare utilization. Although comprised of many variants, the majority of migraineurs have migraine without aura (MoA, previously known as ‘common migraine’). Like most other common disorders, the aetiology of migraine and its variants is multifactorial, with many known genetic components, including genes for calcium and sodium channels, dopamine and insulin receptors, Na+/K+ ATPase pump subunits, and components of mitochondrial energy metabolism (3, 4).
A mitochondrial component to migraine has been postulated. This is supported by the findings in migraine sufferers of lactic acidosis (5, 6), mitochondrial accumulations and cytochrome-c-oxidase negative fibres in skeletal muscle (5), decreased respiratory chain complex activities (5, 7) and reduced in vivo brain phosphocreatine to inorganic phosphate ratio by 31P-magnetic resonance spectroscopy (5). In addition, co-enzyme Q10 and riboflavin, a component and a precursor of a component of the mitochondrial respiratory chain, have shown efficacy in migraine prophylaxis in double-blind, placebo-controlled clinical trials (8, 9). The mitochondrial dysfunction hypothesis of migraine was recently reviewed (10).
Mitochondria are cytoplasmic organelles that produce the bulk of the ATP for cellular energy needs. Mitochondrial proteins are encoded on both the nuclear DNA (chromosomes) as well as the 16-kb mitochondrial DNA (mtDNA). Thus, sequence variants (polymorphisms) that adversely affect energy metabolism and predispose towards migraine pathogenesis theoretically could be on either or both of those genomes. The cytoplasmic-located mtDNA generally is derived solely from the ova without recombination, and individuals related through women carry an identical mtDNA sequence in the absence of a recent mutation.
Pilot studies have suggested a preferential maternal bias in migraine inheritance (11, 12), suggesting the presence of disease-predisposing mtDNA sequence variants. In support of this, about 20 different mtDNA sequence variants have been associated or possibly associated with migraine (10), especially 3243A→G (13). Most of those sequence variants are located in the ‘coding regions’ that comprise 94% of the mtDNA and that code for subunits of the respiratory chain or the transfer and ribosomal RNA molecules needed to translate those subunits. In one study (14), migraine and the related condition of cyclic vomiting syndrome (CVS) were found to be associated with any sequence variation in a 150 base-pair area of the control region believed to regulate mtDNA replication. Furthermore, an entire mtDNA haplogroup (U) was found to predispose towards migraine with occipital stroke (15).
However, migraine is a heterogeneous phenotype in which diagnostic criteria exist but are not always definitive. As a model disorder in which to study migraine genetics, we chose CVS, a condition whose presence or absence is almost always clear by expert application of diagnostic criteria (16). CVS is a disabling condition characterized by recurrent, distinct episodes of nausea, vomiting and lethargy separated by asymptomatic intervals (16–18). Most sufferers encounter recurrent identical episodes that often result in frequent school or work absences and multiple hospitalizations for dehydration. CVS is not rare; it has been reported in nearly 2% of Western Australian (19) and Scottish (20) school children. The condition is more common in children and in girls. CVS is widely believed to be a ‘migraine-like condition’ secondary to a strong family history of migraine headache, as well as frequent prodromic symptoms, headache, abdominal pain, evolution to migraine headache, and a positive response to ‘antimigraine’ medications in CVS patients (21, 22). mtDNA sequence variation is hypothesized to be a substantial risk factor in the pathogenesis of CVS due to the preferential maternal inheritance of functional disorders, including MoA, in the families of most CVS sufferers (17, 23), and the presence of an energy-depleted pattern on urine organic acid measurements in most cases (17). In addition, lactic acidosis, reduced electron transport chain activities and/or heteroplasmic mtDNA sequence variants have been reported in selected cases (23–25). Most mtDNA sequence variants in CVS patients have been reported in the 1-kb mtDNA control region (14, 26), although 3243A→G and large rearrangements have been reported in the 15.6 kb comprising the coding regions (26–30).
In this study, the entire mtDNA was sequenced in 20 individuals with CVS. Any potential disease-associated sequence variants were assayed in an additional 10 CVS cases and in 112 adults with MoA. In order to minimize background mtDNA sequence variability (noise), all patient and control subjects had mtDNA haplogroup H.
Subjects and methods
Subjects
The CVS subjects of whom the entire mtDNA was sequenced included 20 individuals from an earlier study (17) who were ascertained randomly throughout North America based upon postal codes from the database of the Cyclic Vomiting Syndrome Association. Additional CVS subjects who were only assayed for selected mtDNA polymorphisms included six subjects recruited by the above means, and four subjects recruited as part of another earlier study on CVS with comorbid neuromuscular disease (23). Among the 30 subjects, there were 21 girls and nine boys. All subjects were unrelated, met the research definition for CVS (16) as determined by telephone interview, and had mtDNA haplogroup H.
The adult migraineur group consisted of 77 hospital-based patients from a headache clinic near Frankfurt and 35 out-patients recruited from an out-patient clinic in Munich, both from Germany. All subjects were unrelated, met the International Headache Society definition of MoA (31) as determined by a mailed-in questionnaire (32) and had mtDNA haplogroup H. A telephone interview was performed in cases with diagnostic uncertainty.
The control group consisted of 195 full-mtDNA sequences and 213 mtDNA sequences missing only the control region, obtained from published sources (Table 1). In addition, the polymorphisms of interest were assayed for in 36 healthy White children recruited from the first author's institution in Los Angeles. Only haplogroup H sequences from individuals ascertained as part of a population or control study from Europe or North America were included. To reduce potential bias, we excluded samples ascertained due to any illness or symptoms, from self-selected groups (commercial heritage testing), and from islands with small founding and/or geographically isolated populations (Iceland and Sardinia).
Control haplogroup H mtDNA sequences
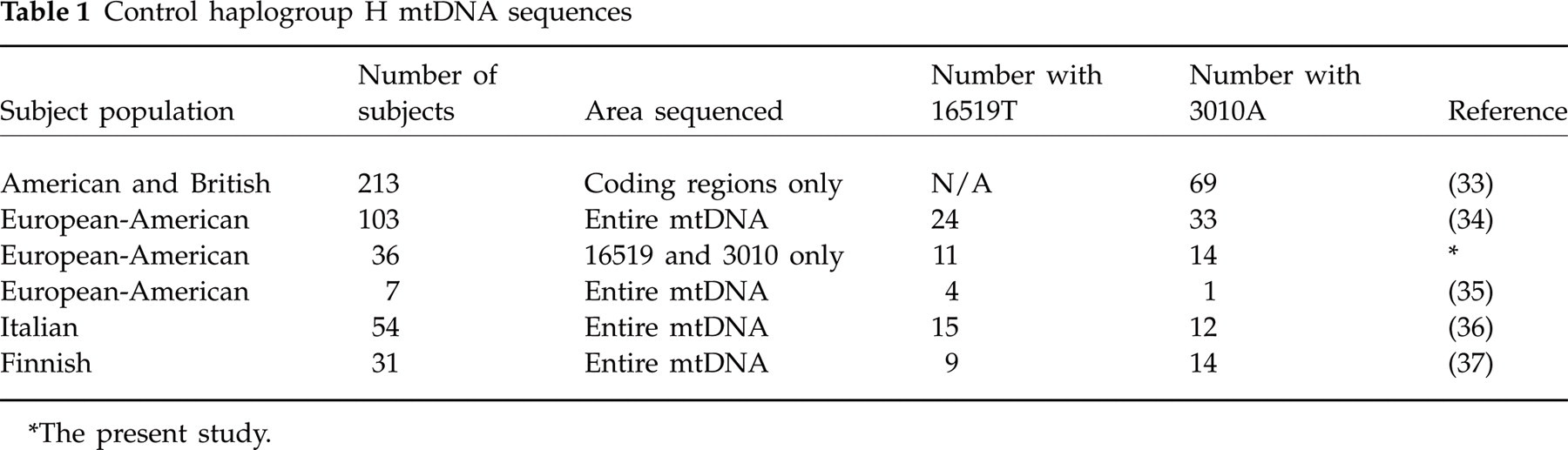
∗The present study.
Informed consent was obtained from each subject or parent, except for de-identified control subjects, and all aspects of the study was approved by the Children's Hospital Los Angeles Institutional Review Board.
Methods
DNA was isolated from blood by standard methods or from saliva using a commercially available kit (Oragene; DNA Genotek Inc., Ottawa, ON, Canada). Haplogroup H was defined in the conventional manner as the presence of a C at position 7028, as listed in published sequence databanks, by cyclosequencing or by polymerase chain reaction-restriction fragment length polymorphism (RFLP)—16519: HaeIII F GGATGACCCCCCTCAGATA, R CTTATT TAAGGGGAACGTG; 3010: BccI F CATGCTAA GACTTCACCA, R TCGTTGAACAAACGAACC.
The entire mtDNA was amplified using 28 overlapping primer sets (26, 14; and additional sets available upon request). Cyclosequencing was performed by SeqWright (Houston, TX, USA), Agencourt (Beverly, MA, USA), or Eton (San Diego, CA, USA). Individual sequences were aligned and compared on Sequencher® software (Gene Codes Corp., Ann Arbor, MI, USA) vs. our reference sequence. Our reference consists of the most common nucleotide among haplogroup H individuals in our control group for each nucleotide position throughout the mtDNA, and is termed the haplogroup H reference sequence (HhRS). The HhRS is identical to the revised Cambridge Reference Sequence (rCRS) (MITOMAP) (38), which corresponds to the actual sequence of one individual of sub-haplogroup H2, with the exception of nine changes correcting for rare and uncommon polymorphisms in the rCRS (see Table 2 legend).
Complete mtDNA genomic sequence data in 20 haplogroup H cyclic vomiting syndrome subjects

∗The HhRS is identical to the revised Cambridge Reference Sequence (rCRS) (38), with the exception of seven rare polymorphisms (263A→G, 310C→T, 750A→G, 1438A→G, 4769A→G, 8860A→G, 15326A→G) and two variants (309C→T and 16519T→C).
†Adult-onset cases.
Pedigree analysis was performed over the telephone by a genetic counselling student (S.W.) in the 14 families that could be located. Families were labelled as ‘probable maternal inheritance’, ‘probable non-maternal inheritance’ or ‘indeterminate’ by our quantitative pedigree analysis method (39). Based on a semistructured family history, the number of all neuromuscular and endocrine conditions among the first- and second-degree matrilineal relatives (who share the same mtDNA sequence as the subject) was divided by the number of subjects. In addition, that resultant quotient was divided by the number of all neuromuscular and endocrine conditions among the analogous non-matrilineal relatives (who have different mtDNA sequences from the subject) to give a ‘maternal inheritance ratio’.
Statistics were performed by WinSTAT Statistics for Windows (Kalmia Co. Inc., Cambridge, MA, USA) and by custom-made software.
Results
All sequence variants relative to the HhRS in our 20 fully sequenced CVS subjects are listed in Table 2. Excluding insertions in the ultravariable 302–315 region, 20 different mtDNA variants were identified in two or more CVS subjects (Table 3), all of which are single nucleotide polymorphisms (SNPs) listed on MITOMAP (38). Most prominently, one of the SNPs, 16519C→T1, was found to be highly associated with CVS vs. controls, and another, 3010G→A1, was found to be highly associated with CVS in subjects with 16519T vs. in controls with 16519T (Table 4).
mtDNA single-nucleotide polymorphisms (SNP) found in ≥ 10% of the 20 fully sequenced haplogroup H cyclic vomiting syndrome (CVS) or 195 fully sequenced haplogroup H control subjects
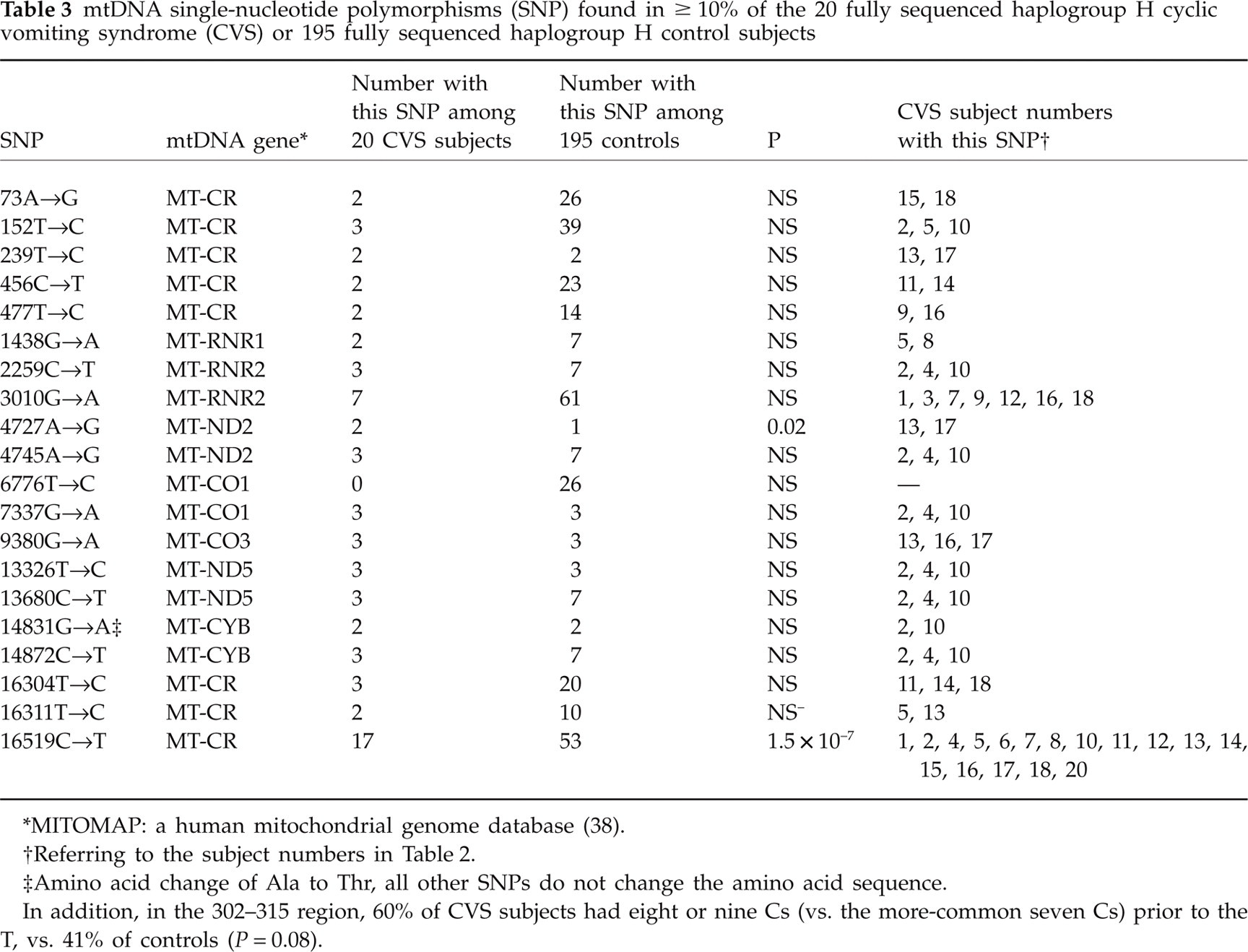
∗MITOMAP: a human mitochondrial genome database (38).
†Referring to the subject numbers in Table 2.
‡Amino acid change of Ala to Thr, all other SNPs do not change the amino acid sequence.
In addition, in the 302–315 region, 60% of CVS subjects had eight or nine Cs (vs. the more-common seven Cs) prior to the T, vs. 41% of controls (P = 0.08).
Prevalence of 16519T and 3010A in haplogroup H cases with cyclic vomiting syndrome (CVS) and migraine without aura
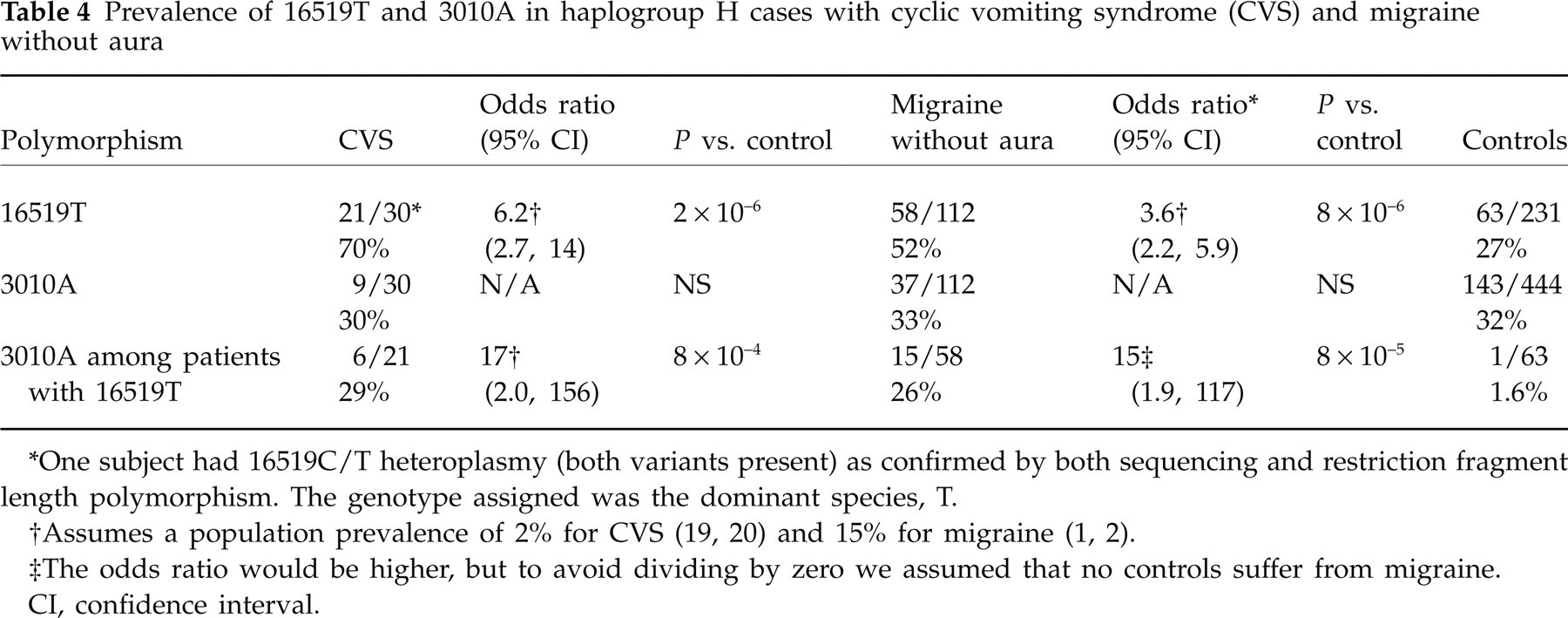
∗One subject had 16519C/T heteroplasmy (both variants present) as confirmed by both sequencing and restriction fragment length polymorphism. The genotype assigned was the dominant species, T.
‡The odds ratio would be higher, but to avoid dividing by zero we assumed that no controls suffer from migraine.
CI, confidence interval.
Twenty-five of our 30 subjects reported paediatric onset of vomiting episodes, in each between early infancy and age 14 years. Among the five adult-onset subjects, three had 16519T and one had 3010A (a subject with 16519T). Among the paediatric-onset subjects that underwent pedigree analysis, the 16519T polymorphism was found in 7/9 with probable maternal inheritance, 1/2 with indeterminate pedigrees and 1/3 with probable non-maternal inheritance (P = 0.20 for probable maternal inheritance vs. others). All four paediatric-onset CVS subjects with the 16519T+3010A genotype had probable maternal inheritance. There were no gender-based differences noted in our data.
Excluding 16519C→T and 3010G→A, there was a mean of 5.2 mtDNA SNPs per individual among the fully sequenced 20 CVS subjects and 4.9 mtDNA SNPs per individual among the 195 fully sequenced controls (P = NS).
The 16519C→T polymorphism was found to be highly associated with MoA (Table 4). Furthermore, among the subset with 16519T, the 3010G→A polymorphism was found to be highly associated with MoA.
Discussion
Complex multifactorial conditions, usually influenced by multiple genetic and environmental factors, are the cause of the vast majority of human disease. These conditions are becoming better understood as (nuclear) genetic polymorphisms that confer an increased risk toward disease pathogenesis are rapidly being identified. Although the mitochondrial genome is small, it is present at high copy number and has a very high polymorphic density (40). Thus, mtDNA sequence variation is likely to affect an individual's risk for the development of some multifactorial conditions in a manner analogous to nuclear DNA polymorphisms (41). Migraine (including MoA and CVS) may follow such a hypothesis, since a genetic component in its pathogenesis, preferential maternal inheritance and mitochondrial dysfunction are all well established. Furthermore, in contrast, migraine is very common in patients with maternally inherited mitochondrial dysfunction (42). Establishing disease-associated mtDNA sequence variant(s) in migraine, or in any other condition, indicates that energy metabolism is a factor in disease pathogenesis, since the 37 mtDNA genes are exclusively involved in oxidative phosphorylation (40). In addition, disease-associated mtDNA sequence variation constitutes an important justification for the use of mitochondrial-directed therapies, some of which have demonstrated efficacy by double-blind clinical trials in migraine, and by retrospective studies and anecdotal observation in CVS (25, 43).
In the present study, we have demonstrated that the common mtDNA polymorphisms 16519T and 3010A are highly associated with the most common form of migraine and CVS. Given the lack of chronic disability in most MoA cases, we assume that our control group, ascertained for forensic or evolutionary studies, contains about the same proportion of migraineurs as does the general population. With this assumption, the 16519T polymorphism alone was found to have an odds ratio (OR) of 3.6 in MoA and 6.2 in CVS (Table 4). This corresponds to predicted prevalence rates of 28 and 11% for migraine in individuals with 16519T and 16519C, respectively. Even more striking is the combined effects of the two polymorphisms, which define evolutionarily separate branches of haplogroup H and thus are very rarely seen together in control populations (Fig. 1). Although the 3010 polymorphism alone does not appear to confer risk for developing migraine, on a 16519T background the 3010A polymorphism has an OR of 15 for MoA and 17 for CVS. Thus, the data predict that migraine is present in 74% of individuals with the combined 16519T/3010A genotype, and that these mtDNA polymorphisms are likely to be acting synergistically.

The figure shows that the 3010A polymorphism occurred on the ancestral 16519C background early in the phylogeny of haplogroup H. The 16519 nucleotide later mutated (mostly C to T, but also T to C) in several of the sub-haplogroups. Although 16519T is rarely seen in sub-haplogroup H1 (the most common H sub-haplogroup, defined by 3010A), this study demonstrates that it is statistically far more common in individuals with cyclic vomiting syndrome (CVS) or migraine without aura. Although 16519C and T are common in control individuals with sub-haplogroups H4 and H13, the data suggest that T may be the predominant nucleotide among patients with CVS. H7 in controls is almost exclusively 16519C, yet our one CVS subject had 16519T. The figure was derived in part from reference 36, and is not to scale.
In addition, our data suggest that there may be other mtDNA polymorphic modifiers on a 16519T background, but the numbers are too small for significance in most cases (Table 3). In one case that did reach statistical significance, the 4727A→G polymorphism was found in 2/20 CVS cases vs. in 1/195 fully sequenced controls. Furthermore, three CVS subjects with 16519T and 3010G share six other polymorphisms (2259C→T, 4745A→G, 7337G→A, 13326T→C, 13680C→T and 14872C→T) in common, a combination not present in any of the 195 fully sequenced controls (P = 7 × 10−4). Those three subjects are not closely related as their mtDNA sequences vary at other polymorphisms.
Although the numbers are too small for statistical significance, our data suggest, as expected, that the prevalence of 16519T and 16519T+3010A may be elevated only in the maternally inherited subset of CVS. Recently, it has been suggested that CVS in adults is different in many ways from that in children (44), although our small number of adults is insufficient to comment as regards the present data. Future studies in CVS should perhaps be conducted by restricting analysis to cases with probable maternal inheritance, and/or analysing paediatric and adult-onset cases separately.
The overall genetic component to the pathogenesis of migraine without aura is moderate, with first-degree relatives being at about twofold increased risk for being affected themselves (45). Thus, 16519T and 3010A probably constitute a substantial proportion of the overall genetic factor in MoA. As we have full mtDNA sequences in 20 of our CVS subjects, in CVS the data do not support that these variants are in linkage disequilibrium with other deleterious mtDNA sequences (Tables 2 and 3). More likely, it is the 16519T and 3010A nucleotides themselves that affect energy metabolism in a manner that predisposes towards the development of MoA and CVS. Furthermore, excluding 16519T and 3010A, all other mtDNA SNPs were found at essentially the same frequency in our CVS subjects and controls. Although we certainly cannot state that these other polymorphisms among our CVS subjects are not disease-predisposing or modifying, it does not appear that any of them are singularly important in a sizable proportion of CVS patients, at least not in haplogroup H.
The 16519T polymorphism is located in the 1-kb non-coding mtDNA control region (often referred to as the ‘D-loop’), not far from the origin of heavy-strand replication and putative membrane-attachment site (38). Although located in an area of the control region with relatively low sequence heterogeneity, 16519T is considered to have one of the highest mutation rates of any mtDNA position and has arisen multiple times in human evolution (see Fig. 1 for occurrences within haplogroup H), including among Americans of West Eurasian (46), East Asian (47), African (48) and Hispanic/Native American (49) origins. Interestingly, 16519T was recently shown to be associated with diabetes and a poorer prognosis in individuals with pancreatic cancer (50). A physiological effect of this polymorphism is also suggested by its complex effects on human exercise physiology (51).
The 3010A polymorphism is located in the 16S-ribosomal RNA gene, whereas it rebuilds a base pair in a stem of the ribosomal A-site (38). Bacterial mutations in this stem confer resistance to certain antibiotics such as chloramphenicol. To complicate matters, the 16S-RNA is probably involved in protein refolding (52) and has a fourfold higher gene expression in women (53). 3010G demonstrates evolutionary conservation in primates, although A is the nucleotide in mice and frogs. 3010A has appeared in human evolution at least 15 times on 10 different mtDNA haplogroups, and may be under positive selection in humans. It defines the sub-haplogroups/clusters of H1, J1, U3 (West Eurasian), D4, C (East Asian) and L2a (African) (38). J1 and D4 are over-represented in centenarians (54). Recently, 3010A was reported to be protective against stroke (55). A heteroplasmic mutation, 2839C→A in the same mtDNA ribosomal gene has been reported in most maternally inherited cases of Dupuytren's disease (56), a nodular palmar fibromatosis that causes digital contractures, which is of interest because ‘arthritis’ was found to show comorbid maternal inheritance with migraine in some families (42).
The mtDNA haplogroups denote sets of ancient matrilineal ancestry tens of thousands of years old. The West Eurasian haplogroup H is well suited for genetic association studies due to the relative lack of intragroup sequence variability and high prevalence. Among our fully sequenced control group of 195 European-derived individuals with haplogroup H, the mean number of nucleotide changes throughout the entire mtDNA genome relative to the HhRS is only about eight nucleotides (data to be published separately), vs. several times this number for unrelated individuals of mixed haplogroups. Thus, limiting our study to subjects with haplogroup H substantially decreases background sequence variability and correspondingly increases statistical power. It is also practical, as haplogroup H is the most common among European-derived populations, including prevalence rates of about 45% in the native population of Germanic countries, and of about 33% in North Americans of apparent-European ancestry (41). Thus, we chose to sacrifice a larger number of subjects for substantially greater genotypic homogeneity by limiting this study to subjects of haplogroup H. Unlike some haplogroups, the 16519 and 3010 nucleotide positions are quite heterogeneous among individuals within haplogroup H. For example, had we chosen to study haplogroups J or T, whereas 3010A and 16519C are nearly universal, respectively, we would have missed the association with either polymorphism. Of course, this reasoning states that we may have missed migraine-associated polymorphisms that are not heterogeneous among haplogroup H individuals, and research in individuals with other haplogroups is needed.
One potential problem with our study is the lack of a German control group. However, the polymorphic frequency of 16519T appears to be highly homogeneous among continental European-derived haplogroup H populations, including 27% from the USA, 28% from Italy and 29% from Finland (Table 1). Furthermore, considering that the migraineurs were ascertained from two large cities in the centre of Europe, Frankfurt and Munich, it is quite unlikely that our results are due to a local increased prevalence of 16519T. Our results are also unlikely to be due to systematic errors in our procedures, as most of our subjects' genotypes were confirmed by both cyclosequencing and RFLP. Furthermore, in our own control group, 31% have 16519T, although the total is limited to 36 haplogroup H subjects. The polymorphic frequency for 3010A is also highly similar among the two larger control groups, 33% in Americans and 32% in mixed British/Americans (Table 1). The proportion does vary in our smaller groups, 22% in Italians and 45% in Finns, but these are non-significant differences (P = 0.18), probably due to the small numbers of subjects in those groups (Table 1). Furthermore, since 3010A is present in only 1/63 controls from diverse locations with 16519T, it is highly unlikely that our findings of 3010A in 6/21 CVS and 15/58 MoA subjects with 16519T is due to an artefact of local polymorphic differences.
We conclude that the common mtDNA polymorphisms 16519T and 3010A are strongly associated with migraine and CVS among individuals with the common West Eurasian haplogroup H, and are likely to predispose to disease. Although only haplogroup H was studied, 16519T and 3010A are found in individuals with a multitude of haplogroups and within all major races. In fact, among Americans of European origin, 16519T is actually slightly more common among non-haplogroup H individuals (34). The effect of these polymorphisms may or may not be dependent upon the background haplogroup, and further study is needed. The mechanism by which these non-protein-coding polymorphisms exert physiological and clinical effects is unknown, but based upon their locations a potential effect on mtDNA replication and translation is suggested. Our data strengthen the hypothesis that there is a component of mitochondrial dysfunction in migraine and provide a additional rationale towards the use of mitochondrial-directed therapies in this condition.
Footnotes
Competing interests
The Children's Hospital Los Angeles (CHLA), R.G.B. and E.A.Z. of CHLA hold patents and/or patent applications on the detection of mitochondrial DNA polymorphisms and on the relationship of these polymorphisms with specific human diseases set forth in this study. Both CHLA and the authors may benefit financially from the commercialization of technologies, and other intellectual property, derived from these patents and/or patent applications.
Acknowledgements
This study was supported in part by a grant from NARSAD.
1This standard convention for mutations is read as 16519C and 3010G being the usual sequences in humans, and 16519T and 3010A being the altered (mutated) sequences. 16519T is one of nine uncommon or rare SNPs that are present in the single individual whose mtDNA was first sequenced and thus comprises the established reference sequence (rCRS) (![]() ). Thus, while 16519C is the ancestral (present in chimpanzees) and the most common nucleotide among humans in general, 16519T is the rCRS nucleotide.
). Thus, while 16519C is the ancestral (present in chimpanzees) and the most common nucleotide among humans in general, 16519T is the rCRS nucleotide.
