Abstract
Mutations in the NOTCH3 gene cause cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL). Exons 3 and 4 are mutation hotspots. Migraine is a clinical hallmark of CADASIL. The objective of this study was to investigate whether genetic variants in exons 3 and 4 of the NOTCH3 gene are associated with migraine. Exons 3 and 4 of the NOTCH3 were analysed for mutations and polymorphisms by direct DNA sequencing in 97 migraineurs and the same number of control individuals. No mutations in exons 3 and 4 of the NOTCH3 gene were found in 97 patients with migraine. However, association analysis revealed significant association of the single nucleotide polymorphism (SNP) rs1043994 with migraine.
Keywords
Introduction
Migraine with (MA) and without aura (MoA) are in most cases genetically complex, although rare monogenetic forms exist, e.g. autosomal dominant familial hemiplegic migraine, for which two defective genes (CACNA1A and ATP1A2) are currently known (1). A number of studies have shown that MA and MoA are associated with juvenile stroke in women and subclinical brain lesions affecting mainly the cerebellum, the brainstem and the white matter of the cerebrum (2–5). Migraine, in most instances with aura, and juvenile ischaemic stroke affecting predominantly the white matter of the cerebrum and the brainstem, are hallmarks of cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL) (6). CADASIL is an autosomal dominant disease caused by mutations in the NOTCH3 gene, while migraine is a multifactorial disorder. The heritability of migraine has been estimated between 40% and 55% in a number of twin studies (7, 8). Migraine is a genetically complex condition and the heritability is higher in women than in men (7, 8). Some recent studies suggest that MA and MoA might not be genetically separate entities, but could represent different severities of the same disease caused by genetic variations in an overlapping or identical set of genes (9). In this study, we address the question whether genetic variants in exons 3 and 4 of the NOTCH3 gene might play a role in the pathogenesis of non-familial, non-hemiplegic migraine. Exons 3 and 4 were chosen because they represent mutation hotspots containing more than 65% of all CADASIL mutations (10).
Patients and methods
Patients and controls
Ninety-seven consecutive, unrelated patients with MA and MoA were included in the study after giving informed consent. The diagnosis of migraine was made according to the criteria of the International Headache Society (11). Every patient received a full neurological and cursory neuropsychological examination excluding patients with neurological deficits or dementia. None of the patients had a family history suggestive of CADASIL. Seventy patients suffered from MoA, 27 patients suffered from MA, of whom 19 had MA as well as MoA. Fifteen male patients and 82 female patients participated in the study. The mean age of the patients was 39 years (14–54 years, SD 13 years, and median 39 years). Ninety-seven age- and sex-matched controls were recruited from a control cohort of healthy blood donors. Patients, as well as controls, were of caucasian ethnicity.
DNA sequencing
Genomic DNA was prepared from peripheral blood lymphocytes. Exons 3 and 4 of the NOTCH3 gene were amplified on genomic DNA by polymerase chain reaction (PCR) using the oligonucleotide primers NOTCH3-Ex3-4F (5′-CGTGTTTCTTGCCT GTCTTGTG-3′) and NOTCH-Ex3-4R (5′-ATTCTGACCCTCAAACCCTAGC-3′). The PCR product was sequenced using the same primers and the primers NOTCH3-Ex4F (5′-AAGAGTCTG GAGGGGAGGTAG-3′) and NOTCH3-Ex3R (5′-GATCTGGCAGGGAGCAGTCAG-3′). Sequencing was performed using the ABI BigDye terminator cycle sequencing kit v3.1, and sequence electrophoresis was performed on an ABI3700 automated DNA sequencer (Applied Biosystems, Foster City, CA, USA). Sequence electropherograms were examined visually using the SeqMan® module of the Lasergene® program package (DNAstar Inc., Madison, WI, USA). The alleles of the two SNPs rs3815188 and rs1043994 were determined from the sequence electropherograms.
Statistical analysis
Allele and genotype frequencies were compared with the χ2 test. If one allele or genotype occurred less than six times, Fisher's exact test was used. Odds ratios (OR) and confidence intervals (CI) for alleleles were calculated for comparisons with a significant difference after correction for multiple testing. Hardy–Weinberg equilibrium and linkage disequilibrium between the two polymorphisms was assessed using the program Haploview 3.0 (12).
Results
Mutation analysis of exons 3 and 4 of the NOTCH3 gene in 97 patients with migraine did not reveal any novel mutations or SNPs. Two known SNPs with the dbSNP accession numbers rs3815188 (C381T with respect to cDNA NM_000435, Thr101Thr) and rs1043994 (G684A with respect t cDNA NM_000435, Ala202Ala) were tested for association with migraine in comparison with 97 healthy age- and sex-matched controls. Genotype and allele distributions and statistical measures of association are shown in Table 1. The genotypes are in very good agreement with Hardy–Weinberg equilibrium for both SNPs in the patient and in the control group (patients: HWpval = 1.0 for rs3815188, HWpval = 1.0 for rs1043994; controls: HWpval = 1.0 for rs3815188, HWpval = 0.74 for rs1043994). There is no strong indication that both SNPs stand in significant linkage disequilibrium with each other (D′= 0.56, 95% CI 0.05, 0.96, R 2 = 0.0). In the unstratified migraine cohort no association was found for genotypes (P = 0.059) of the SNP rs3815188 but weak association was detected for the alleles (P = 0.032), which did not remain significant after correction for multiple testing. Stratification for aura showed that the weak association signal is due to the group of migraineurs without aura (P = 0.029). For SNP rs1043994 we observed significant association with the complete migraine sample for genotypes (P = 0.008) and alleles (P = 0.005). The GG genotype and the G allele of rs1043994 are significantly more common in patients with migraine than in controls and the calculated OR for the G allele is 3.6 (95% CI 1.4, 9.5). Stratification for aura showed again that the association signal is caused by the patients without aura (P = 0.006, OR = 4.5, 95% CI 1.6, 12.9).
Genotype and allele frequency distributions for the NOTCH3 polymorphisms∗
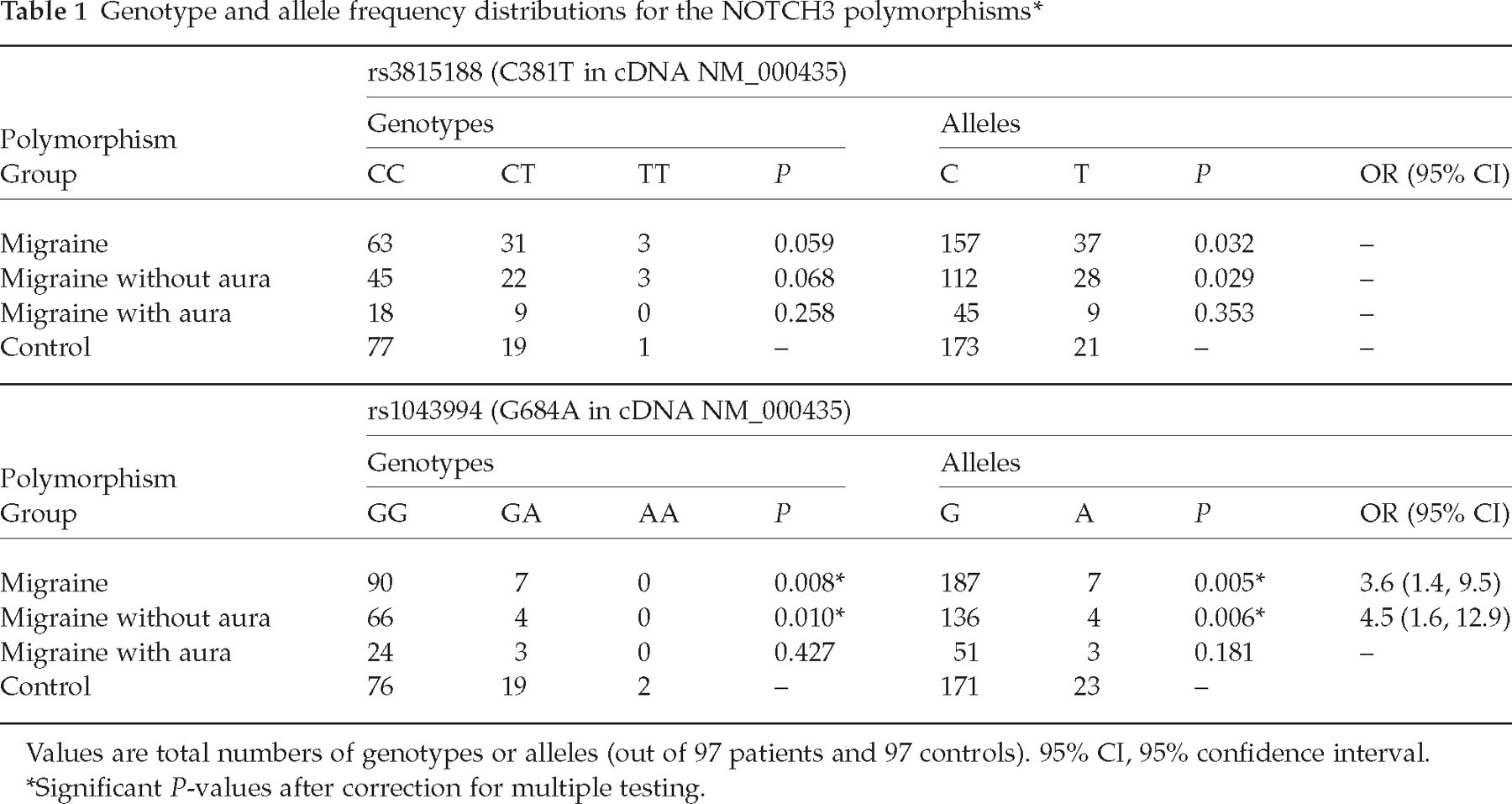
Values are total numbers of genotypes or alleles (out of 97 patients and 97 controls). 95% CI, 95% confidence interval.
Significant P-values after correction for multiple testing.
Discussion
We analysed exons 3 and 4 of the NOTCH3 gene for mutations and association of common genetic variants with migraine, because mutations in NOTCH3 cause CADASIL, a disease sharing some clinical features with migraine. Our study shows that mutations in exons 3 and 4 of the NOTCH3 gene obviously do not play a major role in the pathogenesis of migraine. Novel polymorphisms in exons 3 and 4 of NOTCH3 in 97 migraineurs and 97 controls were not found. However, association analysis with two known SNPs showed significant association of genotypes, as well as alleles, of SNP rs1043994 with migraine and particularly in the subgroup of migraineurs without aura. It is unclear why the association signal stems from the subgroup of patients without aura while most patients with CADASIL do have MA, but it is interesting to note that the weak association between SNP rs3815188 and migraine is also due to the subgroup of migraine patients without aura. However, the study size is to small to allow deriving far-reaching conclusions from the very small subgroups, especially the group of migraineurs with aura comprising only 27 individuals. We did not detect any significant linkage disequilibrium between the two SNPs. This finding could be explained by the fact that the rarer SNP rs1043994, which is significantly associated with migraine, is phylogenetically younger than the more common SNP rs3815188 and arose on a genetic background carrying the latter SNP. The fact that the genotype distributions in both the patient and control groups were in very good agreement with Hardy–Weinberg equilibrium argues against significant population stratification. Genotyping errors are also unlikely because all genotypes were determined by the direct DNA sequencing. The significantly associated SNP rs1043994 does not lead to an amino acid change. It is therefore doubtful that this polymorphism itself is functionally significant, although recent evidence suggests that quite a few seemingly innocent exonic polymorphisms affect the splicing process (13). Alternatively, SNP rs1043994 might be in linkage disequilibrium with another functional variant in NOTCH3, or another gene close by, and this yet unidentified genetic variant could play a role in the pathogenesis of migraine.
CADASIL leads to pathological changes of cerebral vascular smooth muscle cells, leading to thinning and other pathological changes in the wall of cerebral blood vessels. It is therefore conceivable that genetic variants in the NOTCH3 gene render migraine patients more susceptible to the trigeminovascular reflex seen as an initiating event in migraine. These issues can only be resolved by further genetic investigations of NOTCH3 in larger samples of patients with migraine employing linkage disequilibrium mapping.
Footnotes
Acknowledgements
We especially thank the patients who participated in this study.
