Abstract
The past decade has seen impressive advances in neuroimaging, moving from qualitative to quantitative outputs. Available techniques now allow for the inference of microscopic changes occurring in white and gray matter, along with alterations in physiology and function. These existing and emerging techniques hold the potential of providing unprecedented capabilities in achieving a diagnosis and predicting outcomes for traumatic brain injury (TBI) and a variety of other neurological diseases. To see this promise move from the research lab into clinical care, an understanding is needed of what normal data look like for all age ranges, sex, and other demographic and socioeconomic categories. Clinicians can only use the results of imaging scans to support their decision-making if they know how the results for their patient compare with a normative standard. This potential for utilizing magnetic resonance imaging (MRI) in TBI diagnosis motivated the American College of Radiology and Cohen Veterans Bioscience to create a reference database of healthy individuals with neuroimaging, demographic data, and characterization of psychological functioning and neurocognitive data that will serve as a normative resource for clinicians and researchers for development of diagnostics and therapeutics for TBI and other brain disorders. The goal of this article is to introduce the large, well-curated Normative Neuroimaging Library (NNL) to the research community. NNL consists of data collected from ∼1900 healthy participants. The highlights of NNL are (1) data are collected across a diverse population, including civilians, veterans, and active-duty service members with an age range (18–64 years) not well represented in existing datasets; (2) comprehensive structural and functional neuroimaging acquisition with state-of-the-art sequences (including structural, diffusion, and functional MRI; raw scanner data are preserved, allowing higher quality data to be derived in the future; standardized imaging acquisition protocols across sites reflect sequences and parameters often recommended for use with various neurological and psychiatric conditions, including TBI, post-traumatic stress disorder, stroke, neurodegenerative disorders, and neoplastic disease); and (3) the collection of comprehensive demographic details, medical history, and a broad structured clinical assessment, including cognition and psychological scales, relevant to multiple neurological conditions with functional sequelae. Thus, NNL provides a demographically diverse population of healthy individuals who can serve as a comparison group for brain injury study and clinical samples, providing a strong foundation for precision medicine. Use cases include the creation of imaging-derived phenotypes (IDPs), derivation of reference ranges of imaging measures, and use of IDPs as training samples for artificial intelligence-based biomarker development and for normative modeling to help identify injury-induced changes as outliers for precision diagnosis and targeted therapeutic development. On its release, NNL is poised to support the use of advanced imaging in clinician decision support tools, the validation of imaging biomarkers, and the investigation of brain–behavior anomalies, moving the field toward precision medicine.
Introduction
Historically, neuroimaging observations have been limited to gross abnormal findings at the macroscopic level detectable by trained radiologists and have focused on imaging acquisitions that are optimized for qualitative and semi-quantitative detection of disease specific abnormalities. However, the past decade has seen impressive advances in neuroimaging, in terms of quality and speed of acquisition. Available techniques include diffusion magnetic resonance imaging (dMRI), functional MRI (fMRI), and high-resolution structural MRI (sMRI) sequences such as T1-magnetization-prepared rapid acquisition gradient echo (MP-RAGE) that now allow for the inference of microscopic changes occurring in white matter (WM) and gray matter, along with alterations in structure and function. Advances in acquisition hardware and sequences have been complemented by the development of advanced analytic techniques that go beyond conventional slice-by-slice radiological assessment, toward the extraction of detailed volumetric information, number and relationship of WM hyperintensities, insights into connectivity between brain regions, and functional activation maps, which contain valuable information related to patient brain health at a level of abstraction that exceeds human-vision-aided detection. 1 Together, these advances enable a comprehensive interrogation of clinical imaging to identify key biomarkers that indicate presence and severity of injury, as well as markers of prognosis and recovery trajectories that have the potential to provide unprecedented capabilities in discovering and validating diagnostic, prognostic, predictive, and monitoring biomarkers that could be utilized in patient care settings as decision tools for clinicians. Consequently, there has been a proliferation of patient data with advanced neuroimaging acquisitions, along with comprehensive cognitive and behavioral assessments in disorders spanning neuropsychiatry, neurology, neurosurgery, radiology, and rehabilitation.
To fulfil this promise of clinical utility, there is a need for normative data of matching quality. Advanced imaging consists of data that are largely numerical and somewhat abstract compared with conventional imaging and advanced imaging biomarker development calls for the establishment of a normative range of patterns and values, against which patients could be considered. Without a norm by which to compare potentially abnormal scans, clinicians cannot use the information to help in making clinical decisions on diagnosis, prognosis, or treatment pathway options. Normative ranges, in turn, require imaging data on healthy participants spanning a wide demographic, along with a comprehensive medical history and structured clinical assessments, which are logistically challenging to acquire. If comprehensively collected, though, neuroimaging is expected to enhance our knowledge of disease mechanisms in addition to diagnosis, prognosis, and prediction of response to different treatment options. However, the lack of high-quality normative data for brain imaging remains a substantial gap for the field. And the gap is a broad one; the enhanced focus on precision medicine in the past decade has fueled the need for such a normative dataset across brain disorders, including traumatic brain injury (TBI), depression, post-traumatic stress disorder (PTSD), and neurodegenerative disorders such as Alzheimer’s disease and Parkinson’s disease. A number of publicly available neuroimaging databases (see Table 1, including expansion of all dataset name abbreviations) exist that consist of neurologically typically developing individuals. While these databases share the intent of acquiring age-stratified, normative data to provide a contextual framework for the diagnosis of neurological disease and are useful for the purpose for which they were collected, they each have their limitations. These include a limited age range (e.g., primarily youth in Adolescent Brain Cognitive Development [ABCD] or older adults in Alzheimer’s Disease Neuroimaging Initiative [ADNI]) and acquisition of specialized sequences that do not align with typical acquisition parameters in clinical practice (e.g., Human Connectome Project [HCP]) or lack adequate characterization of cognitive and psychological functioning, particularly in cases of brain injury (e.g., Philadelphia Neurodevelopmental Cohort [PNC] and UK Biobank [UKBB]). Existing large healthy control datasets (e.g., HCP and UKBB; see Table 1) are also not well curated in terms of medical history and clinical assessments, making their selection for the purposes of a control cohort challenging (e.g., the methods used to rule out a TBI are not comprehensive and sometimes not stated at all; other comorbidities will be challenging to rule out). Importantly for TBI, screening for or thorough assessment of TBI is not included in many of these datasets. For example, in the Nathan Kline Institute–Rockland Sample dataset, if the participant is currently having symptoms and thinks of it as a medical condition, a TBI might be recorded in response to the question, “Does the participant have any illnesses or health problems at present?”. Otherwise, no specific question about a history of TBI is asked. Additionally, normative cohorts seldom have blood sample collection. These gaps are very relevant to TBI where MRI and, in particular, advanced quantitative MRI techniques, such as diffusion-tensor imaging (DTI) and fMRI, are considered important noninvasive tools that could elucidate TBI disease mechanisms, yield diagnostic biomarkers, and/or serve as a basis for patient stratification or endpoints in therapeutic clinical trials. Furthermore, because of the heterogeneity of TBI, it is likely that the optimal tool for assessing TBI will involve multimodal components—specifically, blood-based and physiological biomarkers coupled with advanced neuroimaging obtained at multiple time points.
Datasets That Have Structural, Diffusion, and Functional MRI Sequences
ASL, arterial spin-labeling; BOLD, blood-oxygen-level-dependent; EPI, echo-planar imaging; FLAIR, fluid attenuated inversion recovery; fMRI, functional MRI; MP-RAGE, magnetization-prepared rapid acquisition gradient echo; MRI, magnetic resonance imaging; NKI-RS, Nathan Kline Institute–Rockland Sample; SWI, susceptibility-weighted imaging; TBI, traumatic brain injury. DSI, diffusion spectrum imaging; DWI, diffusion weighted imaging; GRE, gradient recall echo; IR/SPGR, inversion recovery/spoiled gradient recall echo; MRA, magnetic resonance angiography; MT, magnetic transfer; NITRC, neuroimaging Tools & Research Collaboratory; PASL, pulsed arterial spin labeling; PD, proton density; SPACE/CUBE/VISTA, 3-D GRE branded software from Siemens/GE/Philips; TOF, time of flight; TSE, turbo spin echo; VASO, vascular-space-occupancy.
Finally, despite an increased use of tools that offer the potential to harmonize neuroimaging data across datasets, the cognitive and psychological assessments vary extensively. This heterogeneity poses a challenge when trying to bring the datasets together to overcome some of these issues, such as age range limitations, or to investigate the relationships between brain structure and function.
In addition to the normative databases, there are existing biomarker discovery datasets in patients (as in Translating Research and Clinical Knowledge Traumatic Brain Injury) that have limited numbers of healthy participants (see Table 1 for details of the datasets). The focus of these datasets was acquisition of patient data, and the normal data are not enough to form the reference data essential for normative modeling to detect disease-specific patterns useful in the clinic or for biomarker validation. The Normative Neuroimaging Library (NNL) has been strategically conceptualized and designed to fulfil these gaps, in addition to laying a solid foundation for multimodal biomarker discovery through the addition of blood sample collection in a subset of this normative population. While applicable across many brain disorders, at its core, the NNL design supports the validation of TBI neuroimaging biomarkers by inclusion of TBI-relevant medical history, clinical outcomes, and neurocognitive function assessments with applicable TBI CDEs and TBI-relevant scan sequences including T1 MP-RAGE, T2 fluid attenuated inversion recovery (FLAIR), susceptibility-weighted imaging (SWI), DTI, resting state fMRI, and perfusion-weighted imaging. In this article, we provide an overview of the NNL, its conceptualization, design and implementation, as well as the current status of data collection, curation, and harmonization.
In August 2013, the National Academy of Sciences Neuroscience Forum convened a workshop focused on developing standards for dMRI including co-chairs from NINDS, Veterans Administration (VA), and Cohen Veterans Bioscience (CVB). 2,3 This prompted the American College of Radiology (ACR) to convene a consensus panel in May 2014, at the American Society of Neuroradiology meeting in Montreal, Canada. 4 At this meeting, 55 experts examined issues related to the current status of normative neuroimaging and identified critical gaps in the field. The group recognized that most normative data exist as comparison samples for research 3 and are recruited for their similarity to the experimental population in question. Furthermore, the group determined gaps in existing normative neuroimaging data including (1) scanner harmonization, (2) rigor of subject screening, and (3) inclusion of standard sequences for advanced imaging. Variables recognized to be influencing the normative data include (1) age, (2) sex and gender, (3) race and ethnicity, (4) handedness, (5) socioeconomic status, (6) general intelligence/cognition, (7) psychological health, and (8) general medical history. Based on the above gaps in normative data and the panel recommendations, the ACR and CVB sponsored the creation of a prospectively established NNL that would serve as an open-source normative database. The NNL has been designed to include demographic information, medical history, and standardized cognitive and psychological assessment measures packaged with multi-site harmonization approaches to align sequence acquisitions, account for vendor-related and hardware-related acquisition differences, and include frequent acquisitions of phantom data from both in vitro (manufactured phantom models) and in vivo (the same human being scanned at each site) models.
In summary, the uniqueness of NNL is highlighted by (1) its comprehensive structural and functional imaging acquisition with state-of-the-art sequences; (2) collection of comprehensive demographic details, medical history, and a structured clinical assessment (comprising evaluation of performance validity, cognition, psychological functioning, and substance use) relevant to multiple neurological conditions with functional sequelae; and (3) data collected across a diverse population, including civilians, veterans, and active-duty service members. The collection of these data was monitored to ensure stratification by age-group, and as broad as possible demographic characteristics and social and economic determinants, with an intent to reflect the population diversity in the regions of data collection sites. Standardized imaging acquisition protocols across sites reflect sequences and sequence parameters often recommended for use in adults with TBI and various neurological and psychiatric conditions, including PTSD, stroke and other cerebral vascular disorders, neurodegenerative disorders, and neoplastic diseases. An additional unique advantage of the NNL is the preservation of raw scanner data, which allows higher-quality data to be derived in the future. For example, compressed sensing would allow the acquisition of images with fewer samples and their reconstruction using sparsity-driven algorithms, reducing scanning and storage needs. Additionally, deep learning techniques applied to raw scanner data may improve image quality through noise reduction, artifact correction, and resolution enhancement, resulting in superior images. As new algorithms are developed, the existing raw data can be reused to apply these advancements, ensuring that the NNL remains a valuable resource. Finally, the rigorously curated NNL data will include imaging-derived phenotypes (IDPs) ready for normative modeling and other analyses.
This article aims to introduce NNL to the TBI research community. We begin by comprehensively detailing the existing normative datasets, placing NNL in perspective. We then describe the study design with details of the sample, imaging acquisitions, and clinical assessments. Finally, we discuss use case examples that demonstrate the broad utility of this dataset.
Materials and Methods
Existing normative datasets and motivation for NNL
There are several existing normative dataset, which are detailed in Table 1. The spelled out names of each database abbreviation are also provided in Table 1. The examination of the strengths and weaknesses of these databases reveals that most of them are cross-sectional (UKBB, HCP, PNC), with the focus being a single age-group (children [ABCD], young adults [PNC, The Pediatric Imaging, Neurocognition, and Genetics], or adults [HCP, UKBB]), with ages 25–40 years being under-sampled. Additionally, they do not have the depth of clinical evaluation needed for them to be a comparative dataset for patient samples with neurological and neuropsychiatric disorders. While PNC and UKBB are exceptions, the former is targeted toward psychiatric disorders in children and young adults, and the latter includes a variety of disorders with no detailed collection of clinical assessment scores for each nor well-defined exclusionary criteria for healthy controls. Additionally, some of these datasets fail to capture adequate representation of demographic and socioeconomic diversity or, for those actively collecting samples in 2020 through today, to have coronavirus disease-19 (COVID-19)-related information. Notably, none of the existing databases actively recruit individuals with a military background. In summary, most of these datasets cannot be used as a normative sample for clinical investigations focusing on discovering brain anomalies.
To confirm that the NNL has clinical utility, CVB convened two virtual roundtable meetings on October 1 and November 17, 2020, across a multidisciplinary stakeholder community. The roundtables included 47 individuals representing prominent academic and private research institutions, biopharmaceutical companies, medical technology and software companies, nonprofit organizations, and government agencies, including the U.S. Food and Drug Administration, VA, Department of Defense, and National Institutes of Health (NIH) (see Supplementary Data S1 for a full list of the attendees).
Attendees provided use cases for the NNL relevant to advancing research or clinical care in their sectors. Example use cases were presented from a variety of perspectives, including a diagnostics company, a health care company, and a contract research organization. The use cases centered on (1) determining eligibility for enrollment in future therapeutic clinical trials, (2) stratifying subjects within a clinical trial, (3) assessing which structural and functional characteristics track with disease progression, (4) enabling assessment of disease trajectory, (5) providing guidance for appropriately powering a study, and (6) providing a control cohort for development of an imaging-based diagnostic. Other potential applications of the NNL include the development, testing, and validation of deep-learning-based algorithms and testing of advanced diffusion processing tools.
Given the time frame of imaging collection, from 2017 going forward, researchers also decided to add evaluations of COVID-19 exposure and any long-lasting symptoms that could potentially impact the assessment of “normal.” Based on the Roundtable recommendations, a collection of bio-samples and additional clinical assessments was added to the NNL design in June 2021.
Study design
Population overview
Based on the recommendations of the ACR Working Group and the gaps identified in existing databases, we endeavored to select a sample with representative diversity in terms of geographic location (including rural areas), racial/ethnic background, gender/sex, education level, and socioeconomic status. In addition to civilian participants, our sample also includes both U.S. active service members and veterans. Recruitment included traditional advertisements (radio announcements, posted flyers), advertising on social media, and word of mouth. A central commercial institutional review board (IRB; Advarra) reviewed and approved study procedures and site-wise institutional IRBs relied on the Advarra IRB for direct study oversight while maintaining awareness of the protocol and all materials shared with participants. Participants consented to their data being used for future research and also to be contacted for longitudinal data collection in the future. All consent forms are archived at the clinical sites.
Sample size determination
Determination of the target sample size was guided by precision analyses of WM-based sequences and was based on the population-based variance of normal values. In the absence of a well-established variance estimate, standard power analyses to determine sample size could not be utilized. Therefore, we set the sample size at the point of clear diminishing returns for the maximization of precision in a given population using a 95% confidence interval. To determine this, we used the Nathan Kline Institute dataset 5 consisting of DTI scans from ∼200 individuals acquired with a 60-direction protocol. We assessed the mean and standard deviation of fractional anisotropy of the whole brain WM (see Fig. 1). Approximately 600 participants maximized the precision of the assessed mean and standard deviation. Assuming a normal distribution of values, we determined a total of 3000 individuals would be required for recruitment into each of five specific age-groups. This resulted in the inclusion of 600 participants in each of the following age cohorts (in years): (1) 18–25, (2) 26–35, (3) 36–45, (4) 46–55, and (5) 56–64. Individuals under the age of 18 years and over the age of 65 years were not included due to existing datasets, such as the ABCD and ADNI-3, which focused on these age-groups, and to increase the feasibility of successful project completion.
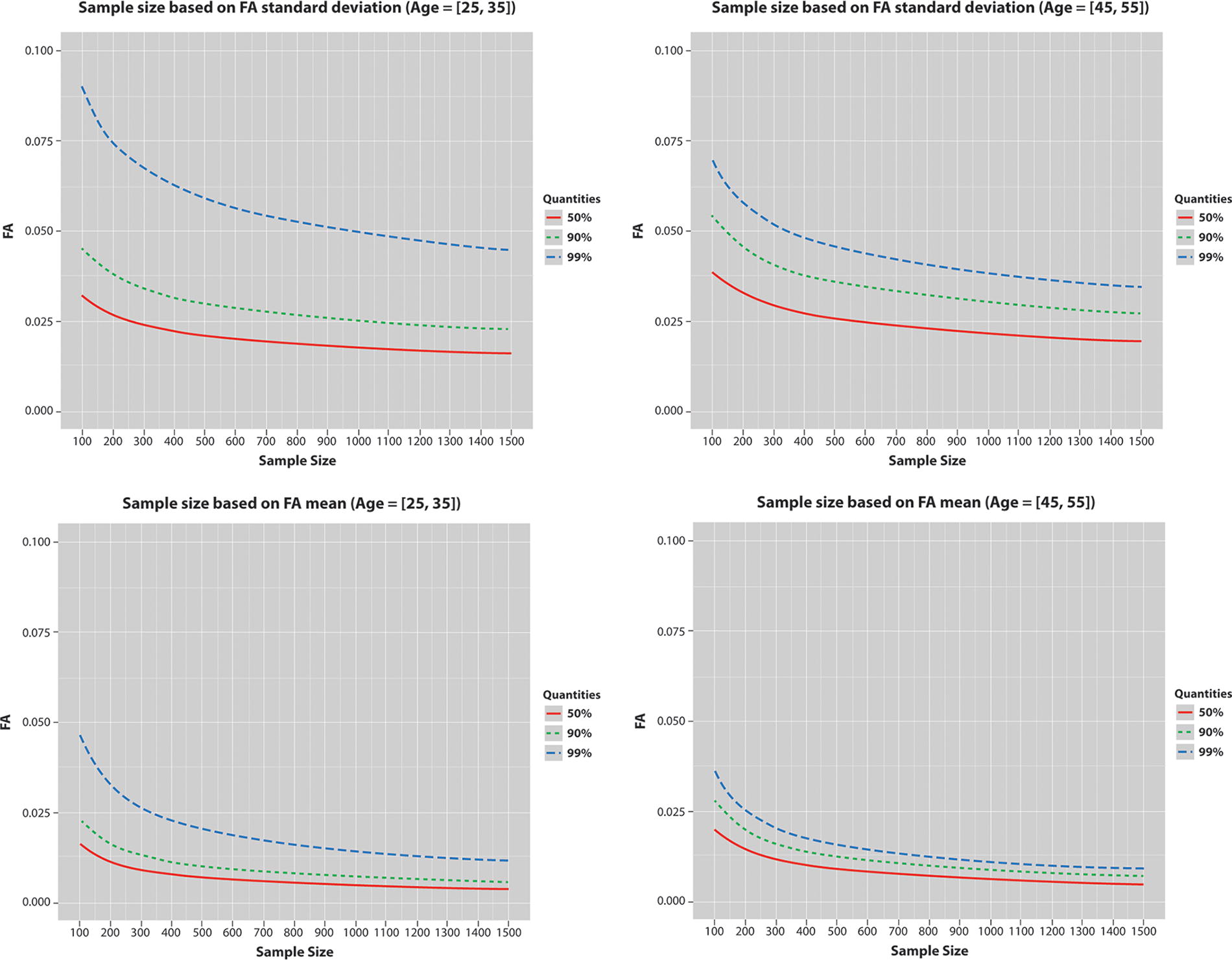
Analysis of the dMRI data from the NKI dataset to determine the sample size for NNL. The top and bottom rows show the variance and mean of the fractional anisotropy (FA) of the whole brain white matter computed in the participants in the age ranges 25–35 and 45–55. The optimal precision is observed starting at around ∼600 participants. dMRI, diffusion magnetic resonance imaging; NKI, Nathan Kline Institute; NNL, Normative Neuroimaging Library.
Inclusion and exclusion criteria
The NNL includes typically developing, healthy adults (18–64 years of age). No exclusions from participation in this study are made based on sex/gender, race, or ethnicity. All data collection was conducted in English due to the standardization and normative data related to the outcome measures. The exclusion criteria were established to reduce any likely confounding variables on MRI related to abnormal development, aging, and brain trauma. Participants were excluded during collection of medical history based on self-report of a TBI with a loss of consciousness of more than 5 min or a memory gap of more than an hour. To further prompt recall on prior brain trauma, participants were queried regarding past accidents and participation in sports to aid in recalling the presence/absence of concussions. They were also queried about whether they ever had any traumatic fractures or other non-fracture-related traumas that resulted in hospitalization. In addition, they were queried about common TBI comorbidities such as PTSD, seizures, multiple types of headaches, or chronic pain. In cases of exclusionary possible TBI, coordinators would review cases with their principle Investigator (PI), and, when needed, all three PI came to consensus on whether a participant would be included or excluded. Exclusion criteria were not designed to restrict the range of symptom reports on the various psychological, behavioral, and cognitive assessments that would be seen in a representative population. See Table 2 for the detailed exclusionary criteria.
Exclusionary Criteria
Probing for any implanted metal objects (pins, screws, clips, shrapnel, etc.).
MRI, magnetic resonance imaging; PTSD, post-traumatic stress disorder; TBI, traumatic brain injury.
Data acquisition
All assessments and procedures were planned to be completed in a single study visit, though, acknowledging real-world situations, participation spanning two or more sessions was allowed. The summary of the total sample to date can be found in Figure 2.
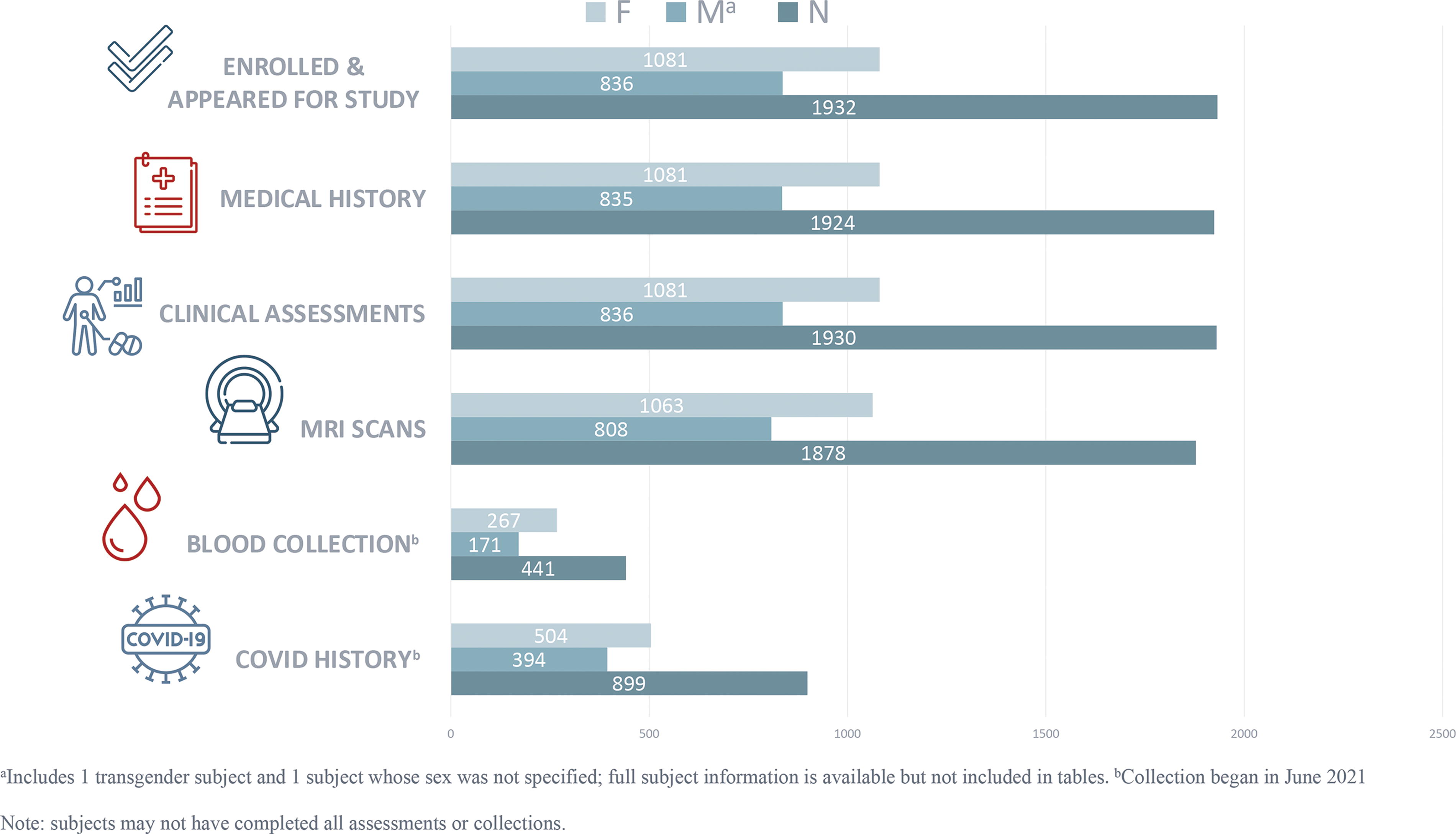
Summary of NNL enrolled subjects’ and assessments’ disposition. NNL, Normative Neuroimaging Library.
Demographic characteristics and medical history
Basic demographic information using recommended National Institute of Neurological Disorders and Stroke common data element variables including age, sex/gender, race, ethnicity, and other characteristics are collected in a standard format for research data (developed by Clinical Data Interchange Standards Consortium; CDISC) ensuring compatibility with other datasets (see Table 3). Additionally, nonexclusionary prenatal and medical history are collected to enable users of the database to filter various medical conditions or exposures and to understand the relationship to any imaging changes (see Table 4). Similarly, all imaging scans were reviewed, and abnormalities and incidental findings recorded for both exclusionary purposes and a better understanding of these findings in a representative normal population.
Basic Demographics of the NNL Cohort
Subjects endorsing multiple races are counted in more than one category; ethnicity information is available for all subjects.
NNL, Normative Neuroimaging Library; SD, standard deviation.
Distribution of Nonexclusionary Medical Conditions
Clinical assessments
Based on Working Group recommendations made at the ACR meeting, we selected a brief battery of widely used measures with validated psychometric properties to assess cognition (including domains of general intellectual functioning, attention, visual and verbal memory, executive functioning, processing speed, and performance validity) and psychological functioning (including symptoms associated with PTSD, depressed mood, anxiety, bipolar disorder, psychotic disorders, and attention-deficit/hyperactivity disorder). We also assessed postconcussive symptoms, pain, sleep, and substance use. The details of the measures can be found in Table 5.
Overview of Clinical Measures Administered to Study Participants
Exclusions:
TOMM < 45 across two trials.
WASI II IQ <80.
MDQ “positive screen.”
ADHD, attention-deficit/hyperactivity disorder.
Objective assessments for biomarker development
Bio-sample collection was not a part of the initial NNL design. However, during the NNL Roundtable, it was agreed that the addition of other potential biomarker modalities could strengthen the ability to validate the imaging biomarkers, as well as provide the ability to derive multimodal biomarker signals that might better describe a disease population subtype than a single biomarker alone. In addition, the collection of a standard blood sample for laboratory value reporting would add to the value of the library by better characterizing the “normal” or “representative” population. To this end, the protocol and informed consent form were amended to allow for collection of a fasting blood sample. Here, ∼55 mL of blood is collected to have sufficient sample for immediate measurement by a single clinical laboratory and the remainder stored at CVB’s biorepository. The stored samples are conserved for future investigations involving the characterization and validation of disease biomarkers, as discovered and validated. Participants entering the study after IRB approval of this protocol amendment provided their consent to the amended informed consent document.
Participants completed a bio-sample collection form that provides information important for both the processing of the potential future candidate biomarkers and their interpretation. These fields on the form include the time of the collection, participant’s vital signs at time of the collection, and, in the instance that they did not come to the clinic in an 8-h fasted state, questions regarding the types of foods they might have consumed in the past 8 h. Finally, information was collected about when they woke up that morning, smoking history, any exercise that day prior to coming into the clinic, and, for female participants, questions about their menstrual cycle. Seven biospecimen tubes were collected by the site’s certified phlebotomist, including serum, plasma, buffy coat, ribonucleic acid, and whole blood samples for proteomic, metabolomic, genomic, transcriptomic, and other analyte studies. Whole blood and serum samples were shipped to the central laboratory the day of the draw and all other samples were kept frozen at −80°C until they were batch shipped on dry ice for longer-term storage at CVB’s biorepository. The clinical laboratory samples were processed within 48 h of collection, and the deidentified results are included as a part of the clinical database.
MRI acquisition and standardization
All participants are screened to determine safety to undergo MRI based on local standard operating procedures. NNL collected data at three sites across the United States: (a) University of Utah using a Siemens 3T Prisma Fit with software version VE11C, VE11E, and XA30 over time and a 64-channel head/neck coil; (b) University of Virginia using a Siemens 3T Prisma Fit 20 with software versions VE11C andVE11E and a 64-channel head/neck coil: and c) Baylor College of Medicine using a 3T Siemens Prisma with software versions VE11C and VE11E and a 64-channel head/neck coil. The goal of the NNL imaging protocol was to capture both the structure and the function of the brain via anatomical, diffusion, and functional imaging in such a way that the data can be integrated across the three sites. To achieve this goal, the NNL instituted a highly standardized and harmonized blend of readily available contrast agent-free imaging sequences and state-of-the-art elements, which are outlined below.
Structural MRI
sMRI sequences provide anatomical information. The following sequences were acquired: High-resolution (1 × 1 × 1 mm3) T1-weighted imaging, optimized for human-brain tissue (WM, gray matter) contrast, is achieved with the MP-RAGE pulse sequence.
6
Using this sequence, a participant’s data can be aligned to a standard template brain (such as the Montreal Neurological Institute brain)
7
and segmented into anatomical regions, to obtain volumetric information. This is also utilized in statistical analysis of cortical ribbon thickness.
8
High-resolution (1 × 1 × 1 mm3 voxels) T2-weighted and CSF-suppressed (i.e., FLAIR) T2-weighted imaging is accomplished with an optimized 3D fast/turbo spin-echo pulse sequence (on the Siemens scanners, SPACE—Sampling Perfection with Application optimized Contrasts using different flip angle Evolutions), which allows for assessment of lesions within the brain.
9,10
The T2-weighted image set is also used in conjunction with the T1-weighted image set to achieve more precise brain segmentation using a multispectral technique.
11
SWI with 0.9 × 0.9 × 1.5 mm3 voxel resolution is also acquired to assess microbleeds while differentiating against calcification.
12
Diffusion MRI
dMRI is a specific MRI sequence that encodes the differential water diffusivity in the brain between the different tissue types, with the flow being maximally hindered in the WM. 13 Given that water diffusion is affected by thermal agitation and highly dependent on the cellular microenvironment, any change in the dMRI-derived measures may indicate pathological change. As WM in the brain has a fibrous structure due to the presence of axons, water will diffuse more rapidly along the direction aligned with the internal structure and slower in a direction perpendicular to it. This property can be exploited to derive measures representative of microstructural integrity and connectivity. dMRI-based analysis is commonly used to assess anomalies in WM in brain connectivity disorders such as TBI, bipolar disorder, and PTSD. In NNL, high-resolution (1.5 × 1.5 × 1.5 mm3), two-shell (b = 1500 s/mm2 and b = 3000 s/mm2), high-angular (90 directions) diffusion-weighted imaging (TR: 4100 ms; TE: 90 ms) was acquired using spin-echo-based echo-planar imaging (EPI). This acquisition allows for a detailed assessment of WM tracts within the brain. The diffusion imaging protocol for NNL matches that for another large consortium, the Lifespan portion of the HCP. 14 The diffusion protocol goes beyond the once-standard single-shell DTI to allow for assessment of non-Gaussian processes 15 and measurement of complex microscopic geometries, such as crossing fibers, 16 that are crucial for studies in TBI and other connectivity disorders. High through-plane (i.e., slice-select direction) resolution was achieved by leveraging simultaneous excitation of multiple slices, known as simultaneous-multi-slice or multiband imaging. 17
Functional imaging
Brain function is assessed through two functional imaging sequences: Resting-state fMRI (rs-fMRI): Rs-fMRI is a specific MRI sequence that measures brain activity by detecting associated changes in blood flow in the brain that creates a blood-oxygen-level-dependent (BOLD) signal and is used to evaluate regional interactions occurring in the resting state (when a task is not being performed). As brain activity is intrinsic, present even in the absence of an external task, any brain region will have spontaneous fluctuation in the BOLD signal. Rs-fMRI is useful to explore the brain’s functional organization, and several networks have been identified, such as the default mode network.
18
The resting-state nature of the acquisition ensures that the data can be collected in a range of patient groups, even those with cognitive and psychological deficits, as in TBI and PTSD. Rs-fMRI is also more generalizable across disorders, unlike task-based fMRI that is disorder specific. In NNL, a 10-min rs-fMRI acquisition is obtained using a state-of-the-art gradient-echo-based EPI pulse sequence that included simultaneous-multi-slice excitation, as also used for diffusion imaging. This allowed for high spatial resolution (2 × 2 × 2 mm3) and a fast volume acquisition time (0.80 sec). The rs-fMRI sequence parameters also match those of the HCP.
14
Arterial spin labeling: Tissue perfusion measurement is collected using a 3D pseudo-continuous arterial spin-labeling pulse sequence. Due to its noninvasive nature, this type of acquisition greatly lowers the barrier for research sites to engage in perfusion measurements. Perfusion allows the assessment of microdamage to arterials caused by shearing forces associated with even mild TBI and assessment of hypoperfusion in both acute and chronic stroke and shows alterations in the absence of visualized structural changes, which may help in subtyping psychiatric conditions such as PTSD and psychosis.
19,20
Additionally, perfusion measurements prove useful when used in conjunction with cortical thickness measurements to better understand the origins of any observed differences in resting-state-derived brain networks between control groups and patient groups.
Data consistency
To confirm scanner performance and accuracy and ensure the viability for retrospective harmonization across diverse neuroimaging datasets, stringent quality control measures were employed, involving the regular collection of phantom data. Phantoms, which are specifically designed objects used as stand-ins for the human body in medical imaging, offer a means of standardization and calibration. They help ensure the consistent performance of the imaging equipment across sites and over time and are important in mitigating potential sources of systematic error.
In this study, data on three phantoms (ACR, ADNI, and Biomedical Information Research Network [BIRN]) were collected monthly at every site. The regular collection of this data facilitates the detection of any subtle changes in scanner performance over time and aids in standardizing images across different scanners. Each phantom provides a different advantage for the possibility of harmonizing across sites. The ACR phantom provides gross indications of geometric fidelity, regular grid pattern, and a contrast array. The ADNI phantom is used for measuring SNR, contrast-to-noise ratio, and geometric stability. The BIRN phantom assesses scanner stability for imaging using BOLD sequences for connectivity analysis.
Additionally, data were also collected annually on a human phantom during the first 3 years across all sites. Here, a human phantom refers to an individual whose brain images are obtained on all scanners within a short period of time, capturing the complexity of the human brain and the variability across and within scanners for harmonization that hardware phantoms do not.
Through these measures, this study aims to ensure that data across all sites can be reliably harmonized retrospectively, thereby improving the consistency, reproducibility, and validity of the imaging data and any subsequent conclusions drawn from it.
Data flow of NNL
Prior to the NNL Roundtable and subsequent protocol amendment, the participating clinical sites had successfully enrolled almost 900 participants. Tables 3–5 present the key demographics and baseline characteristics that have since expanded to include an additional ∼900 participants. A significant portion of these new participants’ data contributes to blood sample collection, additional psychological assessments, and a record of COVID history and symptoms. As shown in the tables, the NNL population comprises ∼56% female, has an average age of 36.5 years (with a range from 18 to 64), and is ∼83% White. Years of education range from 11 to 22, and composite Wechsler Abbreviated Scale of Intelligence, 2nd edition range from 54 to 160. The uniqueness of this dataset is exemplified by the rigorous third-party monitoring of data collection ensuring high data quality and completeness. All imaging incidental findings and nonimaging and non-bio-sample clinical data from all subjects are entered into an electronic database capture (EDC) system (Medidata RAVE®) built to exclusively accept deidentified data and will automatically generate query for out-of-range values. The relative dates for all clinical assessments, biospecimen collections, and MRI acquisitions are meticulously documented to accommodate any scheduling or logistical adjustments, scanner issues, or other influencing factors. A data management monitoring plan ensures prespecified source document verification and manual querying of the data in EDC for missing data entries.
The monitored, deidentified data, along with the MRI scans, will be transferred to the BRAINCommons™ (BC). 21 BC facilitates access to multimodel data, advanced tools, and a secure interoperable system for data sharing and analysis. Only data from participants from whom two main data types (i.e., demographic and imaging, clinical and imaging, demographic, and laboratory values) were acceptably obtained will be transferred. During the ingestion process, BC ensures that submitted data undergo curation and harmonization to the BC Data Model. This model, designed to maintain data and metadata consistency while accommodating the diversity of heterogeneous data types, adopts data standards such as CDISC and CDEs, such as those defined by NINDS, at the nodal level. These standards establish uniform clinical research and terminology practices. Additionally, the imaging data undergo a quality control process. The NNL dataset will be made accessible to all qualified BC users, who can conduct analyses using their preferred analytical tools and pipelines as well as those provided by the BC. Within the BC environment, researchers have the flexibility to analyze the NNL data independently or integrate this normative data with their own data or other available datasets on the platform.
Discussion: Real-World Use Cases of NNL
Large-scale datasets are designed as biomedical resources for exploring scientific questions related to human health and disease. This epidemiological strategy supports both general research efforts as well as those which target specific populations. An example of the former category is the UKBB, which facilitates the exploration of potential correlations between sociodemographic, genetic, and imaging data and led to the establishment of various normative population trends such as correlations of brain structure and general intelligence, 22,23 global neuroanatomical volumetry, 24 and age-related hippocampal atrophy. 25 Relatedly, UKBB resulted in studies demonstrating the neuroanatomical structural effects of lifestyle and environmental factors such as alcohol, 26 cigarette smoking, 27 cannabis, 28 and SARS-CoV-2. 29 Finally, UKBB has been used for the development of novel assessments, such as the Recent Depressive Symptoms score 30 and profiling of modern image analysis techniques. 31 As an example of the latter strategy of data collection to target specific populations, the ADNI study 32 focuses on the development of analysis strategies for early detection of Alzheimer’s disease and supporting its intervention and treatment. ADNI has led to specific clinical benefits (e.g., drug development 33 ) and technological advancements in terms of image acquisition, 34 phantom development, 35 and Alzheimer’s disease-specific image analysis algorithms. 36
The uniqueness of NNL lies in its potential for both general research and for biomarker discovery efforts in specific target populations. Specifically, the following are use cases discussed at the Imaging Roundtable: The combination of rich imaging information with clinical assessments forms a basis for generating reference intervals for biomarker discovery studies and as a comparator dataset for the development of imaging-based diagnostic tools. Several IDPs will be extracted from the rich imaging data acquired as part of NNL. These include brain tissue volumetrics, cortical thickness WM hyperintensity identification (from sMRI), microstructural integrity measures (from dMRI), and structural and functional connectivity (from dMRI and fMRI). These IDPs will prove valuable to clinicians and researchers in need of normative IDP samples and statisticians seeking to address questions of statistical or sampling bias in clinical trials who lack access to a team of sophisticated image analysts. With these IDPs, combined with the comprehensive clinical assessments administered to study participants, NNL has the potential to serve as the basis of power and effect size calculations for research studies and clinical trials where imaging is priority endpoint, as well as for studies investigating brain–behavior and structure–function relationships. With the increasing number of harmonization tools that are available,
37
–39
NNL imaging data can be combined with data from HCP and UKBB and other datasets in Table 1. This will expand the normative database for clinical comparative studies and refine the accuracy of reference interval definitions. With the goal of precision medicine, disorders need to be conceptualized as deviations from normative functioning, enabling a dimensional analysis to study heterogeneous presentations. This requires accurate estimation of “healthy” brains, which can be achieved by normative modeling techniques
40
–45
applied to NNL. The patients can then be quantified as “outliers” based on their degree of pathology. An imaging marker that is gaining prominence is “brain age,” in which information from multiple measures is fused to predict the age of individuals, implicitly defining normative maturation patterns. The deviation of the brain age of an individual from their biological age has been shown to be associated with the extent of pathology in TBI and Alzheimer’s disease. NNL will provide a basis for the training of a brain age predictor combining imaging, clinical assessments and blood markers, and an insight into which modalities are most suited for the same. With the harmonization with other datasets like HCP and UKBB (as mentioned by Institute of Medicine
3
), a normative model of the lifespan can be created. Given that NNL provides medical history and structured clinical assessments for each participant and bio-sample collection in a significant subset, it supports supervised and unsupervised data analysis methods. This allows for the identification of imaging patterns associated with diverse cognitive and psychological scores, as well as comorbidities. As such, NNL can serve as a discovery or validation dataset for developing multimodal approaches of normative modeling. New studies can align their imaging and clinical assessment protocol with those of NNL, allowing the NNL dataset to augment their normative sample by harmonization. This alignment will encourage standardization of study protocols.
These use cases highlight the significant contribution that NNL can make to patient-specific research.
Conclusions and Future Work
This article presents the motivation and design behind the NNL, detailing the breadth of the data that have been collected to date. Harmonization and processing of the data collected to date are ongoing. While several enhancements have been made to the study based on feedback from the 2020 Expert Imaging Roundtables, future plans include having participants return for the collection of longitudinal timepoints ∼1 year from their initial screening, Additionally, there is an intent to extend study population to better represent active-duty military and veteran populations. When employed to establish threshold norms, these enhancements will help elucidate differences, if any, in the biological underpinnings of PTSD due to military-related and non-military-related trauma, differences in TBI due to blast versus other mechanisms of injury, as well as differences in progression over time.
Thus, NNL addresses the significant unmet need for a standardized normative neuroimaging library complemented by clinical assessments. The inclusion of clinical assessments with MRI can advance imaging biomarker validation potentially enabling precision diagnostics and targeted therapies. Depending on the disease population of interest, specific matched controls can be selected for the creation of reference intervals. NNL distinguishes itself by offering well-defined data dictionaries, preserving raw data, an attribute often lacking in other large datasets, and offering the ability to harmonize with other databases. On its release, NNL will be a large, well-curated normative imaging and behavioral dataset. This dataset, by supporting the validation of imaging biomarkers and the investigation of brain–behavior anomalies, is poised to move the field toward precision medicine. These objective markers of disease can help subtype TBI patient populations, for example, by disease mechanisms rather than symptom presentation, and lead to the discovery and validation of therapeutics specific to each TBI subtype. Longitudinal data collection, as well as the addition of new data collection modalities that are relevant to TBI and do not have existing normative databases, (e.g., voice markers and EEG) is being planned based on funding opportunities.
Transparency, Rigor, and Reproducibility Statement
The study design was finalized and signed-off by CVB (sponsor) prior to submission to the IRB. Subsequent changes to the study design were documented in a protocol amendment and a revised protocol that were also signed off by the sponsor and submitted to the IRB. These methods are being registered via this publication. The sample size was set at 3000 healthy volunteers, with ∼600 per age stratification, based on a DTI dataset showing no further meaningful gain in information beyond this number. Approximately 1930 individuals have been screened to date, and of those deemed eligible, 1878 had MRI scans completed. Investigators who performed clinical and biomarker assessments were trained for consistency and rigor protocols at the start of the study, as well as at interim timepoints during the conduct of the study. These included a single detailed imaging protocol, use of monthly phantoms for recalibration, and a blood collection and processing standard operating procedure.
Imaging data were acquired between August 18, 2017, and October 25, 2022, with acquisition times fit into each clinical site’s workflow. Imaging data for the whole population were collected using three different scanners: Siemens 3T Prisma Fit with software versions VE11C, VE11E, and XA30; Siemens 3T Prisma Fit 20 with software versions VE11C and VE11E; and Siemens 3T Prisma with software versions VE11C and VE11E. The scanner manufacturer, model, and software version for each participant are documented in the dataset.
Variability between devices was assessed using both multiple in vitro phantoms monthly at each site and an annual human phantom across all sites. Variability was prospectively reduced using the same standardized imaging protocol and training across all sites. Inadequate acquisition causing imaging data to not be included has occurred to date in a small percentage of participants, most often due to participants having contraindications to MRI discovered after beginning the study, for example, claustrophobia and not fitting in the machine.
Blood samples were collected in a fasted state and, if not possible, a detailed accounting of food consumed in the prior 12 h taken. All collected blood samples were stored at a single biorepository for assaying all at once to remove batch effects.
The key exclusion criteria were conditions or events expected to impact normal brain imaging (e.g., prior traumatic brain injuries, premature birth, or fetal alcohol syndrome) and are established standards in the field. The battery of clinical assessments used to characterize the study population includes established standards in the field, and references are provided in the article. Key inclusion criteria and clinical assessments were conducted by investigators and designated study personnel with professional qualifications, specific training, certifications, and extent of experience as required in the study protocol. The normal distributions of primary clinical assessments were assessed and verified at an interim timepoint when data from ∼800 participants were available.
Once completed, deidentified data from this study will be accessible to qualified researchers in the BRAINCommons™, a data-sharing platform that aligns with FAIR Guiding Principles.
Common Data Elements as defined by the Federal Interagency Traumatic Brain Injury Research Informatics System (source: https://fitbir.nih.gov/chronic-tbi-related-neurodegeneration-cdes); the Phen Toolkit (source: https://www.phenxtoolkit.org/search/results?searchTerm=Common+data+elements&searchtype=smartsearch), and National Institute of Neurological Disorders and Stroke common data element variables (source, NIH Common Data Elements https://cde.nlm.nih.gov/home) are used for more than 90% of data fields. The remaining data fields were designated a unique data element. All fields were collected in a standard format for research data developed by CDISC.
The authors agree or have agreed to publish the article using the Mary Ann Liebert Inc. “Open Access” option under appropriate license.
Footnotes
Acknowledgments
The authors wish to thank the following colleagues for their support in the conduct of this study. For data monitoring and expertise as a clinical trial associate: Saheed Adeyeri (CVB); site clinical coordinators Abby Lyons (University of Virginia), Josephine Dimanche (University of Utah), Elizabeth S. Hovenden (University of Utah), Mathew Spruiell (Baylor College of Medicine), Dr. Maya Troyanskaya (Baylor College of Medicine), and Carmen Velez (University of Utah); for data entry and quality control: Emma Read (University of Utah), Dayna Thayn (University of Utah), Chris Finuf (University of Utah), and Finian Keleher (University of Utah); data collection and training: Paula V. Johnson (University of Utah) and Hannah Lindsay (University of Utah); supervision of/recommendations related to data collection (for University of Utah and San Antonio): David F. Tate; for programmatic oversight from 2017 to 2019: Anne Marie Drennon, Christine Davis, and Alex Norbash; for article review and revisions: Dr. Ragini Verma (CVB).
Authors’ Contributions
A.T.G.: Conceptualization (equal), data curation (equal), and writing—original draft (lead). J.R.S.: Conceptualization (lead), methodology (lead), and writing—reviewing and editing (equal). E.A.W.: Conceptualization (lead), methodology (lead), and writing—reviewing and editing (equal). S.R.M.: Conceptualization (equal) and writing—reviewing and editing (equal). R.C.W.: Methodology (equal), software (equal), and writing—reviewing and editing (supporting). J.P.M.III: Methodology (equal), software (equal), and writing—reviewing and editing (supporting). N.T.: Methodology (equal), software (equal), and writing—reviewing and editing (supporting). B.A.: Methodology (equal) and software (equal). C.T.W.: Writing—reviewing and editing (supporting). L.L.: Data curation (lead) and writing—reviewing and editing (supporting). S.D.B.: Data creation (equal), project administration (lead), and writing—original draft (equal). M.H.: Conceptualization (equal), supervision (lead), funding acquisition (lead), and writing—reviewing and editing (supporting).
Funding Information
This study was supported by
Author Disclosure Statement
The authors have no competing interests to disclose.
Supplementary Material
Supplementary Data S1
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
