Abstract
Ensuring adequate levels of vitamin D3 in the human diet has long been an important objective in crop breeding, as most crops have extremely low levels of this compound. To address this challenge, we have employed the CRISPR-Cas9 gene editing system in tomatoes to induce loss-of-function mutations in one of the two DWARF5 genes, a homologue of the human dehydrocholesterol Δ7-reductase gene. Lines with knocked out SIDWF5A gene exhibited visually indistinguishable phenotypes, yet remarkably accumulated provitamin D3 levels as high as 6 μg/g dry weight (DW) in the red fruits. As the daily recommended intake of vitamin D is 20 μg (800 IU), consuming a single ripe fresh tomato weighing 150 g (equivalent to 15 g DW) has the potential to significantly alleviate widespread vitamin D deficiencies worldwide.
Introduction
Provitamin D3 (ProVitD3) serves a dual role in humans as a protective agent against ultraviolet (UV) irradiation in the skin and as a precursor for the biosynthesis of biologically active vitamin D3 (VitD3), a crucial human steroid hormone. 1 Studies indicate that global levels of circulatory vitamin D (VitD3) in humans often fall below desirable levels, particularly among individuals with limited exposure to sunlight. 2 Dietary supplements represent a viable strategy to address this deficiency; however, only a few animal-derived sources, such as fish, egg yolks, and beef liver, contain significant amount of VitD, 3 whereas vegetables and fruits are generally poor sources due to their limited capacity for ProVitD3 production. 4
In light of this predicament, we propose that metabolic engineering of widely cultivated and consumed tomato plants may offer a solution to enhance ProVitD3 accumulation, thereby addressing widespread VitD deficiencies, especially among populations with limited dietary choices such as the elderly, vegetarians, and individuals residing in nursing homes or hospitals.
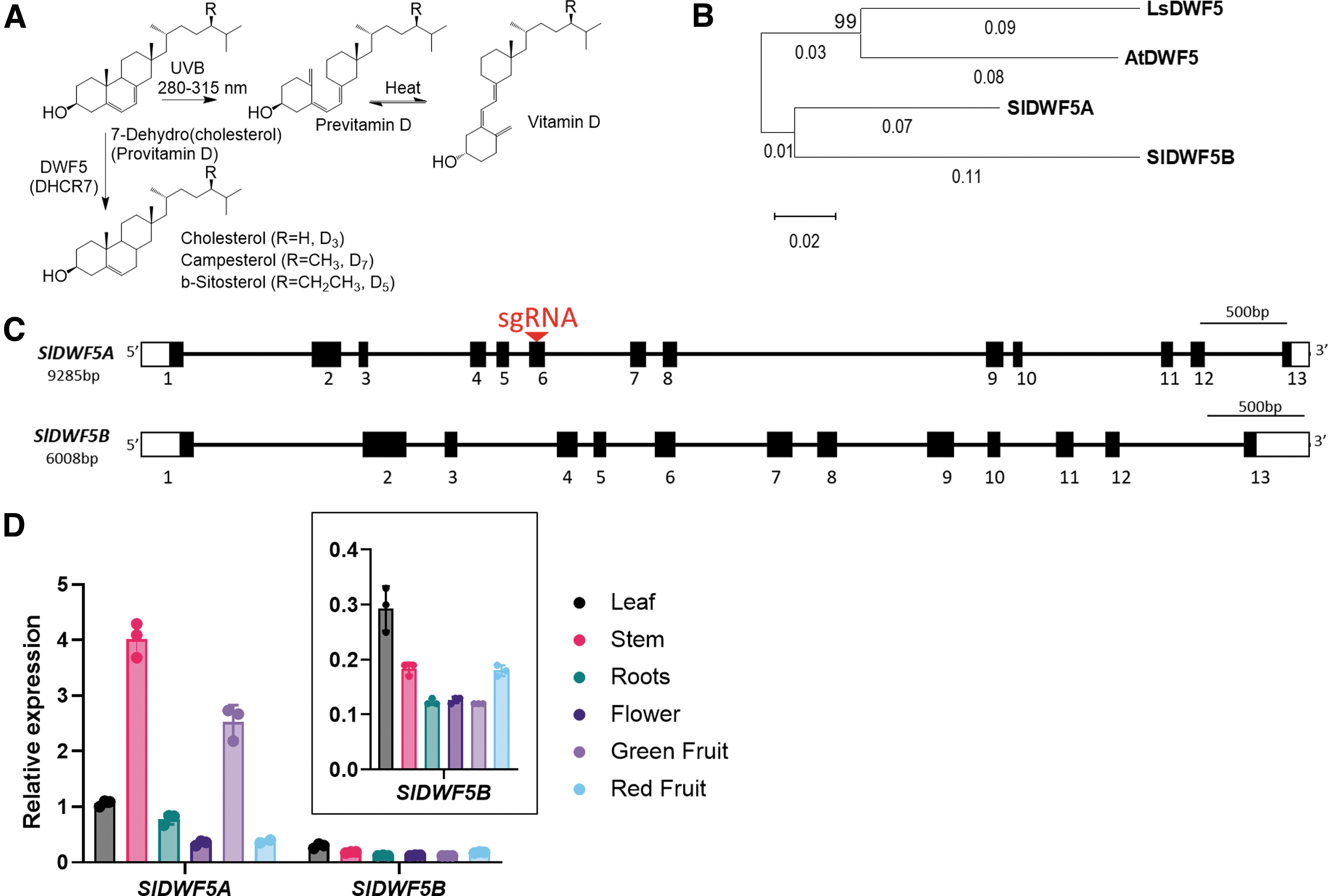
ProVitD3, also known as 7-dehydrocholesterol (7DHC), represents the penultimate intermediate in the cholesterol biosynthetic pathway. The conversion of 7DHC to cholesterol is catalyzed by 7DHC reductase (DHCR7).5,6 Alternatively, exposure to UV-B light (in the 290–315 nm range) triggers the cleavage of the bond between the C9 and C10 carbons of 7DHC, resulting in previtamin D3 (PreVitD3; Fig. 1A). Subsequent enzymatic modification by the liver enzyme CYP2R1 (cytochrome P450 family2 subfamily R member 1) and kidney enzyme CYP27B1 hydroxylate activate PreVitD3 to generate the biologically active form, 1,25-dihydroxy vitamin D3, which plays diverse roles in human metabolism, including calcium and phosphorus absorption.1,5,7
In humans and mice, loss-of-function mutations in the DHCR7 gene lead to a significant increase in serum and tissue ProVitD3 levels. 8 The plant homologue of DHCR7 was initially identified in Arabidopsis as DWARF5 (DWF5). 9 In plants, both campesterol and cholesterol serve as precursors of the plant growth hormones, brassinosteroids, thereby, Arabidopsis dwf5 mutants display dwarfism. Cholesterol is also used as the precursor of steroidal glycoalkaloids in several Solanaceous species such as tomato (Solanum lycopersicum) and potato (Symphytum tuberosum). 10
Previous studies have proposed that the cholesterol biosynthetic pathway in tomato evolved from the existing phytosterol pathway through gene duplication and subsequent specialization. 11 Based on co-expression analysis, researchers suggested that the tomato DWARF5 genes, SlDWF5A and SlDWF5B, act in the phytosterol and cholesterol biosynthetic pathways, respectively. 11 However, given their high sequence identity (83.4%), we hypothesize that these two proteins function redundantly in both the two biosynthetic pathways.
In this report, we demonstrate the successful editing of the SlDWF5A gene using the well-established CRISPR-Cas9 genome editing system, and demonstrate the ensuing accumulation of ProVitD3 in tomato fruits. Our results pave the way for further development of genome-edited nutritious tomatoes, after the recent commercial launch of γ-aminobutyric acid (GABA)-enriched tomatoes in Japan.
Results
Presence of two copies of the DWARF5 gene in tomato
Comparison of the genome databases of Arabidopsis, lettuce, and tomato revealed that whereas Arabidopsis and lettuce possess a single copy of DWARF5, the tomato genome contains two copies (Fig. 1B). The gene organization of both members of this pair of tomato genes displayed similar patterns, consisting of 13 exons and 12 introns (Fig. 1C). As previous studies have indicated that the crucial motif resides in the C-terminus of the protein, 9 our aim was to introduce a single guide RNA (sgRNA) far upstream of the C-terminus to produce a nonfunctional protein (Fig. 1C).
To determine which of the two SlDWF5 genes to edit, we conducted reverse-transcription quantitative PCR (RT-qPCR) analysis to assess the spatial expression patterns of both genes (Fig. 1D). Overall, the transcript levels of SlDWF5A were 2–5 times higher than those of SlDWF5B in all tested tissues (Fig. 1D), particularly in green and red fruits. Assuming that SlDWF5A and SlDWF5B share the same function, we hypothesized that knocking out SlDWF5A would be a more effective strategy for inducing ProVitD3 accumulation in fruits without compromising the agronomic traits of vegetative tissues. This reasoning was based on previous findings that loss-of-function mutations in DWF5 result in severe dwarfism in Arabidopsis dwf5 mutants.
Creation of sldwf5a loss-of-function mutants using CRISPR-Cas9
To generate a knockout mutant for the SlDWF5A gene, we designed an Agrobacterium T-DNA construct harboring Cas9 effector protein and the two sgRNA targeting the exon 6 of SlDWF5A (Fig. 2A); these sgRNAs were specific to SlDWF5A gene. Agrobacterium harboring the Cas9 construct has been transformed into explants of in vitro cultured tomato hypocotyls (a cultivar Seogwang), and 16 plants were regenerated out of the transformed calli (Fig. 2B–E).
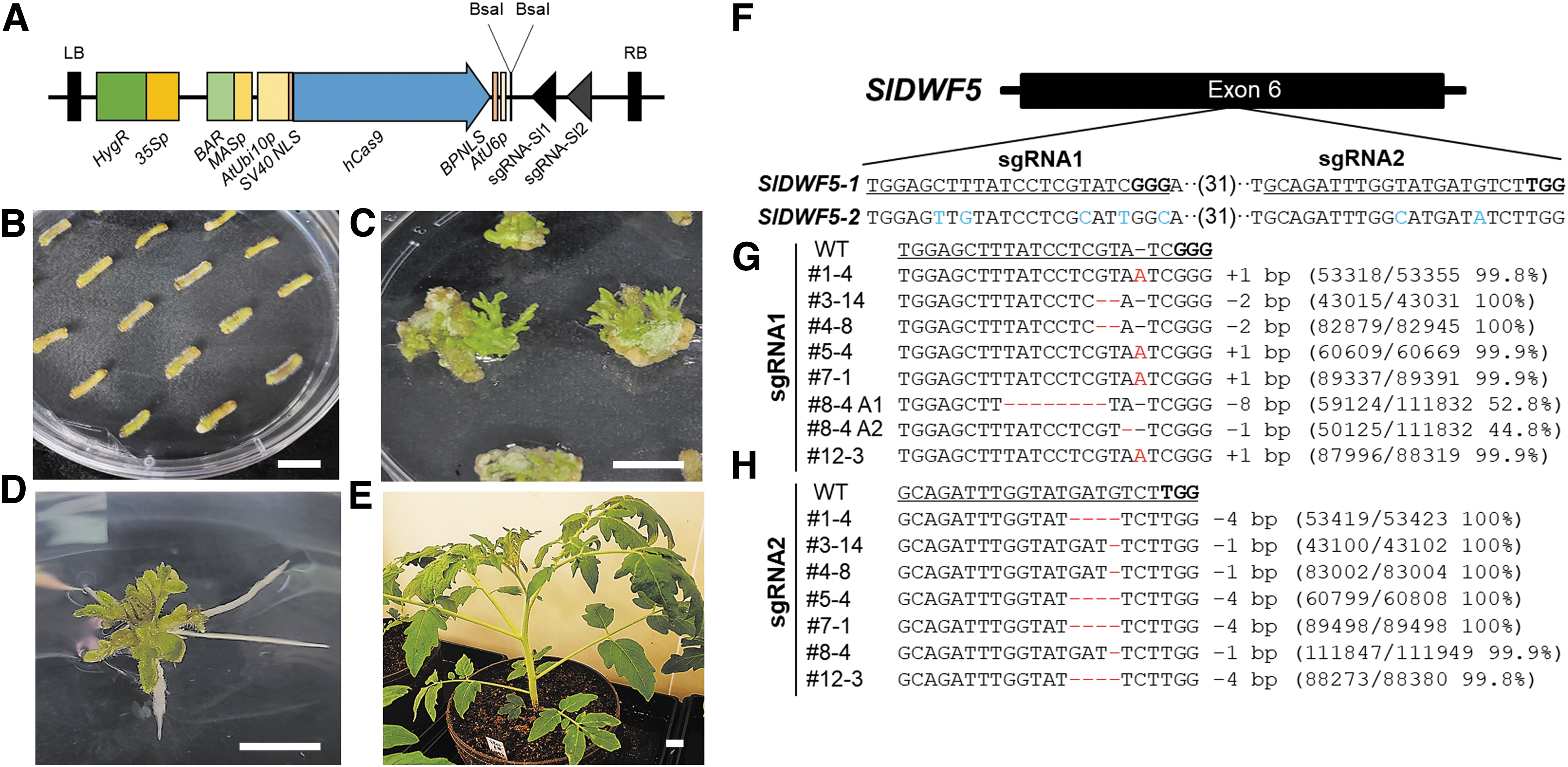
To examine if the Cas9 transgene successfully edited the tomato gene, we extracted genomic DNA from the 16 transgenic plants harboring the CRISPR-Cas9 construct and carried out a T7 endonuclease 1 (T7E1) mismatch detection assay (Supplementary Fig. S1) and amplicon deep-sequencing analysis to identify edited plants (Supplementary Table S1), resulting in the identification of plants #3 and #7 as knockout plants carrying small deletions (#3) (Fig. 2G) or both a small deletion and insertion (#7) (Fig. 2H). The line #3 and #7 had two bp deletion and single bp insertion at the sgRNA1 protospacer site, respectively, and these mutations caused creation of premature stop codon between sgRNA1 and sgRNA2 sites (Supplementary Fig. S2).
To assess the stable inheritance of these changes, we determined the genotypes of their progeny and identified homozygous lines at the T2 generation (Supplementary Table S2; Fig. 1B). All the segregating lines tested displayed near 100% mutations, suggesting that they are homozygous for the mutations in the sgRNA1 and sgRNA2 sites.
Because it is important to examine if our approach resulted in any off-target edits in the tomato genome, we checked indel mutations in possible off-target regions whose sequences are different at only one to four nucleotides (Supplementary Table S3). The frequency of insertion/deletions (InDels) in the off-target sites examined was almost 0%, except the OT1 of sgRNA2, which is located in the SlDWF5B gene, where the rate was as high as 0.4%. This increased off-target effect at the OT1 of sgRNA2 suggests that it would be better to use sgRNA1 when introducing an indel mutation only in SLDWF5A, leaving SlDWF5B unaffected (Supplementary Table S3).
Morphometric analysis of the genome-edited tomato plants
To examine if genome editing resulted in any visible alterations, we performed morphometric analysis (Fig. 3). As shown in Figure 3A–F, the key phenotypes of wild-type (WT), #3, and #7 are all exhibited; seedlings (A), compound leaflets (B), single leaves (C), flowers (D), mature green fruits (E), mature red fruits (F), and three different ripening stages of fruits in the tomato tree (G).
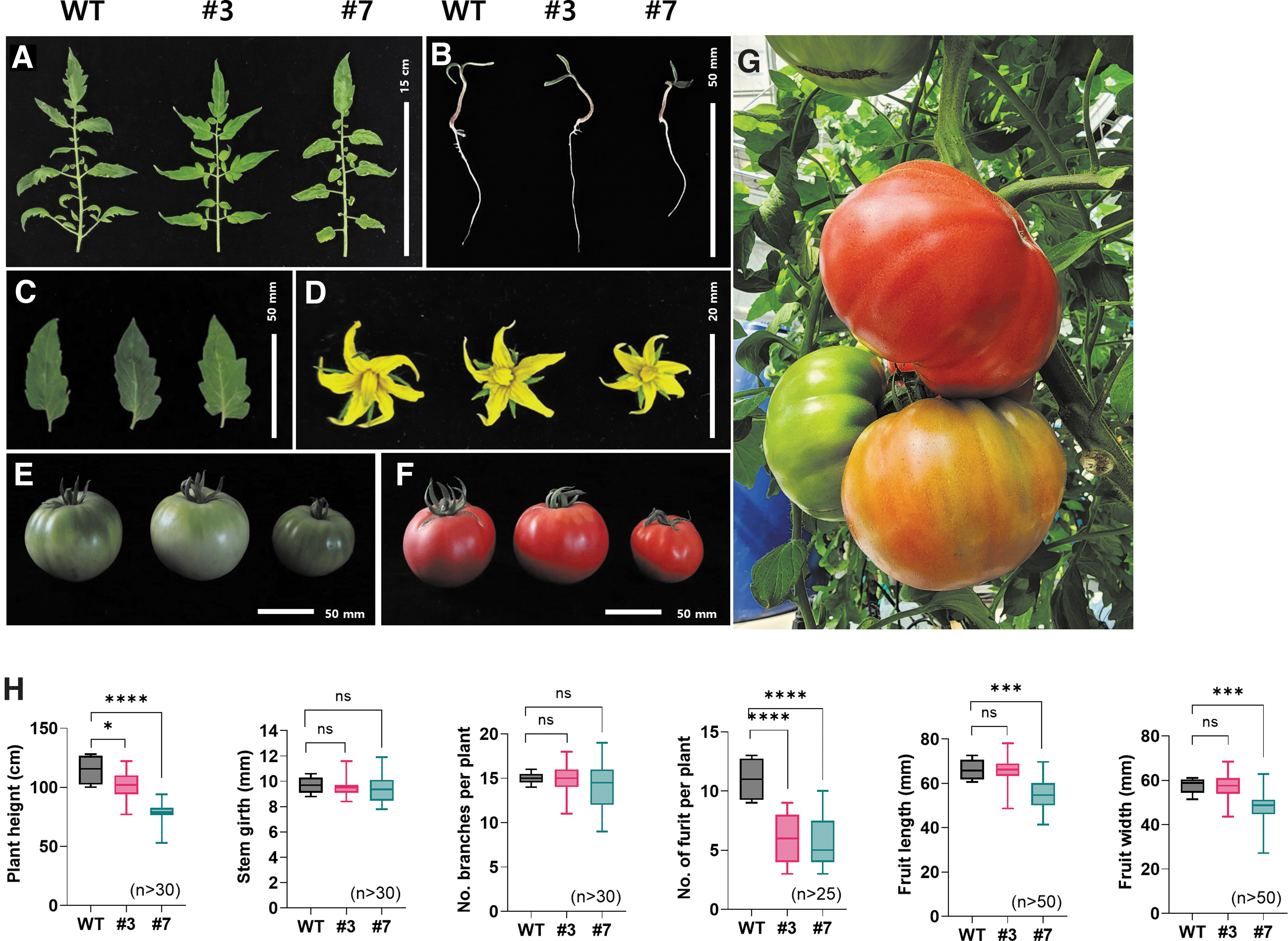
The loss of DWF5 function in Arabidopsis (Arabidopsis thaliana) is caused by a severe reduction in phytosterol and brassinosteroid levels, resulting in dwarf plants as short as only 10% height of WT plants. 9 Our Sldwf5a plants did not exhibit such a dramatic dwarfism, although line #7 was approximately half the height of the corresponding WT. Furthermore, the number of fruits per plant dropped to a 45.7% (#3) and 51.9% (#7) of WT levels (Fig. 3H). Significant differences even between the two edited lines reflect the heterozygosity of the F1 hybrid seeds that were subjected to genome editing.
Accumulation of ProVitD3 in the genome-edited tomato
To show that the knocking out of the SlDWF5A gene successfully resulted in the accumulation of ProVitD3, as intended, we determined the levels of ProVitD3 and phytosterols in leaves, roots, and fruits of Sldwf5a mutants (Fig. 4). Not surprisingly, ProVitD3 accumulated in both green and red fruits, with an average content of 12 μg/g dry weight (DW) in green fruits and 6 μg/g DW in red fruits of #3. Lower levels were seen in #7. Although the ProVitD3 level dropped by half as it turned from green to red, the 6 μg/g concentration in the edible fruits was clearly detectable at repeated experiments in #3 (Fig. 4A). Moreover, ProVitD3 levels were also high in the roots, with 12 μg/g DW in #3 (Fig. 1A).
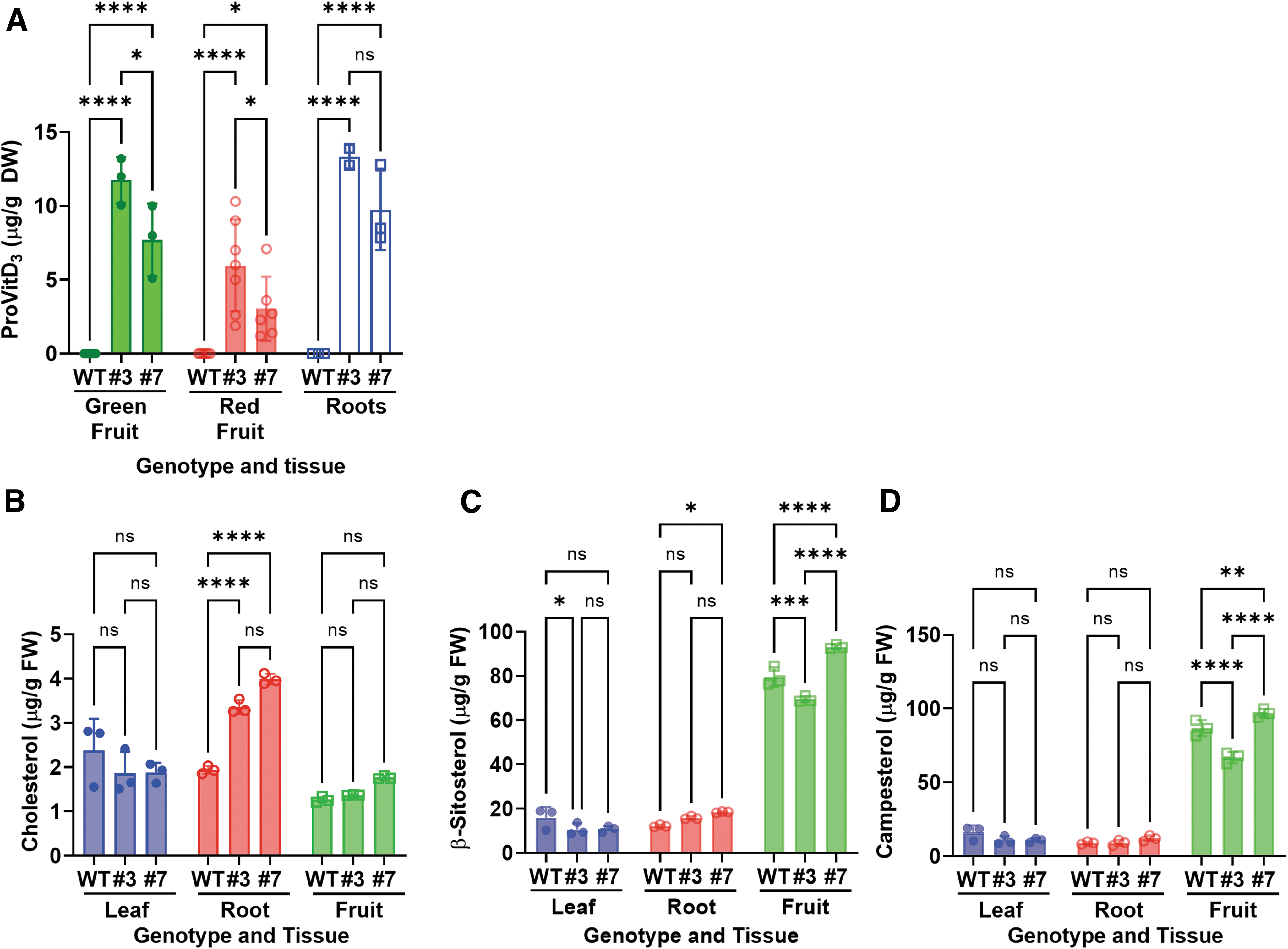
Next, we measured and compared the sterol levels in Sldwf5a mutants, cholesterol levels increased in the roots of both #3 and #7 (Fig. 4B). In the fruits, μ-sitosterol and campesterol were similarly affected by the mutations, in that the levels of the two sterols dropped in #3, whereas they were upregulated in #7 relative to WT control (Fig. 4C, D). Overall, however, the levels of sterols, including cholesterol, β-sitosterol, and campesterol, remained relatively constant (without dropping to zero) in both the WT and Sldwf5 mutants (Fig. 1F), suggesting that SlDWF5B largely compensated for the loss of SlDWF5A function in the genome-edited tomato lines.
In addition, we observed a significant decrease in both β-sitosterol and campesterol in the ProVitD3-accumulating line #3. This suggests we cannot rule out that ProVitD5 and ProVitD7 (Fig. 1A) derived from β-sitosterol and campesterol, respectively, are upregulated in line #3 as well.
Discussion
In contrast to other plants such as Arabidopsis and lettuce, tomatoes possess two copies of the SlDWF5 gene, sharing significant amino-acid sequence identity of >83%. Based on this observation, we hypothesized that knocking out either the SlDWF5A or SlDWF5B gene could lead to the accumulation of the precursor molecule ProVitD3 without causing severe growth retardation.
Our findings support this hypothesis, as we successfully employed CRISPR-Cas9 to knock out the SlDWF5A gene, which exhibits preferential expression in flowers and fruits. This gene manipulation resulted in the substantial accumulation of ProVitD3, reaching levels as high as 6 μg/g DW, while maintaining the overall height of the engineered plants. Notably, our results differ from an earlier study that reported ProVitD3 accumulation in a loss-of-function mutant for the SlDWF5B gene, which exhibited relatively higher accumulation in vegetative tissues such as leaves and stems rather than in red fruits. 12 Furthermore, our analysis revealed that other Solanaceous crops, including eggplant, pepper, and paprika, also possess two copies of the DWF5 genes, suggesting that knocking out the gene preferentially expressed in floral organs could potentially increase endogenous ProVitD3 levels in those plants as well as tomatoes.
Currently, commercial VitD3 supplements are derived from lanolin, a natural wax-like substance secreted by the sebaceous glands of sheep for wool and skin protection. Lanolin contains substantial levels of 7DHC (ProVitD3), which can be chemically converted into cholecalciferol, the active form of VitD3, through exposure to UV-B light and heat. It can be argued that daily intake of VitD3 pills or tablets made from lanolin is more convenient and effective than consuming ProVitD3-enriched tomatoes.
However, consuming a single ProVitD3 tomato fruit per day would offer additional advantages, as tomatoes are already considered a near-ideal food source, containing numerous key vitamins such as A, B, C, and K, as well as essential minerals, including potassium, manganese, copper, magnesium, phosphorus, and iron, in a balanced manner. 13 Production of ProVitD3 from tomatoes, which are producers in our ecosystem, rather than relying on products from primary consumers like sheep, could provide a more sustainable approach to addressing endemic VitD3 deficiencies worldwide.
We acknowledge of course that some of our genome-edited lines displayed less desirable traits, such as a decrease in the number of fruits. We postulate that this variability can be mitigated by introducing the ProVitD3 trait into genetically fixed inbred lines. In our research, we utilized commercially available “Seogwang” seeds obtained from local distributors, resulting in T0 lines of edited genomes that exhibited genome-wide heterozygosity in the F1 hybrid plants. As a result, depending on the background variation, different lines of genome-edited plants displayed different phenotypes, including the number of fruits and the height of the plant. Future research and development should involve repeating our experimental procedures using country- or region-specific inbred lines before producing F1 hybrid seeds.
Future research should focus on quantifying and comparing the levels of PriVitD3 in genome-edited tomatoes with other natural sources of VitD3. Stability and bioavailability studies are also needed to assess the impact of cooking, processing, and storage on the retention of ProVitD3. Evaluating the overall nutritional composition and potential unintended changes in the edited tomatoes is important. Feeding studies and clinical trials can determine the bioactivity and physiological effects of consuming ProVitD3-enriched tomatoes. Finally, comprehensive safety assessments, including toxicological and allergenicity testing, will be necessary to ensure the long-term safety of consuming genome-edited tomatoes.
Recently, the Japanese government approved the commercial availability of GABA tomatoes as food products. 14 Japanese scientists successfully increased GABA levels by deleting the C-terminal autoinhibitory domain of glutamate decarboxylase, a key enzyme in GABA biosynthesis, thus enhancing its enzymatic activity. 15 This exemplifies how gene-edited crop plants can be effectively brought to the market to address public health concerns. Similarly, the introduction of ProVitD3 tomatoes into the consumer market might follow a similar approach, involving regulatory clearance in various countries for commercialization and raising public awareness regarding the benefits offered by genome-edited crops.
Many countries across the Americas, Africa, Asia, and Oceania have relaxed the stringent regulations for genome-edited plants, particularly those developed using site-directed nucleases 1 (SDN-1) technology, where the DNA sequence changes resulting from genome editing are indistinguishable from natural genetic variations. 16 Our ProVitD3 tomato falls under this SDN-1 classification. Consequently, we anticipate that the ProVitD3 tomato will be made available in countries with relaxed regulations for the SDN-1 category soon.
Consumption of this tomato is expected to be beneficial for individuals seeking to supplement their VitD3 intake through plant-based diets, as opposed to relying solely on animal-derived pills. By leveraging the success of the GABA tomato and the changing regulatory landscape for genome-edited crops, we can anticipate the increasing availability and acceptance of innovative plant-based solutions, such as the ProVitD3 tomato, to address nutritional deficiencies and improve public health.
The Bigger Picture
Materials and Methods
Sample preparation
Tomato seeds were planted on culture soil in a growth chamber at 25°C under short-day conditions (8 h light/16 h dark). At 75 days after sowing, approximately four to five leaves were collected from the upper part of the main stem, and stem samples were collected from the upper regions of plants. Flowers were collected from 6-month-old plants, and mature green fruits and red fruits were harvested at an average 140 g fresh weight per fruit. Tomato fruit samples for chemical analyses were prepared by freeze-drying; the other analyses were performed using samples frozen in liquid nitrogen.
Reverse-transcription quantitative PCR
Total RNA was isolated from plant tissues ground to powder using a pestle in liquid nitrogen with an RNeasy® plant mini kit (#74904; Qiagen, Hilden, Germany). Reverse transcription was performed with a RevertAid RT Reverse Transcription Kit (K1691; Thermo Fisher Scientific, Waltham, MA, USA) using 3 μg total RNA and was followed by qPCR analysis of the resulting first-strand complementary DNA (cDNA). The primers used for qPCR are listed in Supplementary Table S4. qPCR was performed in 96-well plates with a Real-Time PCR System (4379216; Applied Biosystems, Foster City, CA, USA) using a KAPA SYBR® FAST qPCR Master Mix Kit (KK4601; KAPA Biosystems, Wilmington, MA, USA) in a volume of 20 μL.
The reactions were performed in three technical replicates per run, with three biological replicates. Absolute quantification was performed using standard curves generated by amplification of a diluted series of cDNA containing individual transcripts. The transcript levels of each gene in different samples were normalized to an internal control, ACTIN (Solyc11g005330.2), using the 2−ΔΔCT method.
Vector construction
A construct used for Agrobacterium (Agrobacterium tumefaciens)–mediated transformation was created harboring an antibiotic selection cassette, the Cas9 gene, and tandem polycistronic transfer RNA (tRNA)-sgRNA repeats for positive selection on medium containing both kanamycin and hygromycin. The human codon–optimized Cas9 gene originating from Streptococcus pyogenes (SpyCas9) was cloned into the pCAMBIA1300 plasmid (#44183; Addgene) and placed under the control of the Arabidopsis UBIQUITIN 10 promoter. To facilitate nuclear localization of the Cas9 protein in tomato cells, simian vacuolating virus 40 (SV40 NLS) and bipartite nuclear localization signal were added at the N and C termini of Cas9, respectively. Using the BsaI restriction enzyme, two sgRNAs were inserted into the pCAMBIA-Cas9 backbone and placed under the control of the promoter of the Arabidopsis small nucleolar U6 RNA gene. The schematic diagram of vector construction is presented in Figure 2A.
Tomato transformation and regeneration
Tomato transformation procedures are summarized in Figure 2. Two-week-old hypocotyls of tomato (Solanum lycopersicum cv. Seogwang) were cut into ∼1-cm2 pieces and cocultured with Agrobacterium strain LBA4404 (OD600 = 0.5) harboring the Cas9 construct for 2 days at 25°C in the dark. The cocultured hypocotyls were transferred to callus-inducing medium (CIM) consisting of full-strength Murashige and Skoog (MS) medium (M0221, Duchefa Farma B.V.) containing 0.5 mg/L nicotinic acid (1414-0130, Showa), 100 mg/L myoinositol (MB-I4715, MB cell), 0.5 mg/L pyridoxine HCl (P-8666, Sigma), 0.1 mg/L thiamine HCl (T0614, Duchefa), 30 g/L sucrose (S0809, Duchefa Farma B.V), 0.25% (w/v) Gelrite (71015-52-1, Duchefa) with 0.1 mg/L NAA (N600, Phytotech Labs), and 1 mg/L BAP (D130, Phytotech Labs) and grown at 25°C in the light for 2 days.
For selection, CIM-grown hypocotyls were transferred to full-strength CIM medium containing 2 mg/L zeatin (Z860, Phytotech Labs), 0.2 mg/L indole-3-acetic acid (IAA; I0901, Duchefa Farma B.V.), 25 mg/L hygromycin B (LPS solution, HYB01), and 200 mg/L ticarcillin disodium (T1090, Duchefa Farma B.V.) and grown at 25°C in the light for 8–12 weeks. The growth plates were replaced by fresh plates at 2-week intervals until shoot generation occurred. The emerging shoots were transferred to half-strength CIM medium with 0.2 mg/L IAA and grown into rooted plantlets in containers. The plantlets were transferred to soil-filled pots and maintained until seed harvest.
T7E1 assay
Genomic DNA was isolated from transgenic plants using a DNeasy Plant Mini Kit (#69104; Qiagen). The target DNA region was amplified and subjected to the T7E1 assay as described previously. 17 For heteroduplex formation, PCR products were denatured at 95°C for 10 min and subjected to an annealing program from 95°C to 85°C (−2°C/s) followed by 85°C to 25°C (−0.1°C/s) for slow cooldown using a thermal cycler. Annealed PCR products were incubated with T7E1 (#m0302, NEB) at 37°C for 20 min and analyzed through agarose gel electrophoresis.
Sanger sequencing of target regions
The sgRNA target regions were amplified from genomic DNA using Q5 Polymerase (#M0491; New England Biolabs) in a 25-μL reaction volume. Then, the PCR products were cloned into a 3′-end T-tailed vector using a PCR cloning kit (VT201-020; Biofact Pharma Ltd., Kildare, Ireland). Twenty clones for each sample were individually sequenced. The primers used for on-target site mutation analysis are listed in Supplementary Table S5.
Targeted deep sequencing
To compare editing events at the target loci, on-target sequencing was performed on 23 sibling lines from seven T1 plants (Supplementary Table S1) and 21 sibling lines of #3 and #7 (Supplementary Table S2). Their sequences were compared with that of the WT. The targeted primers were designed from genomic DNA for 1st PCR, and sequencing adaptors were added to the amplicon. The primer sequences are listed in Supplementary Table S5. High-throughput sequencing was performed using a MiniSeq System (SY-420-1001; Illumina, Inc., San Diego, CA, USA) and analyzed using methods available online. 18
Off-target deep sequencing
Off-target sequencing was performed on seedlings Sldwf5a-1 (#3–14) and Sldwf5a-2 (#7–1) from the T1 generation, and the results were compared with the WT sequence. Potential off-target sites were identified in the S. lycopersicum genome using the Cas-OFFinder (www.rgenome.net/cas-offinder) algorithm. The Sol Genomics Network (https://solgenomics.net) was used as the reference genome to identify homologous sequences that differed from the on-target sequences by up to four nucleotides. From a total of 32 sites, 11 sites were selected for targeted deep sequencing. The primers for the on-target and potential off-target sites were designed from genomic DNA (1st PCR, Supplementary Table S6). Sequencing adaptors were added for the 2nd PCR (Supplementary Table S6). High-throughput sequencing was performed using a MiniSeq System (SY-420-1001; Illumina, Inc.).
Sample preparation at the T2 generation
Samples from the T2 generations of #3 (4 individual lines) and #7 (11 individual lines) as well as WT plants were prepared for analytic samples. Fifteen fruits were picked and weighed to obtain an average of 140 g fresh weight per fruit and were freeze-dried to about 7–10% of the original fresh weight. Roots were prepared from 4-week-old seedlings grown on full-strength solid MS medium (M0221, Duchefa Farma B.V.) containing 0.25% (w/v) Gelrite (71015-52-1, Duchefa).
Gas chromatography–quadrupole mass spectrometry analysis of ProVitD3, cholesterol, and β-sitosterol in tomato
The extraction and analysis of the lipophilic compounds ProVitD3, cholesterol, and β-sitosterol were performed as described previously with several modifications, as specified hereunder. 19 In brief, freeze-dried or powdered tomato samples (10 mg) were mixed with 3 mL 0.1% (v/v) ascorbic acid in ethanol and 0.05 mL 5α-cholestane (10 μg/mL, internal standard, Sigma Aldrich, St. Louis, USA) for crude extraction, and 80% (w/v) aqueous potassium hydroxide was used for saponification. After saponification, lipophilic compounds were purified by hexane extraction, and their derivatization was performed using pyridine (Sigma Aldrich, St. Louis, USA) and N-methyl-N-trimethylsilyl trifluoroacetamide (Sigma Aldrich, St. Louis, USA).
Metabolic analyses were performed using gas chromatography-quadrupole mass spectrometry (GC-qMS; GCMS-QP2010, Shimadzu, Kyoto, Japan). The analytical conditions for GC-qMS were described in a previous study. 19 The identification and quantification of the lipophilic compounds were performed using standard compounds and calibration curves obtained from each standard. Calibration curves were determined for 7DHC (ranging from 0.07 to 8.33 μg/mL), cholesterol (ranging from 0.03 to 8.33 μg/mL), and β-sitosterol (ranging from 0.26 to 66.67 μg/mL) standard and fixed to an internal standard weight of 0.50 μg (Supplementary Fig. S3).
Footnotes
Authors' Contributions
S.H.C. conceived the study. S.M.C., Y.A.J., J.H.K., Y.J.P., J.M.K., and J.J.P. conducted the experiment. S.H.C., M.K.Y., and J.K.K. wrote the article.
Author Disclosure Statement
S.H.C. is a founder of the biotech company G+FLAS Life Sciences. S.M.C., M.K.Y., J.H.K., J.M.K., and J.J.P. were employees of G+FLAS Life Sciences. Authors are inventors on a patent application covering the genome editing method and genome-edited tomato described in this article.
Funding Information
This research was funded by G+FLAS Life Sciences with the control number GFRND-GFC102.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
