Abstract

CRISPR-Cas genome editing has rapidly moved from research tool to therapeutic application, with the first CRISPR Cas based drug, Casgevy (exagamglogene autotemcel; exa-cel), receiving FDA approval in 2024. The advancement of CRISPR-Cas-based genome editing platform technologies for therapeutic application often requires testing in multiple species. Guide RNAs (gRNAs) enable genome-wide target selection, but divergence of target DNA sequences across species presents challenges for Cas9-based platform and therapeutic development. Two distinct challenges are (1) the ability to directly and quantitatively compare editing efficiencies across cell and animal models of different species and (2) the ability to evaluate the activity of therapeutic gRNAs in contexts in which a mutant allele is targeted that cannot be recapitulated in model species required for preclinical research. We sought to identify and demonstrate the feasibility of a tool gRNA that could address the former challenge and support preclinical evaluation of CRISPR-Cas technologies by enabling the evaluation of cross-species editing efficiencies.
We focused on identifying a Streptococcus pyogenes Cas9 (SpCas9) gRNA targeting a genomic region with 100% conservation in all mammalian species commonly utilized for cell-based and animal model experimental systems (e.g., human, macaque, mouse, rat, pig). An ideal tool gRNA for comparative evaluation of editing efficiency across preclinical model systems should be broadly cross-reactive across species of interest while having no effect on cell behavior that could influence observed editing rates. Notably, the genomic sequences of safe harbor loci (e.g., Rosa26, AAVS1), non-coding intronic regions, and therapeutically relevant gRNAs (e.g., exa-cel) are not typically perfectly conserved across mammalian species (Fig. 1A).1–3
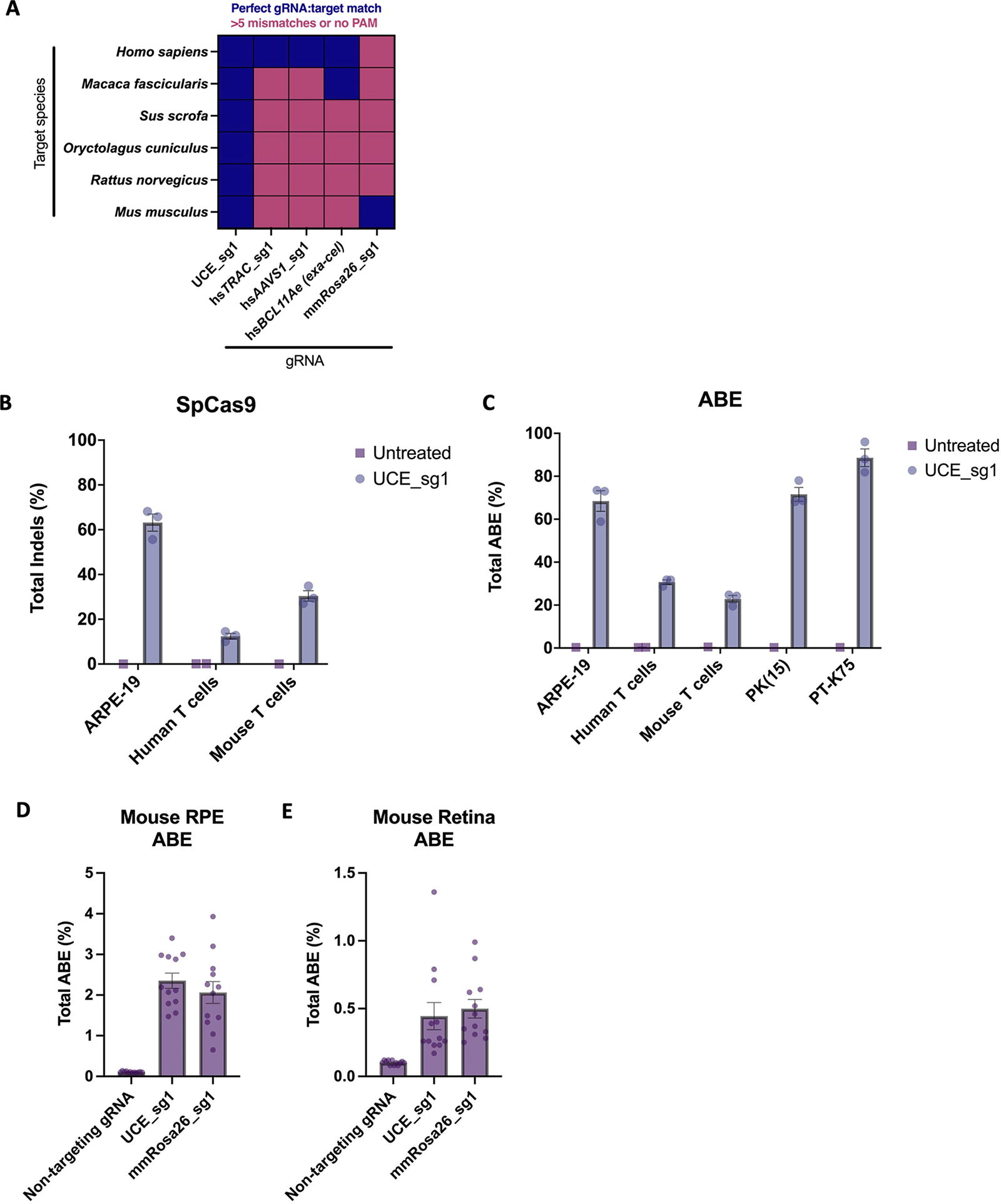
Genome editing with a UCE-targeting guide RNA.
To address this challenge, we identified SpCas9 gRNAs targeting ultraconserved elements (UCEs), which are genomic DNA sequences of 200 or more bp with perfect sequence conservation between human, mouse, and rat genomes. 4 We hypothesized that UCEs could be used to identify species cross-reactive gRNAs. Although UCEs can act as cis-regulatory elements, studies of mice in which one or more UCEs were deleted found surprisingly subtle phenotypes relative to their extreme level of sequence conservation. 5 This led us to hypothesize that UCE-targeting guides would have minimal functional consequences on cellular viability that could skew empirical assessments in the context of assessing genome editing systems. We selected non-exonic elements on autosomal chromosomes from a set of previously reported UCEs. 4 We identified the corresponding sequences from the latest build of the human genome reference sequence (hg38) and further filtered the list of elements by size (≥275 bp), GC content, and confirmation of sequence conservation across mammalian species, yielding a list of 7 UCEs (Supplementary Table S1). 4 We then computationally identified candidate NGG PAM SpCas9 gRNAs within these elements (Supplementary Table S2). 6
As a proof of principle, we evaluated the editing efficiency of a UCE gRNA for indel generation using an SpCas9-NLS nuclease ribonucleoprotein (RNP) and adenine base editing (ABE) activity using an SpCas9-ABE8.20-m RNP 7 (Fig. 1A, Supplementary Table S3). RNPs were delivered by nucleofection in cell systems from multiple species including Homo sapiens (primary T cells and the retinal pigment epithelium-derived ARPE-19 cell line), Mus musculus (primary T cells) and Sus scrofa [PK(15) and PT-K75 cell lines]. We observed nuclease and base editing activity across all cell types evaluated (Fig. 1B–C, Supplementary Fig. S1). We also evaluated other gRNAs that can be used to assess the activity of Cas9 editing technologies (e.g., safe harbor loci) (Supplementary Fig. S2, Supplementary Table S4). The UCE gRNA showed qualitatively similar editing efficiency in comparison to the other gRNAs tested.
We then tested whether the UCE-targeting gRNA has in vivo editing activity. Mice received a single subretinal administration of an ABE delivered as a non-encapsulated RNP. Target tissue layers (i.e., neural retina and retinal pigment epithelium) were dissected from treated eyes followed by Illumina amplicon sequencing of extracted genomic DNA. We observed qualitatively similar levels of base editing for the UCE gRNA and a mouse-specific Rosa26 gRNA (Fig. 1D–E).
Herein, we identify gRNAs cross-reactive across mammalian species and report proof of principle data in human, pig, and mouse experimental systems. We believe that UCE-targeting gRNAs such as those described here can be a useful tool to the research and drug development communities, in particular to support the preclinical evaluation of Cas9-based gene editing platform technologies. For example, a single production batch of a gene editor could be evaluated across experimental systems (e.g., human cells, rodents, and large animals), mitigating guide-intrinsic and test article production variables. A UCE-targeting gRNA could also be utilized to compare the editing rate of multiple variants of an editor to assess whether rank orders are similar across experimental systems. While other variables would remain, such as the possibility that a given UCE is differentially editable in different species, UCE-targeting gRNAs represent a tool of significant utility to the field for qualitative comparisons of gene editing technologies across mammalian experimental systems. We highlight that in addition to the systems tested in this work (human, pig, and mouse), UCE gRNAs are also cross-reactive with other species commonly used to support preclinical drug development (e.g., rat, rabbit, dog, non-human primate), basic research (e.g., Xenopus), and agriculture (e.g., chicken, cow). Thus, UCE-targeting gRNAs may be a broadly applicable and useful tool to the gene editing community. UCE-targeting gRNAs complement other approaches to evaluate editing activities across translational model systems, such as knocking in a humanized target sequence into a model organism or cell line. Furthermore, although we focused on SpCas9 in the context of indels and adenine base editing in this work, the activity of other nucleases (e.g., AsCas12a, GeoCas9) or editing systems (e.g., cytosine base editing, prime editing, homology directed repair) may be evaluated using an analogous UCE targeting approach. In addition, UCE gRNAs may be utilized to further probe the biological functions of UCEs themselves.
Footnotes
Acknowledgments
The authors would like to thank Mary J. Janatpour as well as our current and prior mentors for their invaluable guidance.
Authors’ Contributions
B.G.G., S.C.W., and P.K.: Conceptualization. S.R.M., P.K., J.C., V.J., C.S., and P.B.O.: Methodology, validation, investigation, verification, formal analysis, and visualization. K.M. and P.K.: Software and data curation. B.G.G., P.K., S.R.M., P.B.O., S.C.W., and A.P.S.: Writing (original draft, review, and editing). B.G.G., J.W., R.M., W.L., P.B.O., A.J.C., S.C.W., A.P.S., and M.H.F.: Supervision. R.M., A.P.S., and M.H.F.: Project administration and funding acquisition.
Author Disclosure Statement
All authors are employees of Spotlight Therapeutics, Inc., or were employees at the time the research took place. B.G.G., K.M., A.J.C., J.W., P.B.O., R.M., S.C.W., and A.P.S. are inventors on patent applications related to in vivo and in vitro delivery of Cas9 ribonucleoproteins. B.G.G., K.M., and S.C.W. are inventors on patent applications related to guide RNA sequences. S.C.W. is also a prior consultant for BioEntre and an inventor on a patent for a mouse model of autoimmune adverse events.
Funding Information
The disclosed work was supported in part by NIH grant 1U19NS132296-01.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
