Abstract
Wildlife diseases are a considerable threat to human health, conservation, and the economy. Surveillance is a critical component to mitigate the impact of animal diseases in these sectors. To monitor human diseases, CRISPR-Cas (clustered regularly interspaced short palindromic repeats-CRISPR-associated protein) biosensors have proven instrumental as diagnostic tools capable of detecting unique DNA and RNA sequences related to their associated pathogens. However, despite the significant advances in the general development of CRISPR-Cas biosensors, their use to support wildlife disease management is lagging. In some cases, wildlife diseases of concern could be rapidly surveyed using these tools with minimal technical, operational, or cost requirements to end users. This review explores the potential to further leverage this technology to advance wildlife disease monitoring and highlights how concerted standardization of protocols can help to ensure data reliability.
Introduction
Wildlife diseases are a considerable threat to human health, conservation, and the economy.1,2 Surveillance of these wildlife diseases can help to support the prevention, management, and mitigation of their impacts to society and the environment. 3 For example, zoonotic pathogens of infectious diseases, such as SARS-CoV-2, can have significant impacts, especially to underdeveloped regions with limited financial and logistical resources for diagnostics and treatment. 4 However, surveillance of new and emerging pathogens associated with wildlife diseases is challenging for a plethora of factors. Vulnerabilities apply to wildlife disease pathogen surveillance in regions where monitoring is compounded by the lack of information available for emerging diseases, the minimal standard surveillance of endemic pathogens in wildlife populations, and the challenge to rapidly develop a reliable diagnostic test. 5
Current DNA Detection Tools and Approaches
Direct pathogen detection in wildlife disease surveillance can be conducted using microbiological assays, tissue examination, and molecular methods such as polymerase chain reaction (PCR). 6 However, these methods tend to be time consuming and require specialized setups that could prove challenging for many entities with limited financial and logistical resources to conduct their own pathogen surveillance efforts. Further, culturing methods for the detection of microbes are limited to those few species that are culturable.7,8
Portable quantitative PCR (qPCR) devices are commercially available and have been used for point-of-care detection of numerous pathogens in aquaculture and humans including, the malaria-causing parasite Plasmodium falciparum, the bacterial coldwater disease-causing Flavobacterium psychrophilum, and the COVID-causing viruses SARS-CoV-2.9–13 Although these devices address the portability hurdle of benchtop PCR instruments, the investment necessary to procure and maintain portable PCR devices is challenging for many entities. Technological advances could lead to the development of lower-cost portable devices for more widespread adoption in the future. 14
Isothermal Amplification of Nucleic Acids
An alternative to PCR is recombinase polymerase amplification (RPA), a rapid and cost-effective isothermal amplification molecular method that uses two primers for the amplification of target nucleic acid sequences. 15 RPA can be further improved as a detection tool through the integration of an optional DNA probe that, like qPCR, enables the assay to further discriminate targeted sequences. 16 RPA operates at temperatures between 37°C and 42°C and amplifies the target nucleic acid by employing proprietary recombinase and single-stranded binding proteins capable of internalizing primers into DNA strands without the temperature cycling used in traditional PCR. 17 The previous step enables the use of a DNA polymerase to complete replication of the target. RPA has been widely used as a DNA and RNA detection tool for human viral (e.g., SARS-CoV-2 and HIV-1) and bacterial (e.g., Mycobacterium tuberculosis and Leptospira sp.) agents. 18 RPA can be developed to detect nucleic acids as a stand-alone assay but may be less target-specific because of its innate tolerance to sequence mismatches when amplifying targets. 17 An alternative to RPA for isothermal amplification of target DNA is Loop-mediated isothermal amplification (LAMP), a DNA polymerase-based method that relies on the use of 4–6 primer pairs to recognize multiple sites within the pathogen target sequence.19,20 Similar to RPA, LAMP, which is also patented, does not require temperature cycling, can provide simple-to-read colorimetric (i.e., color change visible to the naked eye) results, and has also been adapted as a stand-alone point-of-use (POU) and field tool for a variety of targets related to wildlife and human diseases.21,22 However, the use of LAMP diagnostic tools is often compounded by the complexity to design a suitable group of primers to target a specific locus, which often requires multiple rounds of optimization and does not have well-defined primer design rules.20,23 Moreover, LAMP primer design for targets with rapidly evolving DNA or RNA sequences, such as viral variants, has proven to be a challenge due to the difficulty of ensuring that the multiple primers used in this assay remain specific. 24 The development of new LAMP assays to match the divergence of highly variable sequences is not a trivial undertaking—a matter that is further compounded by biased, and often nonrepresentative, reference sequences deposited to public databases.24,25 LAMP can also have nonspecific amplification of off-target nucleotides due to circumstances arising from primer-primer interactions, carryover contamination, and the use of a DNA polymerase capable of nontemplated synthesis.26–28 LAMP has been successfully incorporated as a pre-amplification module for clustered regularly interspaced short palindromic repeats CRISPR-associated (Cas)-based biosensors to detect pathogens such as SARS-CoV-2 and the bacterium Neisseria meningitidis.29,30
CRISPR-Cas Detection of DNA and RNA
Novel, field-applicable DNA and RNA detection methods have gained traction that could overcome some of the sensitivity, logistical, and time limitations that are associated with standard diagnostic approaches discussed above. 5 The discovery of the CRISPR and their Cas nucleases has not only provided an understanding of bacterial adaptive immunity and response to repeated exposures, but has also revolutionized the field of targeted gene editing even in organisms lacking robust genetic resources.31,32
Briefly, CRISPR-Cas systems consist of sequence repeats intercalated with nucleic acid sequences, known as spacers, that are derived from pathogens that previously infected the bacterial host. 32 This array of sequences is accompanied by cas genes that encode the Cas nuclease components. 32 The spacers in CRISPR-Cas arrays encode the crRNA component that hybridizes with the Cas nuclease to form a complex that recognizes a target sequence and enables targeted protein-mediated cleavage. 33 In some CRISPR-Cas systems (such as in CRISPR-Cas9), the crRNA requires an additional RNA component, known as the trans activating CRISPR RNA (tracrRNA), to enable processing of the former RNA and generate a functional guide RNA (gRNA) for target recognition. 34 The gRNA requires the recognition of a short, conserved sequence known as a protospacer adjacent motif (PAM) or protospacer flanking site (PFS) adjacent to the target DNA or RNA, respectively. 35 The PAM and PFS are indispensable for the discrimination of self from nonself-DNA or RNA during CRISPR-Cas target recognition and cleavage. 36 Cleavage by Cas nucleases is conducted by a menagerie of domains, namely HNH and RuvC-like, that can cleave double-stranded DNA, single-stranded DNA, or single-stranded RNA depending on the class of CRISPR-Cas enzyme. 37 For example, CRISPR-Cas9 relies on a nuclease domain containing both HNH and RuvC-like nuclease domains for the cleavage of double-stranded DNA, whereas CRISPR-Cas12a only contains a single RuvC-like nuclease domain.37,38 Conversely, CRISPR-Cas13a relies on a higher eukaryotes and prokaryotes nucleotide-binding domain for its RNA cleavage activity. 36 For more in-depth reading regarding the mechanics, including additional details into the structural and functional attributes, of CRISPR-Cas systems and the adaptive immunity process see: Doudna et al. 32 regarding the history of CRISPR-Cas systems and the functional and mechanistic properties of CRISPR-Cas9, Karvelis et al. 33 pertaining to the detail processing of CRISPR-RNA (crRNA) and tracrRNA to modulate CRISPR-Cas9 targeting, Paul and Montoya 33 and Chen et al. 39 for an overview of the structure and function of CRISPR-Cas12a, and Zhao et al. 40 and Gootenberg et al. 41 for an in-depth understanding on the properties of CRISPR-Cas13a.
CRISPR-Cas systems allow for programmable, facile gene editing in myriad organisms and has also gained prominence as a DNA and RNA detection method that is well suited for POU and field applications as they are rapid and do not require specialized equipment (Fig. 1).42,43 The use of CRISPR-Cas for the detection of DNA or RNA associated to pathogens does not raise the same ethical concerns that this technology raises in fields related to gene editing of live organisms or for genetic biocontrol applications.44–47 However, when collecting samples or data, researchers need to understand the necessary permits and/or agreements with the land stewards. Detection of DNA or RNA does not induce any type of genetic change directly into an organism, living or dead, and all reagents are manipulated in vitro and isolated. This, together with the absence of any reagents that would promote delivery of purified CRISPR-Cas complexes into an organism (e.g., lipofectamine), provides a safeguard against any of the components from being introduced into the environment. 48
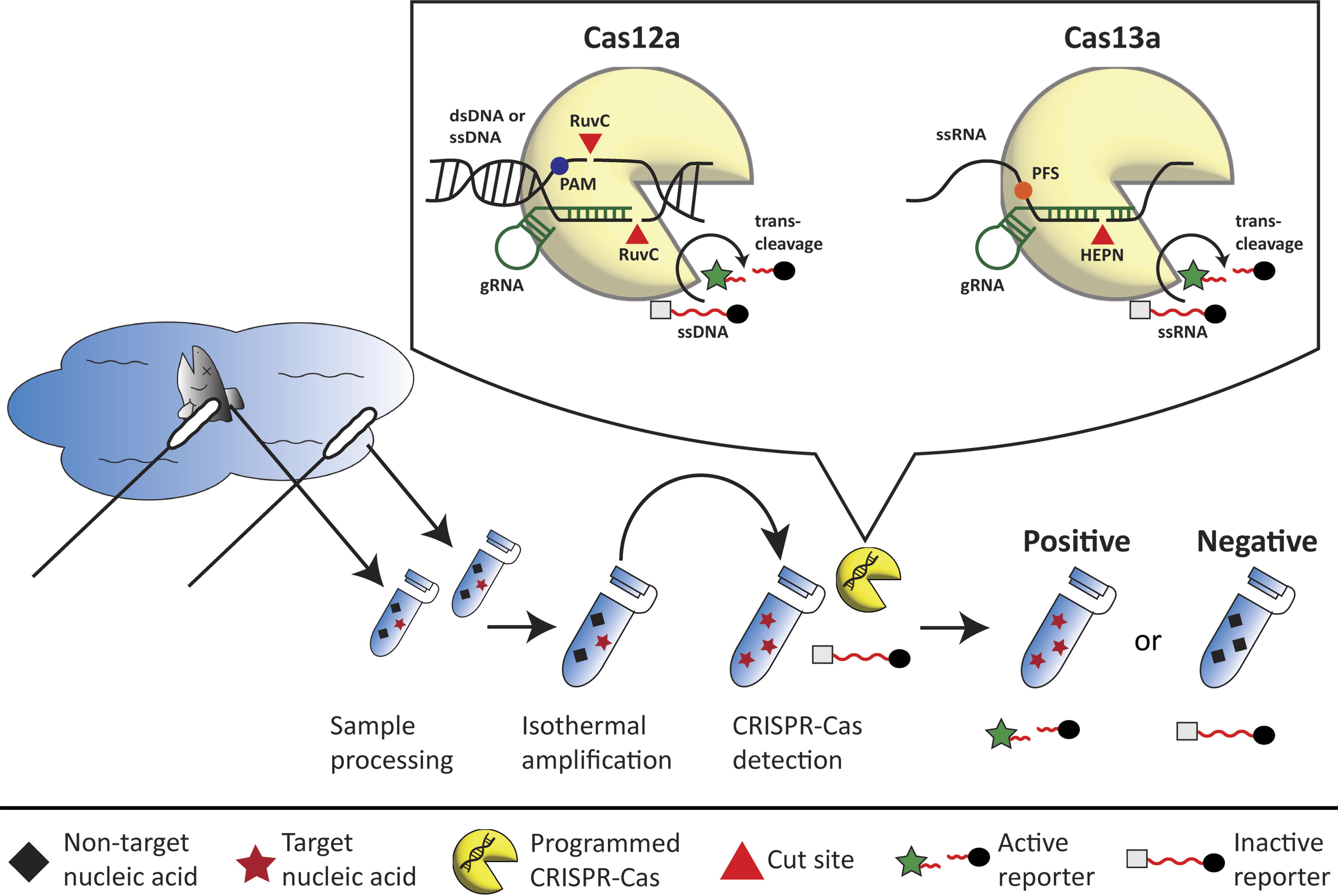
Overview of the mechanism of CRISPR-Cas (clustered regularly interspaced short palindromic repeats-CRISPR-associated protein) biosensors for DNA or RNA detection. DNA or RNA can be collected from myriad substrates and reservoirs using a variety of methods (e.g., dermal swabs, filters, liquid collection, etc.). Sample processing is the act of isolating and purify nucleic acids, this step is dependent on the sample source and might not be required. Isothermal amplification is the process of DNA or RNA acid amplification, which is normally carried out at a single temperature (i.e., isothermally) to augment the sensitivity of the CRISPR-Cas biosensor assay. Methods such as loop-mediated isothermal amplification (LAMP) or recombinase polymerase amplification (RPA) are commonly used for this step. Samples with low quantities of the target DNA or RNA (red stars) are enriched using these amplification methods, whereas samples without the target nucleic acid (black diamonds) are not. After isothermal amplification of the sample, a CRISPR-Cas component and its cognate guide RNA (yellow sector with embedded double helix) are assembled to identify the target DNA or RNA. CRISPR-Cas Detection is determined based on if a signal (i.e., fluorescent or color-based) is produced. Samples with no detectable target nucleic acid (i.e., negative) do not generate any measurable signal, whereas detectable target nucleic acid (i.e., positive) do. Inset: The guide RNA (gRNA) in CRISPR-Cas systems allows for site-specific DNA or RNA recognition and cleavage. Targeting by the gRNA in CRISPR-Cas12a requires a short, conserved protospacer adjacent motif (PAM) site in the target DNA. In CRISPR-Cas13a, recognition of the target RNA requires a protospacer flanking site (PFS). CRISPR-Cas12a relies on a RuvC domain to cleave its target DNA (double or single-stranded) whereas CRISPR-Cas13a utilizes a higher eukaryotes and prokaryotes nucleotide-binding (HEPN) domain to cleave target RNA. In addition to target DNA or RNA cleavage, CRISPR-Cas12a and CRISPR-13a possess indiscriminate, trans-cleavage activity of any single-stranded DNA (ssDNA) or single-stranded RNA (ssRNA). The latter property is the basis for the development of CRISPR-Cas tools as these short ssDNA and ssRNA molecules can serve as reporters to indicate whether a CRISPR-Cas tool detected its intended target or not.
In 2016, Pardee et al. reported the first instance of using CRISPR-Cas for the detection of the RNA of a human infectious agent using a Zika virus biosensor and demonstrated that the introduction of a CRISPR-Cas9 module enabled the discrimination between viral strains up to a single-base resolution. 49
Real-time RNA detection utilizing a CRISPR-Cas platform was further optimized in 2017 by Gootenberg et al. by capitalizing on the discovery of collateral, and indiscriminate, cleavage of single-stranded RNA upon binding of the target RNA locus by the CRISPR-Cas13a complex.41,50 The researchers incorporated a single-stranded RNA fluorescent reporter that provided a readable signal after targeted identification of several pathogenic bacteria (Escherichia coli and Pseudomonas aeruginosa and Zika and Dengue virus). In 2018, Chen et al. reported the discovery that, just like CRISPR-Cas13, the CRISPR-Cas12a complex harbors indiscriminate collateral cleavage activity upon target binding, but of single-stranded DNA instead. 38 CRISPR-Cas12a and 13a can be combined with colorimetric, lateral flow, or fluorometric readouts thanks to their indiscriminate single-stranded DNA or RNA cleavage activity once these CRISPR nucleases associate with their target. CRISPR-Cas9 is not used widely as a detection tool as it lacks the secondary ability of CRISPR-Cas12a and CRISPR-Cas13a to cleave single-stranded DNA or RNA. 38 These discoveries together led to the establishment of CRISPR-Cas-based detection tools of RNA (SHERLOCK or specific high-sensitivity enzymatic reporter UnLOCKing) and DNA (DETECTR or DNA endonuclease-targeted CRISPR trans reporter). In both methods, isothermal amplification through RPA is employed to increase the sensitivity of CRISPR-Cas sensing without requiring significant changes to the protocol such as temperature cycling or the use of more advanced instrumentation. 51 Thanks to these advancements, CRISPR-Cas diagnostic tools became prominent for the low-cost and user-friendly detection of human infectious pathogens that have been reported to be as sensitive and specific as laboratory-based qPCR-based assays. 52 Although these tools have become ubiquitous for pathogen detection in human diseases, their use to support wildlife disease management is limited. Many wildlife diseases of concern could be quickly surveilled using the CRISPR-Cas method with minimal technical or cost requirements for end users.
To further increase the applicability of CRISPR-Cas-based detection tools in the field setting, Myhrvold et al. developed the HUDSON (heating unextracted diagnostic samples to obliterate nucleases) protocol. 53 This protocol simplifies the sample processing pipeline by inactivating inhibiting factors (e.g., nucleases) from complex samples such as bodily fluids. 53 Furthermore, it enabled SHERLOCK diagnostics directly from HUDSON-treated samples without the need for nucleic acid extraction and isolation. Other advances in sample processing for CRISPR-Cas DNA and RNA detection include S-PREP (SHERLOCK parasite rapid extraction protocol), which facilitates the screening of the malaria microbial agent Plasmodium falciparum from blood samples by using a stronger chelating agent than the HUDSON protocol. 54
In SHERLOCK and DETECTR, pre-amplification of the target template to increase assay sensitivity is achieved using RPA or reverse transcription-RPA (RT-RPA), a patented assay that is commercialized as pre-mixed proprietary reaction components. 55 Moehling et al. highlight the proprietary nature of RPA as a hurdle for novel applications and adaptations of the aforementioned CRISPR-based diagnostic tools. 25 CRISPR-Cas sensing coupled to LAMP pre-amplification has been shown to lead to increased specificity when compared with stand-alone RT-LAMP assays targeting SARS-CoV-2, although it does not show a significant increase in sensitivity.56,57 A notable drawback of combining LAMP and CRISPR-Cas is the need to compartmentalize the two components due to their distinct temperature requirements of 65°C and 37°C, respectively, although a modified variant of CRISPR-Cas12a was reported to be active at temperatures optimal for LAMP. 58 The increased uptake of LAMP is in part due to the reagent having a larger pool of vendors and its protocols being freely published. Moreover, many of the early LAMP patents have expired or will soon expire, which can facilitate the access and distribution of this technology for pre-amplification in CRISPR-Cas biosensors. 59
Discussion
CRISPR-Cas biosensors—promising tech for noninvasive DNA and RNA detection
The ability to detect DNA and RNA from abiotic (e.g., water) and organic (e.g., feces) reservoirs in the environment constitutes an attractive strategy to survey for the potential presence of wildlife pathogens with minimal to no disruption of the host organisms or their surroundings. Less invasive techniques, such as swabbing or brushing of the animal, and noninvasive techniques, such as environmental DNA (eDNA) sampling, both demonstrate promise to survey various targets, including cryptic invasive and imperiled species. 60 CRISPR-Cas assays are robust in complex mediums, such as samples contaminated with feces, without the need for DNA or RNA extraction or isolation.53,54 The ability to amplify DNA and RNA in the presence of traditional inhibitors that can interfere with PCR is also a strength of RPA and LAMP, the two most common pre-amplification methods used to increase the sensitivity of CRISPR-Cas.61–65 Additionally, the components in CRISPR-Cas biosensors have been reported to endure a wide range of storage and operational conditions.41,66 Modified reagents could include the lyophilization of reagents and lateral flow strips to enable this technology to be used in a variety of field and on-site settings outside the laboratory.67,68
Limitations of CRISPR-Cas biosensors
The main hurdle for the adoption of CRISPR-Cas biosensors for DNA or RNA detection is the lack of sensitivity by the nuclease component. This limitation makes it impractical to use this technology directly for the reliable detection of targets that could be present in low concentrations, such as eDNA in aquatic mediums. 69 eDNA is genetic material sloughed into the environment through feces, mucus, saliva, gametes, skin cells, and decomposing carcasses. 70 Without the filtering of water to concentrate eDNA material in a membranous filter, it is unlikely that a CRISPR-Cas biosensor could detect its target directly.71,72 Almost ubiquitously, a template pre-amplification step (e.g., RPA or LAMP) is required for CRISPR-Cas biosensors to effectively detect their intended target. This component is generally conducted separately from the CRISPR-Cas reaction. However, researchers have combined both reactions into a single tube in a process commonly termed as “one-pot.” Many options have been employed to address this critical limitation, ranging from the addition of CRISPR type III effector proteins, such as Csm6, to induce cascade signal (e.g., fluorescence) amplification, to the use of microfluidic platforms.73,74 Another significant challenge of this technology is multiplexing, the detection of multiple targets within a single reaction, due to the constraints of relying on collateral cleavage to generate a visual readout. Gootenberg et al. developed SHERLOCK version 2, a system using orthogonal CRISPR-Cas complexes with unique substrate preferences to detect multiple targets simultaneously. 75
Enhancing the potential of CRISPR-Cas biosensors through microfluidics
Microfluidics deal with the motion of small amount of liquids through a series of chambers and channels. This science has seen relevance in the development of CRISPR-Cas biosensors by ensuring that all the pertinent reagents of this detection method can be compartmentalized into a small device. This property obviates the need for using multiple reaction receptacles, incorporating steps such as DNA extraction and pre-amplification into one tube system, and even promoting the possibility of multiplexing the detection of various targets. 76
Integration of CRISPR-Cas biosensors with diverse microfluidic chips 76 are comprised of paper-based analytical devices, 77 centrifugal chips, 78 digital chips, 79 electrochemical chips, 80 and wearable devices—which can provide real-time monitoring of DNA and RNA. 81 Many of these devices use a small reaction volume, which results in a higher template concentration to promote CRISPR-Cas detection without the need for a pre-amplification step. 82 Centrifugal chips compartmentalize the components necessary for DNA and RNA detection to minimize manual manipulation by the user, lessening the likelihood for contamination and allowing for the transfer of samples to separate reaction chambers via centrifugation. A centrifugal chip for the detection of SARS-CoV-2 reliant on reverse transcription, recombinase-based amplification, and a CRISPR-Cas component could detect as little as 1 copy/µL of purified, full-length viral RNA. 78 Electrochemical chips rely on an electric field for myriad purposes, including the separation and concentration of the DNA or RNA from a sample and then focusing on the isolated samples with the CRISPR-Cas components necessary for detection. This type of microfluidics was demonstrated to detect SARS-CoV-2 in about 35 min, inclusive of RNA extraction, with a sensitivity of detection of 10 copies of viral RNA/µL. 80 Digital microfluidics (e.g., digital chips) allow for detections and quantification of amplification-free CRISPR-Cas biosensors. 83 A proof-of-concept CRISPR-Cas-coupled microfluidic device targeting SARS-CoV-2 is capable of RNA quantification in a linear dynamic range of 0.6–2027 copies/µL. 84
Digital microfluidics can provide automation of fluid handling for multiple reaction steps within CRISPR-Cas biosensors. Lu et al. provide an example of this capability with the design of an automated CRISPR-Cas12a-based biosensor to detect bacterial pathogens, such as Staphylococcus aureus. 79 In this study, detection from a complex medium, such as milk and urine, was completed in under an hour and required only 6.6 µL of reagents. 79 Global advantages of microfluidics devices include minimized contamination thanks to the use of a closed system and further decreases in operating costs per reaction due to the minute amounts of reagents required. Overall, microfluidics provides solutions to the challenges related to quantification, multiplexing, and automation that limits traditional POU DNA and RNA detection approaches.
Translating the technology for the identification of pathogens causing wildlife diseases
Molecular detection of DNA and RNA associated to pathogens is not indicative of the disease in question and additional diagnostic approaches may be required. Many pathogenic bacteria can be found on a host without resulting in disease. For example, Pseudogymnoascus destructans (Pd), the fungal causative agent of white-nose syndrome in bats has been identified using qPCR markers in bat species that never develop any symptoms. 85
The CRISPR-Cas component can be programmed to target practically any DNA or RNA sequence by means of introducing a synthesizable “guide” RNA. 32 A guide RNA coupled to a target template pre-amplification method like RPA, only requires the design and validation of two oligonucleotides, and renders CRISPR-Cas biosensors adaptable with relative ease to target a suite of wildlife disease pathogens. 51 To date, some CRISPR-Cas biosensors have been developed that target pathogens associated with human or domestic animal health that are also relevant in the context of wildlife diseases (Table 1). In humans, CRISPR-Cas diagnostic tools have gained prominence in the detection of viral, bacterial, and microbial pathogens including Zika, Dengue, Ebola, SARS-CoV-2, M. tuberculosis, P. aeruginosa, and Plasmodium spp.95,75,196–199Some of these tools have been developed for animal and plant pathogen diagnostics to support the agricultural sector. Some examples in animals include CRISPR-Cas biosensors for the detection of White Spot Syndrome Virus in shrimp, 96 African swine fever virus, 97 porcine reproductive and respiratory syndrome virus, 98 canine parvovirus type 2, 99 white-nose syndrome, 90 and feline calicivirus. 100
Examples of available CRISPR-Cas biosensors for pathogen detection relevant to wildlife diseases
AIOD, all-in-one dual; FAM, Fluorescein; CRISPR-Cas, clustered regularly interspaced short palindromic repeats-CRISPR-associated protein; FINDeM, field-deployable isothermal nucleotide-based detection method; LAMP, loop-mediated isothermal amplification; Lw, Leptotrichia wadei; RPA, recombinase polymerase amplification; SHERLOCK, specific high-sensitivity enzymatic reporter unlocking; STOP, SHERLOCK testing in one pot.
Considerations to the broad application of novel technologies
New emerging technologies such as CRISPR-Cas biosensors are attractive tools for enabling widespread surveillance of wildlife disease-associated pathogens due to their relatively low cost to produce and operate and their ability to be programmed to detect a wide range of targets. Most users with basic laboratory equipment are capable of following published protocols to generate these biosensors in-house thanks to the commercial availability of most reagents and synthesize custom DNA and RNA components needed for targeted programming. However, the assembly of CRISPR-Cas biosensors can affect the reliability, sensitivity, and accuracy of a specific test and complicate the integration of this data into broader context, such as management decisions. Nonexpert molecular biologists, practitioners, or managers are encouraged to work with well-established entities, such as commercial, biotech labs, and expert research labs on the production of CRISPR-Cas kits. While proper design is necessary it can slow access of the technology, and capacity building and training are important, especially in underdeveloped economies.
CRISPR-Cas biosensors for the detection of wildlife disease-associated pathogens
CRISPR-Cas biosensors can assist with the rapid analysis of multiple mediums for the detection of DNA or RNA associated with wildlife disease pathogens. These tools enable users with limited resources to conduct their own tests at their facilities or the field with the caveat that detection of DNA and RNA does not necessarily correlate to disease and additional diagnostic approaches, such as histopathology, are required. Here, we provide examples of the CRISPR-Cas biosensors that have been developed to date for humans or wildlife or those that have been repurposed for wildlife. We also list a few examples of diseases that could benefit from the development or modification of CRISPR-Cas biosensors.
African swine fever
African swine fever is a highly infectious viral disease affecting domestic and wild pigs, notorious for its exceptionally high mortality rate of up to 100%. 101 Due to its severe economic impacts and the lack of an effective vaccine, the management of African swine fever virus has relied on rapid detection to control the extent of its impact on the porcine industry.102,103 Detection of the causative agent relies on traditional viral screens, including virus isolation, ELISA tests, and PCR. However, qPCR stands as the gold standard diagnostic method for African swine fever virus due to its high sensitivity and specificity.102,104,105 Several CRISPR-Cas biosensors based on the Cas12a and Cas13a nucleases have been developed that reliably distinguish this dsDNA virus among other swine viruses—including classical swine fever virus.91,106–108 One of these tools was reported to combine the pre-amplification and the CRISPR-Cas detection into a single—“one-pot”-reaction that greatly enhances the utility of these biosensors by end-users. 92 More recently, a similar approach integrating all steps necessary for CRISPR-Cas-based detection was developed within a microchip. 109 Moreover, detection of the various relevant African swine fever virus genotypes has been mostly focused on mitigating the effects in the pork industry. This disease is also relevant as it is also known to affect wild pigs, decimating their numbers in the wild and serving as a significant vector. 110 The virus exhibits remarkable resilience in the environment, allowing it to survive on clothing, footwear, vehicle wheels, and other materials. 111 CRISPR-Cas biosensors for detecting African swine fever virus could greatly aid in the surveillance of the virus in remote settings and potentially provide a quick test to screen wild pig carcasses and/or feces.112,113 This is particularly important, as other POU detection methods, such as lateral flow assays tracking viral antigens, have limited sensitivity (up to 44% when the sample was freeze-thawed) when testing the blood from wild boar carcasses. 112 Lastly, CRISPR-Cas can also be useful for biosecurity measures since the virus is known to persist in pork products, such as sausage and bacon. 114
SARS-CoV-2
A CRISPR-Cas biosensor is capable of targeting the SARS-CoV-2 virus, which has infected numerous species of pet, captive, farmed, and wild animals. 115 Among these, big cats such as African lions (Panthera leo) and a puma (Puma concolor) were reported to have been infected with this virus and display mild symptoms in a private zoo in Johannesburg, Republic of South Africa. 116 Other captive felids with reports of SARS-CoV-2 infections include tigers (Panthera tigris) and snow leopards (Uncia uncia).117,118 In January 2021, the San Diego Zoo Wildlife Alliance reported that their western lowland gorilla (Gorilla gorilla gorilla) cohort was positive for SARS-CoV-2 and displayed the respiratory symptoms characteristic to COVID-19. 119 Although the aforementioned cases were reported in captive individuals, spillage into wild populations of deer, feral minks, feral cats, and otters have been reported in various countries.120–123 Per the SARS-CoV-2 in Animals Situation Report No. Twenty-two (June 2023) of the World Organisation for Animal Health, 29 species have been reported worldwide to have been afflicted by this disease. 124 Spillover into imperiled species, like the critically endangered Florida panther (Puma concolor coryi) could negatively impact their population. Marine mammals can be highly susceptible to SARS-CoV-2 due to their social structure, interactions with humans, and exposure to wastewater that may contain viral particles.125,126 In 2019, four Atlantic bottlenose dolphins (Tursiops truncatus) at the U.S. Navy Marine Mammal Program in San Diego, CA displayed symptoms characteristic of a coronavirus infection. 127 Analyzes of fecal samples confirmed detection of bottlenose dolphin coronavirus (BdCoV). Although BdCoV is not related to SARS-CoV-2, this example highlights the susceptibility of marine mammals to these viruses. 127 SARS-CoV-2 CRISPR-Cas biosensors have been developed for human clinical samples and could serve as a platform for its transfer to target animals. 128
White-nose syndrome
White-nose syndrome is an infectious disease in bats that is caused by the Pd fungus. 129 This disease has killed millions of bats in North America, posing a significant threat to endangered species. 130 User-friendly POU tools based on CRISPR-Cas and LAMP have been developed for the detection of the Pd pathogen.21,90 The CRISPR-Cas assay was capable of detecting Pd DNA from dermal swabs without the need for DNA extraction and was capable of accurately assessing guano samples when coupled with a DNA extraction kit. 90 The LAMP assay demonstrated high species-specificity, discriminating Pd DNA from that of over 150 fungal species, and showed promise in detecting the fungal pathogen from noninvasive sources such as hibernacula walls without the need for DNA isolation and purification. 131 Neither of these methods are capable of quantification in their current iterations nor can be considered as standalone diagnostic tools. Nevertheless, as the technology progresses, POU tools have the potential to be as sensitive and reliable as traditional qPCR methods—the current gold standard for the molecular detection of pathogenic agents. 132
Avian influenza
A CRISPR-Cas13a biosensor was developed in 2019 to detect this low pathogenicity virus subtype to provide an early detection POU/field tool to alert of potential outbreaks in wild and farmed populations. 93 Highly pathogenic avian-origin influenza (HPAI) viruses are generally regarded as a major concern for the poultry and cattle sectors; however, these viruses have also been associated with mortality events of wild birds, including endangered species.133–135 The first such event was reported in the Republic of South Africa in 1961 among terns (Sterna spp.), which raised the possibility of infection between migrating sea birds and poultry as the virus was related to a chicken strain identified in Scotland in 1959. 136 After this event, numerous HPAI outbreaks have been reported in wild birds (such as waterfowl) across the world, which have been linked to a HPAI strain first identified in domestic geese from Guangdong, China.137–141 Early detection and rapid response to HPAI outbreaks in domestic poultry has been an effective strategy to mitigate the impacts of disease and has been suggested as a model to follow for wildlife managers. 135 Low pathogenicity avian influenza A (H7N9) affect wild birds, such as sparrows, which could serve as vectors to transmit the disease to poultry. 142 Avian influenza, also, poses a significant threat not only to bird populations but to various mammals. Strains like H5N1 and H7N9 have demonstrated the ability to cross the species barrier, infecting mammals such as rats, mice, weasels, ferrets, pigs, cats, tigers, dogs, and horses.143,144 As of June 2024, the U.S. Department of Agriculture reported that 118 dairy cow herds in 12 States have confirmed cases of highly pathogenic H5N1. 133 While birds are the primary carriers, transmission to mammals can result in illnesses ranging from mild to severe, occasionally leading to fatalities. 143 This ability of avian influenza viruses to affect mammals is concerning due to the potential for mutations or reassortment, which might yield strains more transmissible among mammals. Therefore, efforts to rapidly detect and monitor avian influenza among wild bird populations, and mammal carriers, with the help of CRISPR-Cas biosensors could help mitigate the potential devastating effects of avian influenza on wildlife, poultry industry, and ecosystems.
Chytridiomycosis disease
Aquatic chytrid fungi continues to be a major threat to amphibians since 1993 when it was first indicated as the pathogenic agent responsible for high mortality events in countries like Australia and Costa Rica.145–147 Chytridiomycosis is characterized by an epidermal infection in keratinized tissues that leads to mortality by affecting the osmotic balance of the host.148,149 These pathogens have been connected to the decline of at least 500 amphibian species globally, including the extinction of roughly 90 species. 150 Two water-borne species of chytrid fungi, Batrachochytrium dendrobatidis and B. salamandivorans, are responsible for chytridiomycosis and have been the target of eDNA detection approaches using traditional qPCR methods. 151 In 2023, field-deployable isothermal nucleotide-based detection method, a CRISPR-Cas biosensor was developed for the identification of B. dendrobatidis from epidermal swabs that had comparable sensitivity to the standard qPCR method. 86 At the time of publication, effective treatments for this disease were still in development, but once available, early treatment during low pathogen loads could increase treatment effectiveness. Therefore, early detection and rapid removal of individuals from infected water bodies could prove as a mitigation procedure.152–154 Early detection is of utmost importance in North America where a comprehensive survey of wild amphibians suggested the absence of one of the pathogens, B. salamandrivorans. 155
Sylvatic plague
Yersinia pestis, the causative bacterial agent of bubonic, pneumonic, and sylvatic plague, was likely introduced in the 1900s to the United States from rat-infested ships traveling from Asia, as supported by molecular studies indicating a similarity between North American Y. pestis strains to the latter region.156–158 Sylvatic plague, the variant of the disease when it occurs in rural wildlife, afflicts a wide range of mammals, including prairie dogs that are a staple prey of the endangered black-footed ferret (Mustela nigripes) and has the potential to affect the population of these wild mustelids even when there is no associated die-off of their prey items.159–161 In addition to DNA detection for Y. pestis, a rapid field serology method testing for sylvatic plague-related antibodies is available to survey for the prevalence of the bacterium across a wide host of species.162,163 The disease is controlled through pesticides targeting the fleas that transmit it and the vaccination of black-footed ferret through injection or oral vaccine deployed in food pellets. Serological blood tests require an invasive approach, whereas molecular methods can rely on minimally invasive use of flea vectors as a proxy to determine the presence of Y. pestis. 162 The application of DNA detection could be greatly enhanced with the development of a field-deployable method and could improve positive detection of the sylvatic plague agent. 164 Sylvatic plague vaccines are available and have been administered in the field to mitigate the incidence of outbreaks in keystone species like the prairie dog. 165 Early detection tools, such as CRISPR-Cas biosensors could help in the passive surveillance of Y. pestis for more rapid targeted applications of prophylactic methods.
Whirling disease
Whirling disease is a disease caused by the protozoan parasite Myxobolus cerebralis that affects the cartilage of wild salmonids.166,167 DNA detection methods for M. cerebralis based on PCR are available and eDNA from river water and sediments can be collected to survey for this parasite in a noninvasive manner.166,168 More recently, in a study by Lisnerová et al., eDNA metabarcoding was explored as a tool to survey myxozoa diversity, a group of endoparasites that mainly target fish, from freshwater sediments. 169 This pilot eDNA metabarcoding study identified over 50 different myxosporean operational taxonomic units, denoting high diversity of these endoparasites at each surveyed site. The rapid and targeted detection of eDNA associated to the M. cerebralis pathogen or Tubifex, its required intermediate host, is plausible with a CRISPR-Cas system. 167 Detection of fish pathogens can benefit both the wildlife conservation and the aquaculture sectors. A targeted CRISPR-Cas biosensor has been developed for the detection of Atlantic salmon (Salmo salar L.) eDNA from water sources. 71
Plant diseases
In plants, CRISPR-Cas-based diagnostics have mainly focused on the detection of pathogens that affect the agricultural sector. In tobacco plants, a CRISPR-Cas12a nuclease system was coupled with an RT-RPA pre-amplification step for the detection of tobacco mosaic virus. 170 In the same work, the same CRISPR-Cas system was designed and tested for the identification of potato viruses X and Y—two RNA viruses that pose a significant risk potato crops. 171 Similar CRISPR-Cas12a-based DNA or RNA detection systems, coupled to a LAMP pre-amplification step, have been developed for detecting tomato pathogens, such as the tomato yellow leaf curl and tomato leaf curl New Delhi DNA viruses and the tomato mosaic and tomato brown rugose fruit RNA viruses.172,173 CRISPR biosensors have been tested for the detection of nonviral agricultural pathogens such as the fungus Magnaporthe oryzae that affects rice and wheat and Candidatus Liberibacter asiaticus, the gram-negative bacterial causative agent of citrus greening disease.174–176 Targeting of transgenic rice containing a gene from the soil bacterium Bacillus thuringiensis conferring pest-resistance was possible using a CRISPR-Cas12a biosensor and the ongoing development of a CRISPR tool for the detection of banana bunchy top virus exemplifies the increased relevance of these tools for plant pathogen detection.175,177 CRISPR-Cas detections tools could prove useful for the POU detection of wild plant pathogens—an underdeveloped research area. One such example would be for the early detection of the pathogenic fungus, Cryphonectria parasitica, which is responsible for chestnut blight—a disease responsible for the obliteration of mature American chestnut trees (Castanea dentata) from the U.S. landscape. 178 To date, only a qPCR method exists for the detection of this fungal pathogen. 179 The native keystone ‘ōhi’a plant (Metrosideros polymorpha) of Hawai’i is threatened by a disease known as rapid ‘ōhi’a death (ROD) caused by two fungal pathogenic agents: Ceratocystis lukuohia and C. huliohia. 180 In addition to a qPCR assay, a standalone RPA POU DNA detection method was developed to enable for the early survey of these pathogens onsite. 181 Atkinson et al. reported that their RPA POU method also detects nontarget Ceratocystis species, making the RPA assay a candidate for adding a CRISPR-Cas-based assay as a checkpoint to promote species specificity. 181
Conclusions
CRISPR-Cas biosensors provide a robust system for diagnostics and detection as exemplified by the wide range of applications in diverse fields spanning medicine, conservation, and agriculture. These low-cost tools greatly enhance the potential for rapid and accurate screening of pathogens affecting underserved communities that lack the logistics and infrastructure to conduct their own molecular diagnostics.182,183 For example, a large percentage (68%) of the diagnosed Hepatitis B, an infectious disease of the liver, cases were reported in underdeveloped Pacific and African countries with relatively limited access to diagnostic tools.184,185 The application or low-cost, user-friendly CRISPR-Cas-based detection of Hepatitis B virus to this region could offer a solution to better manage the disease. 186
Benefits of CRISPR-Cas biosensors include their ability to provide POU/field testing in a user-friendly manner and the ease to be programmed to target new DNA or RNA targets. Other POU/field detection methods, such as LAMP and portable PCR, rely on target amplification as the sole determinant for detection. This approach is prone to false positives if amplification of nontarget DNA or RNA occurs. The introduction of the CRISPR-Cas component to POU/field detection methods provide additional specificity for targeted amplification methods that can improve the reliability of detection. 187 This property is key toward integrating data from novel pathogen detection tools into wildlife disease surveillance. Furthermore, the adoption of these low-cost, user-friendly tools can also promote passive surveillance by wildlife managers, irrespective of their technical capabilities, to improve early detection and rapid response efforts. 188
Conversely, the facile screening of the pathogens responsible for wildlife diseases could assist in the early detection of endemic and neglected zoonotic diseases and help reduce their impact in underdeveloped communities.189,190 Lastly, these underserved regions overlap with indigenous communities where industrialization is minimal, and biodiversity is rich.190,191 Strong surveillance of wildlife diseases in rural and high biodiversity areas can help to support global conservation efforts, which are intrinsically intertwined with our own wellbeing.
CRISPR-Cas for wildlife disease detection: Challenges and opportunities
CRISPR-Cas biosensors for DNA and RNA detection can, in theory, be programmed to recognize a novel target with relative ease if genomic sequences of the target are available. In this scenario, a new assay can be developed with the minimum requirements of designing a novel gRNA for CRISPR-Cas targeting and two oligonucleotides for the pre-amplification step if RPA is used (Fig. 1). An experienced scientist could develop a new, preliminary CRISPR-Cas assay in a matter of days to respond to emerging threats. However, as with any scientific assay, to ensure that this CRISPR-Cas biosensor is robust and specific to the intended target requires extensive validation and optimization, often testing of numerous samples encompassing both targets and nontargets and comparing results against a similar assay based on a more established technology such as qPCR.
CRISPR-Cas biosensors have proven to be reliable and accurate POU tools in fields such as human medicine. Nevertheless, the broad suite of methods available for this technology can complicate widespread adoption and data integration into repositories for data sharing and effective surveillance. Efforts by the scientific community could help to standardize CRISPR-Cas biosensor assays, especially regarding how data are interpreted. Extensive intercalibration and proficiency testing and careful interpretation of potentially dead or nondisease-causing organisms are important to improve reliability and help alleviate concerns of quality assurance.
CRISPR-Cas, renowned for its precision in cleaving target DNA or RNA, presents outstanding accuracy, sensitivity, and specificity. Its advantages over the other detection methods for pathogenic organisms’ genetic material is evident, highlighting its capability as a POU tool. 192 CRISPR-Cas could help to inform wildlife disease detection, carrying immense potential to enhance the effectiveness of surveillance and decision-making in managing these diseases. Nevertheless, there are still some challenges for this technology such as oversensitivity, off-target recognition, and false positives arising from the intrinsic capacity of the CRISPR system to tolerate mismatched nucleotides. 193
Beyond the complexities inherent in detecting wildlife diseases by CRISPR-Cas-based biosensing per se, the foremost challenge for the field lies in its standardization. Standardizing CRISPR-Cas-based biosensing involves meticulous consideration of numerous variables. Factors such as sample type and preparation, reaction conditions (buffers, duration composition, and temperature), inhibitors and contaminants, uniform signal output interpretation, and validation can significantly influence results and need to be carefully tested and validated in situ and in vivo. 87 A limit of detection (LOD) threshold is often calibrated for a fluorescence signal, below which any molecule is not detected with 95% certainty.70,194 However, the hand-held portable instruments that read the signal are not always equipped with quantitative functionality. Comparison and calibration of the assay’s LOD and limit of quantification with the established qPCR assay is highly recommended. Additionally, the standardized quantification metrics (i.e., signal measurements used to determine a positive detection) for CRISPR-Cas detection methods that do not rely on a microfluidics or digital component are needed for binary “negative” or “positive” detection. For example, CRISPR-Cas detection assays that rely on a visible fluorescence intensity (e.g., using a UV transilluminator to visualize fluorescence) can be subjective when it comes to discriminating between positive and negative detections.
Proper experimental study design in addition to the use of negative and positive controls are critical components of any study. Positive controls at various concentrations can assess the dynamic range of the assay and the lower concentration limits for which it can detect the target sequence. Negative controls are necessary to assess the potential for contamination. Further, negative controls from areas without the target organism present and blank samples containing only laboratory reagents and no nucleic acids can be used to assess inhibition or cross reactivity to nontarget sequences or innate compounds in the samples. For example, the additional DNA extraction step was necessary to overcome inhibitory compounds effecting the RPA or CRISPR-Cas activities in guano. 90 In that same study, it was found that compounds within bat guano also fluoresced with a UV transilluminator even when the target P. destructans was not present but did not fluoresce when using a portable isothermal fluorometer.
Quality control measures, like inter-laboratory calibration or proficiency testing, can effectively address detection accuracy and sensitivity concerns. Collaborative efforts among laboratory networks, official agencies, and ultimately the World Organisation for Animal Health to establish proficiency testing, standards, reference material, and products, could notably expedite the practical deployment of CRISPR-Cas for wildlife disease detection in the future. Standard reference materials are needed for shifting the design of CRISPR-Cas biosensors from an “artisanal” to industrial scale. The efforts toward standardization can mirror those of other consortia such as the international eDNA standardization task force (https://iestf.global/), which works closely with various International Organization for Standardization (https://www.iso.org/) working groups for the effective development of standards for a variety of applications.
While there have been strides in CRISPR-Cas sensors, these have been primarily designed for DNA and RNA detection. This highlights opportunities for pioneering strategies to develop CRISPR sensors capable of activating a more diverse range of targets beyond DNA and RNA. 195 Expanding the scope of these sensors could enhance wildlife disease detection and improve overall diagnostic capabilities.
Footnotes
Acknowledgments
We appreciate the expertise and insights within the U.S. Geological Survey Ecosystems Mission Area Wildlife Disease Community of Practice in writing this review article. We thank Dr. M. Camille Hopkins, the U.S. Geological Survey Wildlife Disease Specialist for the Ecosystems Mission Area, and Dr. Jessica M. Rieder at the U.S. Geological Survey Wetland and Aquatic Research Center for their comprehensive review of this article and providing thorough constructive feedback.
Authors’ Contributions
A.A.P., G.V.M., and M.E.H.: review and editing (equal). M.E.H.: conceptualization (lead). A.A.P.: writing—original draft (lead). A.A.P., G.V.M., and M.E.H.: writing—review and editing (equal). M.E.H.: supervision and resources.
Author Disclosure Statement
The authors declare no conflicts of interest.
Funding Information
We thank the
