Abstract
The genome-editing efficiency of the CRISPR-Cas9 system hinges on the recognition of the protospacer adjacent motif (PAM) sequence, which is essential for Cas9 binding to DNA. The commonly used Streptococcus pyogenes (SpyCas9) targets the 5′-NGG-3′ PAM sequence, which does not cover all the potential genomic-editing sites. To expand the toolbox for genome editing, SpyCas9 has been engineered to recognize flexible PAM sequences and Cas9 orthologs have been used to recognize novel PAM sequences. In this study, Abyssicoccus albus Cas9 (AalCas9, 1059 aa), which is smaller than SpyCas9, was found to recognize a unique 5′-NNACR-3′ PAM sequence. Modification of the guide RNA sequence improved the efficiency of AalCas9-mediated genome editing in both plant and human cells. Predicted structure-assisted introduction of a point mutation in the putative PAM recognition site shifted the sequence preference of AalCas9. These results provide insights into Cas9 diversity and novel tools for genome editing.
Introduction
Clustered regularly interspaced short palindromic repeat (CRISPR) and CRISPR-associated protein 9 (Cas9) in the prokaryotic immune system are important for genome editing.1–4 Cas9 forms a ribonucleoprotein (RNP) complex with two RNAs, CRISPR RNA (crRNA) which is derived from a prokaryotic genome-encoded CRISPR array and a trans-activating crRNA (tracrRNA) which forms crRNA–tracrRNA duplex.5–7 crRNA comprises a variable spacer region, which specifies the cleavage sequence, and a constant repeat region, which hybridizes to the tracrRNA. In genome-editing applications, crRNA and tracrRNA can be linked through a tetraloop to form a single-guide RNA (sgRNA). 5 The genomic target sequence is determined by complementarity to the spacer sequence of the crRNA or sgRNA.
Cas9-mediated recognition of a target site requires Cas9 binding to a protospacer adjacent motif (PAM) immediately downstream of the target sequence.5,8 PAM recognition is mediated by the PAM interacting domain (PID) of Cas9.9–11 The PID–PAM interaction leads to the local unwinding of the target sequence and base pairing with the spacer sequence in crRNA. 12 Each Cas9 has a PAM preference that is influenced by its PID. The widely used Streptococcus pyogenes Cas9 (SpyCas9, 1368 aa), which is type II-A, exhibits the highest activity toward the 5′-NGG-3′ PAM sequence. 5 Although Staphylococcus aureus Cas9 (SauCas9) and Streptococcus thermophilus Cas9 (St1Cas9) are both type II-A, they recognize different PAM sequences. SauCas9 recognizes the 5′-NNGRRT-3′ PAM sequence (where R represents a purine [i.e., A or G]), 13 whereas St1Cas9 recognizes the 5′-NNRGAA-3′ PAM sequence. 14 The diversity in the recognition of PAM sequences by Cas9 orthologs from various bacterial species is thought to be a result of the specificity shaped through competitive coevolution with viruses in the CRISPR-Cas9 system.15,16
The CRISPR-Cas9 system allows for precise genetic modifications, including base editing and prime editing, as well as gene activation or repression.17–22 To achieve optimal efficiency, precise targeting of genomic sites by Cas9 RNP at a one-base pair resolution is essential. Recognition of the PAM sequence restricts the sequence space that can be targeted by Cas9. Although engineered Cas9 variants with a flexible PAM have broadened the targeting abilities of the system, this involves a tradeoff, which reduces on-target activity. 23 Flexible PAMs increase the probability of off-target effects and Cas9 cleaving its own sgRNA coding sequence, 24 which could hinder the safe therapeutic application of Cas9.
To overcome this constraint of PAM sequences, the diversity of PAM sequences provided by naturally occurring Cas9 orthologs was utilized. Large-scale PAM screening of 79 Cas9 orthologs from various organisms identified A-, C-, T-, and G-rich PAM sequences ranging from 1 to ≥4 bases. 16 Mining of Cas9 orthologs in metagenomic databases has revealed 96 Cas9 orthologs (from 41 subgroups) with in vitro cleavage activity and diverse PAM compatibilities. 25 Although extensive mining of Cas9 orthologs has widened the scope of PAM sequences, only a subset of Cas9 orthologs with diverse PAM sequences has been effective for genome editing in mammalian cells and plants.
Here, we identified several PAM sequences from uncharacterized Cas9 orthologs smaller than SpyCas9 and found that Abyssicoccus albus Cas9 (AalCas9, 1059 aa) recognizes a unique PAM sequence (5′-NNACR-3′) distinct from that of known Cas9 orthologs. 16 Optimization of sgRNA length and sequence improved the genome-editing efficiency of AalCas9 in Arabidopsis thaliana protoplasts. AalCas9 with a modified sgRNA exhibits genome-editing activity in HEK293FT cells and A. thaliana, causing indels near the targeted 5′-NNACR-3′ PAM sequence. A point mutation in the putative PAM recognition site on AalCas9 PID, predicted by comparing its AlphaFold2 26 structure with the SauCas9–sgRNA–DNA crystal structure, expanded the PAM sequence preference. These results provide insights into Cas9 diversity and novel tools for genome editing.
Materials and Methods
Identification of type II Cas9 nuclease orthologs
Cas9 orthologs with smaller size than SpyCas9 were identified through a homology-based search of the NCBI nonredundant protein sequence database using the amino acid sequence of SpyCas9 as query. The accession numbers and amino acid sequences of the Cas9 orthologs are listed in Supplementary Table S1.
Identification of crRNA and tracrRNA
The crRNA sequence for each Cas9 ortholog was predicted using CRISPRCasFinder 27 based on the direct repeat sequence annotated in GenBank. We identified tracrRNA sequences by searching for complementary crRNA sequences near cas9. The tracrRNA and sgRNA sequences used are listed in Supplementary Table S1. sgRNAs were designed by connecting crRNA and tracrRNA with a GAAA tetraloop, and their secondary structures were predicted with Mfold. 28
Recombinant Cas9 expression and purification
Escherichia coli codon-optimized genes encoding the Cas9 orthologs, AalCas9, Acidiphilium sp. 21-60-14 Cas9 (AspCas9), Caulobacterales bacterium Cas9 (CbaCas9), and Rhodopila globiformis Cas9 (RglCas9) were synthesized by Integrated DNA Technologies and used to express recombinant Cas9 in E. coli. Cas9 orthologs with N-terminal SV40 nuclear localization signal (NLS) and C-terminal NP NLS-His6 were expressed under the T7 promoter in a pET28 plasmid backbone in the E. coli strain Rosetta 2(DE3) pLysS cells (Novagen). The plasmid for AalCas9H990N expression was generated by site-directed mutagenesis. The His-tagged Cas9 orthologs and a variant were purified as described for SpyCas9 expression with minor modifications 29 (see Supplementary Data S1). The purified Cas9 orthologs were used to determine the PAM sequences and cleave DNA in vitro. Supplementary Table S2 contains plasmid sequences for the expression of Cas9 orthologs and AalCas9H990N (pTOP18, pTOP19, pTOP22, pTOP23, and pTOP321).
In vitro synthesis of sgRNA
The sgRNAs used in this study were synthesized by in vitro transcription using T7 RNA polymerase, as previously described. 29 The transcribed sgRNA products were purified by gel extraction after 8% denaturing urea-polyacrylamide gel electrophoresis, and used to prepare the Cas9 RNP complex. The sgRNAs and primer sequences used for preparation of the DNA templates are listed in Supplementary Tables S1 and S3.
PAM sequence determination with RNPs
PAM sequences of Cas9 orthologs were determined by PAM depletion assay using the 7N-PAM plasmid library that contained a target sequence with 5′-NNNNNNN-3′ (7N) degenerate sequences. To generate the 7N-PAM plasmid library (Supplementary Table S2), the 7N-containing oligonucleotide was converted into double-stranded DNA using ReverTra Ace (TaKaRa Bio) and the following primers: 5′-GGTCTTTGCTCCTGCAG-3′ and 5′-AGCTATGACCATGATTACGAATTCNNNNNNNCTGCAGGAGCAAAGACC-3′. The resulting dsDNA fragments were assembled into a pUC18 plasmid with the target-and next-generation sequence (NGS)-adapter sequences using the NEBuilder HiFi DNA Assembly Cloning Kit (NEW ENGLAND Biolabs). E. coli DH5α cells were transformed with the reaction mixture and incubated at 37°C on an Luria–Bertani plate supplemented with 100 mg/L ampicillin. More than 100,000 colonies were washed off the plates, and the plasmids were purified using the NucleoBond Midiprep Kit (TaKaRa Bio) to prepare the 7N-PAM plasmid library. Prior to the digestion of the 7N-PAM library, the Cas9 RNP complex, with an sgRNA targeting the 5′-GAGAGGATCGTTGGTCTTTGCTCCTGCAG-3′ sequence in the 7N-PAM plasmid, was prepared by mixing 2.5 μM Cas9 and 12.5 μM sgRNA in a Cas9 buffer (20 mM HEPES-NaOH [pH 7.5], 100 mM NaCl, 5 mM MgCl2, and 0.1 mM EDTA) and incubated at 25°C for 20 min. The RNP was incubated with 20 nM of the 7N-PAM plasmid library in the Cas9 buffer at 30°C for 2 h. The reaction was stopped by incubation with 60 mM EDTA at 60°C for 5 min. Undigested plasmid DNAs were purified by gel extraction after agarose gel electrophoresis. The 0.4-kb DNA fragments containing the NGS-adaptor sequence and the target sequence followed by 7N sequence were amplified by PCR using the purified undigested plasmid as template with the following primer set: 5′-GCGATCGGTGCGGGCCTCTTCGCTATTACGC-3′ and 5′-ATTAGGCACCCCAGGCTTTACACTTTATG-3′. Subsequently, the RNPs were incubated at 37°C for 30 min with 200 nM of DNA fragments amplified from undigested plasmid DNAs. The undigested DNA fragments were purified by gel extraction and used in a second PCR to add Illumina barcodes for NGS analysis.
Sequencing was performed using HiSeq X Ten (Macrogen) to obtain fastq sequence data. We extracted 7-bp random bases from the quality-filtered reads and calculated the frequency of particular sequences from Cas9 RNP-treated samples and untreated control. The fraction (read counts for a particular 7-bp sequence/total read counts) was calculated in each type of 7-bp read, and sequences that were >90% depleted in Cas9 RNP-treated samples compared to the untreated control were extracted. The consensus sequence of 131 depleted sequences was visualized using WebLogo. 30
In vitro dsDNA cleavage assay
The 0.35-kb dsDNA fragments containing five target sequences in A. thaliana PHYTOENE DESATURASE 3 (AtPDS3-1–5) were amplified by PCR using A. thaliana genomic DNA as a template with the appropriate primer sets (Supplementary Table S3, for the substrate DNA sequences, See Supplementary Table S4) and purified for use as substrate DNAs for the dsDNA cleavage assay with Cas9 RNP. Before the dsDNA cleavage reaction, the AalCas9 RNP complex was prepared by incubating 1 μM AalCas9 and 2 μM sgRNA in the Cas9 buffer for 10 min at room temperature. The dsDNA substrate (10 nM) was mixed with 100 nM AalCas9 RNP complex containing AtPDS3-targeting sgRNA in 10 μL of Cas9 buffer and incubated at 25°C for 30 min. The reaction mixture was quenched by incubation with 50 mM EDTA at 60°C for 5 min. The reaction products were analyzed using a MultiNA microchip electrophoresis system (Shimadzu Inc.). To analyze the PAM preference of the AalCas9H990N protein, dsDNA substrates targeted by AtPDS3-3 sgRNA_v2, in which the PAM sequence, 5′-TAACG-3′, was replaced with 5′-TAACA-3′, 5′-TAACT-3′ and 5′-TAACC-3′, respectively, were used (see Supplementary Table S4). To prepare AalCas9- and AalCas9H990N-RNP complexes, 2 μM AtPDS3-3 sgRNA_v2 was incubated with 1 μM AalCas9 or AalCas9H990N in the Cas9 buffer for 10 min at room temperature. The dsDNA substrate (10 nM) was incubated with 100 nM AalCas9 or AalCas9H990N RNP complex in 10 μL of Cas9 buffer at 37°C for 5 min. The reaction products were analyzed as described above. To address the temperature dependency of dsDNA cleavage by AalCas9, 10 μL reactions containing 10 nM substrate dsDNA in the Cas9 buffer were pre-heated from 15°C to 60°C, and then the cleavage reactions were started by adding 100 nM AalCas9 RNP. After 2 min, the reaction was stopped by adding 50 mM EDTA and 2 mg/mL proteinase K. The reaction products were analyzed using a MultiNA microchip electrophoresis system. For the time-course experiment, 10 nM of dsDNA substrate was incubated at 22°C with 100 nM AalCas9 RNP in 40 μL Cas9 buffer. Then 10 μL aliquots were withdrawn from the reactions at 5, 15, and 30 min, and the reaction was stopped by incubating with 50 mM EDTA at 60°C for 5 min. The amount of the cleaved dsDNA was quantified with MultiNA as described above. All in vitro cleavage assays were performed at least three times.
Evaluation of genome-editing efficiency in HEK293FT cells
HEK293FT cells (ATCC) were incubated with Dulbecco’s modified Eagle medium (Wako) including 10% fetal bovine serum (Biosera, lot 017BS856) in 5% CO2 condition. To coexpress AalCas9 and sgRNA in human cultured cells, we constructed plasmids encoding CAG promoter-driven human codon-optimized AalCas9 with bipartite NLS and U6 promoter-driven sgRNA targeting AAVS1, CXCR4, EMX1, KRAS, or VEGFA (pTOP266-pTOP273, pTOP308-pTOP317; Supplementary Table S2). Plasmids for AalCas9H990N expression were generated by site-directed mutagenesis using primers and pTOP266, pTOP308, pTOP311, pTOP314, and pTOP315 as templates (Supplementary Table S2). As a control, the cultured cells were transfected with plasmids (pTOP324 and pTOP364-pTOP367) for expression of SpyCas9 and sgRNAs. Each plasmid was mixed with the TransIT-X2 Dynamic Delivery System (Mirus) as 2:1 ratio and 15 μL aliquots added to HEK293FT (2 × 104) cells in a 96-well plate. The cells were cultured in D10 medium for 4 days and total cellular DNA was extracted with 0.05N NaOH. Cellular DNA was used as a template for amplifying the editing target sequence using KOD One PCR Master Mix (TOYOBO) and primers (Supplementary Table S3). PCR amplicons were used for subsequent amplicon-seq analysis using iSeq100 (Illumina) (see Supplementary Data S1).
In silico analysis of three-dimensional structures of Cas9 orthologs
Structural models of AalCas9, Massilibacterium senegalense Cas9 (MseCas9), and Bacillus niameyensis Cas9 (BniCas9) were calculated using AlphaFold2 26 or ColabFold. 31 Superposition of the PIDs of Cas9 orthologs with the crystal structure of the SauCas9–sgRNA–DNA complex (PDB ID: 5AXW) was performed using TM-align. 32 The structure-based multiple sequence alignment was generated using UCSF Chimera. 33 Structural figures were drawn using PyMOL (The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC.).
Statistics and reproducibility
Quantified genome-editing data were analyzed and visualized using R version 4.1.1 (R Core Team). The source data and R scripts used for analysis are available on GitHub (https://github.com/aist-pgrrg/AalCas9). Hiseq X Ten output data was deposited to Short Read Archive (PRJNA1072618).
Results
Identification of PAM sequences for Cas9 orthologs
Four CRISPR-Cas9 systems with smaller Cas9 than SpyCas9 were randomly selected from the database to expand the genome-editing toolbox of CRISPR-Cas9 (Supplementary Table S1). Recombinant Cas9 orthologs fused with an NLS and histidine tag were expressed in E. coli and purified. To identify the tracrRNA and crRNA sequences of Cas9 orthologs, crRNAs were predicted with CRISPRCasFinder 27 using the direct repeat sequence annotated in GenBank. Then, tracrRNA sequences were searched in the region closely located to cas9 based on sequence complementarity with crRNA. We designed sgRNA sequences in silico based on the predicted tracrRNA and crRNA sequences (Supplementary Fig. S1). The sgRNAs for Cas9 orthologs with the same target sequence were transcribed in vitro and purified by urea-page gel extraction. To determine the PAM sequences of Cas9 orthologs, we performed the PAM depletion assay 34 using in vitro assembled Cas9–sgRNA RNP complexes and the 7N-PAM plasmid library containing a target sequence with 7N degenerate sequences at the 3′ region (Fig. 1A). We identified the putative PAM sequences for AspCas9, CbaCas9, RglCas9, and AalCas9 (Fig. 1B). A previous study classified Cas9 orthologs based on the PID sequence and revealed weak correlations between PID sequences and their corresponding PAM sequences. 16 Compared to the classification of Cas9 orthologs based on PID sequence, three putative PAM sequences of AspCas9, RglCas9, and CbaCas9 could be aligned to the PID classification (Supplementary Table S5). 16 Notably, AalCas9 recognizes the PAM sequence 5′-NNACR-3′, which was unexpected based on PID classification and to our knowledge, has not been reported in experimentally validated Cas9. In the following sections, we analyzed the AalCas9–sgRNA RNP because of the uniqueness of the PAM recognition by AalCas9.
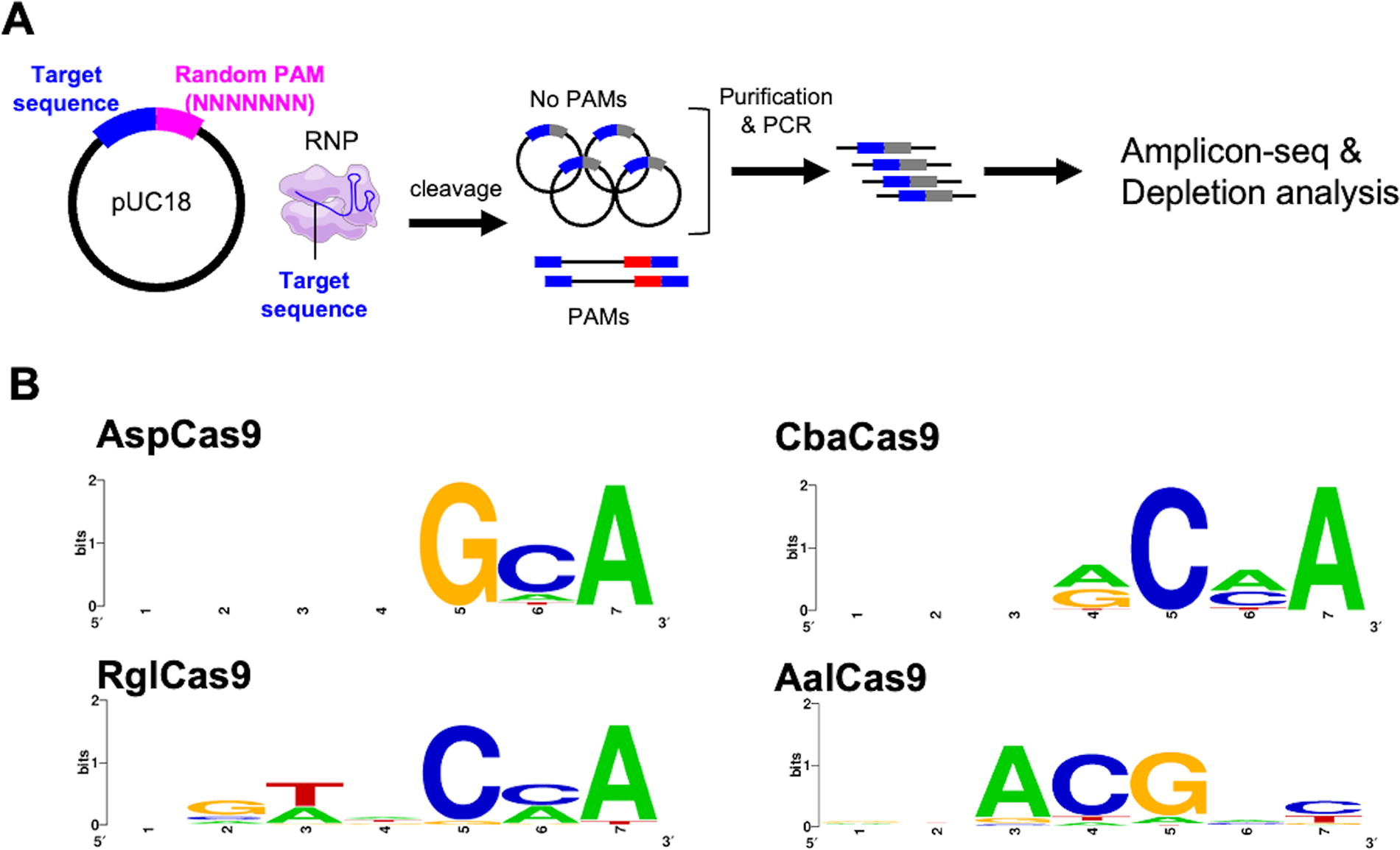
Identification of the PAM for Cas9 orthologs.
Cleavage of dsDNA by AalCas9 in a PAM-dependent manner
To confirm the PAM preference of AalCas9 determined by the PAM depletion assay, we tested the in vitro DNA cleavage activity of AalCas9 RNP for five target sites in A. thaliana PHYTOENE DESATURASE 3 (AtPDS3), which harbors the following PAM sequences in the 3′ region: 5′-NNACG-3′, 5′-NNACA-3′, 5′-NNTAA-3′, and 5′-NNAAG-3′ (Fig. 2A). We detected DNA cleavage activity in three out of five targets that harbored the 5′-NNACG-3′ or 5′-NNACA-3′ PAM sequence, consistent with the results of the PAM depletion assay (Fig. 2B).
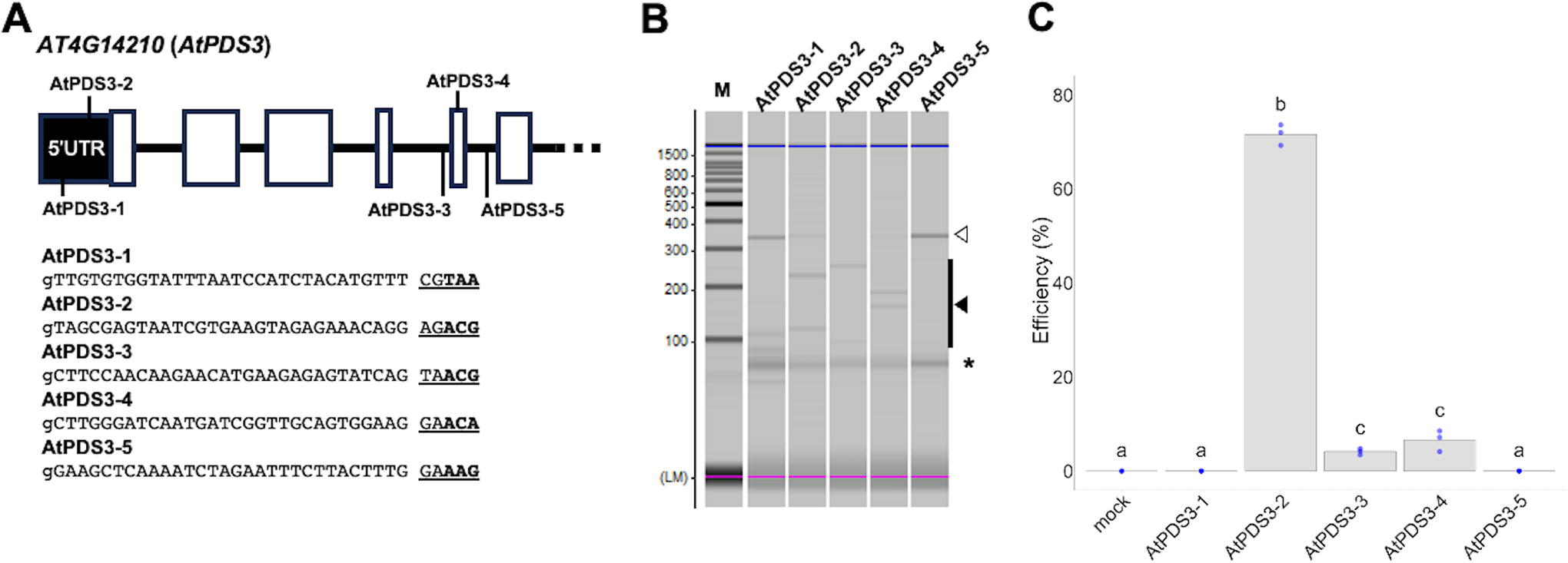
Enzymatic characterization of AalCas9.
We also tested the genome-editing activity of AalCas9 RNP in A. thaliana protoplasts (Fig. 2C). Consistent with the results of the in vitro DNA cleavage assay, genome editing in A. thaliana protoplasts by AalCas9 RNP was detected for the three sgRNAs that were also active in vitro. Although all the sgRNAs targeting with 5’-NNACR-3’ PAM showed high nuclease activity in vitro, the genome-editing activity in A. thaliana protoplasts did not have a similar trend. The discrepancy between the gRNA activity in vitro and in vivo has been reported in CRISPR-Cas9-based genome editing. 35 Remarkably, the RNP harboring AtPDS3-2 sgRNA showed higher genome-editing activity than SpyCas9 RNP harboring the sgRNA, which was reported to edit AtPDS3 (Supplementary Fig. S2). Taken together, we concluded that AalCas9 RNP recognizes the 5′-NNACR-3′ PAM sequence and can be used for genome editing in cells.
Improved efficiency of AalCas9 with modified sgRNA
In the A. albus CRISPR-Cas9 system, a potential tracrRNA–crRNA duplex consists of three stem loops (SLs) and a 30-nt target sequence (Supplementary Fig. S1). Genome-editing activity was detected by using the 134-nt sgRNA harboring these sequences (Figs. 2 and 3A). The clustering of PIDs among Cas9 orthologs by Gasiunas et al. (2020) 16 showed that AalCas9 belongs to cluster 4–8 (Supplementary Table S5). This cluster includes SauCas9, which is widely used in genome editing. Consistently, the sgRNA of AalCas9 has a high similarity to the sgRNA of SauCas9 (Fig. 3A, B, >60% identity). In order to evaluate the impact of sgRNA sequence on AalCas9 RNP activity in vivo, we analyzed several sgRNA variants that exhibit greater similarity with SauCas9 sgRNA. Especially, SL1 and SL3 in AalCas9 sgRNA are dissimilar to that in SauCas9 sgRNA. Deleting the 7- or 10-bp duplexes in the SL1 improved genome-editing activity (Fig. 3C). In contrast, deletion of a 3-bp duplex in the SL3 abolished genome-editing activity, suggesting that SL3 is essential for the function of the AalCas9–sgRNA complex (Fig. 3C). Genome-editing activity was more pronounced when target sequence length was shortened from 30- to 20-nt compared to a longer (>24-nt) target sequence. However, shortening it to 18-nt abolished genome-editing activity (Fig. 3C), which is consistent with previously reported characteristics of SauCas9. 36 The sgRNA variants with truncated target sequence and shortened SL1, 105-nt long in a total length, exhibited higher genome-editing activity than the original 134-nt sgRNA (Fig. 3D). Additive or synergistic effects of a combination of the two mutations on genome-editing activity were not observed (Fig. 3D and Supplementary Table S6). The effect of sgRNA mutations on genome-editing activity was supported by an in vitro DNA cleavage assay (Supplementary Fig. S3). The 105-nt sgRNA of AalCas9 has high (>80%) similarity to the sgRNA of SauCas9 (Fig. 3E and F). We designated the 105-nt long sgRNA as sgRNA_v2 (Fig. 3E), a modified version of the sgRNA for AalCas9, and the sgRNA_v2 scaffold was used in the following experiments.
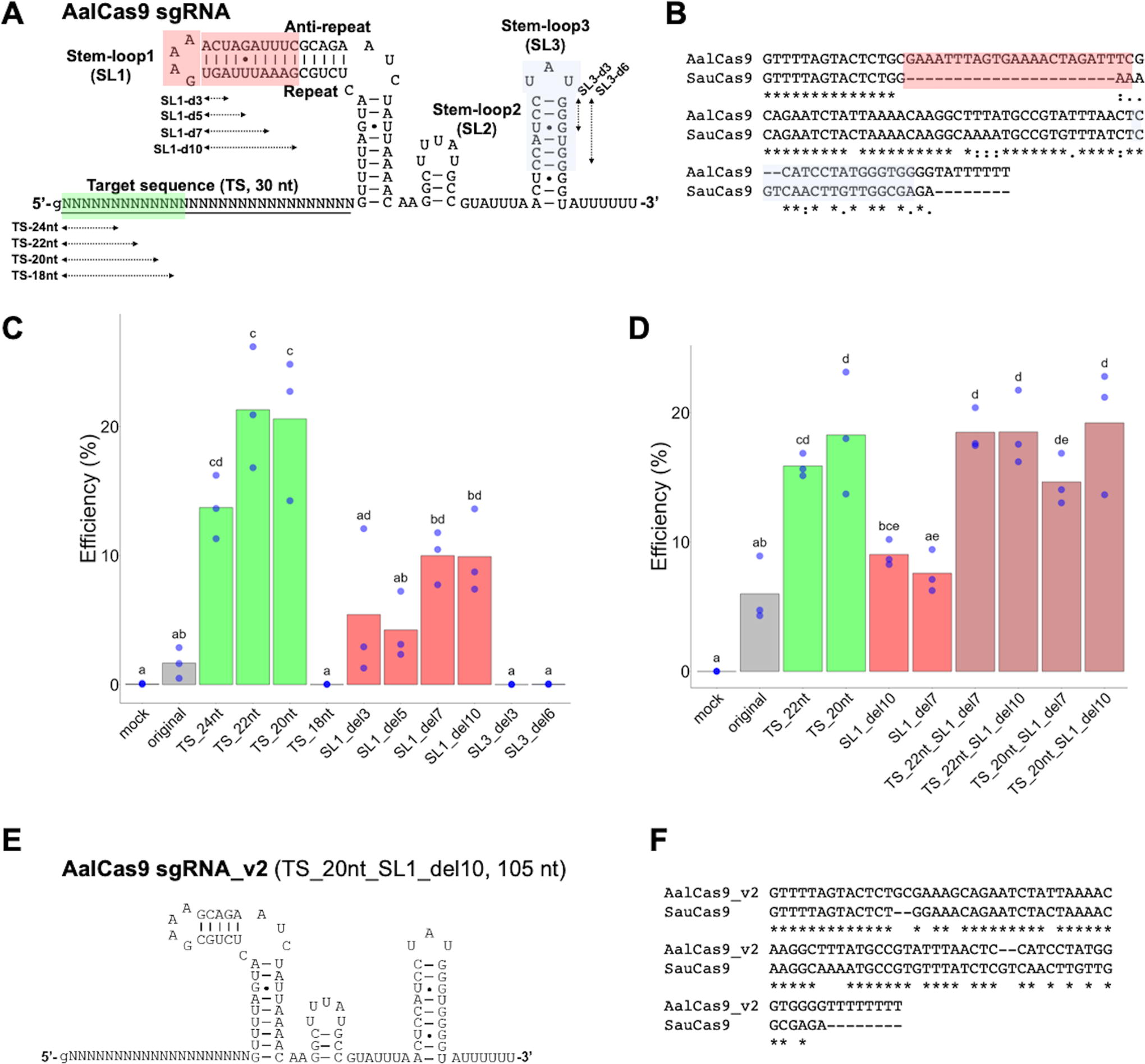
Effects of AalCas9 sgRNAs variants on genome-editing activity.
AalCas9-mediated efficient genome editing in planta and in human cell lines
As all our data were obtained using RNPs, we constructed vectors to express both AalCas9 and sgRNA_v2 in plants and animals, and their genome-editing activities were tested in A. thaliana and HEK293FT cells (Fig. 4). By using the vector harboring human codon-optimized AalCas9 with the bipartite NLS (NLS–AalCas9–NLS) driven by the A. thaliana RIBOSOMAL PROTEIN S5A (RPS5A) promoter, genome editing was achieved in A. thaliana protoplasts with an efficiency of approximately 6% in AtPDS3–gRNA2, although the vector harboring SpyCas9 instead of AalCas9 showed genome editing with an efficiency of approximately 4%, which is not statistically significant to the efficiency of AtPDS3–gRNA2 by AalCas9 vector (Fig. 4A). For the assessment of gRNA target sequence dependency, we tested 11 sgRNAs and found the clear tendency that SpyCas9 is superior to AalCas9 in genome-editing activity in A. thaliana protoplasts (Supplementary Fig. S4). A transgenic approach via Agrobacterium-mediated transformation was used to generate A. thaliana genome-edited mutants with mutations in AtPDS3 using sgRNAs, AtPDS3–gRNA2 or AtPDS3–gRNA4. Biallelic homozygous mutants of AtPDS3 were isolated in the T1 generation, which were transformed with AalCas9 expression plasmids as a binary vector (24 biallelic mutants in 25 transgenic lines expressing AtPDS3–gRNA2 and seven mosaic mutations in 12 transgenic lines expressing AtPDS3–gRNA4 (Supplementary Fig. S5A). Among the sgRNAs, only AtPDS3–gRNA4 targets the coding sequence of AtPDS3 (Fig. 2A). Some leaves of the T1 plants transformed with AalCas9 and AtPDS3–gRNA4 exhibited variegated leaf phenotypes, likely due to the malfunctioning of AtPDS3 (Fig. 4A).
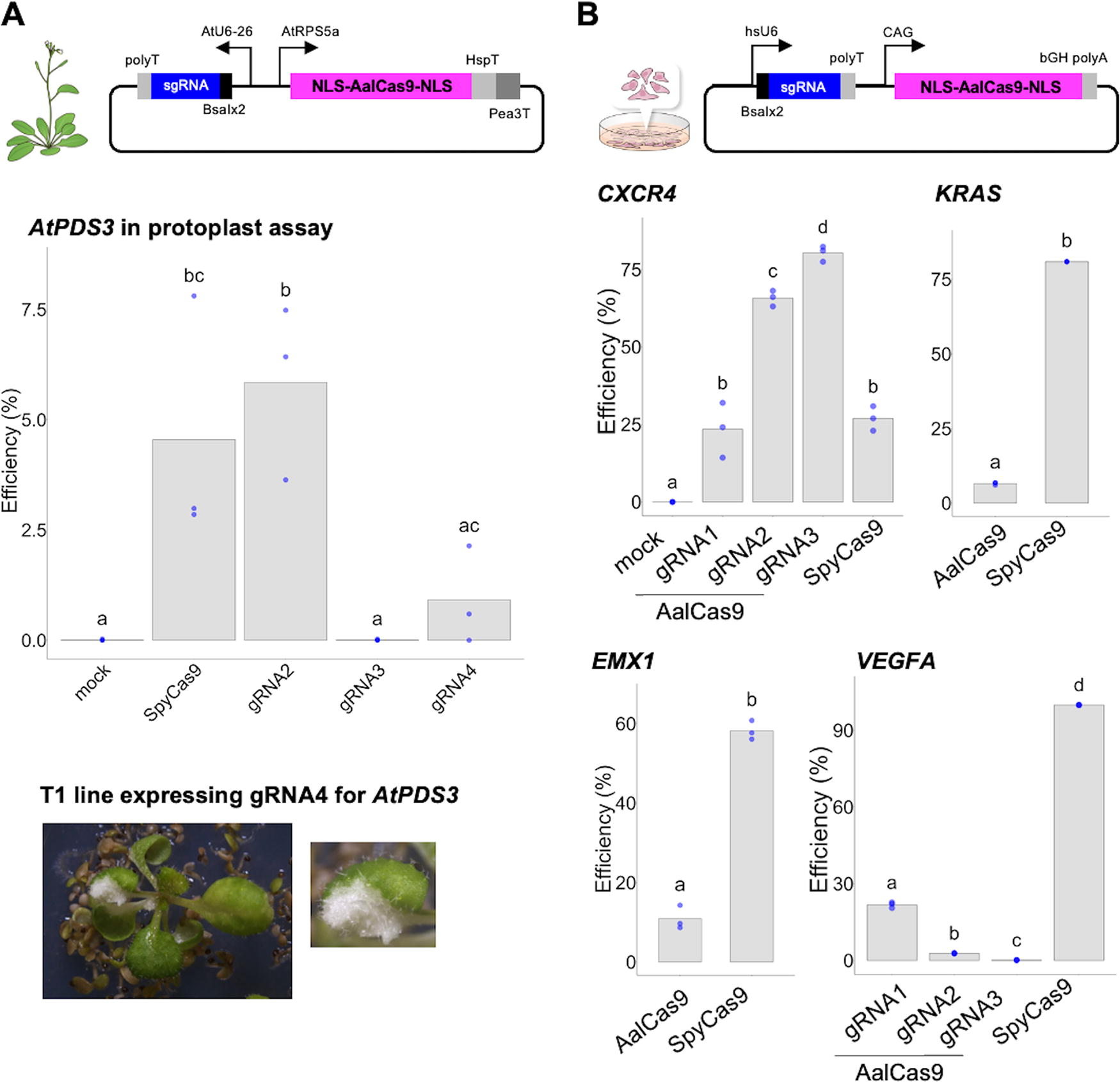
Genome editing in A. thaliana and a mammalian cell line was performed using vectors that harbored expression cassettes of nucleus-localized AalCas9 and its sgRNA_v2.
Considering the relatively low genome-editing efficiency in A. thaliana, we tested temperature-dependent nuclease activity of AalCas9 with sgRNA_v2 complex in vitro (Supplementary Fig. S6). As expected, we detected relatively low nuclease activity around 20°C, where A. thaliana protoplasts incubated. Interestingly, AalCas9–sgRNA_v2 complex showed high nuclease activity (>80% digestion) at 55°C, that is unexpected since the A. albus is derived from deep sea sediments, the environment with relatively lower temperature. From these results, it is expected that the genome-editing activity using AalCas9 could be applicable to human cell lines. A vector harboring the CAG promoter-based expression cassette of NLS–AalCas9–NLS was constructed and tested in HEK293FT cells (Fig. 4B). This vector demonstrated successful introduction of genome editing at target sites with 5′-NNACG-3′ PAM sequence in CXCR4, EMX1, KRAS, and VEGFA (Fig. 4B). The genome-editing efficiency of AalCas9–sgRNA_v2 system was basically lower than that of SpyCas9. The mutation patterns induced by AalCas9 were similar to those previously reported for SpyCas9 (Supplementary Fig. S5B). 13 To check the variance in genome-editing efficiency in various target sequences, eight target sequences harboring 5′-NNACG-3′ PAM in the AAVS1 gene were tested in HEK293FT cells (Supplementary Fig. S7). As in the case of A. thaliana, the genome-editing efficiencies of these sgRNAs exhibited large differences depending on the target sequence. The variation in genome-editing efficiency across different target sequence has been reported. 37 While the present AalCas9-based genome-editing system showed relatively low efficiency for practical usage, there is a need for improvement in on-target activity in the future.
Assessment of AalCas9 PAM preference by analysis of predicted structure of its PID
AalCas9 belongs to cluster 4–8 CRISPR-Cas9 system classified in Gasiunas et al., 16 although it has unique PAM preference compared with other orthologs in cluster 4–8 (Supplementary Table S5 and Supplementary Fig. S8A). To investigate how the PID of AalCas9 recognizes the unique PAM sequence, the predicted structure of AalCas9 generated using AlphaFold2 38 (AalCas9-AF2) was superposed onto the crystal structure of the SauCas9–sgRNA–DNA complex, which is previously experimentally validated 10 (PDB ID: 5AWX, Fig. 5A and B). Furthermore, a structure-based multiple sequence alignment of PIDs from four Cas9 orthologs in the cluster 4–8 (SauCas9, AalCas9-AF2, MseCas9-AF2, and BniCas9-AF2) was generated using TM-align and UCSF Chimera (Fig. 5C and Supplementary Fig. S8B). The PID of AalCas9-AF2 (residues 915–1059) was found to align well with that of SauCas9 (TM score = 0.93, and root mean square deviation = 1.22 Å for 142 residues) (Fig. 5A). SauCas9 recognizes the 5′-NNGRRT-3′ PAM sequence through specific residues in the β5-7 sheet region of PID, 10 and substitution of amino acids in the β5-7 sheet region alters PAM preferences 39 (Fig. 5B left). Comparing PAM sequences of Cas9 orthologs in cluster 4–8 suggests that the 4th cytosine of 5′-NNACR-3′ is particularly unique in the PAM recognition by AalCas9. In SauCas9, Asn985 is crucial for recognizing the N7 atom of the 4th and 5th purines in 5′-NNGRRT-3′ PAM. 10 His990, as observed in AalCas9-AF2, corresponded to Asn985 in SauCas9 (Fig. 5B right). The sequence alignment of PID revealed that this residue was diverse among Cas9 orthologs in the cluster 4–8, which is consistent with the PAM sequence diversity of the 4th and 5th bases (Fig. 5C). These data suggest that His990 in AalCas9 is a key residue that recognizes the 4th and 5th bases in the 5′-NNACR-3′ PAM sequence. To examine the role of His990 of AalCas9 in the 4th and 5th base recognition of the 5′-NNACR-3′ PAM sequence, we altered the His990 to Asn as SauCas9 (AalCas9H990N) and performed PAM depletion analysis. The PAM depletion assay revealed that AalCas9H990N prefers 5′-NNAC-3′ as the PAM sequence. This result suggests that the PID of AalCas9H990N loses the interaction with the 5th purine of the wild type 5′-NNACR-3′ PAM sequence (Fig. 5D). Consistently, recombinant AalCas9H990N showed higher cleavage activity in the 5′-NNACT-3′ or in the 5′-NNACC-3′ target than wild-type AalCas9 (Fig. 5E). Finally, we assessed the AalCas9H990N activity with sgRNA_v2 in HEK293FT cells (Fig. 5F). Interestingly, the AalCas9H990N activity was significantly reduced in comparison with the wild-type AalCas9. This implies that rigid 5′-ACR-3′ PAM recognition is necessary for maintaining the stability of the AalCas9–sgRNA–DNA ternary complex and His990 has significant contribution to it as expected.
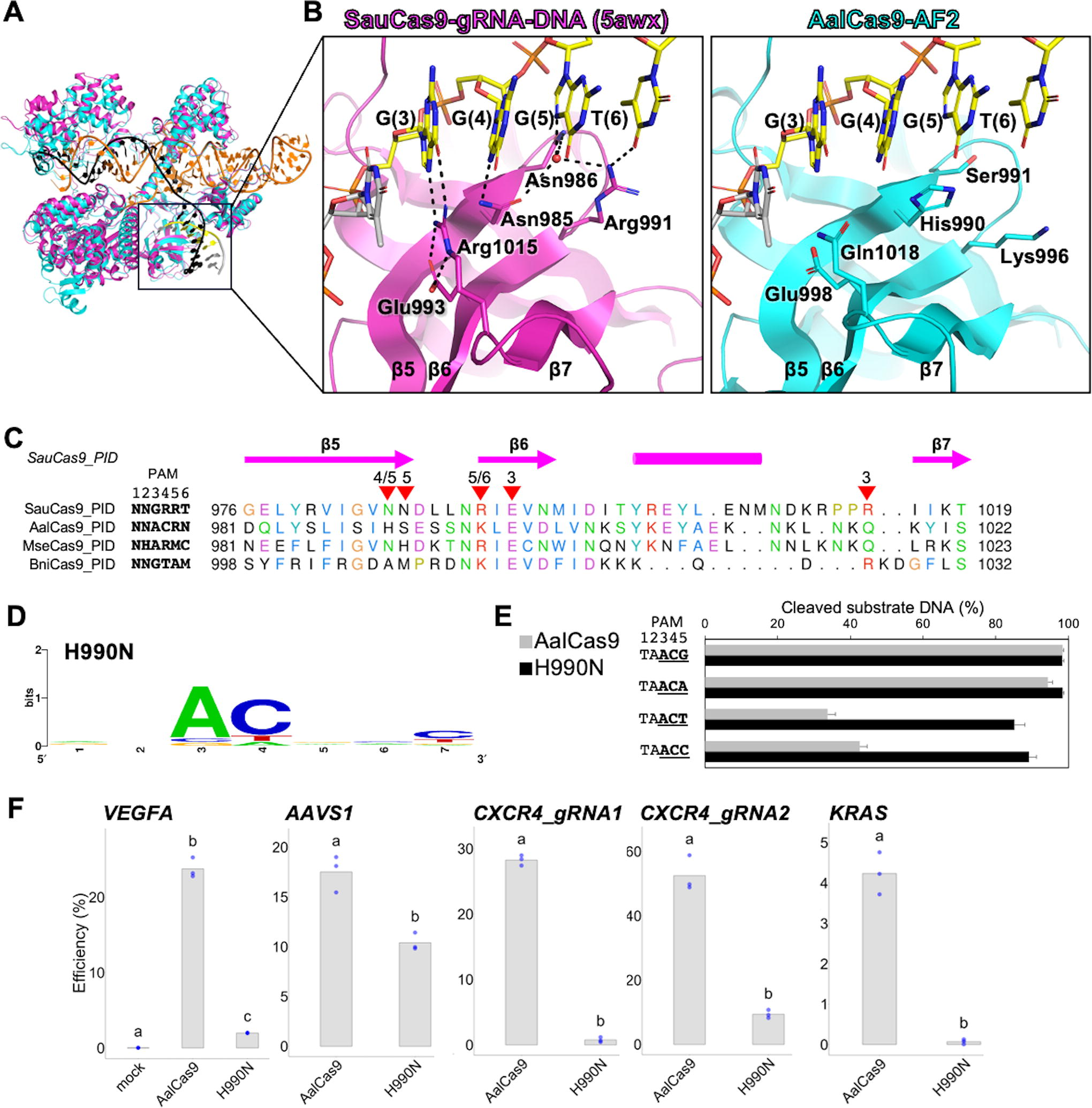
Prediction and analysis of the PAM sequence recognition site in AalCas9.
Discussion
This study identified the PAM sequences of four Cas9 orthologs. AalCas9 was found to recognize the unique PAM sequence 5′-NNACR-3′, which other closely related Cas9 orthologs do not recognize (Fig. 1 and Supplementary Fig. S8). Based on the structural comparison of AalCas9 and SauCas9, which is an ortholog of AalCas9 and works with sgRNA highly similar to that of AalCas9, we found that His990 in the β5-7 sheet region of AalCas9 could participate in recognizing the unique 5′-NNACR-3′ PAM sequence. Consistently, the PAM sequence preference of AalCas9 was successfully expanded from 5′-NNACR-3′ to 5′-NNAC-3′ by introducing a point mutation at His990, although the genome-editing activity of AalCas9H990N in vivo was considerably low (Fig. 5D and F). The structural comparison suggests that the substitution of His990 with Asn could result in the loss of the interaction with the 4th and 5th bases of the PAM sequence. However, AalCas9H990N still showed a preference for the 4th base, indicating that additional residues in AalCas9 are involved in the interaction with the 4th base. Because the structure predicted by AlphaFold2 was less accurate in the nucleic acid-binding site, 40 structural information of the AalCas9–sgRNA–DNA complex would be worth examining for the better understanding of PAM sequence recognition of AalCas9. Interestingly, the amino acid residues predicted to influence PAM preference in AalCas9 were less conserved than those in the sgRNA interaction region (Supplementary Fig. S9). Based on this variation, we hypothesized that the evolution of PAM interaction region is faster than that of sgRNA interaction region. 16 Accordingly, the findings of this study indicate that PAM preference in Cas9 rapidly evolves under selective pressure in the arms race between phages and bacteria. 39
The sgRNA of AalCas9 could be shortened to 105-nt. This engineered sgRNA showed a higher genome-editing efficiency than the original sgRNA (Fig. 3). Although SL1, derived from the conjugation of crRNA and tracrRNA by a tetraloop, can be deleted to some extent, SL3 is indispensable (Fig. 3). The 105-nt sgRNA sequence was very similar to that of SauCas9 (Fig. 3F), which is consistent with the fact that AalCas9 belongs to the Cas9 orthologs category (cluster 4–8) 16 which SauCas9 belongs to. In combination with the results of the change in PAM preference, this supports the hypothesis that the crRNA–tracrRNA duplex evolves slower than PAM recognition of Cas9.
AalCas9 from A. albus, isolated from deep-sea sediment, showed DNA cleavage activity at a broad range of temperatures exceeding 55°C, although the nuclease activity decreased to less than 50% around 20°C (Supplementary Fig. S6). Consistently, AalCas9 induced genome editing in A. thaliana, which was incubated at 22°C. However, the efficiency was relatively lower than that observed in human cell lines (Figs. 2–4). As the genome-editing activity of AalCas9 is highly dependent on the target sequence of sgRNA, several sgRNAs should be tested for practical use in the present AalCas9-based genome-editing system. Although there could be still much room for improvements of its genome-editing activity, AalCas9 would be a fascinating choice for the target genome locus harboring 5′-NNACR-3′ not only in human cell lines but also in plants. In addition, AalCas9H990N showed broader PAM preference (5′-NNACN-3′), than the wild type, thereby increasing the likelihood of using AalCas9 for broad genome-editing applications targeting 5′-NNACN-3′ sequences (Fig. 5). AalCas9 contains 1059 amino acids and is relatively smaller than SpyCas9, which has 1368 amino acids. AalCas9-based genome editing has a wide range of applications, including delivery using adeno-associated viral vectors. In conclusion, our findings highlight the potential of AalCas9 as a valuable addition to the genome-editing toolbox, expanding the possibilities of genetic manipulation in various biological systems by taking advantage of its unique PAM preference.
Footnotes
Acknowledgments
We thank A. Kuwazawa, S. Tsujikawa, Y. Taguchi, E. Yokoyama, S. Kobayashi, Y. Kimura, and Y. Yamada for their technical assistance. We also thank Dr. H. Tamaki for the usage of iSeq100.
Authors’ Contributions
A.N.: Conceptualization: developed the foundational ideas and design of the research; investigation: conducted gRNA identification, its improvement, AalCas9 purification, in vitro gRNA synthesis, and in vitro cleavage assay including PAM assessments; performed the analysis of the predicted structure of AalCas9 and designed its mutants; writing—original draft: contributed to drafting the article; writing—review and editing: reviewed and edited the article. H.Y.: Investigation: executed genome-editing experiments on Arabidopsis and human cell lines; performed in vitro translation-based PAM depletion assay; writing—original draft: contributed to drafting the article; writing—review and editing: assisted in drafting and revising the article. T.Y. and R.H.: Investigation: conducted in vitro sgRNA synthesis and in vitro DNA cleavage assay. N.M.: Conceptualization: participated in developing the research’s foundational ideas and design; writing—review and editing: reviewed and edited the article. T.T.: Conceptualization: contributed to the development of the research’s foundational ideas and design. Y.M. and S.I.: Conceptualization: aided in formulating the research ideas and design; investigation: assisted in performing Amplicon-seq analyses including PAM depletion assay; writing—review and editing: reviewed and edited the article. S.S.S.: Conceptualization: developed the foundational ideas and design of the research; investigation: conducted Amplicon-seq analyses including PAM depletion assay; participated in genome-editing experiments on Arabidopsis and human cell lines; writing—original draft: contributed to drafting the article; writing—review and editing: contributed to reviewing and editing the article.
Author Disclosure Statement
R.H., T.Y., and T.T. are employees of Inplanta Innovations Inc. Y.M. and S.I. are employees of TOPPAN Inc. A patent has been filed on behalf of JP2022165722A by AIST, Inplanta Innovations Inc., and TOPPAN Inc.
Funding Information
This work was supported by TOPPAN Inc.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
