Abstract
CRISPR-Cas technology has transformed our ability to introduce targeted modifications, allowing unconventional animal models such as pigs to model human diseases and improve its value for food production. The main concern with using the technology is the possibility of introducing unwanted modifications in the genome. In this study, we illustrate a pipeline to comprehensively identify off-targeting events on a global scale in the genome of three different gene-edited pig models. Whole genome sequencing paired with an off-targeting prediction software tool filtered off-targeting events amongst natural variations present in gene-edited pigs. This pipeline confirmed two known off-targeting events in IGH knockout pigs, AR and RBFOX1, and identified other presumably off-targeted loci. Independent validation of the off-targeting events using other gene-edited DNA confirmed two novel off-targeting events in RAG2/IL2RG knockout pig models. This unique strategy offers a novel tool to detect off-targeting events in genetically heterogeneous species after genome editing.
Introduction
The ability to introduce site specific modifications into the pig genome using the clustered, regularly interspaced, short palindromic repeat (CRISPR) technology has revolutionized the production of gene-edited (GE) pigs for both agriculture and biomedical applications. Advancements in this technology have paved the way for novel methods to produce GE pigs at high efficiencies. While the benefit of the technology is evident, the possibility of causing unintended modifications elsewhere in the genome has remained the main concern. The CRISPR system often utilizes targeted genomic double strand breaks (DSBs), produced by the CRISPR associated protein 9 (Cas9), to introduce site-specific genetic modifications. 1 A 20-nucleotide single guide RNA (sgRNA) directs the Cas9 nuclease to a specific location in the genome, and Cas9 produces DSBs if a protospacer adjacent motif (PAM) containing an NGG site is present. 2 The DSBs activate the cell’s DNA repair mechanism; however, small changes to the genomic sequences, that is, insertions-deletions (InDels), can be introduced as an outcome. Since the DSB relies on the short 20 nucleotide guide sequence to direct the endonuclease, Cas9, unintended modifications, known as off-targeting events, to the genome have been considered to be a potential side effect of the technology. In addition, if utilized in embryos, there is a chance for a high rate of mosaicism to occur. 3 This creates a challenge when studying founder animals that may contain more than two alleles, confounding phenotyping data.
Pigs are an ideal animal model for human diseases 4 and are the second most consumed meat globally. 5 Therefore, the use of gene editing technology in pigs for promoting human health research and identifying ways to improve pig production for agriculture purposes has increased immensely. 6 Unlike the genome of traditional laboratory animals such as the mouse, the pig is generally not inbred, and therefore, a high level of natural variation exists in the genome among different breeds. The high level of natural variation markedly complicates the detection of true off-targeting events as there is not an ideal way to filter only the modifications originated from CRISPR-Cas system when natural variation exists against a reference genome. In addition, during GE pig production, unknown genetic background is often introduced maternally as oocytes are provided by abattoir-derived ovaries, and therefore, the genetic background of founder GE pigs is not fully characterized. Various prediction software and biochemical methods have been developed to predict off-targeting events and shown to be effective;7–9 however, to the best of our knowledge, there is not a system that can rapidly identify off-targeting events in gene-edited organisms carrying unknown genetic background. As the CRISPR-Cas system is now routinely used to introduce targeted modifications into nonconventional animal models such as pigs, which carry a heterogeneous genome and the parental genetic information is not necessarily available, it is important to develop a strategy that allows us to identify potential off-targeting events. Previously, we detected off-targeting events in gene-edited pigs by searching for targeted loci. 10 While screening a handful of genes by targeted PCR has been informative, our priority is to identify and screen novel off-targeting events on a global level.
In this study, we utilized whole genome sequencing (WGS) paired with an off-targeting prediction software to evaluate the level of off-targeting events in the genome of three GE pig models carrying different levels of off-targeting events. The system can confirm known off-targeting events in the gene-edited pigs while also detecting novel off-targeting events, thus building an effective approach to overcome the difficulty in screening off-targeting events in heterogeneous animal models.
Methods
sgRNA design
Design of single guide RNAs (sgRNAs) to generate pigs lacking functional immunoglobulin heavy locus (IGH) and carrying modified recombination activating 2 (RAG2) and interleukin 2 receptor subunit gamma (IL2RG) was previously published.11,12 The sgRNAs targeting amyloid beta precursor protein (APP) were designed using CRISPOR 13 as previously described 10 (Fig. 1). Potential sgRNAs were aligned to the Sus Scrofa 11.1 reference genome using the Basic Local Alignment Search Tool (BLAST) to determine specificity. The designed sgRNAs were then in vitro transcribed to produce RNA that was coinjected with Cas9 RNA into zygotes as previously described. 14
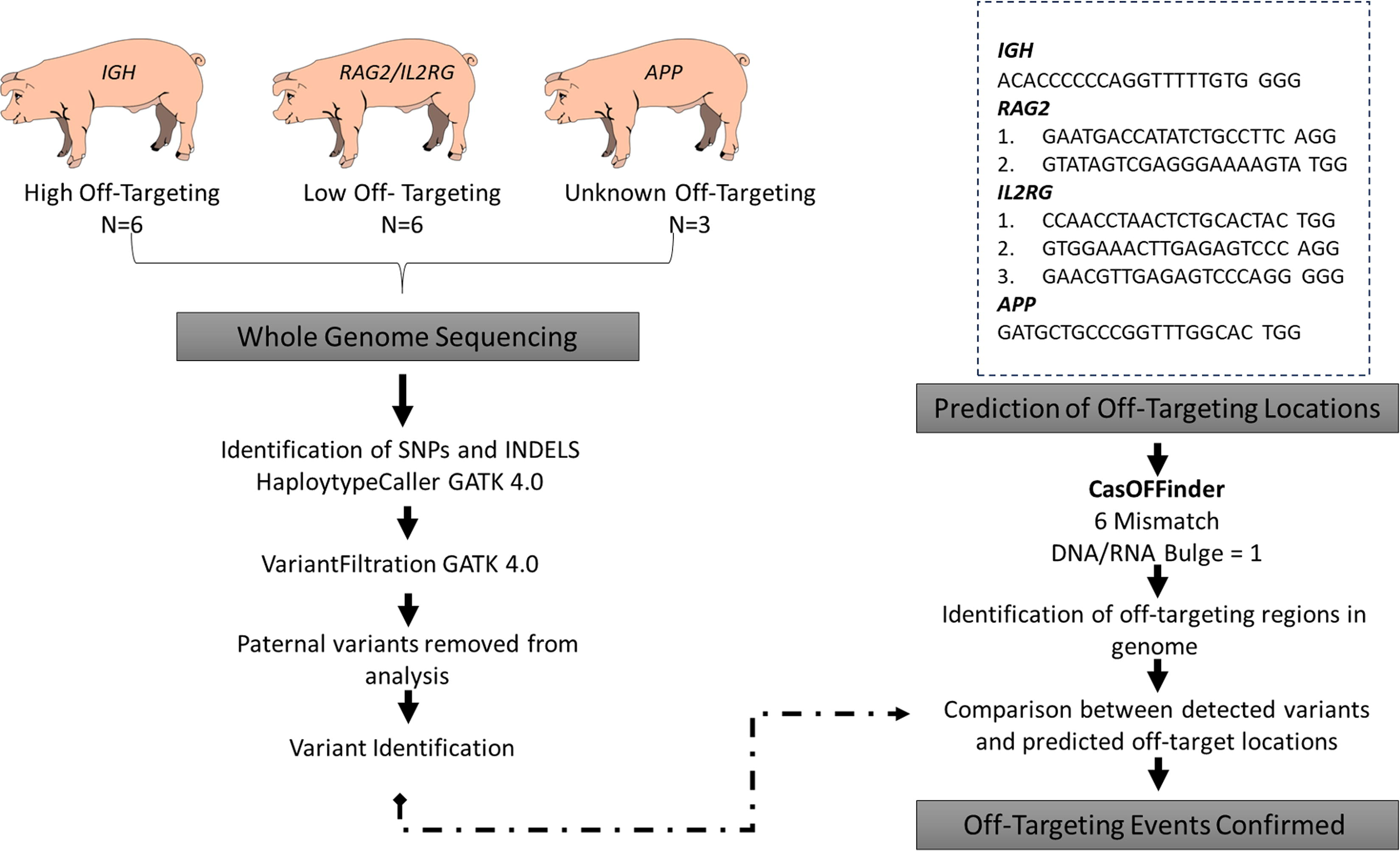
WGS and prediction workflow of identifying off-target modifications in GE pigs.
GE embryo and pig production
All animals were maintained according to the approved protocol and standard operating procedures by the Institutional Animal Care and Use Committee of the University of Missouri. Mature oocytes were coincubated with wild-type boar semen for 4 h at 38.5°C in an atmosphere of 5% CO2 in air as previously described. 15 Mature oocytes were fertilized and then injected with 10 ng/µl APP sgRNA + 20 ng/µl Cas9 RNA and then incubated in PZM3-MU3 medium16–18 at 38.5°C in an atmosphere of 5% O2, 90% N2, 5% CO2 in air until day 6 postfertilization. Individual blastocysts were screened by PCR to determine whether the sgRNA was producing high rates of on-target modifications. Once efficiency was validated, 40 to 50 injected blastocysts were transferred to a surrogate on day 5–6 after estrus was detected. The resulting APP founder piglets were collected by cesarean section on day 116 of gestation. Multiple tissues were collected from three APP knockout pigs to identify on- and off-target events and mosaicism on a global level.
To validate off-targeting events identified in WGS analysis, sgRNAs targeting IGH + Cas9 RNA or RAG2 and IL2RG sgRNAs + Cas9 were injected into zygotes as described above. On day 7 after fertilization, at least 20 single embryos for each model were collected to assess off-target modifications using targeted PCR and subsequent Sanger Sequencing. PCR products from selected presumable off-targeting locations were used for cloning to detect all alleles present in the developing embryos. Off-target modifications were then assessed by aligning the sequence to WT and the SScrofa 11.1 genome.
Whole genome sequencing and on- and off-target identification
For all GE pig models, genomic DNA was isolated from knockout founder pigs (IGH n =6, RAG2 and IL2RG n = 6, and APP n = 3, from brain, lung, muscle, spleen, and white blood cells [WBCs]) and from the boars that were used to create the GE embryos. These samples were submitted to the University of Missouri DNA Genomics Technology Core for WGS using NovaSeq flow cell technology. Bioinformatic analyses were conducted on the “Lewis” HPC cluster at the University of Missouri. Variant calling was accomplished using a custom Nextflow workflow to highly parallelize the processes. 19 Reads were mapped to the Sscrofa11.1 reference genome 20 using Minimap2. 21 Next, duplicated reads were marked and removed using MarkDuplicates in GATK (v4.2.6.1), and variants for each sample were called using HaplotypeCaller in GATK (v4.2.6.1). 22 Finally, hard filters for both single nucleotide polymorphisms (SNPs) and InDels were applied with VariantFiltration in GATK (v4.2.6.1) following GATK best practices, 23 and only InDels were used for further analyses.
The CRISPR off-target sites at the genome level were estimated by Cas-OFFinder 24 using the same reference genome and the sgRNA with allowed mismatch number = 6, DNA bulge size =1, and RNA bulge size = 1. Potential off-targeting loci from the Cas-OFFinder were matched to genome locations from the variance identified in the whole-genome sequencing; identities encompassing 50 bp on both sides of the InDel were analyzed to account for larger deletions or insertions to also be identified. To further filter the off-target sites for each sample, the CRISPR edited pig’s parental line was sequenced. The off-target sites for each sample were excluded if they also appeared in its parental line: Pumba was the paternal line for IGH and RAG2/IL2RG GE pigs, while W60 was the paternal line for the APP GE pigs. Finally, the remaining candidate off-target sites were identified as the off-target sites for each sample with its specific CRISPR sgRNA.
Candidate off-target sites were further analyzed by aligning each guide to the specific location in the genome of the GE pigs where off-target modifications were predicted to occur. Potential off-targeting loci carrying sequence identity to sgRNA and the presence of a PAM were suspected to be genetic variation, that is, off-targeting, introduced by the CRISPR-Cas system. Suspected off-targeting driven variations were searched on all pig genomic database available on the NCBI to identify if the variation existed in other pig breeds naturally.
Results
WGS of GE founder pigs
To investigate whether off-target modifications exist on a global level in GE pigs, we analyzed the genome of 15 founder pigs representing three different pig models. The genomic DNA from IGH and RAG2/IL2RG pig models generated in previous reports11,12 and the genomic DNA of APP knockout pigs were generated using sgRNAs that presented high editing efficacy (100%). In previous off-target analysis of the IGH GE pigs, two off-target genes (RBFOX1 and AR) were found to contain modifications at the frequency of 70% and 80%, respectively. No off-targeting event(s) were identified in the RAG2/IL2RG pigs from the previous study. To investigate the off-targeting potential of the guides on a global scale, WGS was completed from genomic DNA of the pigs. The WGS output was aligned to the pig reference genome (Sscrofa 11.1) and found to have an average of 96% coverage. Using the Genome Analysis Toolkit, regions harboring variations to the Sscrofa 11.1 were filtered to generate variant call format (VCF) files. The number of SNPs and InDels from each pig/sample are listed in Table 1. The average number of SNPs identified in our pig models compared to the reference genome ranged from 4,356,183 to 5,277,150. While lower, the number of InDels identified ranged from 1,360,695 to 1,551,824. The level of variation was similar in wild-type genomic DNA paternal line, Pumba, who is the paternal line for both IGH and RAG2/IL2RG, as the GE founder animals; however, the paternal line of APP, W60, had fewer total variants than the other pigs (66,654 InDels and 224,654 SNPs). The significantly lower level of variance is presumably due to the presence of Duroc genetic background in W60; Sscrofa 11.1 reference genome originated from a Duroc pig.25,26 On-target modifications were identified as expected, that is, all on-target modifications matched Sanger sequencing-based genotypes of each founder pig.
WGS Analysis of gene edited pigs and paternal lines to identify variation compared to Sscrofa 11.1 reference genome
Off-target prediction
The high level of variants in our founder pigs compared to the reference was expected as pigs are a heterogeneous animal model; however, the level of variations masked off-targeting events from the dataset and was impossible to extract off-targeting information. Therefore, a list of potential off-target sites was generated using the Cas-OFFinder software 24 using PAM NGG sites and allowing up to 6 mismatches with one RNA and/or DNA bulge compared to the on-target sequence. The WGS variants that were present in the Cas-OFFinder output predicting the off-target location, including 50 bp on either side of the variant location, were selected for further evaluation. There were a similar number of potential off-target locations identified from each sgRNA with an average of 8,608 ± 116 off-target locations (Table 2). However, as expected, the sgRNA designed to target IGH had the highest predicted off-target locations (9,071). The IGH sgRNA was designed within an exon that was less than 100 bp; thus, the sgRNA was not projected to have high specificity and have the potential to produce off-target modifications. 10 From the list generated by Cas-OFFinder, the VCF files were filtered to identify regions obtained from Cas-OFFinder and then the sorted variants were evaluated to determine whether they were a result of unintended editing by the CRISPR-Cas system (Fig. 1). Since the genetic background of the paternal genome was known, variations that originated from the paternal genome were also excluded.
Predicted off-target sites identified by Cas-OFFinder
This approach significantly reduced the number of potential off-targeting locations. For example, over 1 million locations were found to carry InDel variations against the reference genome (Table 1). By utilizing Cas-OFFinder, potential off-targeting sites were reduced to <100 locations carrying InDels in the six IGH knockout pigs (Fig. 2). The sequence identity of the potential off-target locations against sequences of CRISPR-Cas system was verified using the alignment tool in VectorBuilder to determine if the variations could be derived from the CRISPR-Cas system (Fig. 2). We analyzed whether there was an adjacent PAM sequence that could induce Cas9 to introduce off-target event(s). The variations detected included on-target allele modifications that were previously confirmed from genotyping; however, in one APP GE pig, we identified a different composition of alleles in different tissues, indicating mosaicism in the animal.
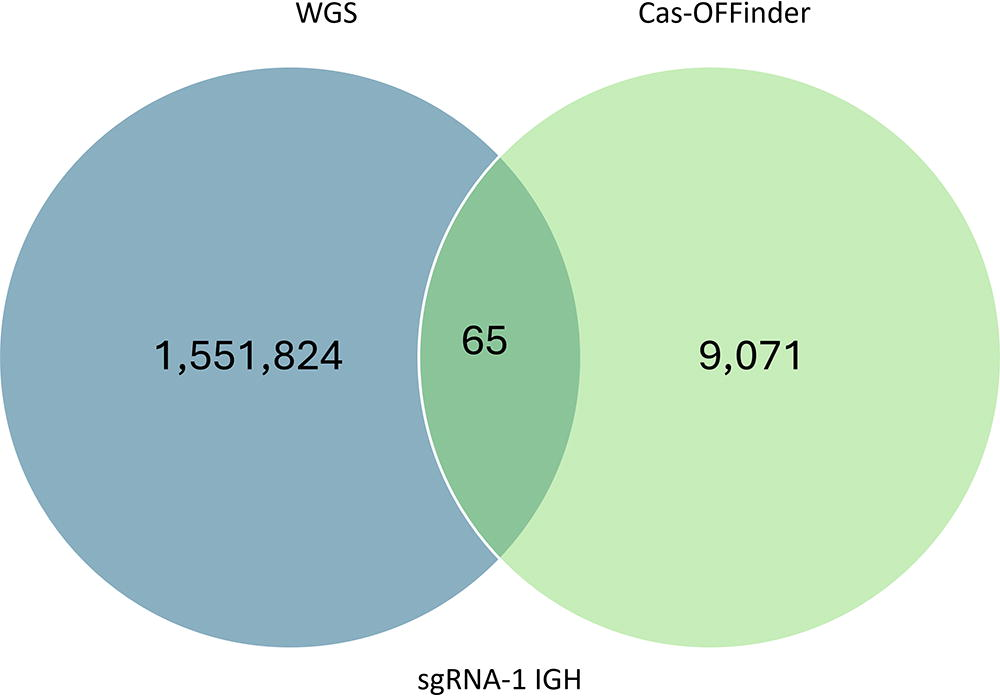
Representative Venn diagram illustrating the number of overlapping off-target locations between WGS and Cas-OFFinder for sgRNA-1 used to create IGH 1–2 pig.
Identification of GE sequence variation of founder pigs
The pipeline confirmed previously detected off-targeting events in IGH knockout pigs, that is, AR and RBFOX1, and generate a list of novel off-target sites in the genome (Table 3). Each location identified, whether in an intron/exon or unannotated, was next to a PAM sequence, which could make them more susceptible to endonuclease activity by Cas9. Two additional prospective off-target locations were identified in IGH knockout pigs and eight potential off-targeting locations were discovered in the genome of RAG2/IL2RG pigs (Table 3). Each sgRNA used to generate RAG2/IL2RG pigs may have contributed to introducing 1–4 off-targeting events. For instance, sgRNA designed to target IL2RG gene may have introduced an unintended modification to DDRGK1; two out of six edited pigs carried variation in the gene compared to the reference sequence.
List of novel off-target genomic locations identified by screening WGS variants paired with Cas-OFFinder prediction software
Validation of the off-targeting events in two pig models
To validate whether these identified sites were true off-targeting events, knockout embryos were generated independently by injecting designed CRISPR-Cas system targeting IGH or RAG2/IL2RG into fertilized oocytes. Genotyping of the embryos revealed that most of the potential off-targeting sites were either natural variations existing in other pig breeds or the frequency was too low to detect. For example, two embryos injected with sgRNA for IGH contained a 4 bp deletion in the intron of ribosomal protein S6 kinase c1 (RPS6KC1) compared to the Sscrofa 11.1. However, the variation was also observed in the genome of Ningxiang pig breed, thus determined as a natural variation rather than an off-targeting event by the Cas9 nuclease. Similarly, most of the potential off-targeting locations identified from the WGS were confirmed as natural variations or no off-targeting events were identified from the independent screening.
The independent off-targeting detection identified two locations in the genome of RAG2/IL2RG pigs that were suspected to be off-targeting events introduced by Cas9. Two genes, epithelial cell transforming 2 like (ECT2L) and DDRGK domain containing 1 (DDRGK1), were found to contain locus-specific modifications in at least one of the 20 embryos analyzed (Fig. 3). These modifications occur next to a potential PAM sequence and have not been shown in other pig breeds; therefore, the modifications were classified as true off-targeting events brought on by Cas9. Analysis of the APP knockout pigs did not detect any potential off-targeting events.
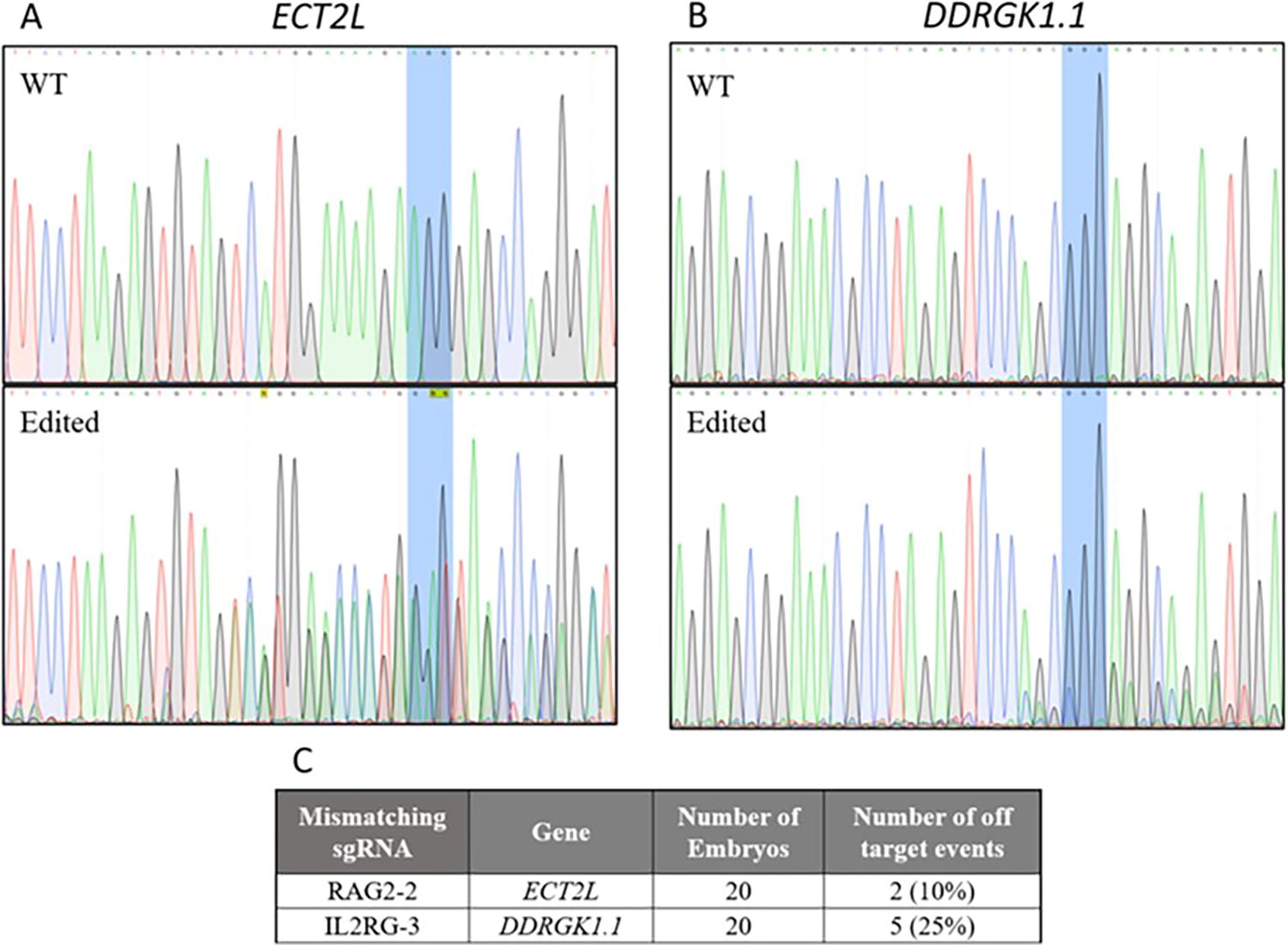
Sanger sequencing of individual blastocysts for novel off-target genes DDRGK1 and ECT2L in RAG2/IL2RG sgRNA injected embryos. Representative chromatograms for wild-type control embryos and injected embryos assayed for ECT2L
Genome integrity in various tissues derived from genome-edited pigs
Multiple tissues from APP knockout pigs were used for WGS to identify any tissue-specific genome variation or targeting efficiency. No differences in the frequency of off-targeting were detected in the tissues. In fact, no noticeable off-targeting event was identified in the tissues, indicating high specificity of CRISPR-Cas system used to target APP. However, the use of WGS was able to detect mosaicism that can occur by injecting the CRISPR-Cas system in developing embryos in one of our GE pig models. From the APP knockout pigs, we analyzed genomic DNA from lung, brain, muscle, skin, spleen, and WBCs. Only pig #2–3 possessed a combination of four different alleles for the on-target locus in the brain, muscle, and spleen (Table 4). The lung tissue from pig #2–3 did not contain any modifications, and only wild-type APP sequence was identified. All other pigs contained the expected on-target modifications and were not mosaic in any other tissue.
Identification of mosaicism in one APP genome edited pig
Discussion
With the rise in use of the CRISPR-Cas system, interest in developing methods for detecting off-target sites associated with Cas9 is being increased. The possibility of producing unintended modifications in unknown regions of the genome is a major concern of using genome editing technology, especially with clinical application of CRISPR-Cas system to cure genetically linked diseases. Many sgRNA design tools, such as CRISPOR, 13 predict the off-target possibilities of a given guide. However, the possibility of producing an unknown or unintended modification still exists. To further gain understanding of off-targeting events introduced by the CRISPR-Cas system, numerous in vitro methods have been developed to identify off-target sites in purified genomic DNA (SITE-Seq, Circle-SEQ, Digenome-Seq, among others).7–9 While these systems reflect cleavage of the genome by the designed CRISPR-Cas system, the system cannot calculate actual level of CRISPR-Cas system in target cell types or organisms. In fact, different cell types may have various genome editing efficiencies presumably due to different amounts of CRISPR-Cas to interact with the genome. 27
Detection of off-targeting system is often based on monitoring changes in genomic sequences after genome editing.28–31 For instance, genome variances compared to a reference genome or following genome editing events can reveal on- and off-targeting events introduced due to genome editing systems. However, detection of off-targeting events in animal models with an unknown genetic background is difficult to determine because of existing natural variations to the reference genome. Unlike conventional rodent models, most pig breeds used in research have heterogeneous genetic backgrounds. In addition, during genetic engineering to establish novel pig models, unknown maternal genetic background is likely to be introduced because the oocyte source for producing GE embryos is normally abattoir-derived ovaries. Because of the unknown genetics, detection of off-targeting in GE pigs is extremely difficult. The reference genome of the pig, that is, Sscrofa 11.1, is also derived from a single pig with a specific genetic background and, therefore, may not capture genetic variation or changes caused by CRISPR-Cas system accurately. 32 The heterogeneity in the genome may also create a challenge to generate a comprehensive list of potential off-targeting candidates using software such as Cas-OFFinder. However, because sgRNAs used in this study were designed to recognize exon sequences which are likely to be conserved amongst different breeds and we allowed for up to six base pair mismatches when generating the off-targeting list, we should have captured potential off-targeting candidates on a global level.
The high level of natural variation that exists among different pig breeds complicates the detection of true off-targeting events. Most of the off-targeting research published previously has been on mouse models derived from inbred mouse strains, 33 homogeneous cell populations, or using genome mapping of background cell cultures. 34 In this study, WGS was used to detect the level of off-targeting events on a global scale. The level of variation in all three gene-edited pig models compared to the pig reference genome was very similar, and even one of our paternal wild-type pig lines contained a high level of variation to the reference genome. This method of using WGS and Cas-OFFinder to find overlapping variants provided a means to rapidly identify and filter for variances that are likely to be off-targeting events. It is possible that the lack of parental genome information still hinders identification of all off-targeting events and labeling variations to be CRISPR-Cas origin. However, our ability to generate gene-edited embryos using the same sgRNAs and analyzing the resulting embryos validated effectiveness of the pipeline analysis and conclude some of variations to be introduced by CRISPR-Cas system. Our pipeline detected two novel off-targeting locations in our gene-edited pig models without having to install any complicated coding to dissect off-targeting events versus natural variation. Since technology to perform whole genome sequencing in blastocysts is available, 35 our pipeline can be used to identify the realistic level of off-targeting from designed CRISPR-Cas systems. This would allow the selection of the safest CRISPR-Cas system prior to performing embryo transfers, which is a significant investment in large animal species such as pigs.
The number of total variations in our gene-edited and wild-type control pigs indicate the heterogenous nature of the pig genome and underscore the safety of the CRISPR-Cas system, that is, no noticeable genome instability at the global level. The WGS was performed using Illumina NovaSeq based sequencing, and the genome was assembled from 150 bp reads. The sequencing reads were mapped to the pig reference sequence. While the genome assembly was successfully used to detect overall variances in the genome, using short reads has some limitations as gene inversions or certain regions on chromosomes may have not been captured from the approach. Combining our strategy with a longer read application such as PacBio or Nanopore sequencing36,37 may give a more accurate assessment of off-targeting events and impact of genome editing on genome integrity.
The level of off-targeting events identified in the gene-edited pigs in this study was low. Most pigs carried no detectable off-targeting events in the genome, and independent validation of those off-targeting events in newly generated gene-edited embryos revealed that off-targeting frequency was low; only two out of eight off-targeting locations were confirmed. The detection rate may be increased if more gene-edited pigs/embryos were screened; however, the low frequency emphasizes that these off-targeted sequences are not likely to be carried to next generation after breeding and phenotypes observed in founder gene-edited pigs and its progeny should be rooted from the on-target modification.
Mosaicism has been identified as a potential problem brought on from the direct injection of the CRISPR/Cas system into oocytes or zygotes. 38 In this study, we show that only one APP knockout pig derived from oocyte injections, out of three pigs, was mosaic. The other two pigs had on-target modifications likely introduced prior to the first embryo cleavage event and, therefore, did not possess more than two alleles. Furthermore, we did not detect any tissue specific bias in genome editing or off-targeting events in this pig model. However, it is difficult to draw more conclusions with the current sensitivity as no off-targeting events were detected in these APP knockout pigs because of the low sample number used in this study.
Conclusions
The method presented here offers a simplified approach to detect off-targeting events in an unknown genetic background. Using this method, we were able to identify on-target modifications, mosaic tissues in GE pigs, known off-targeting modifications, as well as novel off-targeting events. Pigs are considered a suitable preclinical animal model, 39 and the ability to monitor off-targeting events following application of genome editing tools will increase their value as a preclinical model when genome editing technology is already entering the clinic.40,41 Our finding will offer a guideline to detect off-targeting events in pigs for biomedical applications and imply the level of off-targeting events that could be encountered while applying genome editing technology in the clinics.
Footnotes
Acknowledgments
The authors would like to acknowledge the oocyte microinjection and fertilization team, Emily Eitel, Melissa Fudge, and Lee Spate. The authors also like to acknowledge Melissa Samuel for the daily pig care and collection of pig samples.
Data Availability
All data generated during this study are published within this article and its supplementary information file. All other data are available from the corresponding author upon reasonable request. Raw files are available in the NCBI Sequence Read Archive (SRA) repository—BioProject:
Authors’ Contributions
K.L., B.K.R., J.Y., and H.A. designed the study and collected data. H.A. analyzed all whole genome sequencing data and Cas-OFFinder sites for each guide. K.U. and E.R. generated gene-edited pigs. ER analyzed and filtered resulting WGS/Cas-OFFinder data. J.Y. and P.R.C. genotyped and analyzed the injected embryos and assisted with article. B.K.R., R.S.P., and K.L. provided all oversight for the study. All authors reviewed and approved the final article.
Author Disclosure Statement
The authors declare that they have no competing interests.
Ethics Approval and Consent to Participate
All animals were maintained according to the approved protocol and standard operating procedures by the Institutional Animal Care and Use Committee of the University of Missouri. All methods were carried out in accordance with an approved Institutional Biosafety Protocol from the University of Missouri.
Consent for Publication
Not applicable
Funding Information
This research was supported, in part, by USDA-ARS project number 5070–31320-001-00D, R21AG079292, and R01OD035561, and funding for the National Swine Resource and Research Center is from the National Institute of Allergy and Infectious Disease, the National Institute of Heart, Lung and Blood, and the Office of the Director (U42OD011140).
