Abstract
Allergy to domestic cat affects up to 15% of the population, and sensitization to cat allergen is associated with asthma. Despite the pervasiveness of cat allergic disease, current treatments have limited impact. Here, we present a bioinformatics analysis of the major cat allergen, Fel d 1, and demonstrate proof of principle for CRISPR gene editing of the allergen. Sequence and structural analyses of Fel d 1 from 50 domestic cats identified conserved coding regions in genes CH1 and CH2 suitable for CRISPR editing. Comparative analyses of Fel d 1 and orthologous sequences from eight exotic felid species determined relatively low-sequence identities for CH1 and CH2, and implied that the allergen may be nonessential for cats, given the apparent lack of evolutionary conservation. In vitro knockouts of domestic cat Fel d 1 using CRISPR-Cas9 yielded editing efficiencies of up to 55% and found no evidence of editing at predicted potential off-target sites. Taken together, our data indicate that Fel d 1 is both a rational and viable candidate for gene deletion, which may profoundly benefit cat allergy sufferers by removing the major allergen at the source.
Introduction
Domestic cat (Felis catus) is the most common source of mammalian allergen, with allergy to cat affecting up to 15% of adults and children and producing symptoms that range from rhinoconjunctivitis to severe asthma.1–3 Recent data from the U.S. National Health and Nutrition Examination Survey attributed more than 500,000 asthma attacks per year and ∼350,000 emergency care visits to patients who were sensitized and exposed to cat allergen. 4 Despite the prevalence and potential severity of cat allergic disease, current treatments merely address the allergic symptoms (e.g., antihistamines) or have demonstrated inconsistent results (e.g., cat allergen immunotherapy5,6).
The major cat allergen, Fel d 1, is a 35 kD tetrameric protein composed of two heterodimers, each of which consists of two chains: chain 1 (70 aa, 8 kD protein) and chain 2 (92 aa, 10 kD).7,8 The genes CH1 and CH2 (human gene nomenclature used here for cat), encoding chains 1 and 2, respectively, are located in a genomic span of <10,000 bp (chromosome E2, base pairs ∼18,487,507–18,496,763; NCBI assembly Felis_catus_8.0). The crystal structure of recombinant Fel d 1 shows that the protein binds Ca2+ ions and has two amphipathic internal cavities that may bind steroid ligands. 9 Fel d 1 is a secretoglobin 10 produced by cat salivary, 11 sebaceous, 12 perianal, 13 and lachrymal 11 glands. Natural levels of the allergen vary significantly between cats (>100-fold) and even within the same cat. 14
Fel d 1 causes IgE antibody-mediated sensitization in ∼95% of cat allergic patients and accounts for 60–90% of the anti-cat IgE.15–19 High-level exposure to Fel d 1 drives modified Th2 immune responses that elicit IgG4 antibody production in allergic and nonallergic individuals. 20 Though several other cat allergens have been identified (e.g., Fel d 4), they are of less allergenic and clinical significance.21,22
While the biologic function of Fel d 1 is unknown, studies suggest that the protein may play a role in chemical communication, epithelium defense, or immune regulation.7,23–26 To date, there have been no documented knockouts of Fel d 1 in cats. The Fel d 1 genes are not expressed in species beyond the family Felidae, though homologous sequences with ∼25–50% identity have been identified in rabbit (uteroglobin protein 7 ), human (Clara cell 10 kD secretory protein 7 ), mouse (androgen-binding protein 27 ), and the slow loris primate (brachial gland secretion protein 24 ).
Genome engineering presents a unique opportunity for the unequivocal deletion of Fel d 1 from cat cells and tissues and may provide the critical step in establishing the definitive function of the allergen. CRISPR in particular offers considerable improvements in throughput, target specificity, editing efficiency, and precision compared to prior technologies. 28 Additionally, CRISPR has demonstrated promise in therapeutic applications (e.g., sickle cell disease and β-thalassemia 29 ) and offers profound clinical potential. Given the advantages of CRISPR gene editing and a specific genomic target in Fel d 1, CRISPR is a practical approach for tackling cat allergic disease, which may greatly improve the lives of cat-allergic individuals.
Here, we report a structural bioinformatics analysis of Fel d 1 to optimize single-guide RNA (sgRNA) site selection for CRISPR editing of the allergen. This approach has enabled the evolutionary relationships between Fel d 1 and exotic cat orthologs to be established and provides novel insight into the biologic function of the allergen. The results indicate that Fel d 1 is amenable to gene editing using CRISPR and provide the first step in creating Fel d 1 knockout cats.
Methods
Comparative analyses of domestic cat Fel d 1 and exotic cat orthologs
Domestic cat DNA extraction and Fel d 1 sequence analysis
Discarded tissue samples (feline testes, ovaries, uteri) from the spay or neuter procedures of 50 domestic cats (98% domestic short- or long-haired cats of mixed ancestry) were generously provided for the study (Charlottesville Albemarle SPCA).
Genomic DNA was extracted from the samples (DNeasy® Blood & Tissue Kit; Qiagen), and CH1 and CH2 were polymerase chain reaction (PCR) amplified (Supplementary Table S1), purified (QIAquick® PCR Purification Kit; Qiagen), Sanger sequenced (Eurofins Genomics), and translated (ExPASy). Pairwise and multisequence alignments of the Fel d 1 DNA or protein sequences with the GenBank references (DNA: M74952.1 and M77341.1; protein: AAC37318.1 and AAC41616.1) were performed, and percent identity was determined (CLUSTAL Omega, Geneious Prime).
Sequence analysis of exotic cat Fel d 1 orthologs
The DNA reference sequences for full-length (including introns) Fel d 1 chain 1 (NCBI accession X62477) and chain 2 (NCBI accession X62478) were used to perform a basic local alignment search (BLAST) of 24 exotic (nondomestic) cat genomes available via the NCBI sequence read archive database (Supplementary Table S2). Eight big or wild felid species were analyzed (n = 3 cats per exotic species), including Chinese mountain cat (Felis silvestris bieti, a potential subspecies of Felis silvestris30–33 ), black-footed cat (Felis nigripes), leopard cat (Prionailurus bengalensis), fishing cat (Prionailurus viverrinus), cougar (Puma concolor), Iberian lynx (Lynx pardinus), Bengal tiger (Panthera tigris tigris), and African lion (Panthera leo).
The resulting whole-genome sequencing fragments were assembled to the Fel d 1 references to yield DNA consensus sequences, which were then translated (Geneious Prime). Pairwise and multisequence alignments of the exotic cat DNA or protein orthologs with the GenBank Fel d 1 references were performed, and percent identity was determined (CLUSTAL Omega, Geneious Prime). Unrooted, neighbor-joining phylogenetic trees of Fel d 1 chains 1 and 2 were built from multisequence alignments of the 50 domestic and 24 exotic cat sequences (Geneious Prime).
Selective pressure analysis of Fel d 1 orthologous sequences
Selective pressure was estimated by comparing the GenBank DNA references for Fel d 1 CH1 or CH2 with the exotic cat DNA orthologs (one representative cat per exotic species). The numbers of synonymous sites (S), synonymous substitutions (DS), nonsynonymous sites (N), and nonsynonymous substitutions (DN) were determined, and the synonymous and nonsynonymous substitution rates (dS and dN, respectively) were calculated. 34 The ratio of the nonsynonymous and synonymous substitution rates (ω = dN/dS) was determined for each exotic species (Supplementary Table S3). If no synonymous substitutions were present, the dN/dS ratio was estimated with the alternative: ω = dN/((DS +1)/S).
Gene editing of Fel d 1 using CRISPR-Cas9
CRISPR sgRNA design
Conserved regions of domestic cat CH1 and CH2 were identified by sequence analysis (Geneious Prime). CRISPR sgRNAs with sequences complementary to conserved coding regions of Fel d 1 and adjacent to CRISPR-Cas9 specific protospacer adjacent motif (PAM) sequences (“NGG”) and with few predicted potential off-target cleavage sites were designed (CRISPOR 35 ). Ten Fel d 1–specific sgRNAs were selected and synthesized (Invitrogen; Supplementary Table S4).
Feline cell culture
Immortalized feline kidney epithelial cells (Crandell–Rees Feline Kidney [CRFK]; ATCC CCL-94) were grown in Eagle's Minimum Essential Medium (EMEM) supplemented with 10% horse serum (Thermo Fisher Scientific) and 1% penicillin-streptomycin (ATCC). The cells were maintained at 37°C with 5% CO2 and passaged every 2–3 days using 0.25% Trypsin-EDTA (ATCC).
CRISPR transfection of feline cells
Feline CRFK cells were seeded in antibiotic-free, serum-rich EMEM at 250,000 cells/well on six-well plates ∼24 h prior to transfection. The CH1- or CH2-specific sgRNAs and Cas9 nuclease (TrueCut Cas9 Protein v2; Invitrogen) were delivered to the cells in a 1:1 molar ratio by lipid-based transfection (Lipofectamine CRISPRMAX; Invitrogen), and the transfected cells were incubated at 37°C with 5% CO2 for 48 h. The editing efficiencies of each of the 10 Fel d 1 sgRNAs were evaluated individually, with three biological replicates per sgRNA. A noncoding sgRNA was used as a negative control, while sgRNAs targeted to feline hypoxanthine phosphoribosyltransferase 1 (HPRT1) served as positive transfection and CRISPR controls (Invitrogen; Supplementary Table S5).
Enzymatic detection of CRISPR edited Fel d 1
Control and edited CRFK cells were lysed 48 h post transfection (Genomic Cleavage Detection Kit; GeneArt). Partial CH1 or CH2 sequences were PCR amplified around the predicted CRISPR cleavage sites (Supplementary Table S6). The resulting DNA fragments were denatured and randomly re-annealed to form heterogeneous DNA duplexes, which were incubated with T7 endonuclease 1 (T7E1, Genomic Cleavage Detection Kit, GeneArt; Alt-R Genome Editing Detection Kit, IDT) and run on a 2% agarose gel at 100 V for 1 h.
The gel was imaged (DigiDoc-It) and analyzed (ImageJ) to estimate editing efficiency from the gel band intensities given editing efficiency = 1 – [(1 – fraction cleaved)1/2], where fraction cleaved = sum of cleaved band intensities/(sum of cleaved and parent band intensities).
Sequence decomposition of CRISPR edited Fel d 1
Control and CRISPR edited CRFK cells were lysed 48 h post transfection (Genomic Cleavage Detection Kit; GeneArt). Partial CH1 or CH2 sequences were PCR amplified around the predicted CRISPR cleavage sites (Supplementary Table S6), and the resulting DNA fragments were purified (QIAquick® PCR Purification Kit; Qiagen) and Sanger sequenced (Eurofins Genomics).
The control and edited DNA chromatogram traces were uploaded to the Tracking of Indels by Decomposition (TIDE 36 ) and Inference of CRISPR Edits (ICE; Synthego) bioinformatics platforms. TIDE and ICE determined all possible edits for the specified control trace, and used regression analysis to estimate sgRNA editing efficiency based on the relative abundance of each indel (insertion/deletion) in the mixed DNA trace of the CRISPR edited sample. 37
CRISPR off-target identification and detection
Potential sites of off-target CRISPR editing were identified for sgRNAs C1G1 and C2G1 (CRISPOR 35 ). Four predicted potential off-target sites (C1G1: ENSFCAG00000030374-AQP4 [intergenic], OSMR-LIFR [intergenic]; C2G1: KIZ [intron], TEX264 [exon]) with three nucleotide mismatches from the C1G1 or C2G1 target sequences were evaluated. Control and edited CRFK DNA was PCR amplified around the predicted off-targets (Supplementary Table S7). Editing efficiency of C1G1 or C2G1 at the off-target sites was evaluated by sequence decomposition (TIDE, ICE) and T7E1 enzymatic detection.
Results
Sequence and structural analysis of domestic cat Fel d 1
CH1 and CH2 were sequenced from 50 domestic cats to identify protein variants or amino acid substitutions, as well as conserved DNA target regions for CRISPR sgRNA design. From the domestic cat protein sequences, 37 unique amino acid substitutions were identified, including 18 novel, natural Fel d 1 variants resulting from seven single nucleotide polymorphisms (SNPs), four multiple nucleotide polymorphisms (MNPs), or seven rare mutations (polymorphisms observed in a single cat; Fig. 1A and Supplementary Table S8).
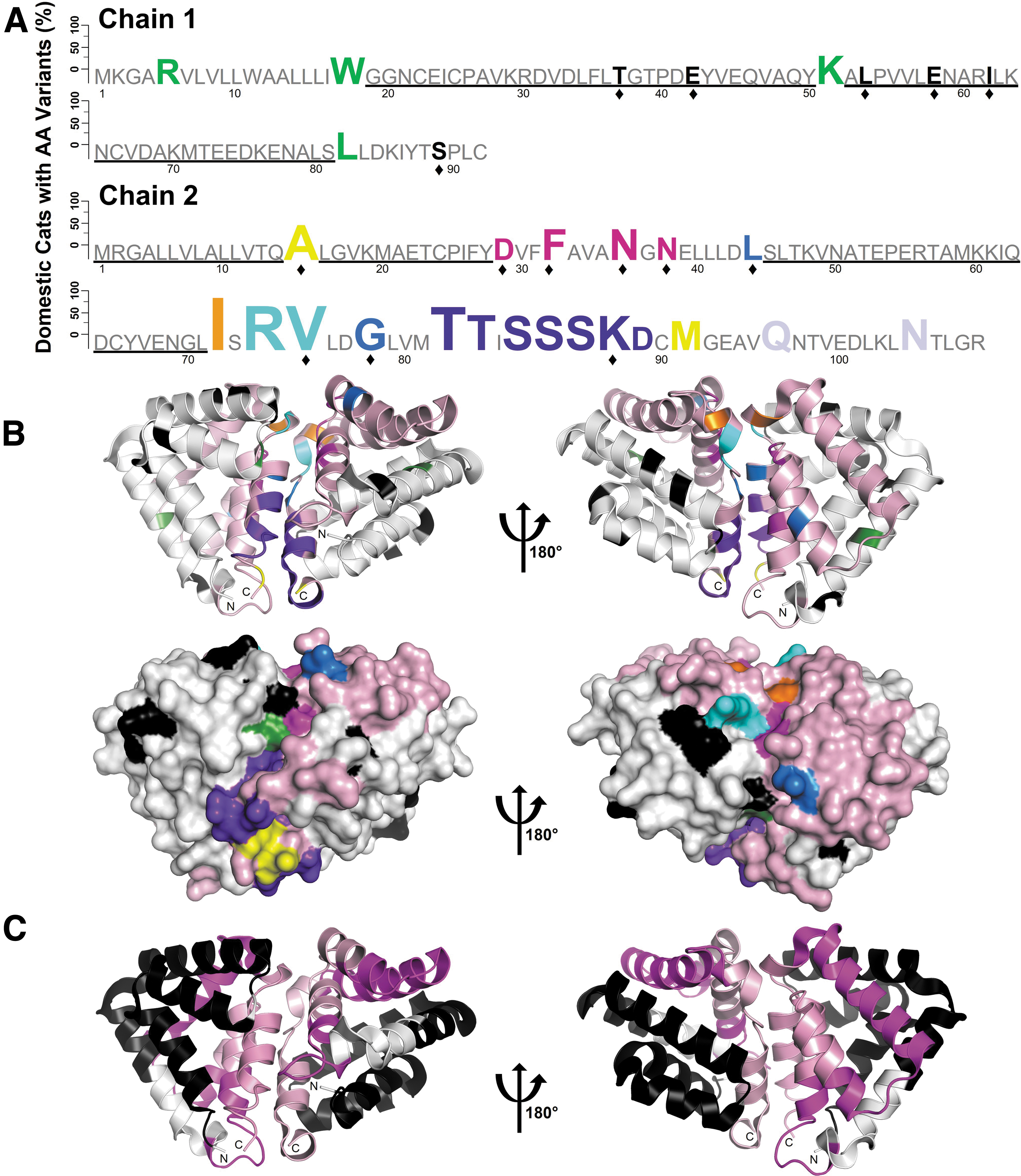
Comprehensive sequence and structural analyses of domestic cat Fel d 1. Fel d 1 chains 1 and 2 were sequenced from 50 domestic cats.
The domestic cat variants were mapped onto the tetrameric crystal structure of Fel d 1 (PDB 2EJN 9 ) and were found to be primarily concentrated at the interface of the two dimers (Fig. 1B). Several patterns of correlated Fel d 1 variants were revealed, and notably, 42% of the domestic cats encoded a noncanonical isoform, CH2S, 8 or a similar novel variant at residues Thr82-Asp89 (Fig. 1A and B and Supplementary Table S8). The domestic cat Fel d 1 variants are highlighted in an accompanying interactive visualization (www.feld1viz.com).
Multiple conserved coding regions in the Fel d 1 genes with suitable target sequences for CRISPR editing were revealed. Fel d 1 sequence fragments with more than 20 well-conserved (>98% identity), consecutive nucleotides adjacent to a CRISPR-Cas9 PAM sequence (“NGG”) were deemed appropriate targets for CRISPR editing. Given these criteria, we identified two conserved CRISPR target regions in CH1 (Gly19-Tyr50, Ala52-Ser81) and three regions in CH2 (Met1-Gln14, Leu16-Tyr28, Ser45-Leu71; Fig. 1A and C). These conserved sequences guided the design of 10 Fel d 1–specific CRISPR sgRNAs (Supplementary Table S4).
Multisequence alignments of the domestic cat coding DNA sequences produced identities of 99.5% and 96.4% for CH1 and CH2, respectively (Supplementary Figs. S1 and S2). Pairwise alignments of the DNA sequences with the GenBank Fel d 1 references (M74952.1 and M77341.1) yielded identities ranging from 96.2% to 100% for CH1 and 93% to 100% for CH2. Multisequence alignments of the corresponding protein sequences resulted in identities of 97.9% and 90.1% for chains 1 and 2, respectively (Supplementary Figs. S3 and S4), while pairwise alignments of the protein sequences with the GenBank references (AAC37318.1 and AAC41616.1) yielded identities of 90.3–100% for chain 1 and 84.6–100% for chain 2.
Evolutionary significance of exotic cat Fel d 1 orthologs
Twenty-four exotic (nondomestic) cat genomes representing eight big or wild felid species were examined by BLAST search to identify Fel d 1 orthologous sequences (i.e., homologs separated by speciation; Supplementary Table S2).
The exotic cat species included Chinese mountain cat (Felis silvestris bieti; a potential subspecies of the wildcat, Felis silvestris30–33 ), black-footed cat (Felis nigripes), leopard cat (Prionailurus bengalensis), fishing cat (Prionailurus viverrinus), cougar (Puma concolor), Iberian lynx (Lynx pardinus), Bengal tiger (Panthera tigris tigris), and African lion (Panthera leo). The subset of exotic cats spans several lineages of the family Felidae, ranging from members of the same genus as domestic cat, Felis, to Panthera species that diverged from domestic cats ∼11 million years ago (Fig. 2A). 38
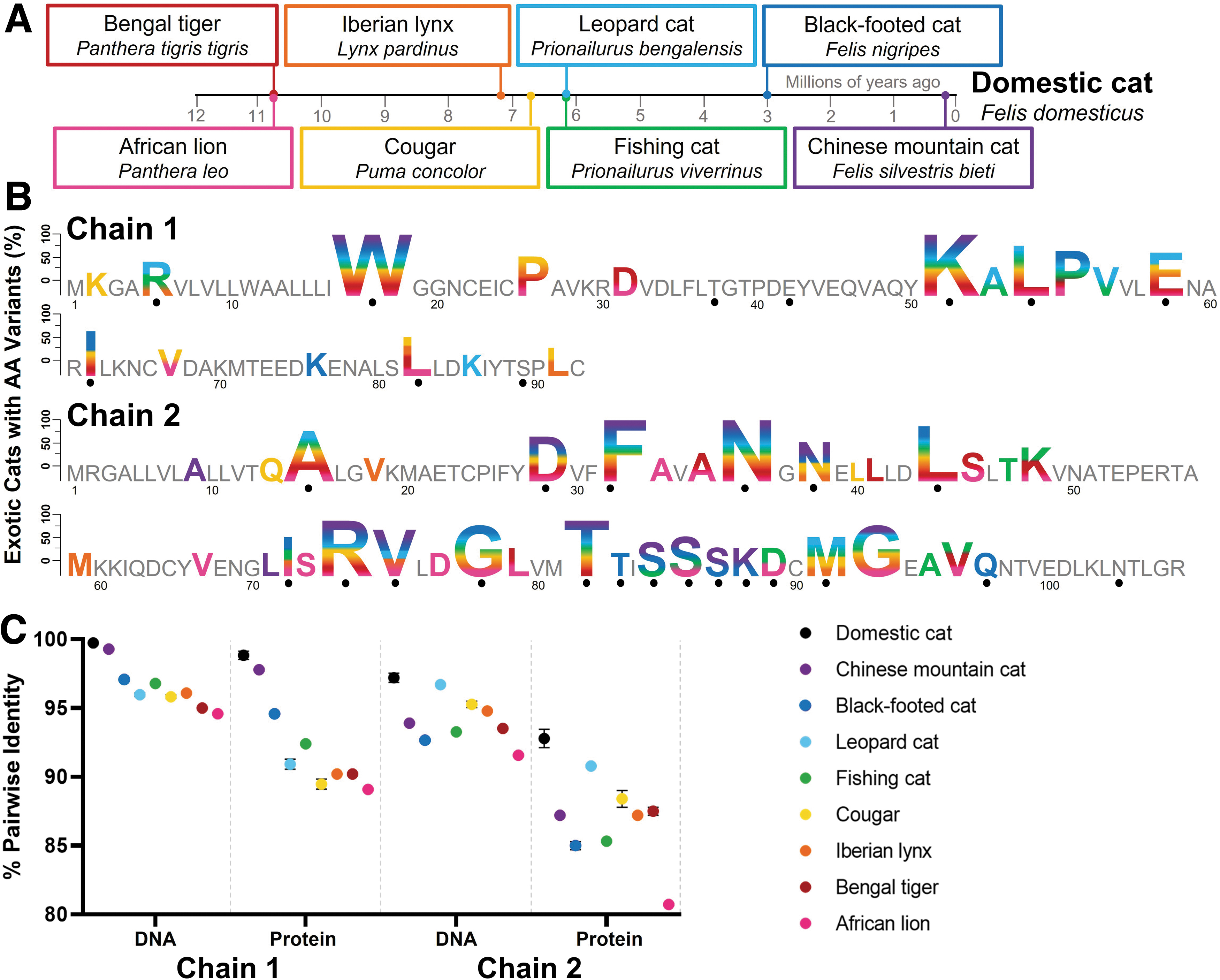
Comparative sequence analysis of exotic cat Fel d 1 orthologs. Fel d 1 orthologous sequences from 24 exotic cats, representing eight felid species, were analyzed.
Fifty-five unique amino acid substitutions, including 29 substitutions specific to the exotic cat species, were identified from the Fel d 1 protein orthologs (Fig. 2B and Supplementary Table S9). The exotic cat variants were mapped onto the Fel d 1 crystal structure, 9 which revealed an accumulation of variants at the dimer interface (Fig. 3A). The exotic cat orthologs and corresponding variants are featured in an accompanying interactive visualization (www.feld1viz.com).
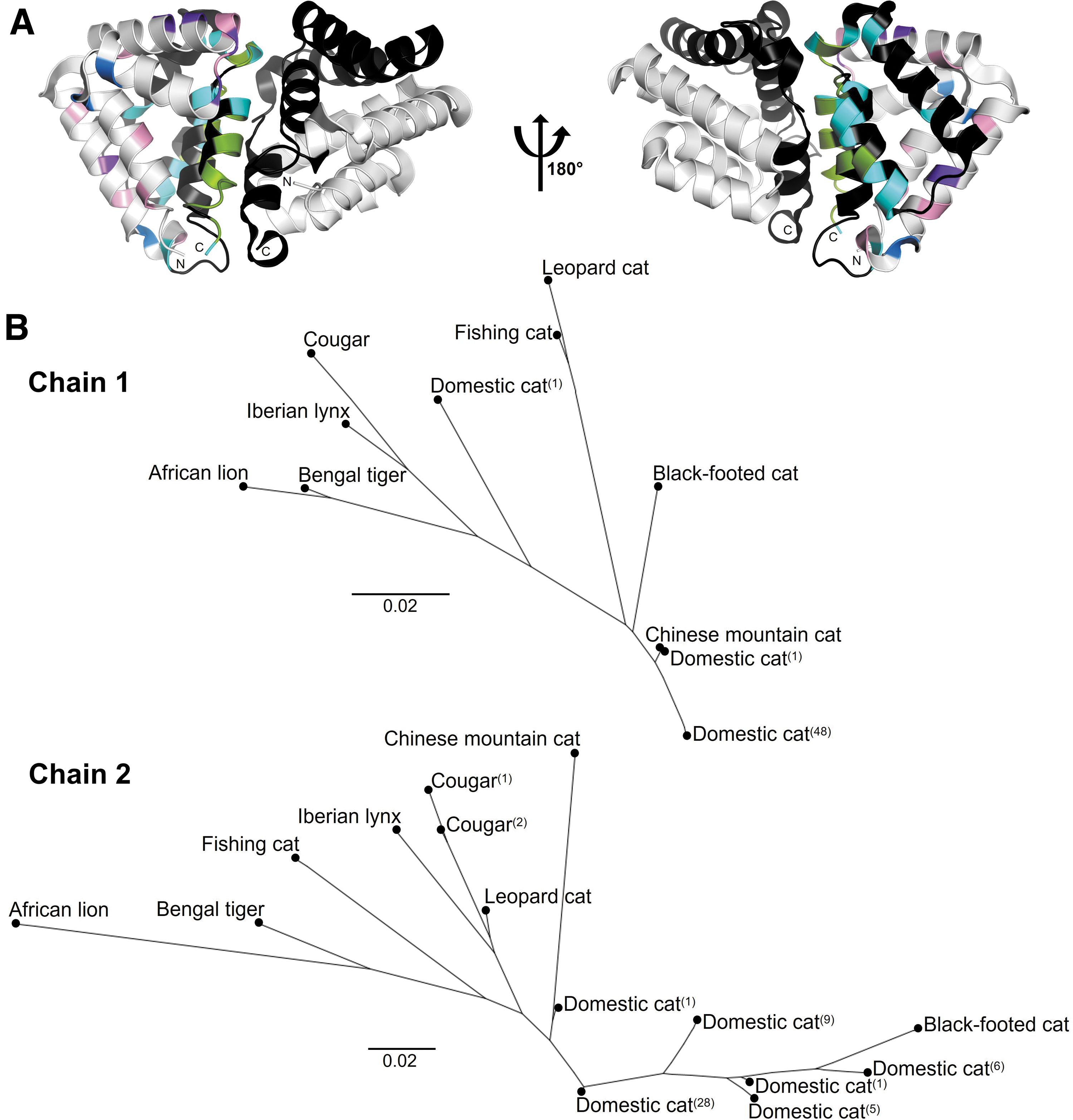
Evolutionary relationships between domestic and exotic cat Fel d 1 orthologs.
Multisequence alignments of the coding DNA orthologs from 24 exotic cats produced identities of 95.9% and 93.5% for CH1 and CH2, respectively (Supplementary Figs. S5 and S6). Pairwise alignments of the DNA orthologs with the GenBank Fel d 1 references yielded identities ranging from 94.6% to 99.3% for CH1 and from 91.5% to 96.7% for CH2 (Fig. 2C). Multisequence alignments of the corresponding protein orthologs yielded identities of 92.3% for chain 1 and 86.2% for chain 2 (Supplementary Figs. S7 and S8), while pairwise alignments of the orthologs with the GenBank references produced identities of 89.1–97.8% and 80.7–90.8% for chains 1 and 2, respectively (Fig. 2C).
A cumulative, multisequence alignment of the DNA orthologs for all domestic and exotic cats analyzed yielded identities of 97.8% for CH1 and 94.8% for CH2. Similarly, a cumulative, multisequence alignment of the protein orthologs produced identities of 94.6% and 87.3% for Fel d 1 chains 1 and 2, respectively. Unrooted, neighbor-joining phylogenies were built from the alignments of the Fel d 1 protein orthologs to evaluate the evolutionary relationships between the domestic and exotic cat species (Fig. 3B). While chain 1 reflected the established evolution of the family Felidae, chain 2 deviated from the standard due to greater variability between the orthologs (Figs. 2A and C and 3B).
Editing Fel d 1 with CRISPR-Cas9
The editing efficiencies of each of the 10 CRISPR sgRNAs targeted to either CH1 (C1G1-6) or CH2 (C2G1-4) were individually evaluated (Supplementary Table S4). The sgRNAs and Cas9 nuclease were delivered to immortalized feline epithelial cells (CRFK, ATCC CCL-94) as ribonucleoprotein (RNP) complexes using lipid-based transfection. Fel d 1 gene knockout following CRISPR induced frameshift mutations was assessed by DNA sequence decomposition (TIDE 36 and ICE) or by enzymatic detection of nucleotide mismatches (T7E1).
Sequence decomposition found CRISPR editing efficiencies of 3.0–54.9% for the 10 Fel d 1–specific sgRNAs (Fig. 4A). Positive control sgRNAs targeted to feline HPRT1 yielded editing efficiencies of 52.2–77.6% (Supplementary Table S5). The two most efficient Fel d 1 sgRNAs, C1G1 (mean editing efficiency 52.1 ± 8.3% standard error of the mean [SEM]) and C2G1 (mean editing efficiency 54.9 ± 10.1% SEM) targeted to CH1 and CH2, respectively, were identified.
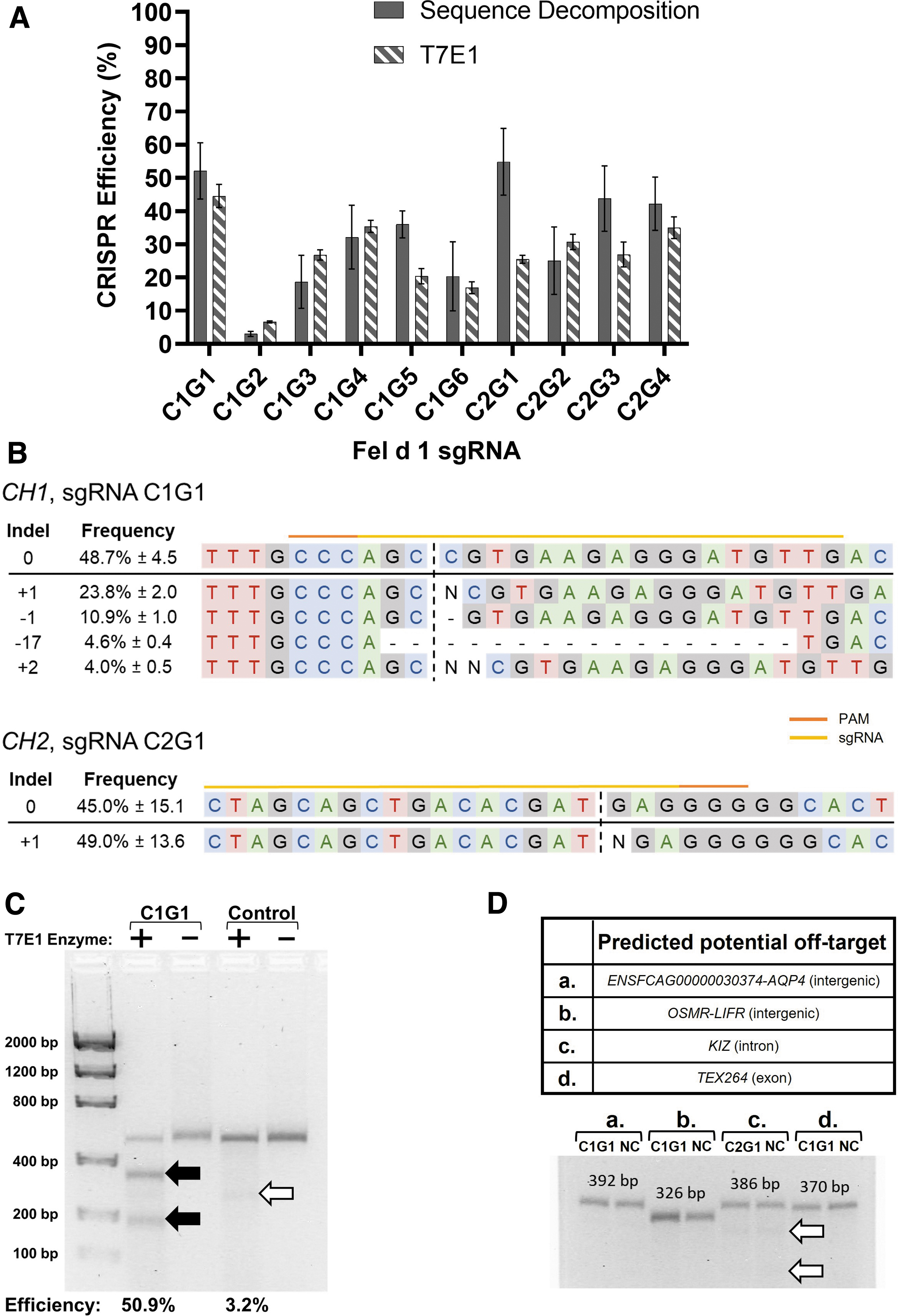
In vitro genome editing of Fel d 1 with CRISPR-Cas9.
The distribution of indels, which range from +2 nucleotide insertions to −17 nucleotide deletions, following editing by CRISPR-Cas9 were evaluated for these two sgRNAs (Fig. 4B). For C1G1, +1 nucleotide insertions or −1 nucleotide deletions accounted for 46% and 21% of all nonzero indels, respectively. By contrast, +1 nucleotide insertions accounted for 89% of the nonzero indels for C2G1. Excluding +1, +0, and −1 nucleotide indels, all other indels were observed at <5% frequency for either sgRNA (Fig. 4B). Indels predicted to produce frameshift mutations in the Fel d 1 protein sequences accounted for 99% and 96% of all nonzero indels for C1G1 and C2G1, respectively.
T7E1 analysis found CRISPR editing efficiencies of 6.6–44.9% for the 10 Fel d 1–specific sgRNAs, with mean editing efficiencies of 44.9 ± 2.5% SEM and 25.5 ± 1.2% SEM for sgRNAs C1G1 and C2G1, respectively (Fig. 4A and C). The T7E1 assay underestimated the efficiency of C2G1 due to its relatively low variability of indel distribution (Figs. 4A and B). Comparison of the sequence decomposition and T7E1 analyses allowed for the identification and subsequent exclusion of several natural Fel d 1 SNPs from the editing efficiency calculations (Fig. 4C).
Finally, DNA sequence decomposition and T7E1 analysis were used to evaluate off-target CRISPR editing at several predicted potential off-target cleavage sites (CRISPOR 35 ), which found no evidence of CRISPR editing at the predicted off-target sites due to Fel d 1–specific sgRNAs C1G1 or C2G1 (Fig. 4D).
Discussion and Conclusions
Though the precise biologic function of Fel d 1 remains unknown, our data indicate that the variants or amino acid substitutions are concentrated at the protein dimer interface, suggesting that this region of Fel d 1 is not well conserved and potentially nonessential for the function of the allergen. By contrast, the conserved regions identified as potential target sites for CRISPR-Cas9 editing may prove to be more functionally relevant for Fel d 1.
The variants identified in our study have not been linked to defined levels of Fel d 1 protein expression. However, several of the variable chain 2 residues (Asp29, Phe32, Gly78, Thr82, and Ser85; Supplementary Table S8) were recently recognized as potential binding sites of steroid ligands. 39 Our results imply that the ligand-binding function of these residues may be nonessential or that the residues may bind alternative ligands. These interpretations rely on the published structure of recombinant Fel d 1, 9 as the definitive structure of the natural allergen has yet to be determined.
On average, the chain 2 orthologs were found to be more variable than chain 1 for most of the cat species analyzed, in part due to variability in residues Thr82-Asp89 of chain 2. The CH2S 8 isoform was preferentially encoded by the subset of exotic species closely related to domestic cat, while the more divergent exotic cats favored the canonical CH2 isoform.
Interestingly, the 50 domestic cat sequences encoded either of the chain 2 isoforms or related variants. A study of RNA derived from domestic cat salivary gland or skin cells proposed that CH2S is preferentially expressed in the skin, while the canonical CH2 gene is largely expressed in feline salivary glands. 8 Tissue-specific expression may partly account for the variability observed in the domestic and exotic cat orthologs, which were sequenced from genomic DNA isolated from blood and tissues that lack Fel d 1 protein expression.
For both chains 1 and 2, the multi- and pairwise sequence alignments of the Fel d 1 orthologs revealed that the DNA sequences exhibit higher identities, or are more similar, than the corresponding protein sequences. More specifically, the data indicate that >70% of the Fel d 1 amino acid variants are the result of nucleotide substitutions in the first or second codon positions, rather than the third position that allows potential redundancy in the subsequent protein sequence.
The ratios of the nonsynonymous and synonymous substitution rates (ω = dN/dS) ranged from 0.5 to 3 for CH1 and from 0.7 to 4.7 for CH2 across the eight exotic species analyzed relative to domestic cat (Supplementary Table S3). This suggests that the Fel d 1 genes have been subject to selective pressure in some species, leading to a greater accumulation of nonsynonymous variants.34,40–42
The multisequence alignments suggest that the Fel d 1 gene sequences are not conserved, with the relatively low sequence identities (95.9% and 93.5% for CH1 and CH2, respectively, in 24 exotic cats) roughly mirroring the noncoding, genome-wide averages between species. 43 This lack of evolutionary conservation implies that the Fel d 1 genes may be nonessential for cats and suggests that CH1 and CH2 are suitable targets for gene deletion.
A protein BLAST search of Fel d 1 chains 1 and 2 in NCBI returns more than 400 significant aligned sequences, ranging in identities from 25% to 100% and representing a broad diversity of mammalian species from the Eutheria, Metatheria, and Prototheria subclasses. One homologous protein specific to the Mus genus, androgen-binding protein (ABP), exhibits 35–60% identity with the Fel d 1 sequences. The structural and binding affinity characteristics of Fel d 1 are similar to those of ABP, which is involved in mate selection and chemical communication among mice.23,27,39
Notably, recent studies observed that deleting mouse ABP produced no measurable adverse effects in ABP-deficient mice, indicating that ABP function is likely not essential. 44 While these studies of a homologous protein suggest that Fel d 1 may function in a similar capacity for cats and may ultimately prove to be nonessential, the definitive test of functionality must be experimental.
Our CRISPR knockouts in immortalized feline cells demonstrate that Fel d 1 is a viable target for gene deletion, which is expected to improve the health of cat allergic patients substantially by effectively removing the allergen at the source. Several recent approaches to cat allergy aim to neutralize Fel d 1 and to reduce cat allergen exposure.
Cat food supplemented with anti–Fel d 1 polyclonal egg IgY antibody reduced the allergen levels in cat saliva, 45 resulting in a 47% reduction in Fel d 1 deposited on cats' haircoats from saliva. 46 Cats immunized with an anti–Fel d 1 vaccine (HypoCat™) exhibited a 50% reduction in Fel d 1 levels detected in cat tear extracts, 47 which resulted in a ∼30% decrease in symptom severity in nine allergic patients. 48 Anti–Fel d 1 monoclonal IgG antibodies administered to cat allergic patients competitively blocked IgE binding to the allergen and reduced symptom severity by ∼60% in 50% of patients. 49
Modest reductions in allergen levels may have negligible clinical impact, given that nearly all cat allergic patients experience symptoms at a proposed threshold of 8 μg of Fel d 1 per gram of house dust, 50 while the allergen may accumulate at levels >1,000 μg/g in house dust. 51
In the future, the CRISPR knockouts will be replicated in Fel d 1–expressing primary feline cells and, eventually, in vivo in cats. Future studies will aim to develop a means for deleting the Fel d 1 genes in adult cats and effectively rendering the cats hypoallergenic, while further analyses of homologous proteins from other mammalian species may provide additional insight regarding the precise biologic function of Fel d 1.
Though the current study evaluated the editing efficiencies of individual sgRNAs targeted to either CH1 or CH2, future studies will investigate the efficiency of simultaneous knockouts of the Fel d 1 genes using multiple sgRNAs. Finally, before transitioning to in vivo studies, the off-target analysis will be expanded to detect and identify genome-wide double-strand breaks using unbiased screening approaches, such as GUIDE-seq or CIRCLE-seq.52,53
Though CRISPR editing in cats is ostensibly novel, genetic engineering of cats (e.g., selective breeding to create interspecies hybrids 54 ) is not unprecedented. Our comparative genomics analysis suggests that Fel d 1 may not be conserved or essential for cats. However, unambiguously deleting Fel d 1 with gene editing technology remains the critical step for identifying the biologic function of the allergen and for determining whether that function is vital for the cats' survival.
To date, this is the most comprehensive bioinformatics analysis of any allergen and the first study to investigate the evolutionary origins of an allergen. Our data illustrate a practical application of CRISPR technology in allergy research or veterinary medicine, and underscore the unique intersection between evolutionary biology, genome engineering, and therapeutic development.
Footnotes
Acknowledgments
The authors thank Nishant Jha, Broad Institute of Massachusetts Institute of Technology and Harvard University, for the design and development of the interactive visualization.
Author Disclosure Statement
N.F.B. and A.P. are employees of InBio. M.D.C. is co-owner and shareholder of InBio. M.A. is a consultant for InBio. B.W.D. declares no competing interests.
Funding Information
This study received funding from InBio.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
