Abstract
Currently, most advances in site-specific epigenetic editing for human use are concentrated in basic research, yet, there is considerable interest to translate this technology beyond the bench. This review highlights recent developments with epigenetic editing technology in comparison with the canonical CRISPR-Cas genome editing, as well as the epistemic and ethical considerations with preemptive translation of epigenetic editing into clinical or commercial use in humans. Key considerations in safety, equity, and access to epigenetic editing are highlighted, with a spotlight on the ethical, legal, and social issues of this technology in the context of global health equity.
Background: A Comparison of Epigenetic Editing to Genetic Editing
The epigenome is defined by the chemical modifications of DNA and the nuclear positioning of higher-order DNA structures, acting in concert with regulatory elements such as noncoding RNA to register, signal, or perpetuate activity states of DNA. 1 Epigenetic editing thus refers to the modification of the epigenome using engineered constructs, with the goal of modulating the chemical states that govern DNA structure or function.2,3 The CRISPR-Cas epigenetic editing technology borrows much of its targeting methodology from its genomic editing predecessor: the type II CRISPR-Cas9 system.2,3 Both systems use an engineered Cas construct with single-guide RNA (sgRNA) to target specific regions of the genome and induce changes in a site-specific manner as opposed to whole-genome modifications of the epigenome. 2
In site-specific epigenetic editing, the catalytically inactive variant Cas is used instead, such that it binds but does not cut DNA. 4 This allows the engineered construct to confer changes to the epigenome itself or to recruit other epigenetic regulatory factors (effectors) in order to generate epigenetic changes at the specific location. 2 In this sense, the Cas construct acts as a targeting mechanism to alter chromatin structure or gene transcription/regulation, with the potential of impacting downstream regulatory effects in an amplified and global manner, such as activating genes via modified epigenetic activity on promoters and modified epigenetic activity on promoters and enhancers (see Fig. 1).5–7
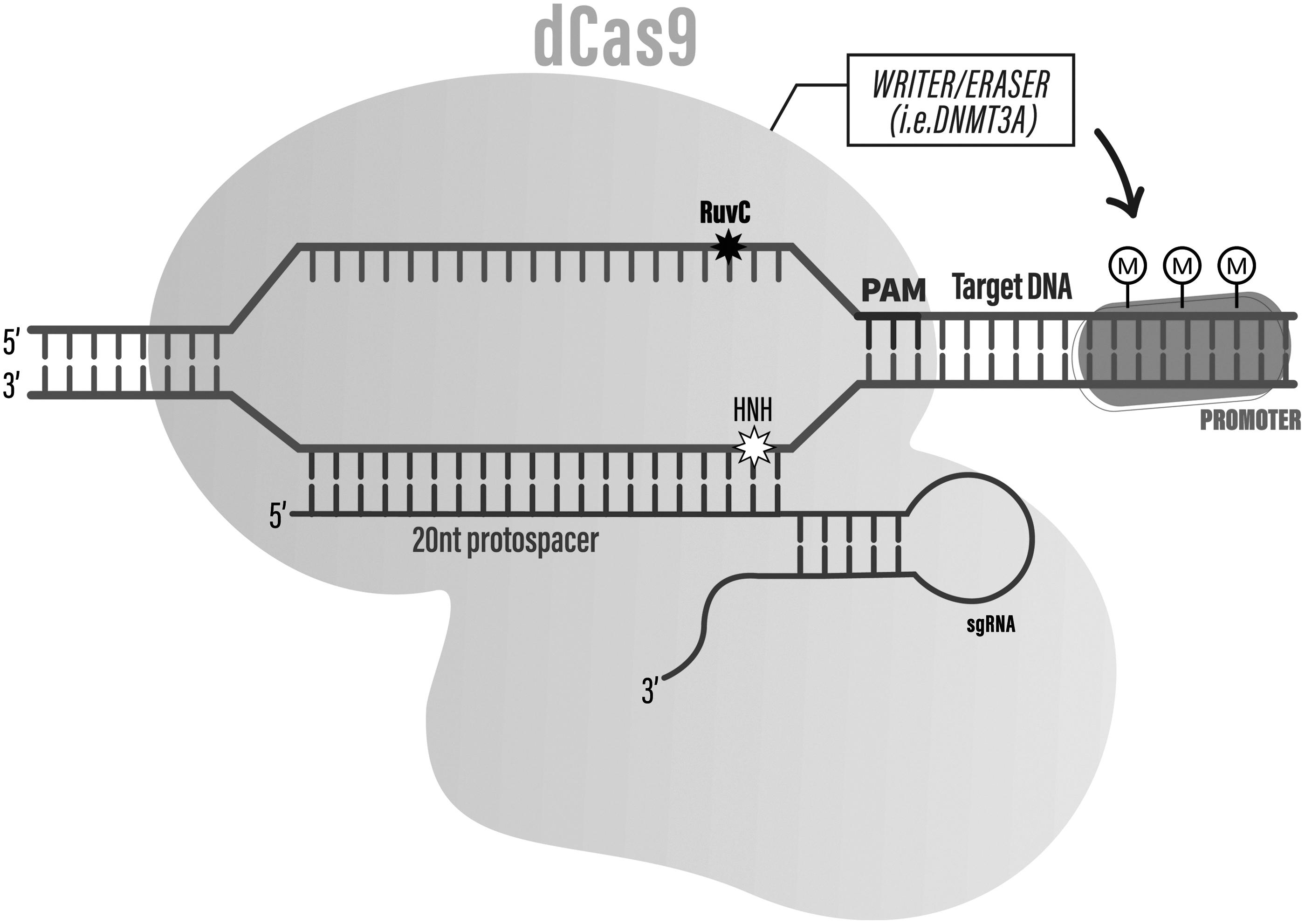
A typical dCas9 enzyme (from the commonly used Streptococcus pyrogenes) situated on the target DNA for epigenetic editing. The stars indicate the site of two mutations (D10A and H840A) on the RuvC and HNH nuclease domains, which renders the Cas enzyme unable to cleave the DNA target. Further pictured is the sgRNA with a 20 nt protospacer region matching to the PAM sequence. Attached to the Cas enzyme are epigenetic writer or eraser domains (effectors), such as DNMT3A, which can add de novo methylation marks on CpG islands of the host DNA. dCas9, nuclease-dead CRISPR-associated protein; RuvC and HNH, sites of Cas9 nuclease mutations; sgRNA, single-guide RNA; PAM, protospacer adjacent motif; DNMT3A, DNA-methyltransferase 3A; M, methylation marks.
Like genome editing, these epigenetic modifications can be performed on somatic or germline cells. However, unlike genome editing which is permanent, epigenetic editing can have transient or persistent effects, accomplished through various engineered constructs such as the FIRE-Cas9 system or the CRISPRon/CRISPRoff system.8,9 Although site-specific epigenetic editing is also achievable through other means such as zinc-finger nucleases (ZFNs) or transcription activator-like effector nucleases (TALENs), and in some specific biological contexts may outperform CRISPR, these techniques possess their own limitations to versatility that predate the increasing popularity of the CRISPR-Cas system, especially when considering their application in therapeutics.10–13
This review will provide an overview of the ethical and epistemic challenges in translating epigenetic editing from the bench to bedside (an overview of the search strategy is provided in the Appendix). The focus will be on Cas-mediated epigenetic technologies, as most tools developed for genetic and epigenetic editing use CRISPR-Cas due to its ease of use and programmability. As these technologies are undergoing the most rapid and widespread changes in development and commercialization, it acts as a hallmark example to discuss the potential issues related to epigenetic editing. It is noted that the considerations raised can be relevant to other forms of epigenetic editing in humans.
Key Knowledge Gaps in Epigenetic Editing Translation between Bench to Bedside
Presently, nonsite-specific genome-wide epigenetic “treatment” has been firmly established in clinics in the form of DNA (e.g., azacytidine) or histone (HDAC inhibitors) modifiers in oncology. It stands to reason that genomics will look toward site-specific epigenetic editing as the next form of precision medicine. Epigenetic editing is currently restricted in use within basic scientific research, where the goal is to alter epigenetic states in model organisms and human cell lines or to improve the efficiency of epigenetic editing technology through the development of new hybrid systems.9,14
Common applications of epigenetic technologies in basic research also include characterizing new regulatory elements in the epigenome or genome-wide forward screens.2,9 However, compared to traditional gene editing, less is known about the impact of epigenetic editing in the context of specific human diseases. 15 In other words, the methodology of Cas as a genomic targeting system can be translated to epigenetic editing, but knowledge regarding the effect of epigenetic editing in the human context is still limited.
For example, although the site-specific epigenetic editing CRISPRoff system has been used to perform genome-wide methylation successfully in human cell lines, its effects on a wide range of tissues have not been studied and nor have its side effects on cellular response and DNA damage been elucidated.9,16 In other words, an epistemic gap exists due to the lack of knowledge surrounding the ability to discern the influence of epigenetic networks. This gap could be attributed to a bottleneck of current technological limitations in delivery and detection of sensitive epigenetic marks or because researchers simply have not established enough quantifiable epigenetic knowledge. 17
Consequently, epigenetic technologies today are restricted to screening and diagnosis purposes in the clinical and commercial sectors, whereas editing of the human epigenome for therapeutic purposes is still under development.
Furthermore, epigenetic editing does not change the underlying DNA, and thus it is perceived to be more easily reversible, less permanent, less heritable, and therefore safer than genetic editing by many scientists and members of the general public.18,19 This perception could create a false sense of security, leading scientists and regulatory stakeholders to be less proactive in considering the potential ethical or safety pitfalls unique to epigenetic editing during the adaptation of this technology for human use.20,21 As a result, it is of utmost importance to highlight the epistemic (or knowledge-making) limitations of the editing process and to call attention to the ethical implications that could arise before clinical or commercial applications of epigenetic editing are pursued.
Epistemic Considerations: Concerns with Knowledge Translation and Feasibility of Epigenetic Editing Techniques
A common goal of epigenetic editing is to improve the human condition. In an ideal world, scientists could use epigenetic editing to change cancer cell gene expression, reset the molecular markers of aging, or cure rare genetic diseases. 2
Research on these topics are already associated with traditional CRISPR-Cas genome editing. 22 However, as epigenetic editing is still in its early developmental stages, several key aspects differ it from genome editing such that reasonable doubt exists regarding the adaptation of epigenetic editing for therapeutic use. 23 The underlying reason behind this doubt can be attributed to the way epigenetic editing is conceptualized. Since discussions of epigenetic editing often rely on comparisons to genetic editing, many translational stakeholders, regulators, and policy makers can find it difficult to articulate the conceptual nuances between the two technologies. 24
The best way to showcase these epistemic conceptual differences is to delineate the physical differences between epigenetic editing and genetic editing. In ideal site-specific epigenetic editing, the DNA sequence is not changed. Rather, any component of the epigenome could be changed. It may result in an alteration of the epigenomic element itself or create a downstream effect based on the alteration, such as a propagating chemical modification in chromatin or other epigenetic elements down the regulatory cascade. 6 This makes epigenetic editing harder to discern, as the success of an epigenetic edit at the molecular level can only be confirmed by measuring the on-target effects of the specific edit by screening for an epigenetic marker such as methyl-seq, ChIP-seq, or ATAC-seq. 25
However, off-target epigenetic regulation or global cellular responses can also affect the edited sequence, which can produce quantifiable changes at the on-target epigenetic site that is different from the intended edit. 26 Conceptually speaking, it thus becomes harder to translate epigenetic knowledge between native, unperturbed states versus new, edited states, such as whether the change is due to on-target epigenomic editing or secondary effects of native epigenetic regulation. 26
This knowledge gap can also be partly attributed to the differences in technological advancement: next-generation genome sequencing techniques are highly refined, whereas comparable sequencing technology for validating epigenetic editing is still in development. 27 As the process of developing precise validation tools is still ongoing, it is important to remember that preemptive efforts to use epigenetic editing knowledge will be hindered by this technical limitation.
Epigenetic editing can create further knowledge gaps regarding the biological response in epigenetic states as the system adjusts to the initial edit. 28 Epigenetic states are tissue and cell-specific: what particular epigenetic profile might be useful for one cell or tissue can be harmful to another, and perhaps more difficult to discern in an edited context. 29
By contrast, a DNA sequence in itself remains unchanged, even if the sequence is interpreted differently by the cell or tissue. 30 There is also much less ambiguity in validating a successful genome edit - one can simply confirm success by sequencing the DNA region of interest. 31 By comparison, a “successful” or helpful epigenomic edit can be harder to determine, depending on the type and scale of epigenetic change that occurred. As epigenetic states in real, in vivo systems can exert many levels of control over a single genomic region, consequently, establishing valid epigenetic editing knowledge adds another layer of complexity.
This reflects the distinct, physical differences between the epigenome and genome. Epigenetic markers are dynamic and often changing in response to the environment of the cell. 32 However, existing epigenome profiling techniques provide only a “snapshot” of what is happening in the epigenome at a given time. Whatever epigenetic state is validated at a given time and in a certain chemical environment may not be corroborated again later on, as environmental changes can affect the edited state. 8 It poses a challenge when comparing editing knowledge between in vivo and in vitro conditions. 33
Much of our in vivo knowledge in humans is translated from research in vitro on artificial systems. Yet, there is a greater knowledge gap between in vivo and in vitro states in epigenetic editing. For example, if sequencing an epigenetic edit revealed a silencing marker on gene X, is it because the in vivo state has truly been edited, or because the in vitro reagents, solutions, and chemicals used throughout the editing and sequencing process somehow produced a silenced chromosome state on gene X? This distinction should be carefully considered as clinical in vivo epigenetic editing is developed from in vitro or in silico knowledge.
The complexities are further compounded by how an edited epigenomic element behaves over a long time. An edited DNA sequence is static and unchanging—it generally stays the same throughout all cellular processes, whether the specific gene is expressed or not, or whether the DNA sequence sits in an in vivo or in vitro context. Moreover, an edited DNA sequence will reflect this edit in future cell divisions and processes. 34 By contrast, an edited epigenetic element in the body could change again in response to another epigenetic edit, or from future cell divisions in response to changing environmental conditions of the cell, tissue, or organism. 8
Therefore, there is more uncertainty to account for in epigenetic editing persistence when adapting epigenetic editing to a large spatial (tissue, organ or organism) or temporal (multiple cell divisions, organismal life span) scale. In general, these considerations make validating a successful epigenetic edit more complicated than genetic editing and should prompt stakeholders to reevaluate the typical translational procedures for epigenetic editing. As the biological peculiarities of epigenetic editing differ greatly in practice from gene editing, a holistic approach to epigenetic editing cannot be accomplished by transcribing the rules of governance for gene editing. 6
Knowledge translation of epigenetic editing: equity, diversity, and inclusion as an epistemic issue
Any adaptation of epigenetic editing to clinical or commercial use requires policymakers, clinicians, and various stakeholders to understand the nuanced, scientific mechanisms behind the technology that stand to separate it from the accepted process of genetic editing.35,36
However, the current method of propagating epigenetic editing knowledge is inaccessible to a broad audience, which creates some epistemic barriers in knowledge translation efforts.37,38 The most obvious barrier to accessibility, which is common to all kinds of emerging biotechnology, is the language barrier. 38 In order for stakeholders to understand the epigenetic editing process, they would require a thorough understanding of complex scientific terms and concepts: regulatory elements, methylation analysis, transcription factors, chromosome structure, the list goes on. This language requirement thus becomes a practical barrier to understanding and propagating epigenetic editing knowledge between stakeholders of different backgrounds. 38
Another barrier stems from the social context that epigenetic editing knowledge is translated from. The use and propagation of Western medical knowledge has historically focused on particular socio-economic and racial classes at the expense of other underrepresented populations.39,40 While this is in itself a diversity issue, it becomes an epistemic issue when we consider how it impacts the kind of epigenetic editing knowledge created.
The development of genomic knowledge such as the Human Genome Project has been permeated with epistemic contingencies, namely that databases of reference genomes consisted primarily of male and white individuals. 41 Previous efforts to diversify the database simply meant including a token person from another race to represent the genomic data of said demographic. 42 As a result, researchers could not establish equivalent predictive power whether the genomics knowledge will apply to these other demographics. 43
Likewise, epigenetic editing knowledge must be used carefully to avoid perpetuating social inequities, especially as emerging research shows that sex, race, or age markers can be correlated to the epigenome. 44 The underrepresentation of particular social demographics translates to epistemic exclusion when generalizing inferences made from insular databases; knowledge that supposedly applies to all demographics is in fact knowledge that only applies to select demographics. 43 In addition, it reinforces the social imbalances already existent in society, including the diversity inequities that still exist today within academic and scientific practices involving epigenetic and genomics research.45,46
For example, taking a gender- or race-blind approach to epigenetic editing studies in clinical research mistakenly assumes that these existent social inequities would not affect the kind or value of clinical knowledge created. This would only further the mistrust in scientific technology within underrepresented populations, such as Black and Indigenous communities, over the supposed utility and universality of Western medicine. 47 When emerging epigenetics research increasingly points to social inequities being discernable as epigenetic states, such as perceived racial discrimination detected in DNA methylation among African American women, clinicians, regulators, and policymakers will need to acknowledge the tangible consequences of these inequities on epigenetic knowledge.46,48
Ethical Considerations: Concerns with Appropriate Uses of Epigenetic Editing
As with any application of scientific technology, ethical issues often arise from the context in which it is used. For epigenetic editing, some issues are specific to the technology itself, some are relevant to gene editing in general, and some are consequences of the existent social inequities described above. 49
Here, we describe some plausible ethical scenarios to consider before epigenetic editing becomes adapted for widespread usage in humans, building on the concerns raised by others in the genomics community.50–55 These concerns serve as a caution to preemptive translation efforts without careful ethical consideration. Although hypothetical, we provide evidence for ethical concerns already occurring within emerging epigenetics research and show how these concerns can perpetuate health and social inequities in the genomics field.
Consequences of reductionism in healthcare, public policy, and health outlook
A common epistemic issue underlying all genomics-based technologies is the tendency to rely on a reductionist view of health states. For the epigenetics field, it means reducing diseases or disorders to a set of epigenetic states inherited by genetics and/or influenced by environmental factors. 53 This view underlies the driving force in epigenetic editing translation efforts: that one can use epigenetic editing to cure a condition by changing the underlying epigenetic state.
In epigenetics literature, several findings correlate epigenetic markers to chronic health issues such as cancers, depression, neurodegenerative diseases, psychiatric diseases, developmental diseases, and imprinting disorders. 6 For example, rare diseases such as Beckwith–Wiedemann syndrome can be diagnosed through an analysis of epigenetic markers alone. 54 Other epigenetic variants, especially in psychiatric conditions, are less clear-cut but show promise to act as biomarkers of disease or confer protection against disease. 55 These discoveries can be insightful, as they shed light on previously unexplained rare diseases, and become a great motivator to develop downstream editing technologies for human use.
However, this epistemic issue becomes an ethical one when the reductionist view of genetics is applied to medicine. Media narratives and scientific research often portray epigenetics as the underlying “cause” of health states, as if epigenetics is the (often sole) reason for the disease. 56 However, it would be a gross oversimplification to see disease this way, even for diseases with specific and measurable epigenetic markers. 53 At best, the correlation of epigenetic states to its actionability on disease is still in the premature stages of clinical validity. 55 In other words, researchers are only beginning to confirm which disease can be detected as epigenetics states, and it is still unclear if these epigenetic differences are truly due to underlying genetic causes or a result of other influences.
On the other hand, one can argue that the conception of “disease” versus “normal” state is even more blurred within the epigenetics field. Disease is multifaceted and contextually defined; how humans experience disease also depends on social norms and practices, with social determinants of disease possibly detected within a person's epigenome. 57 Taking a nuanced view of disease may mean tracing how epigenetic traits manifest differently over time and in response to various environmental changes. This is different from how we conceptualize diseases within a reductionist mindset, with disease seen as genetic variants often determined at birth. 58 In fact, for the majority of health conditions, diagnosing epigenetic issues can be highly subjective if there is no persistent or distinct epigenetic marker that can be measured.
The problem with the reductionist mindset is that it becomes easy to forget the complex and dynamic nature of epigenetics. Outside of cancer research, current epigenetics discoveries are mostly speculative thus far for detecting disease. Yet, these narratives often make highly speculative claims on the future of epigenetic editing to “fix” these issues. This in turn can lead to contentious promises that seek to identify or correct epigenetic states for “disorders” such as violence, alcoholism, or sexual orientation—conditions that are being associated with epigenetic states.59–63
Therefore, regulators and healthcare professionals should carefully assess the relation of epigenetic editing to disease, especially for loosely-defined disorders, psychological disorders, nonterminal physical diseases, or multifaceted diseases with complex causes. Seeking a cure through epigenetic editing should not necessarily be seen as the only way to “cure” a disease, and it is not without concern.
Permissible uses of epigenetic editing for the prevention of chronic and lifelong disability
From a utilitarian perspective, the prevention of chronic diseases and lifelong disability is one of the greatest economic benefits to the healthcare industry. 64 There is increasing research that shows epigenetic misregulation as the underlying cause in rare diseases, cancers, and diseases with previously unknown causes. 17 The use of epigenetic editing could thus become a cost-efficient method to mitigate chronic disabilities before they happen, as preventing a disease or chronic disability can be more desirable than treating an existing condition. 17 However, even if epigenetic editing technology could permanently correct an epigenetic marker of disease, there is no guarantee that the disease itself can be prevented.
“Permanently edited” epigenetic states can always be influenced by shifting environmental factors, or epigenetic reprogramming during early embryo development.65,66 Relying on the supposed permanence or utility of epigenetic editing to prevent disease simplifies the idea of disease on a public health level, while overlooking the social and environmental influences of disease—a problem that epigenetic editing cannot cure.
The greater ethical concern lies in the promise of epigenetic editing as the ideal way to address diseases and disorders (whether permanently or temporarily). This is a common view adopted by stakeholders when speaking about the value of epigenetic editing for humans, but it implies the erasure of disability as a legitimate status of existence. Disability activism, as the acceptance of human bodies that differ from idealized medical and clinical standards, is already an underrepresented viewpoint in healthcare. 57 Epigenetic editing technologies employed in the context of curing chronic diseases exacerbates the idea of “perfect genetic fitness” as the ideal health goal, and ultimately compromises disability activism that seeks to normalize a diversity of physiological, mental, and genetic states in healthcare.
Furthermore, stronger ethical consideration should be given to distinguish the generational impact of epigenetic editing in disability erasure by artificially eliminating a diversity of health states. For example, correcting embryonic epigenetic diseases could be morally permissible if it would otherwise lead to embryo loss, but correcting the epigenetic state of viable embryos with perceived debilitating diseases can lead to erasure of the disability community if it effectively erases the physiological or psychological differences of human bodies. 57
This distinction also implores us to reconsider a distinction often overlooked in public health policymaking: whether disabilities should be prevented altogether, cured through genetic engineering, or supported through acceptance and accommodation. Right now, it is unclear if epigenetic editing would establish a middle ground between health and disability or exacerbate the existing neglect of the disability community.
Consequences of modifying sex/gender-related traits in embryo development
Sex and gender-based discrimination has prevalently been an issue in society. Even without the use of scientific technologies, societies have historically normalized discriminatory practices such as favoring the selection of male babies or promoting feminine traits in women. 67 The reduction of sex, gender, or sexuality characteristics to biological elements can become another method of justifying sex/gender-based discriminatory practices in genetics and embryo development research. 68
Today, epigenetics research is tightly tied to, and actively shapes, the evolving discourse on sex and gender. For example, there is ongoing research that characterizes the differences in epigenetic expression between male and female embryonic development. 69 Epigenetics has also been shown to affect sexual differentiation in the neonatal brain of animal models, while DNA methylation and histone acetylation of the estrogen receptor alpha has been linked to differences in masculine/feminine brain functions.70–72 Furthermore, it is possible through epigenetic regulation to control the sexual differentiation of a male embryo into a female one or selectively regulate some sex/gender-related molecular characteristics. 44
Currently, international ethical guidelines discourage germline gene editing and sex/gender-related embryo modifications for reproduction due to the permanence of these changes on future generations73,74 By contrast, the perceived transient and reversible nature of epigenetic editing could lead to greater acceptance of its usage to enhance biological traits in humans, especially for projects of human improvement unrelated to health. These possibilities are supported by the principlist and individualist mindset of Western healthcare ethics, which promotes individuals to have greater autonomy and freedom of choice in decisions related to personal or child development.75–77
However, there are concerning consequences with allowing the usage of epigenetic editing that is motivated by discriminatory social beliefs, such as the erasure of transgender and intersex persons, or artificial selection for feminine and masculine traits.37,78 Epigenetic editing without nuanced conversations about sex and gender inequities can thus exacerbate existing sex/gender-based discrimination if proper caution is not taken to inform stakeholders and parents about the ethical implications of epigenetic editing.
Issues of justice and access over the use of epigenetic editing technology
Epigenetics has been shown to be highly relevant to biological aging and anti-aging research. 53 One of the biggest breakthroughs in epigenetic editing is the possibility of altering the epigenetic clock (often considered a metric of biological age) by changing the DNA methylation patterns of various human cell and tissue samples. 79 Epigenetic editing has also proven useful in other fields related to human biology, such as cancer research, the skincare and beauty industry, and the food industry. 80 While these milestones should be encouraged in theory, they bring vast and unique concerns specific to each industry regarding the ethical implementation of epigenetic technology. 81
One common ethical belief underlying all applications of epigenetic editing is the applicability of this technology for the greater good. Many people believe new scientific technologies will directly and immediately benefit the general population in the greatest manner.
From a feminist ethics perspective, existing social inequities often overshadow altruistic attempts to implement medical technology for human improvement. People are situated with different privileges in society based on intersecting social identities. 82 These privileges translate to systemic imbalances when they intersect with real situations, such as access to novel medical technologies. 83 For example, it is already an existing issue that those with high socioeconomic status (SES) are more likely to seek out and have easier access to health-promoting technologies, while lower SES demographics face multi-faceted barriers to accessing the same technologies. 84 Applied to epigenetic editing, this could lead to instances where benefits of this technology become primarily accessible only to those with exceptional status in society, such as wealthy funders or private companies.
Basic epigenetic editing research, such as reprogramming telomere regulation or cell-cycle progression, should be encouraged so that scientists can gain a better understanding of the molecular mechanisms of aging. 53 However, the ethical concern is that the product of refined, customizable epigenetic editing, should this goal ever be realized, will only be accessible to those who have the means to obtain it. 85 Thus, stakeholders should first assess the practical social barriers to epigenetic editing to ensure an equitable and fair access for all populations. 81
Future Directions and Potential Policy Implications of Epigenetic Editing
There is reasonable hope that the future of health and well-being will move toward site-specific epigenetic editing for genomic engineering in humans. The widely used non-specific epigenetic modifiers in cancer therapy suggests that epigenetic editing is a key element of concern in serious and chronic diseases. 86
There also exists growing interest in adapting epigenetic editing technologies for commercial use. Start-ups and private companies around the world, such as Inari Agriculture and Epicrop Technologies, are utilizing epigenetic and genetic editing technology to produce transgene-free crops for the food industry. 80 Recently, Chroma Medicine announced $125 million in private financing for the development of epigenetic editing for treating human genetic diseases. 87 In light of these developments, effective, site-specific epigenetic editing is expected to become a widespread technique for fixing human conditions.
Based on the translational habits of past scientific technologies, future applications in epigenetic editing could lead to direct-to-consumer epigenetic editing services, medical tourism for epigenetic editing clinics, and black-market distribution of epigenetic editing kits.
One of the biggest translation concerns is that goals of adapting epigenetic editing technology may quickly surpass the ability of scientists, regulators, and ethicists to establish proper protocols ensuring its safe, efficacious, effective, and ethical use. 49 The past has shown that preemptive implementation of genomic technologies can pose conflicts, such as 23andMe's regulatory issues with the U.S. Food and Drug Administration, in attempts to put genomic sequencing technology at the hands of the global population. 88 The same may happen with efforts to adapt epigenetic editing technology for widespread use in humans.
Recall that many scientists and stakeholders believe the purported transient and reversible mechanism of some epigenetic editing systems could designate it a “safer and easier” way to achieve genomic engineering compared to gene editing. 18 However, it should not be a reason to overlook the development of this technology. 49 Because of the complexity in epigenetic regulation within human biology, our understanding of epigenomic editing may be incomplete, such as off-target effects spanning multiple temporal and spatial domains. 21 In fact, it is not the case that there are fewer safety concerns in the therapeutic application of epigenetic editing; rather, the potential safety issues of this technology are conceptually different and more difficult to measure than those of genomic editing. 89
Thus, the increasingly popular narrative that epigenetic editing is safer is yet to be proven. Although there are seemingly less detrimental consequences in the context of human applications, many of the epistemic and ethical issues of epigenetic editing remain unsolved, and warrants further attention before downstream uses lead to concrete social inequities.
Conclusion
In conclusion, this review highlights some important epistemic and ethical issues regarding the translation of epigenetic editing (see Fig. 2). Epistemic issues arise from the way epigenetic editing is compared to genetic editing, including diversity issues that can influence the kind of epigenetic editing knowledge developed. This in turn leads to ethical issues such as the perpetuation of social inequities that shape the value of this technology.
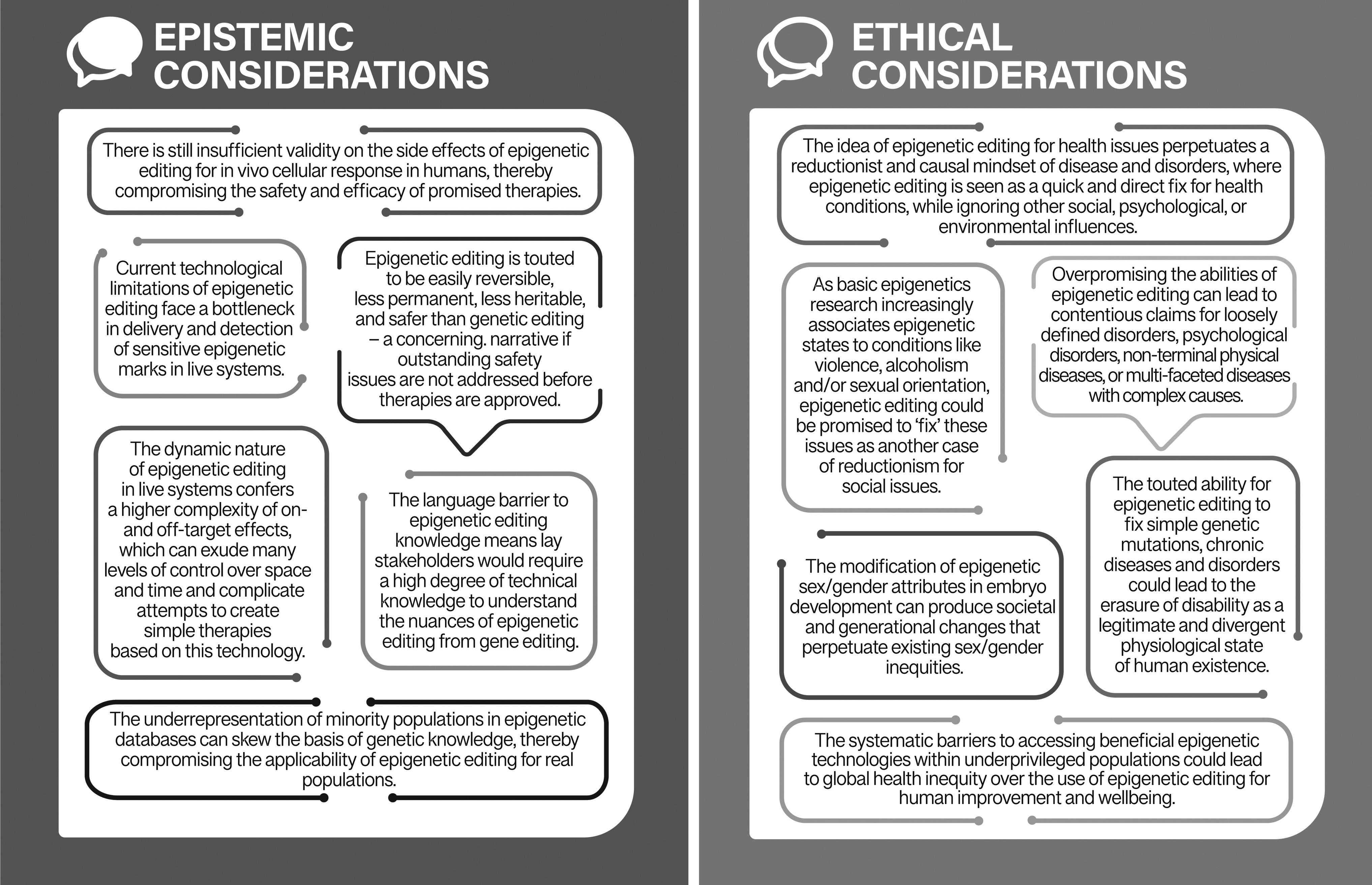

Summary of the epistemic and ethical considerations highlighted by this review in the translation of epigenetic editing for human use.
As a starting point, we make the following recommendations. First, epigenetic editing technologies should be developed with equitable access in mind during every step of the translation process, which will help to combat the issues highlighted in this review and other editorials.90,91 Second, stakeholders should revisit existing guidelines, policies, and governance on gene editing and amend its scope to include the use of epigenetic editing in human bodies, taking into account the raised ethical considerations before epigenetic editing becomes introduced in society. 55 Third, more research is needed to determine how desirable and safe epigenetic edits can be made in a stable manner comparable to genetic edits.
Finally, special care should be taken to communicate the epistemic peculiarities of epigenetic editing technology apart from its gene editing predecessors to avoid misconceptions about the current limits of this technology in a manner that is clear to lay audiences.
Footnotes
Acknowledgments
The authors would like to thank Dr. TE Hébert (McGill University) for his efforts in revising earlier manuscripts, as well as Dr. Martin Hirst (University of British Columbia) and the members of the IHEC Bioethics workgroup for their insightful comments on the manuscript.
Author Disclosure Statement
None of the authors have any competing interests or financial interests in relation to the writing and publication of this manuscript. None of the authors were, are, or anticipate being employed by an organization that may gain or lose financially from publication of this manuscript. To the best of our knowledge, none of the authors have any personal relationship that may inappropriately affect the integrity of the reported research.
Funding Information
The manuscript was funded by the Canadian Institutes of Health Research (CIHR) through the Multidimensional Epigenomics Mapping Centre (EMC) at McGill University (CEE-151618) and the Lo Ka Chung Foundation through the Hong Kong Epigenomics Project.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
