Abstract
Abstract
The CRISPR-Cas9-based multiplexed gene editing (MGE) provides a powerful method to modify multiple genomic regions simultaneously controlling different agronomic traits in crops. We applied the MGE construct built by combining the tandemly arrayed tRNA–gRNA units to generate heritable mutations in the
Introduction
Wheat is the second most important food crop in the developing world. Currently, the genetic diversity of the natural1,2 or chemically mutagenized populations 3 of wheat and its wild ancestors 4 are the main sources of useful allelic variation for wheat improvement. 5 However, the deployment of these alleles in breeding is affected by the distribution of genetic diversity across genes controlling a trait, as well as the local recombination rate around these genes that can be severely suppressed in the peri-centromeric, highly divergent, or structurally rearranged genomic regions. CRISPR-Cas9* technology, with its multiplex genome editing capacity,6–14 holds great promise in overcoming some of these limitations and provides a tool for inducing beneficial modifications in multiple genes controlling major agronomic traits.
Since the introduction of CRISPR-Cas9 for editing mammalian genomes,8,9,15 it has been applied to modify the genomes of a number of model and crop plants, including tobacco,16,17 tomato,6,18 barley, 19 Arabidopsis, 16 wheat,10,20–25 rice,11,14 and maize.7,26 Following the first reports of CRISPR-Cas9-based genome editing in wheat protoplasts,22,25 gene edited plants have been regenerated from the wheat immature embryos transformed with CRISPR-Cas9 in different forms, including plasmids, linear DNA fragments, linear RNA, and ribonuleoprotein complexes.20,21,23
Several strategies have been developed for multiplexed gene editing (MGE). The first MGE reported in human cells mimicked the natural architecture of the CRISPR-Cas9 locus; protospacers targeting different genes were included into a single pre-crRNA array, which was then processed into individual crRNAs by Cas9, tracrRNA, and endogenous RNase III. The resulting ribonucleoprotein complexes, including Cas9, tracrRNA, and crRNA, were capable of inducing mutations at multiple target sites. 9 Another MGE strategy based on combining multiple gRNAs under the control of their own promoters into a single construct was also successfully applied in wheat, rice, tomato, maize, and Arabidopsis.7,11–13,16 However, due to the size of the individual promoter-gRNA units (450 bp), the level of multiplexing achievable using this approach is limited by the insert size capacity of plasmid vectors. The third strategy that used the Csy4 nuclease and its 28 bp recognition site to generate functional gRNAs from the tandemly arrayed units was successfully tested in human cells 12 and several plant species. 10 The small size of the Csy4 recognition site allowed a higher level of multiplexing to be achieved but required the simultaneous expression of Csy4. Finally, effective MGE in rice was accomplished by spacing multiple gRNAs with tRNA under the control of a single promoter, where functional gRNAs are generated by processing a polycistronic gene transcript through the endogenous tRNA processing system. 14 The functionality of gRNAs produced by either Csy4 ribonuclease or endogenous tRNA processing systems was successfully demonstrated in immature wheat embryos. 10
Here, we investigated the frequency and heritability of the mutations generated by the MGE construct expressing the gRNA–tRNA units in allopolyploid wheat. The MGE construct included gRNAs targeting the
Materials and Methods
Plasmids and vector construction
The wheat codon-optimized Cas9 fused with the nuclear localization signals and wheat U6 promoter-gRNA was synthesized by Integrated DNA Technology. Plasmid pA9mRFP, pA9Cas9, and pU6sg were constructed as shown in Supplementary Fig. S1 and Supplementary File 1 (Supplementary Data are available online at www.liebertpub.com/crispr). Plasmid pBUN4217 with the maize codon-optimized Cas9 gene (Supplementary Fig. S1) and pGTR 14 containing gRNA–tRNA units were ordered from Addgene.
To design gRNAs targeting the
Protoplast transformation and DNA isolation
The seedlings of wheat cultivar Bobwhite were grown in the dark for 2 weeks. Shoot tissues were fine-sliced and digested for 2.5 h in an enzymatic solution containing 1.5% Cellulase R10 (from
Next-generation sequencing of PCR amplicons and estimation of gene editing efficiency
To detect CRISPR-Cas9-induced mutations, genomic regions harboring the gRNA targets were PCR amplified and sequenced. To facilitate simultaneous next-generation sequencing (NGS) analysis of multiple genomic regions, Illumina's TruSeq adaptors were added to the both ends of amplicons using two rounds of PCR (Supplementary Fig. S3A). PCR products were purified and sequenced at the K-State Integrated Genomics Facility on the MiSeq instrument (Supplementary Fig. S3B and C). All reads passing quality control were aligned to the wild-type reference sequences. The genome editing frequency at each target site was calculated by dividing the number of mutated reads to the total number of all aligned reads.
Regeneration and genotyping of transgenic plants
Wheat immature embryo transformation and plant regeneration were performed, as previously described. 35 To isolate DNA, leaf tissues were sampled and homogenized in 500 μL of TPS buffer, then incubated for 20 min at 75°C. After centrifugation, 150 μL of the supernatant was mixed with 150 μL of isopropanol and incubated for 20 min at room temperature. DNA was precipitated, washed with 70% ethanol, and re-suspended in 100 μL of deionized water.
The presence of CRISPR-Cas9 constructs in the transgenic plants was validated by PCR using three pairs of primers amplifying different regions of the Cas9 and gRNA expression cassettes (Supplementary File 2). The CRISPR-Cas9-induced mutations were examined only in the plants showing the presence of all three PCR products. The detection of mutations was performed using the NGS-based procedure. Four bar-coding bases between the target-specific primer and Illumina TruSeq adaptors (Supplementary File 2) were added to index multiple plants under each of the Illumina TruSeq barcodes.
Screening of gw2 knockout mutants
To screen the
Screening of the TaLpx-1 gene mutants
The progenies of T1 plant GLM-2-5 were screened for the presence of mutations in the
Plant growth and phenotypic analysis of gw2 plants
Plants were grown in the greenhouse under 12 h of light for 1 month, and then grown until seed harvesting under 16 h of light at a temperature of 24°C during the day and 21°C at night. The MARVIN seed analyzer (GTA Sensorik GmbH) was used to estimate the TGW and grain width, length, and area of the wild-type plants (genotype
Statistical analysis
To evaluate the segregation ratio of mutated alleles in T1 plants, a chi-square test was performed using the CHITEST function implemented in Microsoft® Excel. A two-tailed Student's
Results
Multiplex genome editing in the wheat protoplasts
The editing efficiency of the designed gRNAs targeting the
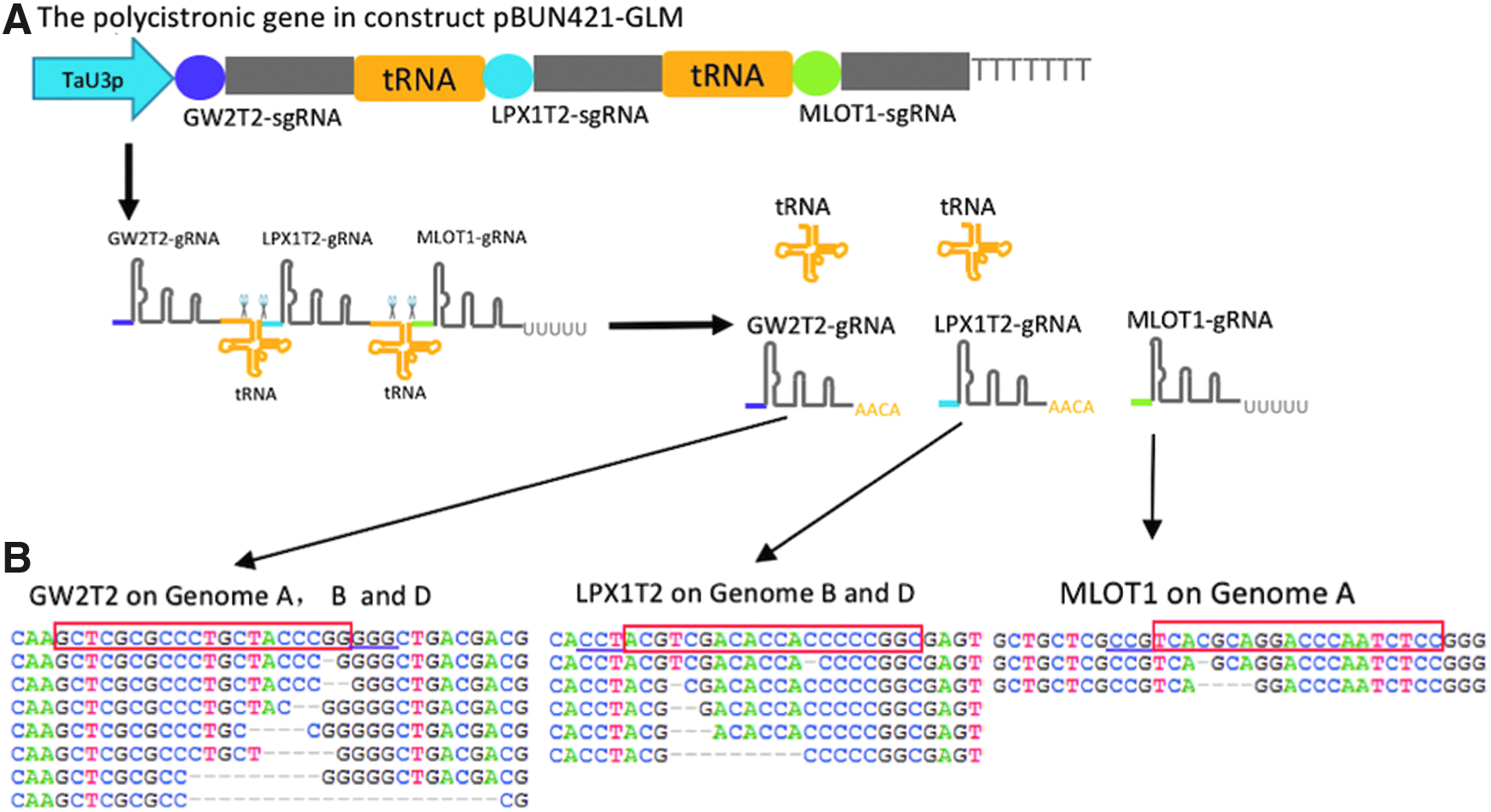
CRISPR-Cas9-based multiplex editing in hexaploid wheat using gRNAs processed through the endogenous tRNA-processing system.
The examples of mutated reads are shown in Supplementary Figures S5 and S6.
Proportion of mutated reads was normalized using the 20% protoplast transformation efficiency estimated for this experiment.
Transmission and phenotypic effects of the MGE construct-induced mutations
The transgenic plants were regenerated from the wheat immature embryos transformed with the MGE construct pBUN421-GLM. The single gRNA construct pBUN421-GW2T2 was used as a control. In total, 102 plants were regenerated for the pBUN421-GLM construct (henceforth GLM plants), and 61 plants were regenerated for the pBUN421-GW2T2 construct (henceforth GT2 plants). Based on the PCR results, 22 GLM plants and 17 GT2 plants had the intact CRISPR-Cas9 expression cassette. Six GLM plants and six GT2 plants carried mutated alleles at the GW2T2 target site, and nine GLM plants carried mutated alleles at the MLOT1 target site (Fig. 2A and B, Supplementary file 5). Five GLM plants carried mutated alleles at the GW2T2 and MLOT1 target sites simultaneously. While mutations at the LPX1T2 target site were detected by NGS in several GLM plants, within the analyzed set of lines, we did not recover fixed heritable mutations at this site (Supplementary File 5). Among the 12 plants that had mutations at the GW2T2 target site, only one plant, GLM-1, carried the mutated version of the
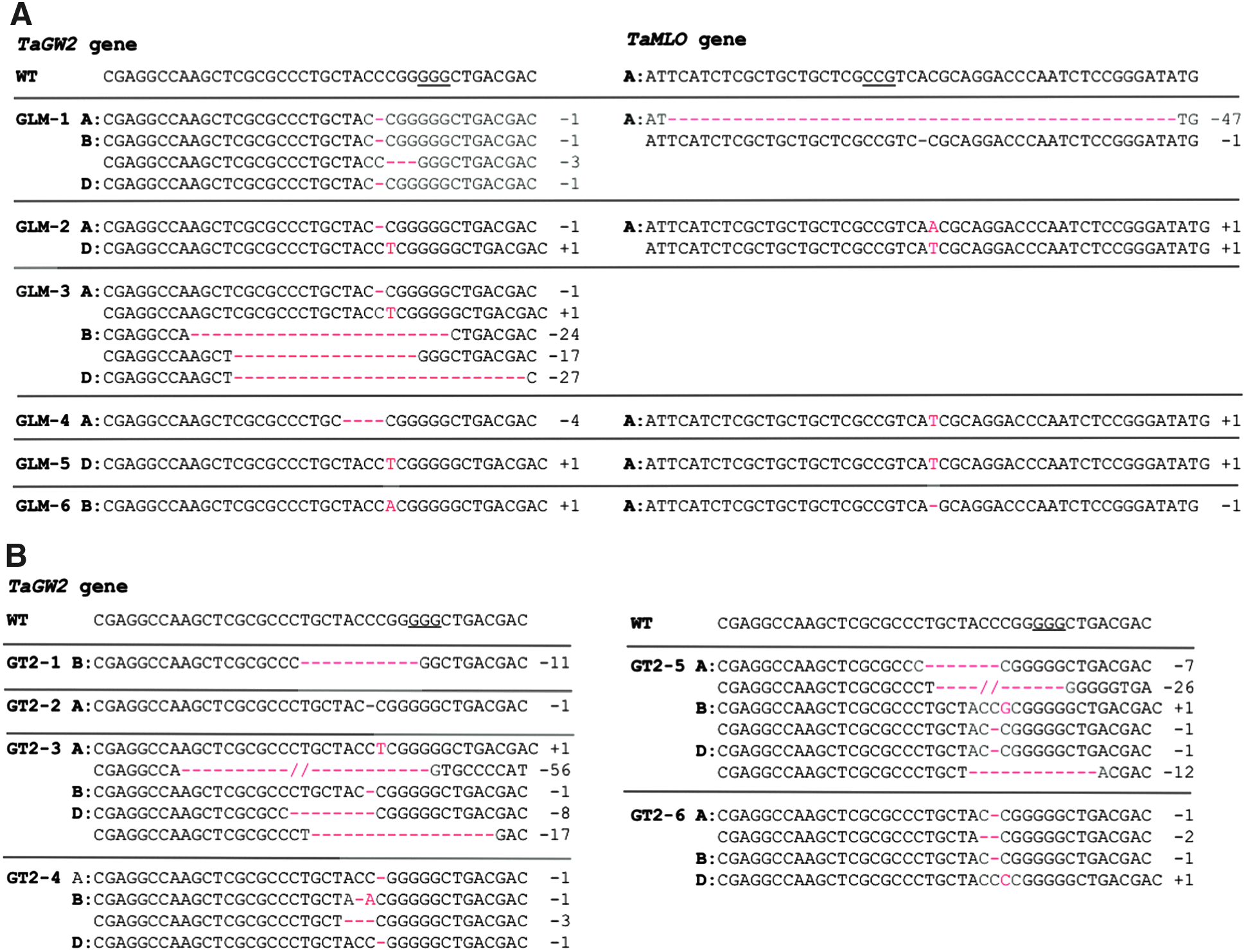
CRISPR-Cas9-induced mutations in the T0 transgenic plants identified by NGS.
The transmission of CRISPR-Cas9-induced mutations to the next generation was investigated in the self-pollinated progeny of two T0 GLM plants (GLM-1 and GLM-2) with the mutant alleles in both the
A/a, B/b, and D/d represent loci in different genomes; the uppercase and lowercase stand for wild-type and mutated alleles, respectively.
In GLM-1, the B genome copy of the
In GLM-1, the A genome copy of the
Plant GLM-2 has wild-type reads and two different types of mutated reads (Fig. 2A) in the A genome copy of the
New
We investigated the effects of MGE construct-induced mutations on the phenotype by measuring easily scorable seed morphology traits in the T1 progeny of GLM-1. T1 plants carrying knockout mutations in all three copies of the
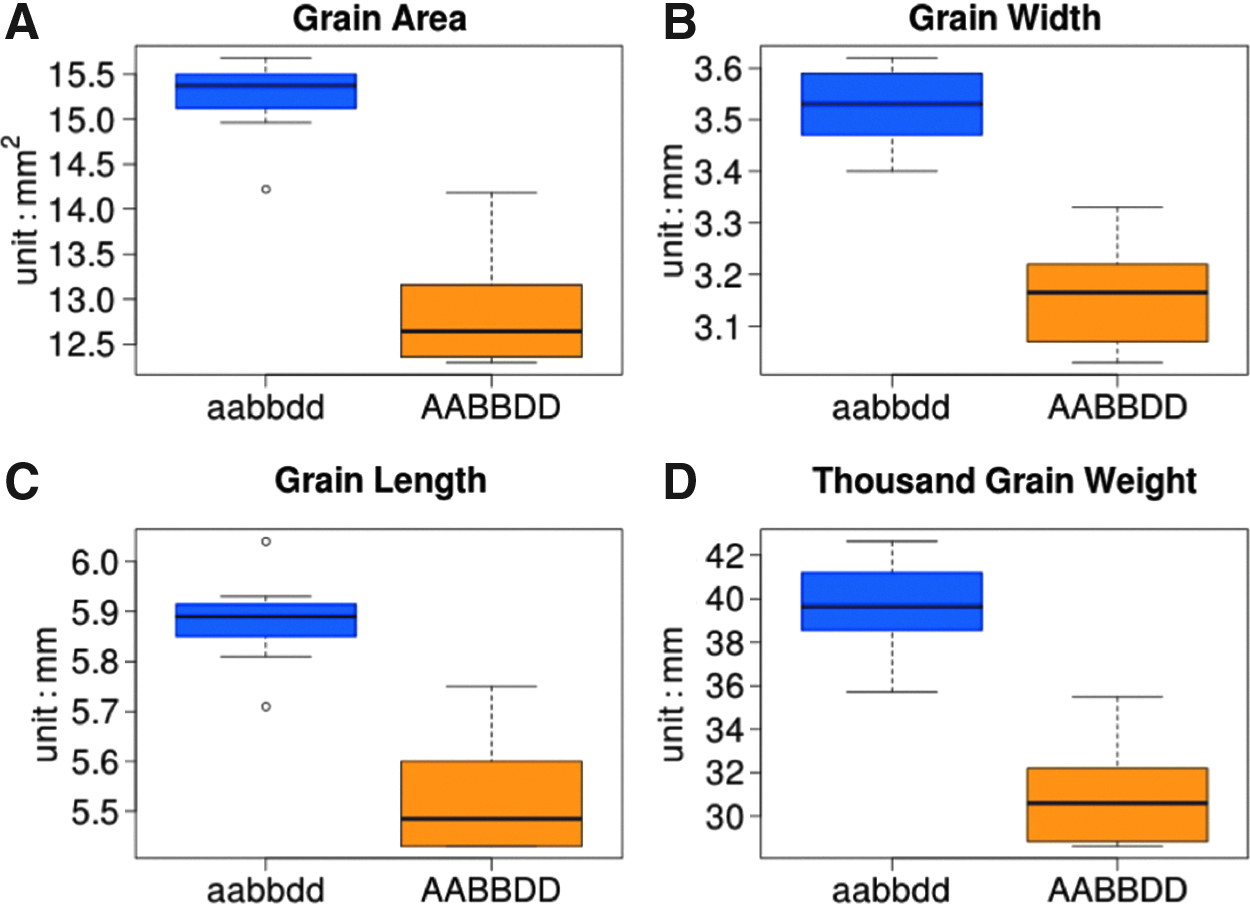
Phenotypic effects of CRISPR-Cas9-induced mutations in the
Generation of novel mutations in the TaGW2 and TaLpx-1 genes by transgenerationally active CRISPR-Cas9
Among the 39 CRISPR-Cas9-positive plants, we detected only one
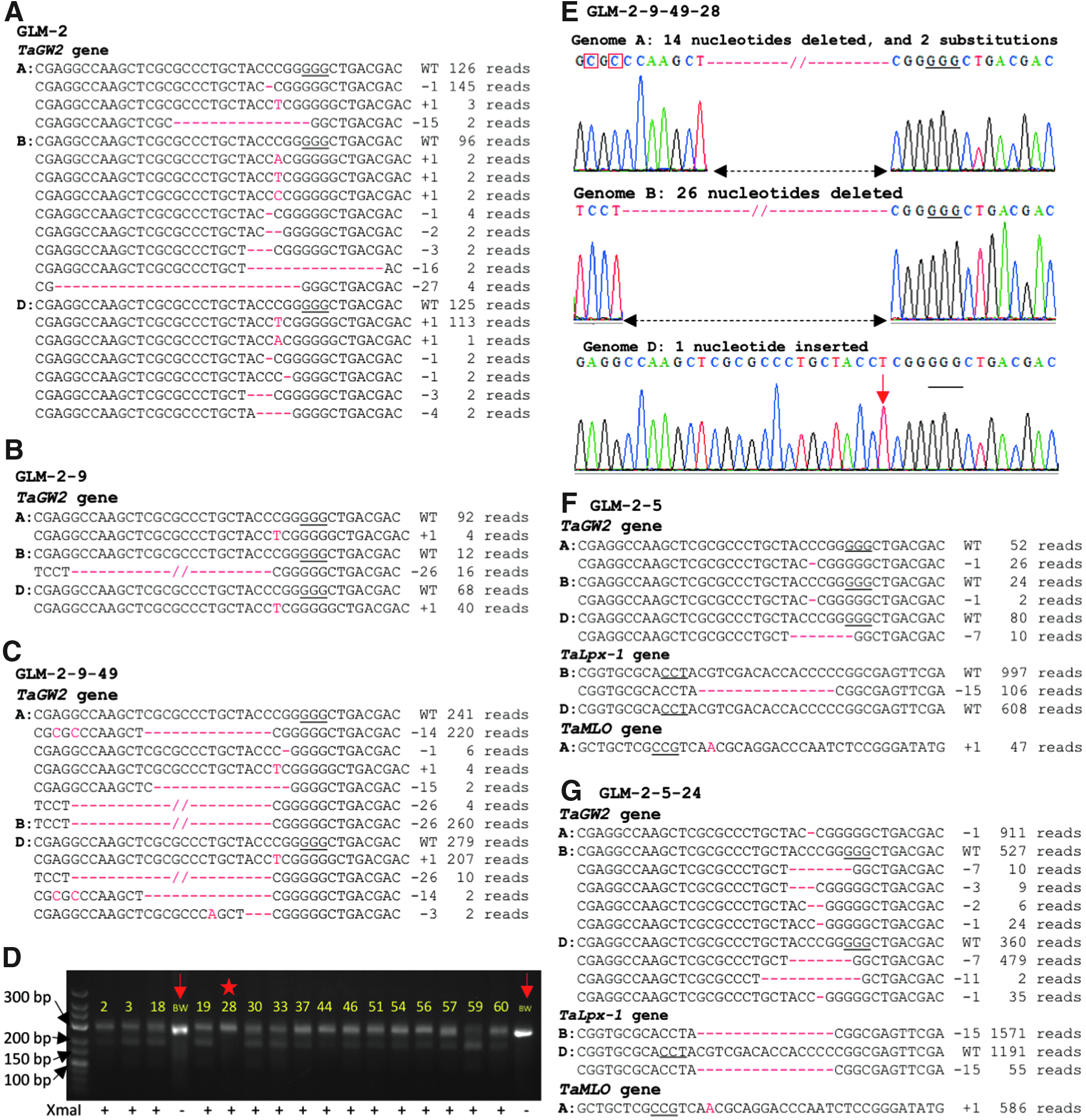
Transgenerational CRISPR-Cas9 activity induces new mutations in the
To investigate the process of gene editing across generations, we screened the progenies of several plants lacking mutated alleles in at least one of the homoeologous genomes at the target sites (Fig. 4 and Supplementary Figs. S7–S9). For the GLM-2 plant that carried the
Discussion
Consistent with the results previously reported in rice and maize,14, 36 we demonstrated that heritable mutations in multiple gene targets in the wheat genome can be induced using a construct with an array of the gRNA–tRNA units driven by a single promoter. The phenotypic effects of the MGE construct-induced mutations were confirmed by measuring the seed morphology traits of the
The MGE approach provides a flexible tool for implementing complex gene editing strategies in wheat where the majority of genes have at least three homoeologous copies. By designing gRNAs to the targets carrying homoeolog-specific sites or to the conserved targets identical in all three wheat genomes, it should be possible either to edit multiple genes from a specific genome selectively, or to introduce modifications into all duplicated gene copies. Thus, multiple beneficial allelic changes across the wheat genome can theoretically be induced by transforming plants with a single polycistronic tRNA-gRNA/Cas9 construct. However, our results suggest that the recovery of T0 lines carrying mutations in all copies of multiple genes for polyploid plants, due to whole genome duplication, could be more challenging than for diploid plants.
The NGS of CRISPR-Cas9-expressing plants allowed us to detect rare somatic gene editing events present in only part of the cells in the leaf tissues, indicative of ongoing CRISPR-Cas9 activity. The successful recovery of novel gene editing events in the progenies of transgenic plants suggested that this CRISPR-Cas9 activity can serve as the source of novel heritable variation. Consistent with this conclusion, we successfully recovered plants with multiple edited targets in the progenies derived from a single plant carrying the active CRISPR-Cas9 MGE construct. Another advantage of the transgenerational CRISPR-Cas9 activity in the MGE construct-expressing plants is that it provides the possibility for obtaining lines carrying different combinations of mutations across multiple targets. For traits with a complex genetic basis, these lines can be used to study epistatic and/or additive interactions among multiple edited gene variants and to select the most optimal combinations of novel alleles.
Recently, the transgenerational CRISPR-Cas9 activity has been shown to induce new functional variants at multiple target sites in tomato F1 plants. 6 This result suggests that beneficial mutations in the wheat breeding lines not amenable to transformation could be induced by crossing these lines with the wheat lines carrying the CRISPR-Cas9 constructs. This strategy provides an effective approach for the systematic transfer of beneficial gene editing events into the breeding programs for further phenotypic evaluation of their effects in different genetic backgrounds.
Conclusion
Our study demonstrated that the MGE construct employing endogenous tRNA processing system can be effectively used to induce heritable mutations in the genes controlling multiple agronomic traits in hexaploid wheat. The transgenerational CRISPR-Cas9 activity demonstrated here suggests that effective Cas9-inducible trait transfer can be accomplished by crossing wheat breeding lines with lines expressing CRISPR-Cas9 constructs. Further evaluation of this strategy will be required to assess its value for wheat improvement.
Footnotes
Acknowledgments
This work is supported by the Agriculture and Food Research Initiative Competitive Grants 2017-67007-25932 from the USDA National Institute of Food and Agriculture and by the Bill and Melinda Gates Foundation grant BMGF: 01511000146. We would like to thank Shichen Wang for assistance with the initial bioinformatical data processing, and Katherine Jordan for the valuable discussions of the presented work. We would like to thank Qi-Jun Chen for providing plasmid pBUN421, and Yinong Yang for providing plasmid pGTR.
Author Disclosure Statement
All the authors declare that no competing financial interests exist.
*
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
