Abstract
The T
INTRODUCTION
The T
When all pairwise distances are observed without error, T
Observed distances are usually noisy due to uncertainty in the measuring equipment (Huang and Dokmanić, 2021). This leads to the N
We propose a new approach that casts the problems N
We empirically demonstrate the performance of our proposed bilevel algorithm and subsequently show that it outperforms state-of-the-art methods in various synthetic and realistic biological settings, such as the partial DNA digestion task (Huang and Dokmanić, 2021). Our algorithm comes to accurate solutions even under extremely noisy observation conditions. We also demonstrate that the proposed algorithm runs more efficiently than previous approaches, capable of handling partial digestion instances with up to a hundred thousand digested fragments. This means that our algorithm scales to genome-sized applications, which is not possible with previous methods. We additionally extend our algorithm to a generalized one-dimensional distance matching problem that accepts partitioned and partially labeled distances along with a set of known missing distances. In summary, our algorithms advance the capacity to address the N
METHOD
Problem setting
Let
Let
Eq. (1) is a bilinear program, as fixing either P or z reduces the problem to a linear program in the remaining variable. This motivates a bilevel MM scheme (Sun et al., 2017) to optimize this objective. We do this by relaxing our objective into two alternating subproblems. At iteration
This implies the permutation that applies P then transposes i and j is no worse than P. Iterating this argument leads to a sorting permutation that is also no worse than P. The initial permutation was arbitrary, so there exists an optimal permutation that sorts x. Finally, note that any two sorting permutations P and
On the other hand, the second subproblem fixes an estimation for
Since the inner product of two vectors is maximized when they are parallel and
As objectives (2) and (4) have closed-form solutions, they motivate a practical bilevel optimization routine described in Alg. 1 and visualized in Figure 1. Note that the unit projection does not affect the permutation in the next iteration, so we omit it until the vector is returned. We avoid storing both the incidence matrix and intermediate distance vector by using implicit matrix multiplication and a problem-specific matching algorithm Alg. 2. The runtime of the optimization inner loop is derived in Proposition 2. In the same proposition, we also derive a memory efficient implementation that avoids storing intermediate values during optimization.
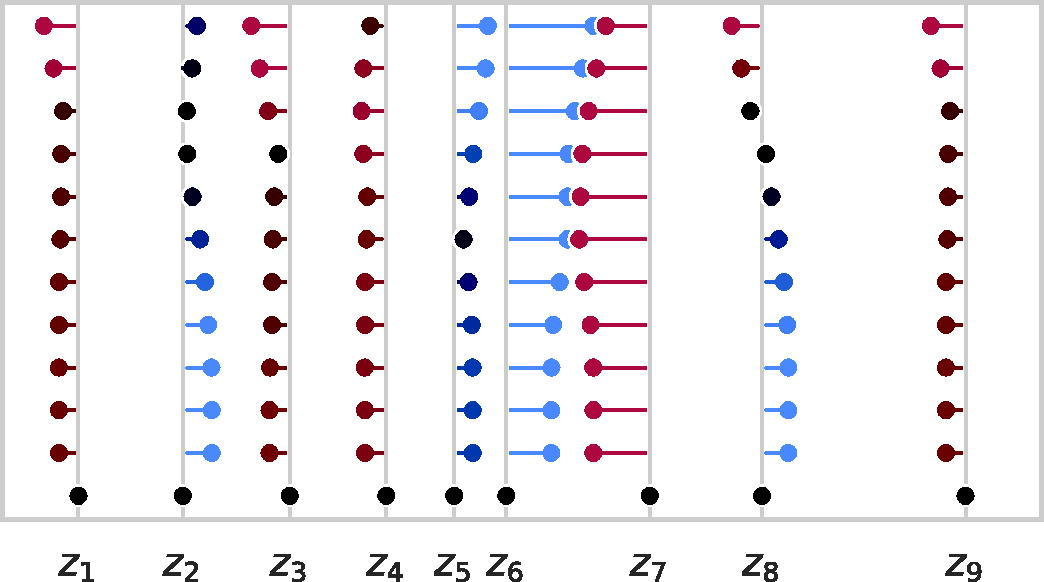
A run of Alg. 1 on a 9-point distance set where the gray lines emanating from the x-axis denote ground truth points. Estimated points for each iteration are drawn from top (the initializer, iteration zero) to the bottom (final iteration, the solution), and an additional line is drawn from each estimated point to the closest ground truth point. These lines are colored red, black, or blue to indicate that the estimated point is too far to the left, nearly matched, or too far to the right relative to the respective ground truth point. Note that, by the final iteration, the estimated points are perfectly matched to the ground truth points, so the estimate lines disappear.
During iteration t, interval
The algorithm performs
For the second claim, notice that steps 5 and 6 of Alg. 1 take
In practice, we observed that Alg. 1 converges quickly but is prone to becoming trapped in local maxima. To prevent this, we further propose a divide-and-conquer heuristic formally described in Alg. 3. In particular, after each pass of Alg. 1, we partition the estimation
The intuition behind our proposed heuristic is that, if
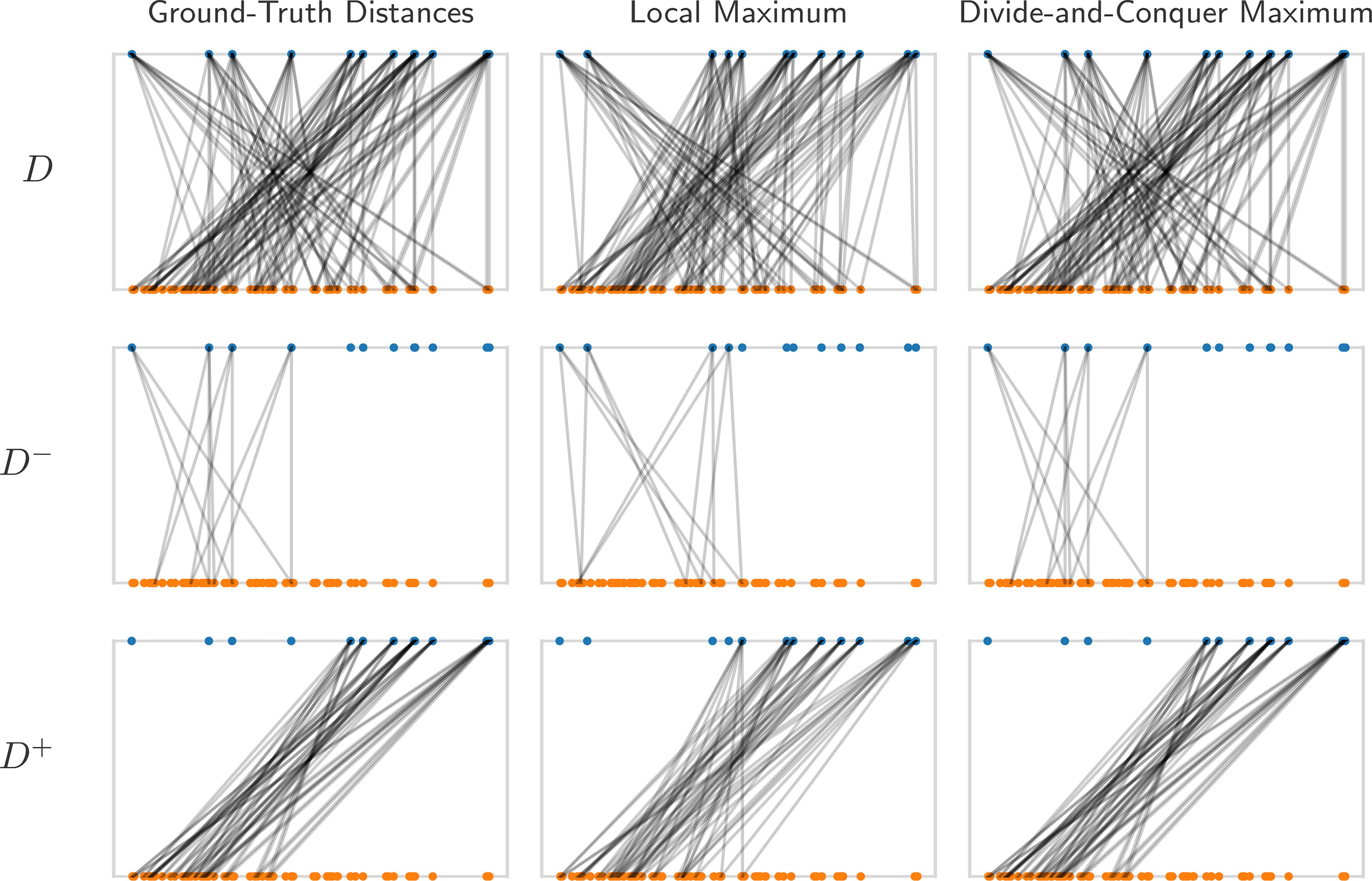
An example of the divide-and-conquer strategy applied to a local maximum, leading to global optimality. The figure presents a series of plots, where the point vector (blue) at the top corresponds to the distance vector (orange) at the bottom. Each distance has two edges to the points that generated it. From top to bottom, the rows represent the full distance set, the distance set associated with negative points, and the distance set associated with non-negative points. From left to right, we see the ground truth matching, local maximum matching, and divide-and-conquer maximum matching. This instance displays the ideal case for the divide-and-conquer approach, where each half of the interval gets nearly its entire distance set.
In the B
The B
We account for the two different arcs between any two points by solving for a 2n dimensional angle vector indexed by integers in
Extension to problem variants
Motivation
The labeled partial digest problem (LPDP) (Pandurangan and Ramesh, 2002) is a variant of the Turnpike problem where we receive both the endpoint distances
Partition formulation
Suppose we receive as input a partitioned distance set
As before, the point vector z is centered and sorted. Suppose that we fix all the permutation matrices except
From Lemma 1 and Proposition 1, an optimal choice of
Note that this is a slight abuse of notation, since we cannot add the permutation matrices; however, this is fine because we can simply lift each permutation to a partial permutation with zeros on all disjoint indices. Thus, an optimal choice is given by the same closed form presented in Alg. 1, i.e.,
Missing distances
Suppose distances
Block-update algorithm
We have a closed-form solution to every subproblem, and the objective is nondecreasing with respect to each closed subproblem. Thus, we can use block updates (i.e., update one subproblem at a time) or parallel updates (i.e., update and aggregate all subproblems) to make progress. For simplicity, the version we present (Alg. 5) is an extension of Alg. 1 that updates all permutations in parallel and then updates the point vector. We stop the optimization once the point vector converges and adapt Alg. 2 to run on the partitioned distance set while preserving the
There are three primary differences between Alg. 3 and Alg. 6.
Before the loop, we initialize indices When a missing distance is relaxed, we simulate the distance using When an interval in
These changes ensure a distance from the correct partition segment is assigned to the current interval, and if it is missing, it is filled in from the current vector
The choice of an initializer for Algs. 3 and 5 is critical to both convergence and reconstruction quality, as a well-chosen initializer leads to rapid convergence, improved stability, and a more accurate solution. Here, we consider three practical initializing schemes. The first scheme samples a random Gaussian vector and sorts it. Though efficient to implement, this scheme is unlikely to produce a good initializer if the ground set exhibits pathological features such as having spread-out point clusters. On the other hand, if the points are well-spread, this scheme often finds a close starting point. The second scheme provides a random permutation
Integer programming formulations
In this section, we develop an integer programming analog of Alg. 5 in order to provide global optimality certificates. To efficiently model our problems, we do not explicitly represent the permutation matrix of Problem 1, as that requires
Recall that a (strong) separation oracle for a convex set
In this formulation, the residual vectors
EMPIRICAL RESULTS
Experimental design
We assessed the performance of our proposed algorithm on synthetic data as a baseline and compared its performance to the backtracking method (Skiena and Sundaram, 1993), the distribution matching method (Huang and Dokmanić, 2021), and our projected gradient descent baseline using the Gumbel-Sinkhorn relaxation (Mena et al., 2018). To evaluate the performance of our method in genome reconstruction, we conduct a series of experiments that aim to reconstruct a DNA sequence from fragments generated by synthetic enzymes. All experiments were implemented in Python 3.10 using a C++20 library implementing the algorithm integrated with Python using PyBind11, and conducted on a computer equipped with 1.0 TB of RAM and two Intel Xeon E5-2699A v4 CPUs.
Synthetic data
Synthetic datasets were generated by sampling n points on the real line from three distributions: the Cauchy distribution, the standard normal distribution, and the uniform distribution on [0, 1]. The uniform distribution was chosen to align with the setting explored by Huang and Dokmanić (2021). The normal distribution was chosen to test point sets with clustered points. The Cauchy distribution was selected to test the effect of contrasting scales, and dispersion of points sampled from the Cauchy distribution presents a challenge for
We examined sample sizes ranging from 50 to 2000 points (in increments of 50) and three additional large sample sizes of 5000, 10,000, and 100,000 to demonstrate the method’s scalability. To simulate measurement uncertainty of magnitude
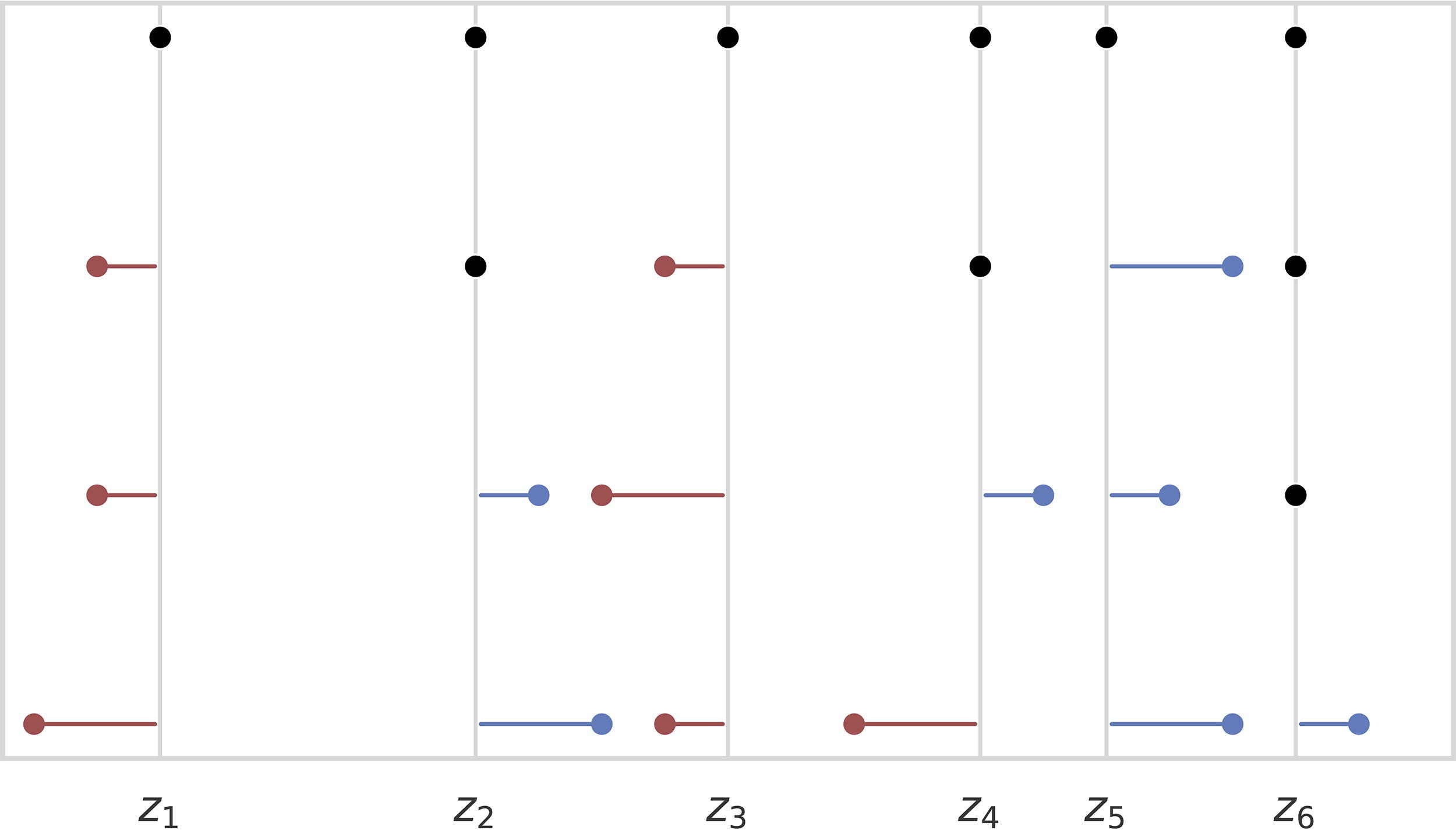
From top-to-bottom, we see the ground truth point set and then point sets that are an MAE of
We investigated the three initialization strategies (Section 2.5) to select one for subsequent experiments. We boosted the Gaussian point vector and permutation point vector initializer by drawing n distinct samples that were scored by solving Problem 2 for each and taking the maximum value. The sample with the maximum score from each strategy was used as the initializer. We used the Gaussian initializer as the starting point for the greedy-search initializer. We tested the strategies across all settings described previously. Figure 4 shows the cosine distances between the estimated and uncertain distance vectors. Among the three approaches, the permutation strategy exhibited the worst similarity scores, with an average magnitude 13 times larger than that of the greedy-search strategy. The Gaussian strategy demonstrated an average error magnitude that was 8 times larger than the greedy-search initialization.
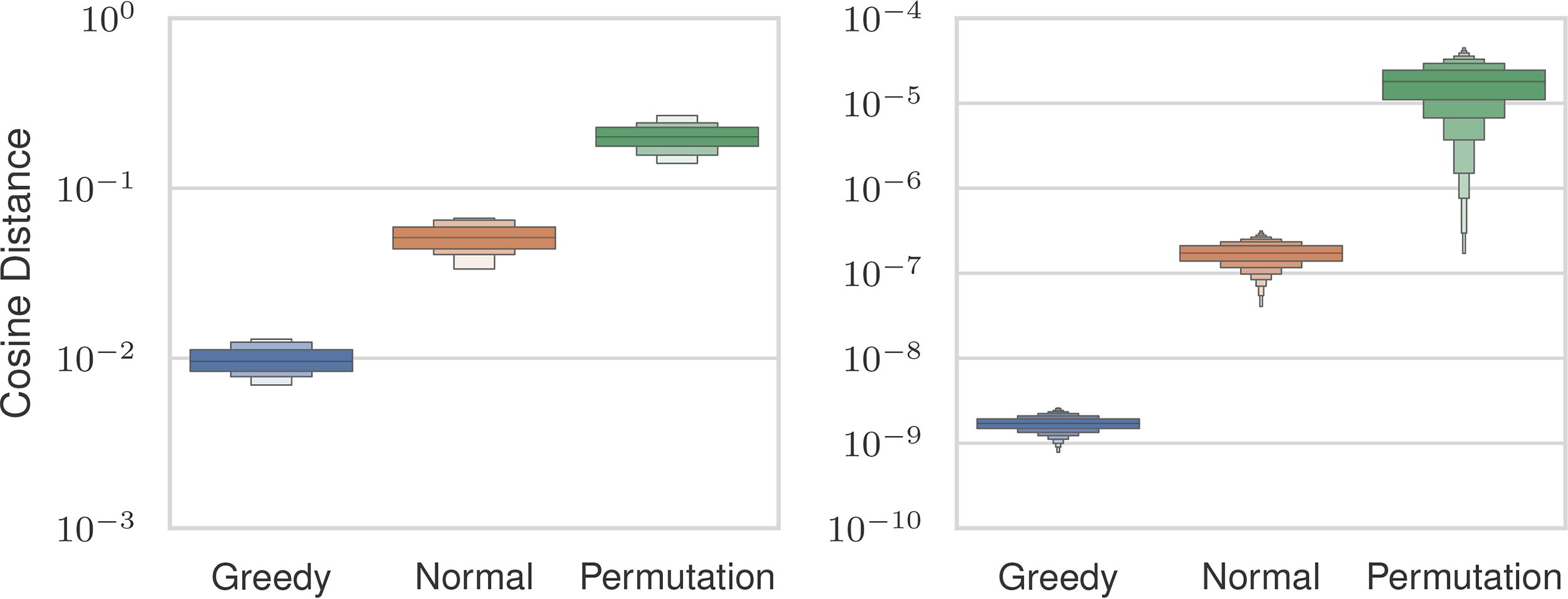
Cosine similarities between estimated and ground distance vectors (
A better initial score does not necessarily guarantee a better reconstruction after optimization. To assess the efficacy of each initializer, we analyzed whether lower pre-optimization errors translated to reduced post-optimization errors. The cosine distance after optimization is also shown in Figure 4. The permutation initialization had the highest errors, and the greedy-search approach had the lowest errors, which is consistent with the pre-optimization cosine distance. We used the greedy-search initializer for our experiments since it exhibited the lowest post-optimization distance. The greedy-search strategy’s lower error comes at a computational cost. Table 1 shows the median runtime for the greedy-search initializer, Gaussian initializer, and optimization loop across a representative set of problem sizes. For all sizes, the greedy-search initialization takes more time than running the optimization, whereas the Gaussian initialization strategy runs in an order of magnitude less time than the optimization. This is due to the inherently serial nature of the greedy-search initializer, which requires all previous steps to be considered first. This is in contrast to the optimization loop, which has a runtime dominated by sorting, which is parallelized.
Median Runtimes (in Seconds) for the Minorization-Maximization Optimizer, Gaussian Initializer, and Greedy Initializer over Different Sample Sizes
We tested how accurately the MM (Section 2.2), backtracking, distribution matching (Huang and Dokmanić, 2021), and gradient descent methods were able to reconstruct point sets. Table 2 shows the median MAE normalized by the uncertainty for a representative set of problem sizes and uncertainties. Each method had 1 hour to solve each instance, with the exception of 10,000 and 100,000 point instances, which were given 90 and 6000 minutes, respectively. The backtracking method was able to solve instances with 1,000 or fewer points, but exhibited larger errors than our method. The gradient descent method solved all instances with 500 or fewer points with residual error that ranged between 10 and 1,000 times higher than the MM approach. The distribution matching method performed similarly to our method but could not scale past 100 points. Our method was able to solve instances with 2,000 points with a median MAE that is 10 times lower than the uncertainty level up to a magnitude of
Median Mean Absolute Error Normalized by the Magnitude of Measurement Uncertainty
across Different Point Set Sizes and Uncertainties
Median Mean Absolute Error Normalized by the Magnitude of Measurement Uncertainty
We compare our method (Minorization-Maximization, MM), Distribution Matching (DM), Backtracking (BT), and Gradient Descent (GD) Approaches. A dash indicates that a method did not finish solving any instances of this size due to either memory or runtime constraints.
Figure 5 shows our method’s MAE and MSE over all settings plotted with respect to the magnitude of uncertainty. We observed that uncertainty in the distances correlated with reconstruction error, but the MAE is an order of magnitude lower than the uncertainty on average when the uncertainty is
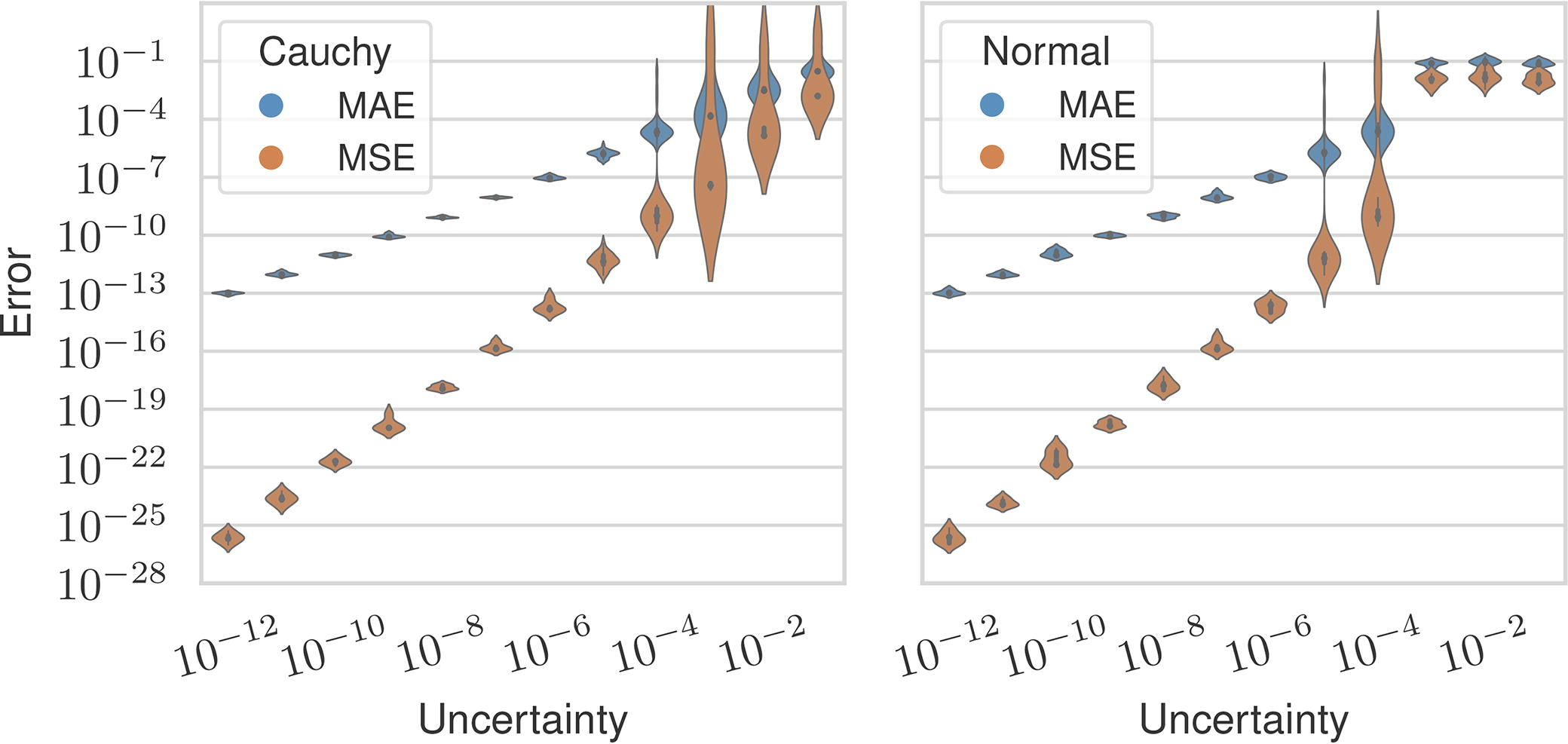
Mean absolute error (blue) and mean squared error (orange) between the estimated and ground vectors across all levels of measurement uncertainty and separated by distribution.
Mean Runtime in Seconds with Standard Deviations across Point Sizes for Various Turnpike Solvers
BT, Backtracking; DM, Distribution Matching; GD, Gradient Descent; MM, Minorization-Maximization.
We test the use of our algorithms for reconstructing genomes in the setting that uses an enzyme to partially digest DNA fragments at restriction sites (Alizadeh et al., 1995). The fragment lengths give the distances between all restriction sites, which are at unknown positions. The genome is assembled from the fragments after inferring the restriction site locations from the distances, a process equivalent to solving the T
Table 4 shows normalized MAE for the T
Normalized Mean Absolute Error for 10 Sizes and 4 Uncertainty Magnitudes Using the Minorization-Maximization (MM) and Gradient Descent (GD) Algorithms on Simulated Partial Digestions of a Linear Genome
Normalized Mean Absolute Error for 10 Sizes and 4 Uncertainty Magnitudes Using the Minorization-Maximization (MM) and Gradient Descent (GD) Algorithms on Simulated Partial Digestions of a Linear Genome
A Dash Indicates That the Size Did Not Finish Due to Memory Constraints or Runtime Constraints.
Normalized Mean Absolute Error for 10 Sizes and 4 Uncertainty Magnitudes Using the Minorization-Maximization (MM) and Distribution Matching (DM) Algorithms on Simulated Partial Digestions of a Circular Genome
Pandurangan and Ramesh (2002) performed a LPDP recovery experiment using the restriction sites of the enzyme HindIII on the bacteriophage
For each distance d, they simulated relative uncertainty of order
Recovery Success by % Relative Error for Our Base Solver (MM), Our Partition Solver (PMM), and the LPDP Solver (Pandurangan and Ramesh, 2002)
Recovery Success by % Relative Error for Our Base Solver (MM), Our Partition Solver (PMM), and the LPDP Solver (Pandurangan and Ramesh, 2002)
LPDP, The labeled partial digest problem; MM, Minorization-Maximization; PPM, Partition Minorization-Maximization.
We tested the integer program of Theorem 1 using three objectives inspired by various distance models. The first uses the support function introduced in Eq. 2.4, where we use Alg. 5 to implement the separation oracle. The second is inspired by the relative error model of Pandurangan and Ramesh (2002), where we use the
Mean Runtime in Seconds across All Scenarios for Point Sets of Size n for Three Support Function Objectives: The Relative-Error Norm
, the Minorization-Maximization Support Function
, and the Partitioning Norm
Mean Runtime in Seconds across All Scenarios for Point Sets of Size n for Three Support Function Objectives: The Relative-Error Norm
Average Mean Absolute Error of Recovered Points for Partial Digestions across All Objective Functions
Average Mean Absolute Error of Recovered Points for Labeled Partial Digestions across All Objective Functions
The N
Footnotes
AUTHORS’ CONTRIBUTIONS
C.S.E.: Conceptualization, Methodology, and Writing—Original Draft. M.H.: Technical Discussion, Baseline Implementation, and Proofreading. M.F.: Technical Discussion and Proofreading. C.K.: Conceptualization, Technical Discussion, Writing—Review and Editing, Supervision, Project Administration, and Funding Acquisition.
AUTHOR DISCLOSURE STATEMENT
C.K. is a co-founder of Ocean Genomics, Inc.
FUNDING INFORMATION
This work was supported in part by the US National Science Foundation [DBI-1937540, III-2232121], the US National Institutes of Health [R01HG012470], and by the generosity of Eric and Wendy Schmidt by recommendation of the Schmidt Futures program. C.S.E. was partially supported by a fellowship from Carnegie Mellon’s Center for Machine Learning and Health.
