Abstract
Understanding the mutational history of tumor cells is a critical endeavor in unraveling the mechanisms that drive the onset and progression of cancer. Modeling tumor cell evolution with labeled trees motivates researchers to develop different measures to compare labeled trees. Although the Robinson–Foulds (RF) distance is widely used for comparing species trees, its applicability to labeled trees reveals certain limitations. This study introduces the k-RF dissimilarity measures, tailored to address the challenges of labeled tree comparison. The RF distance is succinctly expressed as n-RF in the space of labeled trees with n nodes. Like the RF distance, the k-RF is a pseudometric for multiset-labeled trees and becomes a metric in the space of 1-labeled trees. By setting k to a small value, the k-RF dissimilarity can capture analogous local regions in two labeled trees with different size or different labels.
INTRODUCTION
In the realm of evolutionary biology, trees serve as fundamental mathematical concepts, offering a versatile framework for modeling the evolution of various entities, including organisms, species, and genes. Beyond their application in understanding biological evolution, trees find practical utility in medical diagnosis within the health care domain. The diversity of tree models has given rise to the significant challenge of effectively comparing different trees to evaluate various inference methods. This challenge has spurred researchers to define robust measures within the space of targeted trees. For example, mutation/clonal trees are introduced to model tumor evolution. In this representation, nodes denote cellular populations and are labeled with the gene mutations present in those populations (Karpov et al., 2019; Schwartz and Schäffer, 2017).
The growth and metastasis of tumors varies from patient to patient. In addition, such variations are significant for cancer treatment. As a result, dissimilarity measures for mutation tree comparison have become a focus of recent research (DiNardo et al., 2020; Jahn et al., 2021; Karpov et al., 2019; Llabrés et al., 2021).
In earlier work on phylogenetic trees, various measures have been proposed to compare two phylogenetic trees. Some examples of such measures are Robinson–Foulds (RF) distance (Robinson and Foulds, 1981), nearest-neighbor interchange (NNI) (Li et al., 1996; Robinson, 1971), Quartet distance (Estabrook et al., 1985), and Path distance (Steel and Penny, 1993; Williams and Clifford, 1971). They are defined under the assumption that the involved trees share the same label set. Consequently, they may not be useful when applied to trees where all nodes are labeled, especially when using different label sets.
Related work on comparison of labeled trees
To overcome the constraints associated with the above-mentioned measures in the comparison of mutation trees, computational biologists have introduced new dissimilarity metrics for mutation trees. Some of these measures are Common Ancestor Set (CASet) distance (DiNardo et al., 2020), Distinctly Inherited Set Comparison (DISC) distance (DiNardo et al., 2020), and Multi-Labeled Tree Dissimilarity measure (Karpov et al., 2019). Although these distance measures enable efficient comparison of clonal trees, they are defined based on the assumption that mutations cannot occur more than once and mutations will not be lost in the course of tumor evolution. As a result, these metrics exhibit multiple limitations when applied to the comparison of trees used to model complex tumor evolution, wherein mutations may indeed occur multiple times and subsequently be lost.
Apart from the three measures discussed earlier, a few additional dissimilarity metrics have been introduced to facilitate the comparison of mutation trees, including Parent–Child distance (Govek et al., 2018) and Ancestor–Descendant distance (Govek et al., 2018). These measures are metric for “1-mutation” trees, in which nodes are each labeled by one distinct mutation.
There are also measures for mutation trees that are defined through generalization of popular measures that are used for phylogenetic trees. Here, researchers aim to extend the definition of an existing distance, which was mostly used to compare phylogenetic trees with mutation trees. For example, the generalized NNI (Jahn et al., 2021) is defined by some minor modifications of NNI. The other example is the Path distance (Govek et al., 2018). Although these measures are applicable to mutation trees, they are only well defined for mutation trees with the same label sets (Govek et al., 2018; Jahn et al., 2021).
The generalized RF (GRF) distance is another distance introduced recently (Llabrés et al., 2021; Llabrés et al., 2020). This measure is used not only to compare mutation trees or clonal trees but also enables the comparison of species trees and even phylogenetic networks. A useful property of GRF is that its value is significantly contributed by the intersection of clusters or clones of targeted trees. However, the intersection is not quantified in the RF distance, as one only checks whether two cluster or clones of the two involved trees are identical or not when the RF-distance between two trees is computed. As a result, the GRF has a better resolution than the RF distance (Llabrés et al., 2020).
There are some other generalizations of the RF distance, such as Bourque distance (Jahn et al., 2021). The measure is able to compare mutation trees with same or different label sets, and it has linear time complexity. However, like the above distances, it does not allow for multiple occurrences of mutations during the tumor history (Jahn et al., 2021). Other generalizations of the RF distance have also been proposed for gene trees (Briand et al., 2022; Briand et al., 2020).
The aforementioned dissimilarity measures do not apply to some evolutionary models, such as Dollo (Farris, 1977) and the Camin–Sokal model (Camin and Sokal, 1965). This is because mutations may get lost after they are gained in the Dollo model, and the same mutation may occur more than once during the tumor history in the Camin–Sokal model (Llabrés et al., 2020). As far as we know, the only measure introduced to address the problem is the Triplet-based Distance (Ciccolella et al., 2021). The distance allows to compare mutation trees in which nodes have nonempty subsets of mutations as their labels. In addition, it also allows multiple occurrences and losing of mutations during the tumor history (Ciccolella et al., 2021). Despite the applicability of the measure to the larger group of labeled trees, Triplet-based Distance does not apply to labeled trees in which multiple copies of a mutation is observed in the label of a single node.
Our contributions to tree comparison
In this study, we develop the k-RF dissimilarity measures designed for the comparison of labeled trees. They are first defined for 1-labeled trees (Section 3). Subsequently, we extend these measures to multiset-labeled trees (Section 5). We delve into the mathematical properties of the k-RF measures in Sections 4 and 5. In particular, k-RF is a metric for 1-labeled trees. We also assess the validity of the k-RF measures through comparisons with CASet, DISC, and GRF (Section 5), and the evaluation of their performance in the context of tree clustering (Section 6).
CONCEPTS AND NOTATIONS
A (directed) graph consists of a set of nodes and a set of (directed) edges. In graphs, each edge is a pair of distinct nodes. In directed graphs, each edge is a pair of ordered distinct nodes.
Let G be a (directed) graph. V(G) and E(G) are used to denote its node and edge set, respectively. If G is undirected, (u,v) will still be used to denote an edge between u and v with the understanding that (u,v) = (v,u). Let
The degree of v is defined as the number of edges incident to v. In addition, if G is directed, the indegree and outdegree of v are defined as the number of edges (x,y) such that y = v and x = v, respectively. The nodes of degree 1 are called the leaves in an undirected graph, whereas the nodes of indegree 1 and outdegree 0 are called the leaves in a directed graph. We use Leaf (G) to denote the leaf set for G. Nonleaf nodes are called internal nodes.
A path of length k from u to v consists of a sequence of nodes
If G is undirected,
Trees
A tree T is a graph in which there is a unique path between any two distinct nodes. A binary tree is a tree in which every internal node has degree 3. A line tree is a tree in which every internal node has degree 2. The number of leaves in a line tree is 2.
A directed tree is a directed graph that is a tree if we ignore the orientations of edges.
Rooted trees
A directed tree is called a rooted tree if it has a special root node from which the edges are directed away. In a rooted tree, indegree of each nonroot node is 1, which implies that there is exactly one path from its root to any other node.
Let T be a rooted tree,
A rooted tree is called a binary rooted tree if the root has indegree 0 and outdegree 1, and every other internal node has indegree 1 and outdegree 2.
A rooted tree is called a rooted line tree if each internal node has exactly one child. A rooted tree is called a rooted caterpillar tree if the set of children of each internal node contains at most one internal node.
Labeled trees
Suppose L is a set and
Phylogenetic and mutation trees
Let X be a finite taxon set. A phylogenetic tree (respectively, rooted phylogenetic tree) on X is a binary tree (respectively, binary rooted tree) in which only leaves are labeled by the elements of X, and two distinct leaves have different labels.
A mutation tree on a set M of mutations is a rooted tree in which nodes are labeled with nonempty subsets of M.
Dissimilarity measures for trees
Let
THE k-RF MEASURE FOR 1-LABELED TREES
In this section, we first recall the definition of the RF distance and then present k-RF dissimilarity measures for 1-labeled trees for arbitrary k.
The k-RF measure for 1-labeled unrooted trees
Suppose X is a finite set and T is a 1-labeled tree on X. Each
We further define:
The RF distance of two 1-labeled trees S and T is defined as:
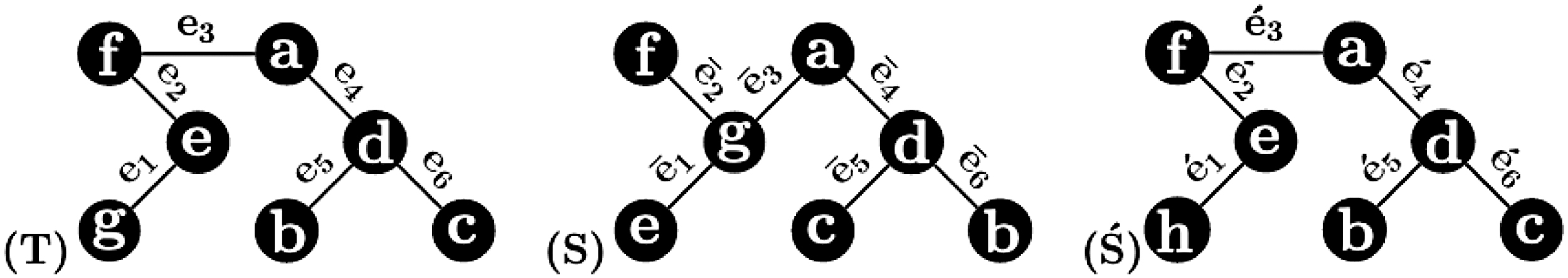
Three 1-labeled trees in Example 1 to illustrate that the Robinson–Foulds distance exhibits a heavy penalty against trees with different labels. Although T and
The above example illustrates that if two 1-labeled trees have different label sets, their local similarity are not captured by the RF distance. One of the widely used measures for the comparison of sets is the Jaccard distance. It is defined as a fraction whose numerator is the size of the symmetric difference of two sets and whose denominator is the size of their union. Two 1-labeled trees are identical if and only if they have the same set of edges with the understanding that each node is uniquely determined by its label. Hence, we aim to use
Let
Clearly,
respectively, and
respectively. We also have
Let k ≥ 0 be an integer and let T be a 1-labeled rooted tree. For a node
For each
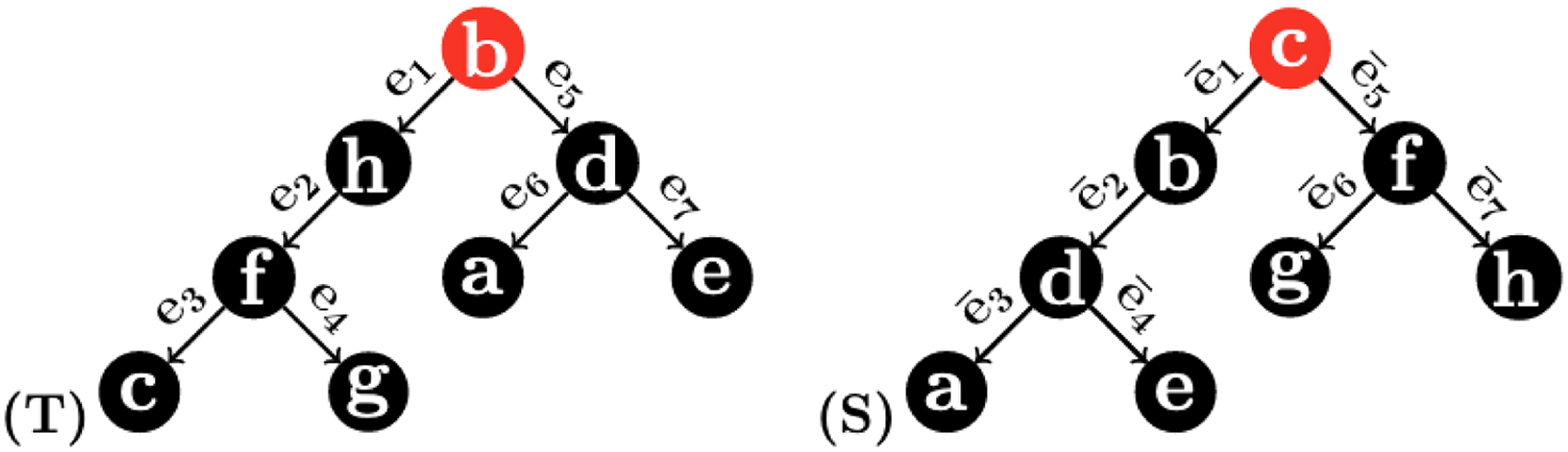
Two 1-labeled rooted trees used to illustrate the 1-RF in Example 3.
This implies that
To assess k-RF measures, we initially give their mathematical properties. Subsequently, we present experimental findings regarding their frequency distribution.
Mathematical properties
(a) For any k ≥ 1, (b) Assume that (c) Renaming each node with its label, we have (d) If
Proof. (a) Note that if k ≥ 1 and
(b) The second inequality follows from that
Let
If
If
In summary, we have proved that there are at least k + 1 edges (x,y) such that either
If
(c) Note that we may represent each node of a 1-labeled tree with its unique label. As a result,
(d) It follows from the definition of the k-RF.
Proof. Let k ≥ 0. The non-negativity and symmetry conditions are trivial. The triangle inequality
Proof. Let S and T be two 1-labeled rooted trees. By Remark 1, it is enough to show that S and T are identical if
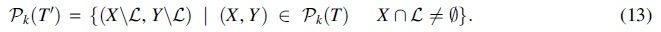
Proof. Since T is 1-labeled, we identify a node of T with its label in the following discussion. With this convention, for any subset S of nodes,
Let
If
For an edge
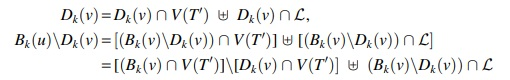
If
This has proved Equation (13).
Proof. Let S and T be two 1-labeled rooted trees. By Remark 1, it is enough to show that
Since
Assume S is uniquely determined by
For a leaf
For
Let V′ be the set of all nodes whose children are just leaves and
For the tree T′ obtained from T by the removal of the leaves of
This concludes the proof.
Proof. If k = 0, the statement follows from the same proof as for Proposition 2. Now, let S and T be two 1-labeled trees and k ≥ 1. By Lemma 1, it is enough to show that if
Proof. Since T is 1-labeled, we can identify a node of T with its label. In this way,
The statement is obvious in the case k = 0, since
Assume that all the
This implies that 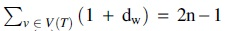 set operations. In total, we can compute all subsets
set operations. In total, we can compute all subsets 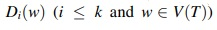 in at most
in at most
Proof. Since T is a 1-labeled rooted tree, we identify a node with its label. In this way, we just need to show that
Let r be the root of T. For any node
Then, we have that
Given the subsets
Proof. We first consider the rooted tree case. Let S and T be two 1-labeled rooted trees with n nodes. Without loss of generality, we may assume that S and T are labeled on the same set L, with
In the unrooted case, we first root the trees at a leaf. In this way, we can compute all the edge-induced pairs of label subsets in the derived rooted trees in
Distribution of k-RF scores
We examined the distribution of the k-RF dissimilarity scores for 1-labeled unrooted and rooted trees with the same label set and with different label sets.
The distribution of the frequency of the pairwise k-RF scores in the space of n-node 1-labeled unrooted and rooted trees for n from 4 to 7 are presented in Figures 3 to 6, respectively. For each n, it suffices to consider
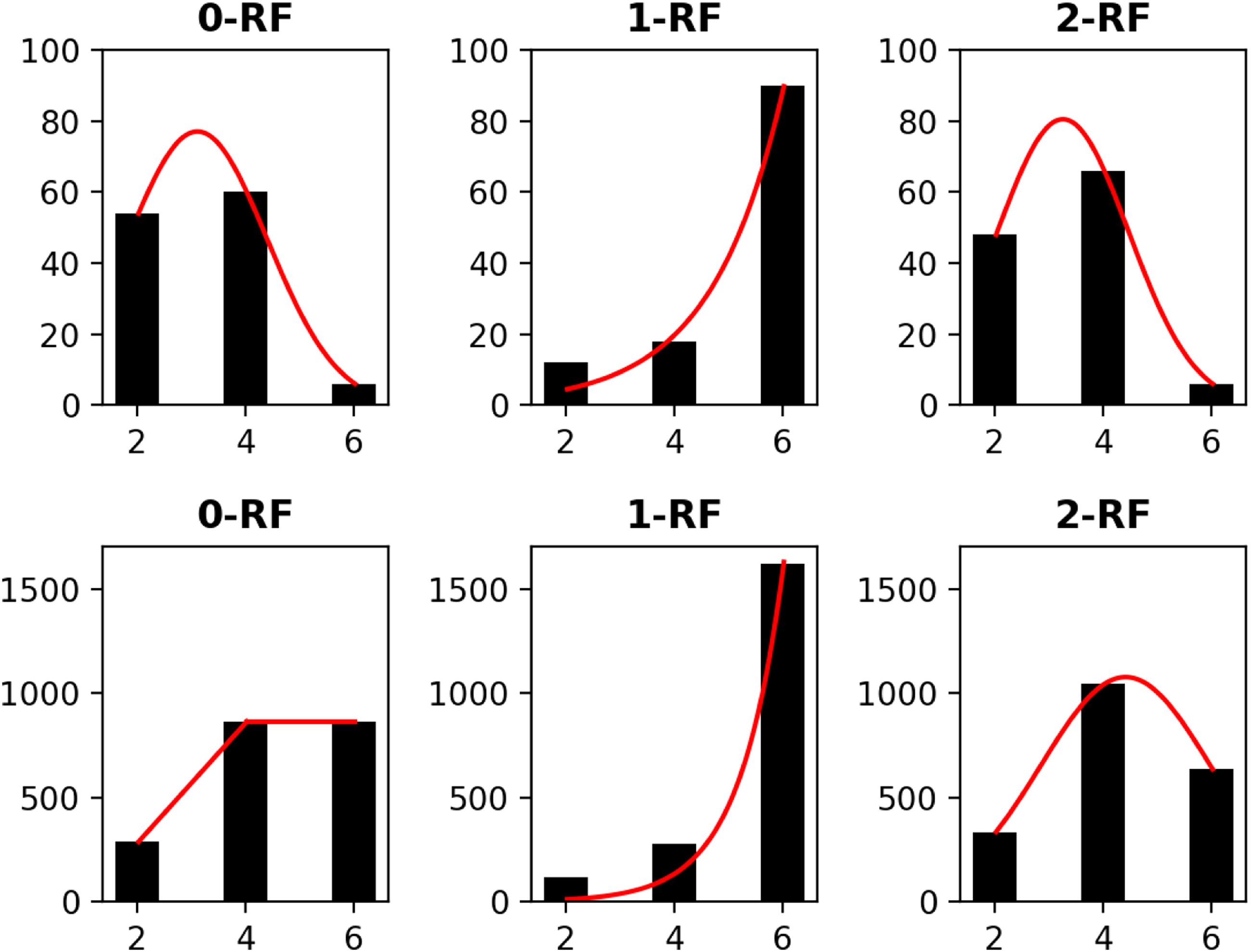
The frequency distributions of all pairwise k-RF scores in the space of 1-labeled unrooted (top row) and rooted (bottom row) 4-node trees for
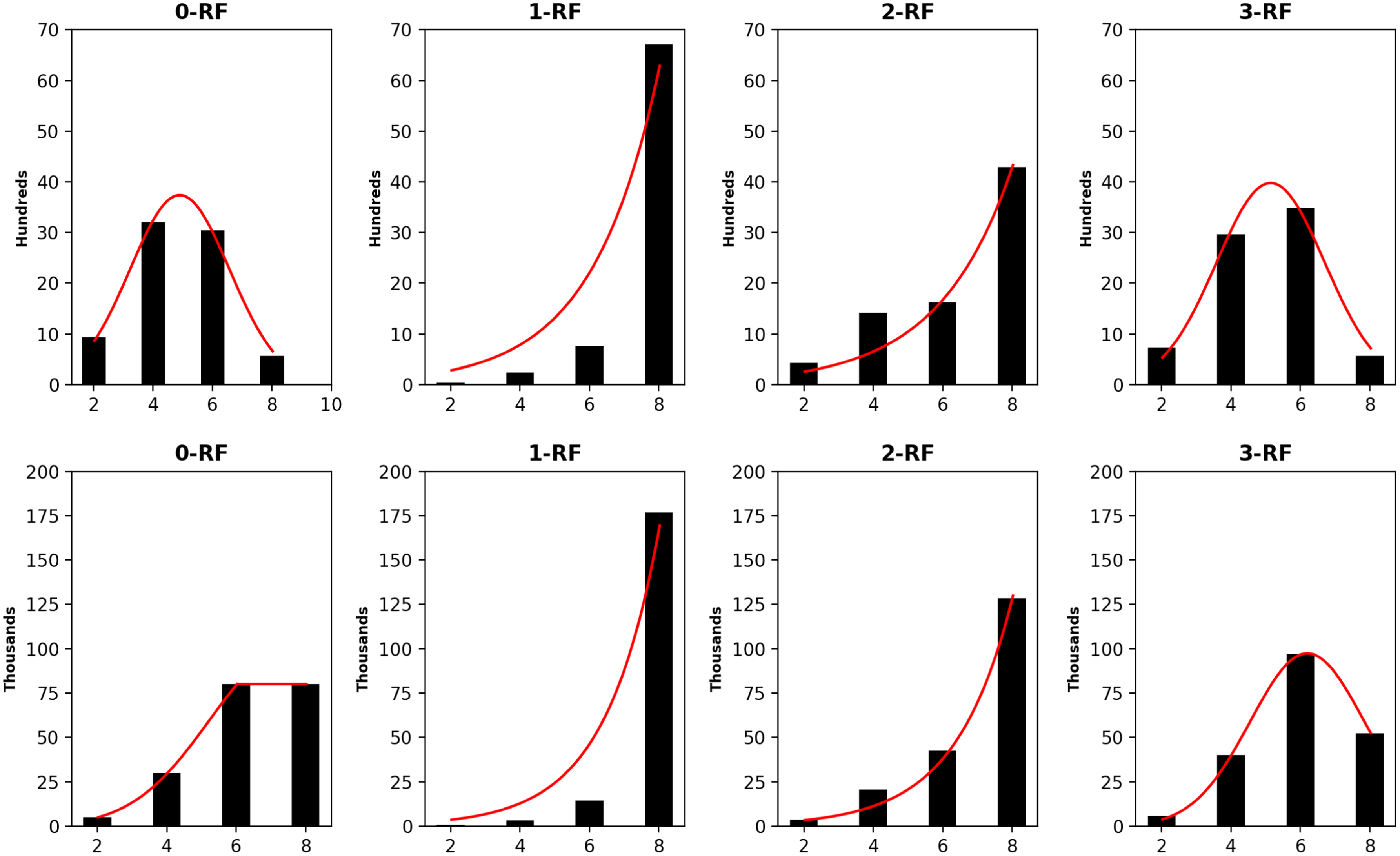
The frequency distributions of all pairwise k-RF scores in the space of 1-labeled unrooted (top row) and rooted (bottom row) 5-node trees, where k ≤ 3. In the bar-charts, the x-axis represents k-RF scores and the y-axis represents the number of tree pairs with a specific k-RF score.
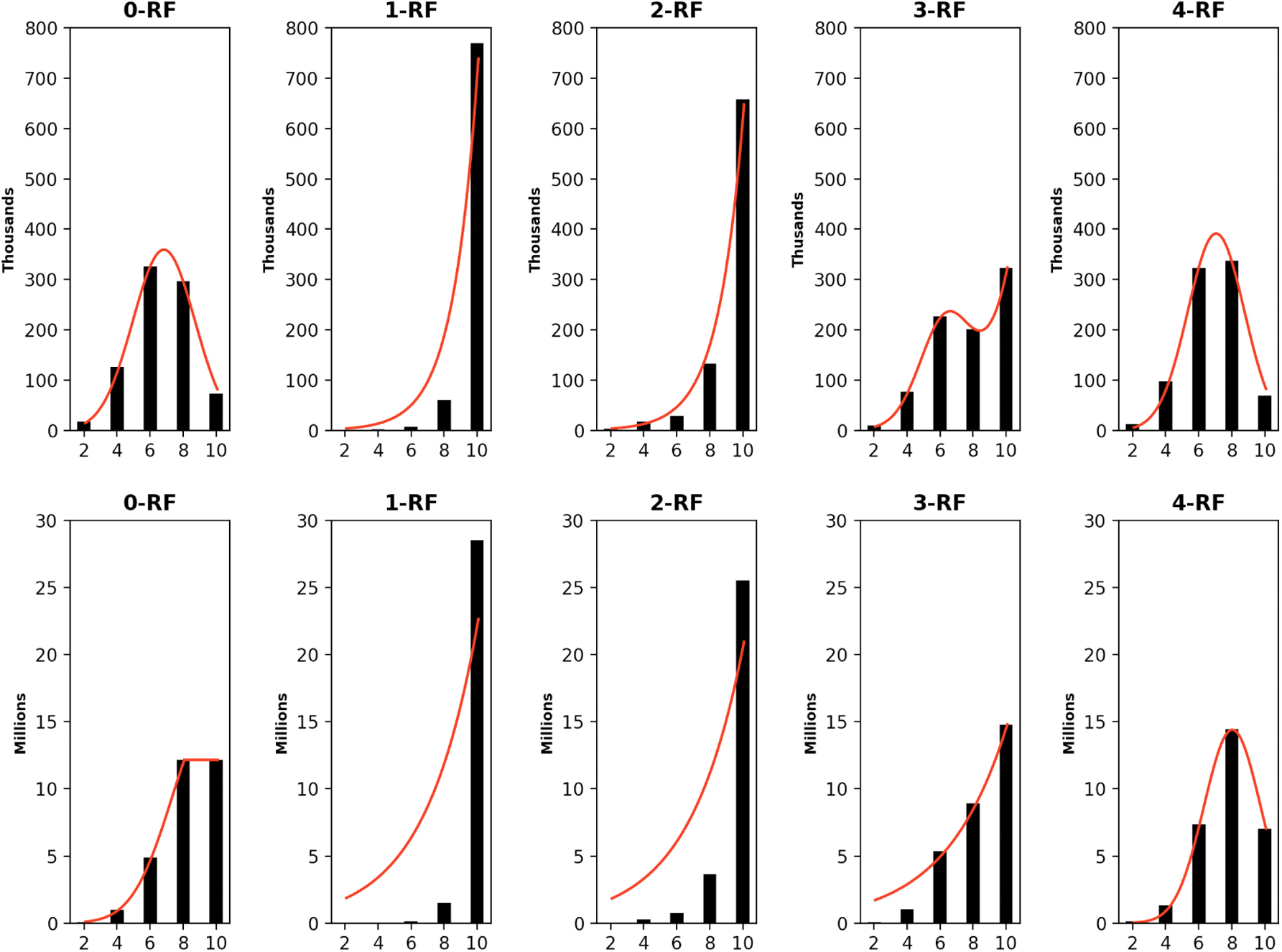
The frequency distributions of all pairwise k-RF scores in the space of 1-labeled unrooted (top row) and rooted (bottom row) 6-node trees for k ≤ 4. In each bar-chart, the x-axis represents k-RF scores and the y-axis represents the number of tree pairs whose k-RF equals a given score.
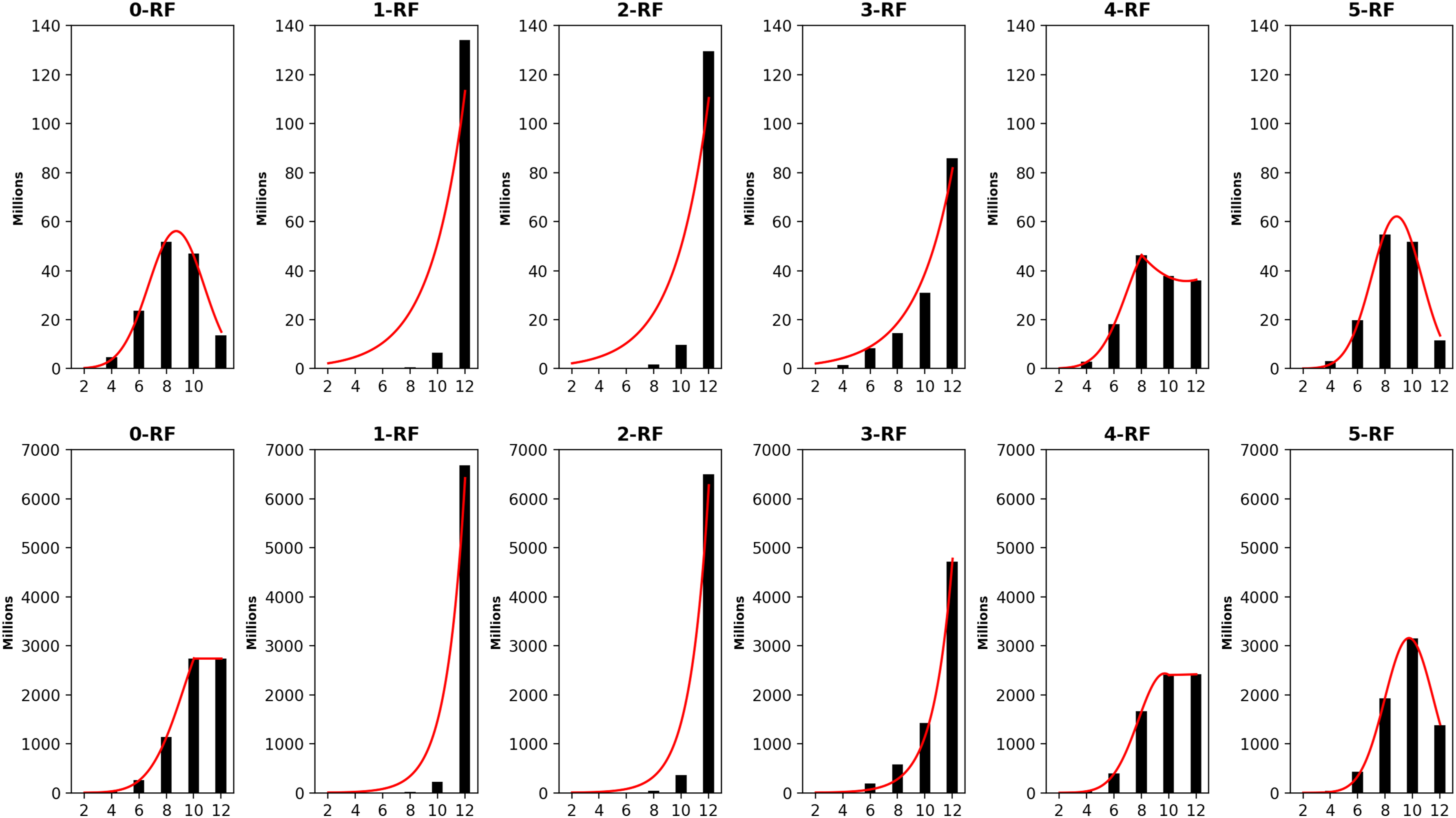
The frequency distributions of all pairwise k-RF scores in the space of 1-labeled unrooted (top row) and rooted (bottom row) 7-node trees for k ≤ 5. In each bar-chart, the x-axis represents k-RF scores and the y-axis represents the number of tree pairs whose k-RF equals a given score.
We examined 1,679,616 (respectively, 60,466,176) pairs of 6-node 1-labeled unrooted (respectively, rooted) trees such that the trees in each pair have c common labels, with
Number of Pairs of 1-Labeled 6-Node Unrooted (Top) and Rooted (Bottom) Trees That Have c Labels in Common and Have 1-Robinson–Foulds Score d for
RF, Robinson–Foulds.
In this section, we extend the measures introduced in Section 3 to multiset-labeled unrooted and rooted trees.
Multisets and their operations
A multiset is a collection of elements in which an element x can occur one or more times (Jűrgensen, 2020). The set of all distinct elements appearing in a multiset A is denoted by Supp (A). In this study, we simply represent A by the monomial
Let A and B be two multisets. We say A is a sub-multiset of B, denoted by
where
Let L be a set and

Let T be a multiset-labeled tree. Then, each edge
where
Two multiset-labeled trees used to show that different edges can give the same label multi-subset pair. Here, 
We use Equations (16), (3), and (4) to define the RF-distance for multiset-labeled trees by replacing
Let k ≥ 0. We use Equations (5), (6), and (7) to define the k-RF for multiset-labeled trees by replacing
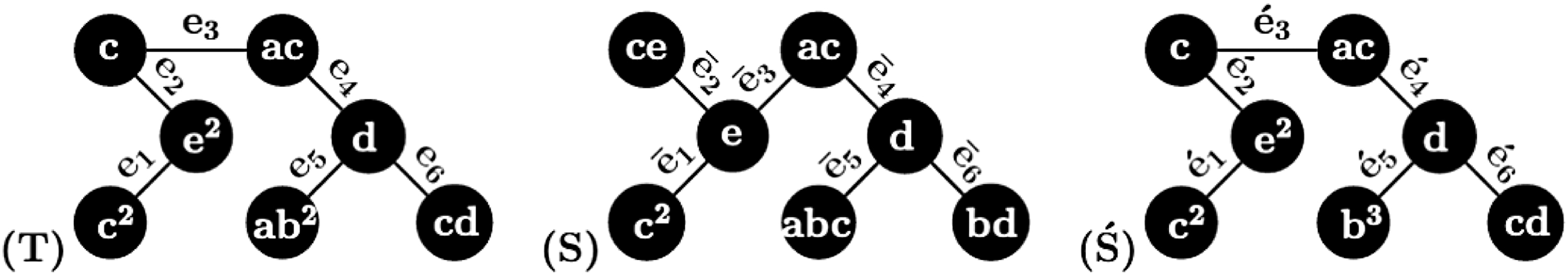
Three multiset-labeled trees in Example 4.
Edge-Induced Unordered Pairs of Multisets in the Three Trees in Figure 8 for
It is not hard to see that both
Let k ≥ 0 be an integer. We use Equations (10), (11), and (12) to define k-RF for multiset-labeled rooted trees by replacing
Proof. The non-negativity and symmetry conditions follow from the definition of the k-RF. The triangle inequality condition is proved as follows.
Let T1, T2, and T3 be three multiset-labeled trees. We need to show:
Note that
If
On the contrary, if
If
Finally, if
In addition, for each
Therefore, we have:
that is, the triangle inequality holds.
For multiset-labeled rooted trees, the proof is similar and hence omitted.
Two distinct multiset-labeled trees S and T satisfy that 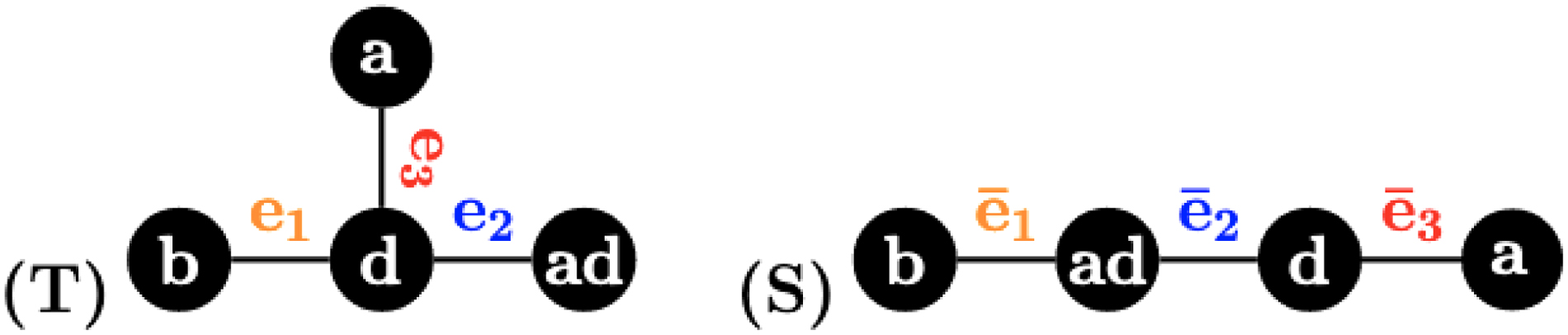
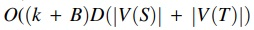 time, where B is the maximum multiplicity of a label appearing in
time, where B is the maximum multiplicity of a label appearing in
Proof. An algorithm for the 1-labeled case can be modified as follows for computing k-RF on multiset-labeled rooted and unrooted trees:
Represent each label multiset as a D-dimensional vector, in which the integer at position j is the multiplicity of the j-th label. Computing all edge-induced pairs in both trees takes Radix-sort all the edge-induced pairs for S and T in Compute the symmetric difference of the set of the edge-induced pairs in the two input trees in
In summary, one can compute 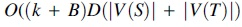 integer operations, as
integer operations, as
Let T and S be two 1-labeled rooted trees with the same label set X. Again, we identify the nodes with their labels in the two trees. For any two subset X′ and X″ of X, we use
Using the Pearson correlation (PC), we compared the k-RF with CASet
First, we conducted the correlation analysis in the space of mutation trees with the same label set. Using a method reported by Jahn et al. (2021), we generated a simulated dataset containing 5000 rooted trees in which the root was labeled with 0 and the other nodes were labeled by the disjoint subsets of
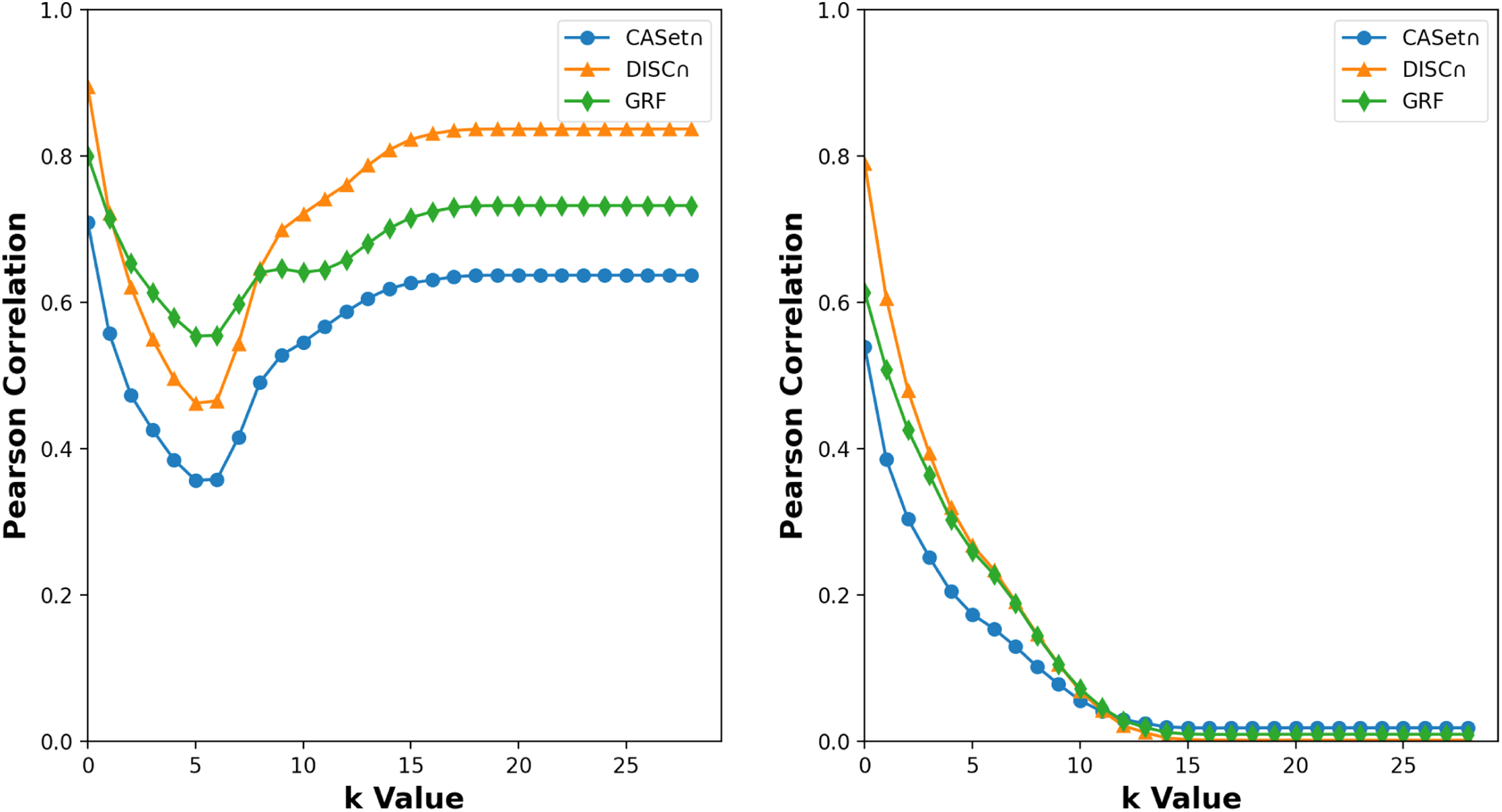
PC of the k-RF with CASet
Our results show that CASet
The GRF and k-RF are highly correlated for each k < 8.
The DISC
The 5-RF and 6-RF were less correlated to CASet
The PC between k-RF and CASet
Next, we performed PC analysis on trees characterized by distinct yet intersecting label sets. The dataset was created using the identical methodology, comprising a union of five sets of rooted trees, each encompassing 200 trees and sharing the same label set. Dissimilarity scores were calculated for each tree within the initial group and each tree in the remaining groups. Subsequently, we computed the PC between different dissimilarity values. Once more, all dissimilarity measures exhibited a positive correlation, although less pronounced than in the initial scenario [refer to Figure 10 (right)}. This observation could be because of that difference in label sets of two trees makes their k-RF score at least k + 1. However, the difference does not strongly contribute to the other distances because DISC
The right dotplot of Figure 10 shows that the k-RF and DISC
A test was designed to compare the k-RF, CASet
We generated randomly 5 tree families each containing 50 trees using the program reported by Jahn et al. (2021). The nodes were labeled by the subsets of a set of size 30 in the trees of each family. The label sets of any two different families were distinct in only one label. We imposed such restriction on the label sets as in each tree, distinct nodes were labeled by disjoint subsets; hence, each different label between the label sets of two trees induces d pairs that only belong to the tree with the label, where d is the degree of the node with the label. Therefore, the more different the label sets are, the more distinguishable the trees could be by the k-RF.
We computed the pairwise dissimilarity scores for all 250 trees in the 5 families via each measure; we then clustered the 250 trees into c clusters using the K-means clustering method, where c ranges from 2 to 57. The clustering results were assessed using the Silhouette score (Kaufman and Rousseeuw, 2009).
As Figure 11 shows, the correct number of tree families was not recognized by either of the CASet
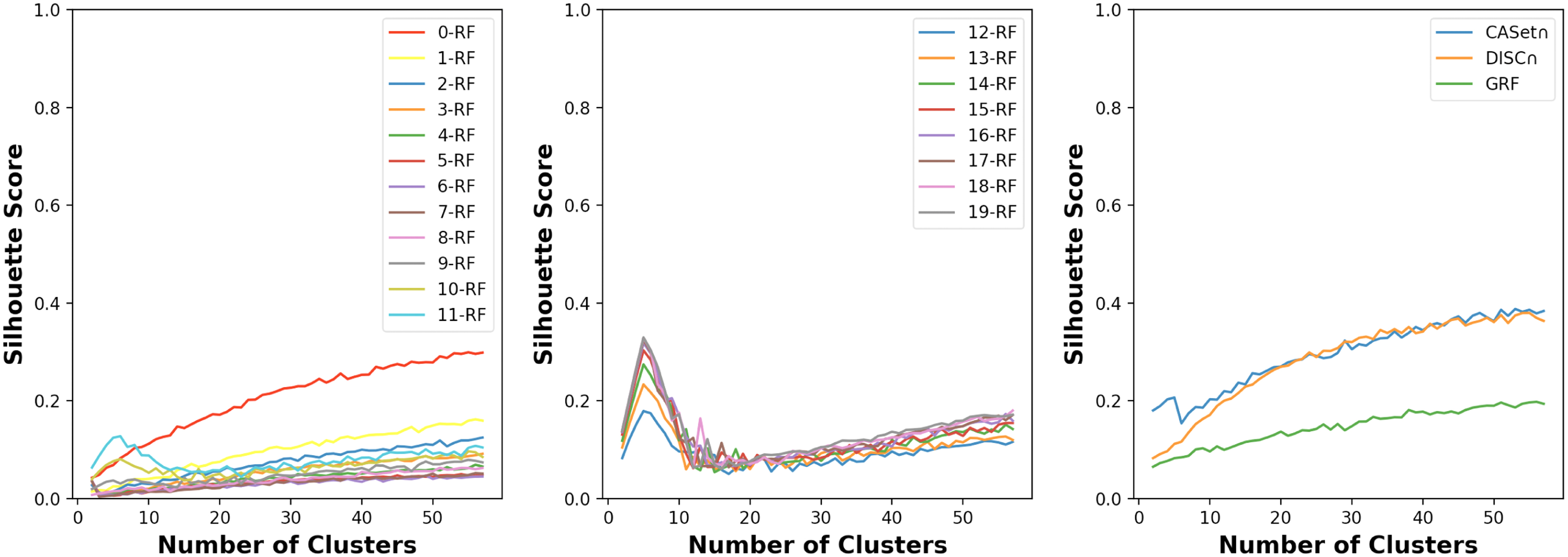
Silhouette scores of clustering 250 rooted trees with k-RF for
The development of an efficient and robust measure for the comparison of labeled trees is important. In this study, we have proposed a novel variant of dissimilarity metrics, namely the k-RF, tailored for labeled trees. The k-RF facilitates the analysis of local structures in labeled trees, accommodating nodes labeled with (not necessarily the same) multisets. Significantly, these metrics find practical applicability in mutation trees used in cancer research.
The RF distance is succinctly expressed as (n − 1)-RF within the space of labeled trees with n nodes. By setting k to a value smaller than n − 1, the k-RF metric can capture analogous local regions in two labeled trees. Of note, for every k, the k-RF is a pseudometric for multiset-labeled trees and becomes a metric in the space of 1-labeled trees. However, the distribution of pairwise k-RF scores in the space of 1-labeled unrooted (or rooted) trees conforms to a Poisson distribution specifically for k = n − 2, and unlikely have the same trend for other values of k ≥ 1.
We verified the k-RF measures through a comprehensive comparison with CASet, DISC (DiNardo et al., 2020) and GRF (Llabrés et al., 2021) on randomly labeled trees generated by a house-made program (Jahn et al., 2021). Our findings revealed a consistent positive correlation between k-RF and each of the other three measures for every value of k. Of note, the correlation values exhibited a tendency to be higher when the measures were applied to assess mutation trees with identical label sets. Furthermore, our study underscored the superior clustering capabilities of k-RF compared with the three mentioned measures.
We would like to emphasize that selecting an appropriate k-RF in practical applications lacks a universal rule of thumb, primarily owing to a shortage of experience in this domain. Perhaps a judicious approach involves choosing a suitable k-RF by carefully considering the topological similarity among the trees under consideration.
Future work includes how to apply the k-RF to designing tree inference algorithms like GraPhyC (Govek et al., 2018) and also how to infer the exact frequency distribution of the k-RF for each k ≥ 1. It is also interesting to investigate the generalization of RF-distance for clonal trees (Llabrés et al., 2020).
The code for computing the pairwise k-RF scores of a group of multiset-labeled trees can be downloaded from https://github.com/Elahe-khayatian/k-RF-measures.git.
Footnotes
ACKNOWLEDGMENTS
This work is an extended version of a conference article that was presented at the RECOMB-CG 2023, held at Istanbul, Turkey. The authors to thank the anonymous reviewer for providing helpful suggestions and comments to our first submission of the work.
AUTHOR DISCLOSURE STATEMENT
The authors declare they have no conflicting financial interests.
FUNDING INFORMATION
This research was partially supported by the Ministerio de Ciencia e Innovación (MCI), the Agencia Estatal de Investigación (AEI) and the European Regional Development Funds (ERDF) through project METACIRCLE PID2021-126114NB-C44, also supported by the European Regional Development Fund (FEDER), by the Agency for Management of University and Research Grants (AGAUR) through grant 2017-SGR-786 (ALBCOM), and by Singapore MOE Tier 1 grant R-146-000-318-114.
