Abstract
Minimizers and syncmers are sketching methods that sample representative k-mer seeds from a long string. The minimizer scheme guarantees a well-spread k-mer sketch (high coverage) while seeking to minimize the sketch size (low density). The syncmer scheme yields sketches that are more robust to base substitutions (high conservation) on random sequences, but do not have the coverage guarantee of minimizers. These sketching metrics are generally adversarial to one another, especially in the context of sketch optimization for a specific sequence, and thus are difficult to be simultaneously achieved. The parameterized syncmer scheme was recently introduced as a generalization of syncmers with more flexible sampling rules and empirically better coverage than the original syncmer variants. However, no approach exists to optimize parameterized syncmers. To address this shortcoming, we introduce a new scheme called masked minimizers that generalizes minimizers in manner analogous to how parameterized syncmers generalize syncmers and allows us to extend existing optimization techniques developed for minimizers. This results in a practical algorithm to optimize the masked minimizer scheme with respect to both density and conservation. We evaluate the optimization algorithm on various benchmark genomes and show that our algorithm finds sketches that are overall more compact, well-spread, and robust to substitutions than those found by previous methods. Our implementation is released at https://github.com/Kingsford-Group/maskedminimizer. This new technique will enable more efficient and robust genomic analyses in the many settings where minimizers and syncmers are used.
INTRODUCTION
Minimizers (Roberts et al., 2005; Schleimer et al., 2003) and syncmers (Edgar, 2021) are methods to deterministically sample k-mers from a sequence at approximately regular intervals. These sketching methods preserve sufficient information about the sequence identity in the set of sampled k-mers for comparison purposes, and they are widely used to reduce run-time and memory consumption in bioinformatics programs such as read mappers (Jain et al., 2022; Li, 2018), k-mer counters (Deorowicz et al., 2015; Erbert et al., 2017), high-throughput sequencing (Ben-Ari et al., 2021; Nyström-Persson et al., 2021), and genome assemblers (Ekim et al., 2021).
The k-mer sampling minimizer scheme is derived from a k-mer ordering. That is, the minimizer scheme selects the lowest-ranked k-mer (e.g., minimizer) from each window (substring with fixed length greater than k) in the input sequence. While the minimizer sampling rule is dependent on other k-mers in the same context window, the syncmer sampling rule trades off this window sampling mechanism for other useful properties, such as better robustness when sketching homologous sequences (Edgar, 2021; Shaw and Yu, 2021).
In particular, k-mer sampling syncmer schemes are derived from s-mer orderings, where
The parameterized syncmer scheme (Dutta et al., 2022) generalizes these syncmer variants using a subset parameter that encodes the selection rule. Specifically, given some subset
This flexible encoding of sampling rules offers a practical handle on the performance of syncmers, where subsets that correspond to neither open-syncmer (i.e.,
The quality of the k-mer sketches obtained by these schemes can be quantified by various metrics. Schleimer et al. (2003) uses the density metric (i.e., the sketch size proportionate to the sequence length) to estimate the degree of cost savings in downstream applications. More recently, Edgar (2021) proposes the conservation metric (i.e., the likelihood of sketched k-mers to be persistently sampled across homologous sequences) and argues that high conservation is preferable when comparing sequences that might have diverged due to mutations and/or sequencing error.
Shaw and Yu (2021) subsequently demonstrated that syncmers have better expected conservation than minimizers when both the input sequence and the ordering parameter are randomly drawn from uniform distributions. In this work, we consider the coverage metric that measures the spread of the sketch across the input sequence. We show that these three metrics are often adversarial to one another, and consequently propose a more holistic generalized sketch score (GSS) to evaluate sketching performance (Section 3).
Previous studies have established expected density guarantees for minimizers (Marçais et al., 2017; Schleimer et al., 2003) and syncmers (Edgar, 2021) with uniformly random input sequences. These results support the use of a fixed ordering for general sketching applications. However, when dealing with scenarios involving multiple query sequences being aligned against a single reference string, such as genome assembly, it is often more desirable to have an ordering that is optimally configured based on the reference.
For instance, Zheng et al. (2021) and Hoang et al. (2022a) have developed practical algorithms to optimize the k-mer ordering in the minimizer method. These studies have demonstrated that sequence-specific minimizer sketches generally achieve much lower density compared with non-optimized minimizer sketches.
Nonetheless, these optimization methods cannot be directly applied to configure low-density syncmer sketches, because they explicitly leverage the minimizer window sampling mechanism to construct their respective learning objectives. For example, the polar set method adopts a heuristic that selects as many k-mers as possible from the set of k-mers that are w (i.e., window size) bases apart (Zheng et al., 2021), whereas the
In addition, syncmers have no minimum density guarantee, unlike minimizers that derive this property from the window sampling mechanism. As such, optimizing the syncmer method for a specific sequence can potentially result in a vacuous sketch with zero coverage (Section 8.1). Finally, extending previous density optimization methods to account for the conservation metric is also challenging, as conservation and density are adversarial metrics (Section 3.4).
To address these challenges, we adapt the parameterized syncmer framework (Dutta et al., 2022) such that the pattern-aware sampling rules are applied in conjunction with the window sampling rule of minimizers. We call this adaptation masked minimizers. Specifically, given a subset v (or equivalently a binary mask variable in our formulation), the masked minimizer scheme selects all minimizers that are found at some offset position in v (with respect to the windows they minimizer).
Similar to the parameterized syncmer framework, the pattern-aware sampling rules give masked minimizers the ability to balance the trade-off between density, conservation, and coverage. However, our formulation differs from that of Dutta et al. (2022) since the selection of a masked minimizer depends on k-mers around it, whereas a parameterized syncmer is selected in a context-free manner. This distinction is identical to how minimizers and syncmers differ and allows us to leverage and extend density optimization algorithms developed for minimizers (Section 5).
In particular, we develop a sequence-specific optimization algorithm for masked minimizers that extends the
We show that the optimized masked minimizer sketch of various human and bacterial genomes consistently achieves better GSS than previous optimization approaches, such as
We also discover a specific class of complement mask patterns (i.e., masks that include most offset positions except one) that combines desirable properties from minimizers (i.e., high coverage) and open-syncmers (i.e., tolerance to low-complexity sequences).
In summary, our contributions include: (1) an adaptation of the parameterized syncmer method (Dutta et al., 2022) that generalizes minimizers, which we call masked minimizers, (2) a novel sketching metric that combines and reflects the trade-off among density, conservation, and coverage, which we call the GSS, and (3) a bi-level optimization algorithm for the masked minimizer scheme, which jointly selects the optimal mask and ordering with respect to the GSS metric.
Our sequence-specific sketching framework combines the strength of sequence-specific minimizers and parameterized syncmers to improve sketching performance in a holistic manner.
SUBSTRING SAMPLING SCHEMES
Notation
Let
Finally, given an arbitrary k-mer scoring function
Minimizers
The k-mer sampling minimizer scheme is characterized by a tuple of parameters
where
The k-mer sampling open-syncmer scheme (Edgar, 2021) is specified by a tuple
Finally, the qualifying offset position
Based on the syncmer concept, Dutta et al. (2022) introduces the parameterized syncmer scheme, which replaces t by a subset of qualifying offset positions
Setting
While sequence-specific optimization of minimizers with respect to the density metric has been well addressed (Ekim et al., 2020; Hoang et al., 2022a; Zheng et al., 2020), the same capability has not been developed for either open syncmers, parameterized syncmers, or for other metrics than density.
To overcome this challenge, our goal is to extend the minimizer method with pattern-based sampling rules similar to that of parameterized syncmers. This extension allows us to incorporate desirable properties from the syncmer family, yet fully retain access to density optimization algorithms developed for minimizers.
To this end, we introduce the masked minimizer scheme specified by the tuple
Density
Let
Let
When too few k-mers are selected, the sketch will not sufficiently cover the sequence and is therefore not useful in practice. Although the minimizer scheme ensures that every
For example, an open syncmer scheme with offset t can theoretically select an empty sketch if the lowest scoring s-mer is always found within the first
To formally quantify this property, we introduce the notion of w-coverage. The w-coverage metric computes the fraction of
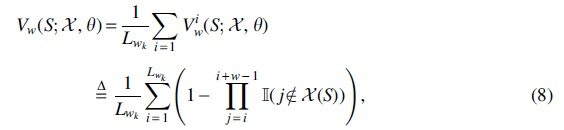
where
Generalized sketch score
It is straight-forward to see that:
The above derivation implies that conservation is upper-bounded by density for any arbitrary sketching scheme
Intuitively, the GSS metric encourages striking a balance between high conservation, low density, and high coverage. This is achieved by measuring the trade-off ratio between conservation/density, and normalizing this value by the w-coverage score of the sketch:
As a consequence of Eqs. (8) and (9), Gw is guaranteed to be in
This section provides an analysis of the change in performance of the masked minimizer scheme as v varies in the power set of
Proof. Let
Proof. By definition of density:
where the inequality follows directly from Proposition 1.
Proof. Let
where the inequality follows from Proposition 1, and the fact that the indicator variables take values in
where the inequality follows from Eq. (12) and linearity of expectation. Rearranging the above result concludes the proof.
These results imply that any masked minimizer scheme can improve conservation by adding more locations to its qualifying subset, or improve density by removing locations. However, as density upper-bounds conservation (Section 3.4), it is difficult to simultaneously improve both metrics by varying the mask, and hence it is necessary to formulate the optimization in terms of their trade-off ratio (e.g., the GSS metric), and with respect to the mask variable.
Given a choice of
Our greedy pruning step (outer loop) starts with the complete qualifying set
To address the inner loop optimization, we construct a differentiable loss function that extends the
Density optimization
Following Hoang et al. (2022a), we employ a pair of collaborating neural networks to model a hash function with low density on S:
The first neural network, 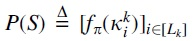 and implicitly defines
and implicitly defines
The second neural network, 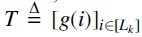 and implicitly defines a positional k-mer scoring function
and implicitly defines a positional k-mer scoring function
Intuitively, these networks, respectively, ensure the validity of a minimizer scheme and the ideal low density. A low-density sketch, thus, can be viewed as a consensus sketch
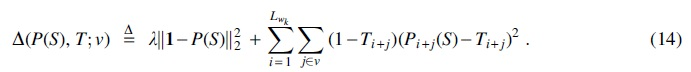
The first term
The second term differs from that of Hoang et al. (2022a) by the introduction of the inner summation over the offset positions in v, which is specific to the masked minimizer method. This sum represents an aggregation of weighted
The weight at each k-mer location is, therefore, jointly determined by its template value (i.e., how likely it is that this position will contribute to the sketch) and whether it can be found in the qualifying subset of some window (i.e., how relevant this is positioned to the current sampling rule).
Conservation optimization
The above objective only focuses on minimizing density. To account for conservation, our loss function
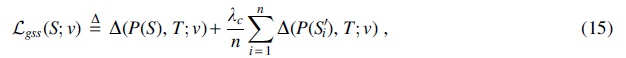
where
The first term of
Remark
Although we do not have direct results on the NP-hardness of optimizing GSS with respect to a target sequence, many problems adjacent to it, such as finding a sequence-specific universal hitting set (UHS; Orenstein et al., 2017) or the smallest size polar set (Zheng et al., 2021), have been shown to be NP-hard. A direct, brute-force strategy that performs the inner loop optimization on every possible mask would scale exponentially with the inner loop cost, and thus is prohibitively expensive.
Our pruning heuristic instead enables a worst case complexity of
EMPIRICAL RESULTS
In this section, we demonstrate the effectiveness of our optimization algorithm in learning high conservation, low density, and high coverage masked minimizer sketches. We also explore various ablation scenarios to confirm the practical usage of various specific masks (qualifying subsets).
Experimentation details
We compare the following baselines to construct the k-mer ordering for masked minimizers: (1) random ordering; (2) training with variants of our objective, including the
All experiments are conducted on the human chromosome 1 (labeled
The details of these sequences are given in Section 8.2. The gradient-based loss functions are computed per batch of sampled subsequences since it is not possible to fit the entire sequence on GPU memory. Our PyTorch implementation is available at https://github.com/Kingsford-Group/maskedminimizer. Other implementation details are given in Section 8.2.
Adversarial relationship of density and conservation
This experiment demonstrates that density and conservation are, indeed, conflicting objectives and confirms our argument in Section 3. Specifically, we train two masked minimizers using the minimizer mask
We note that the masked minimizer scheme with vo employs the window-based sampling mechanism, hence it does not recover exactly the open-syncmer scheme (or equivalently the parameterized syncmer scheme with mask vo), and only emulates its sampling pattern in the context of minimizers. We, respectively, denote these schemes by The vanilla The conservation loss given by the second term in Eq. (15),
Our loss function
Figure 1 plots the density, conservation, coverage, and GSS metrics on the sequence
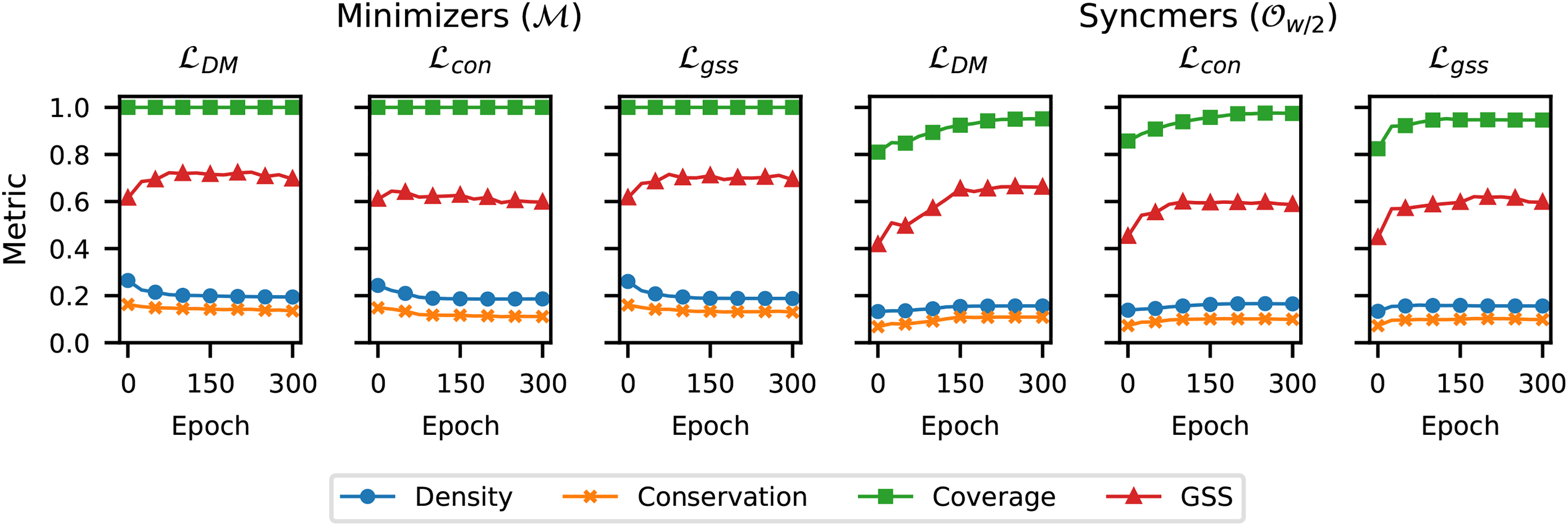
Comparing density, conservation, coverage, and GSS versus number of training epochs using different training losses and masks v on the bacterial genome
In addition, we observe that neither the density nor conservation metric reflects the drop in coverage when moving from the minimizer mask
This section demonstrates that our loss function
We denote the complement mask by
Figure 2 plots the GSS of the masked minimizers
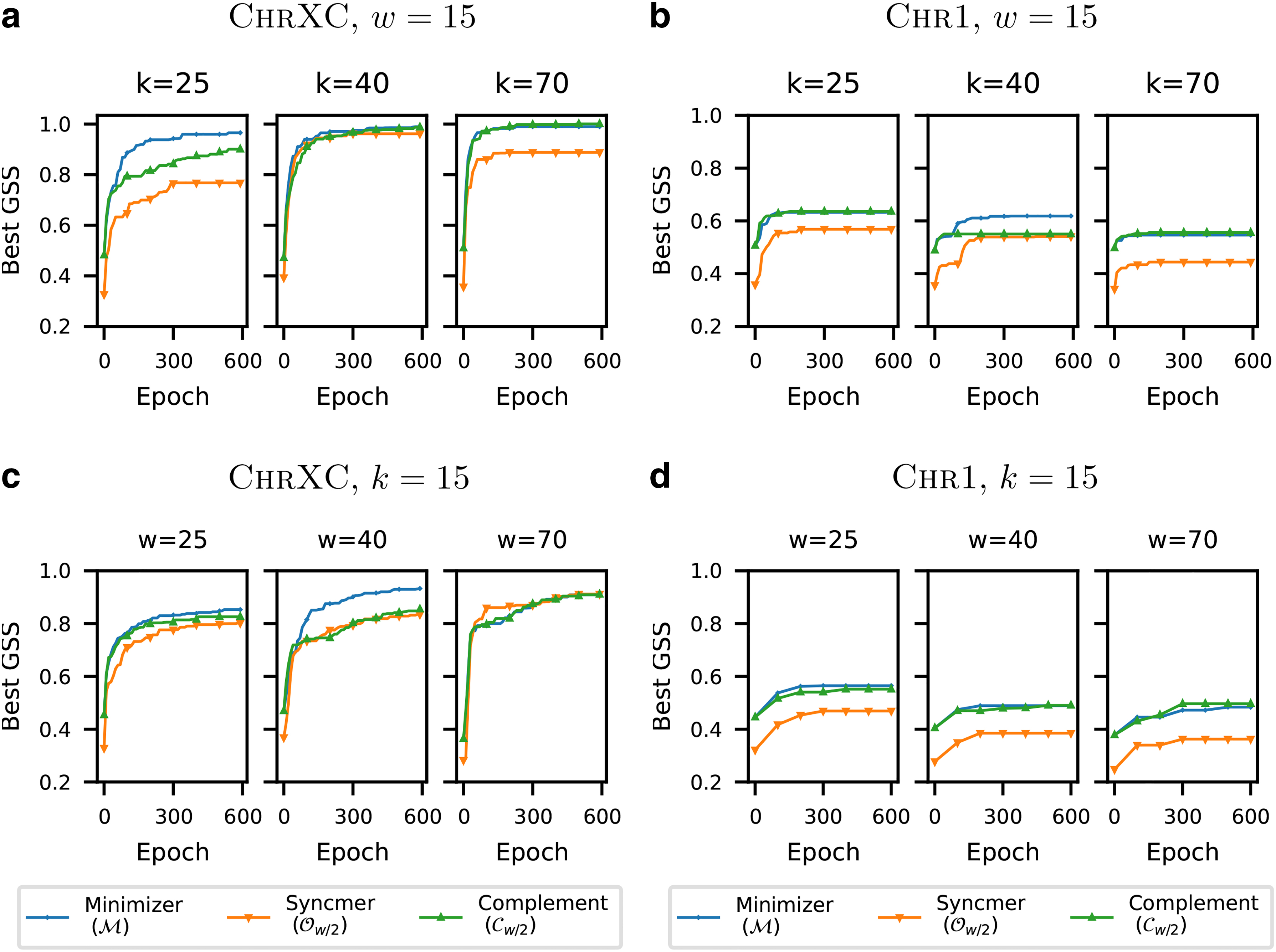
Comparing GSS of different masked minimizer variants versus number of training epochs on
We observe that the performance of the minimizer mask (
We further observe that both the minimizer mask (
In this section, we demonstrate the importance of optimizing for the mask variable. Specifically, we compare the GSS performance among methods that optimize for the k-mer ordering alone with respect to some fixed mask, and our method (Algorithm 1) that jointly optimizes both variables.
We, respectively, denote the optimized mask and its induced masked minimizer scheme by
For random ordering and the UHS-based methods, which only select the ordering once, the inner-loop optimization is simply replaced by evaluating the GSS metric with respect to the current v. We repeat our experiment for
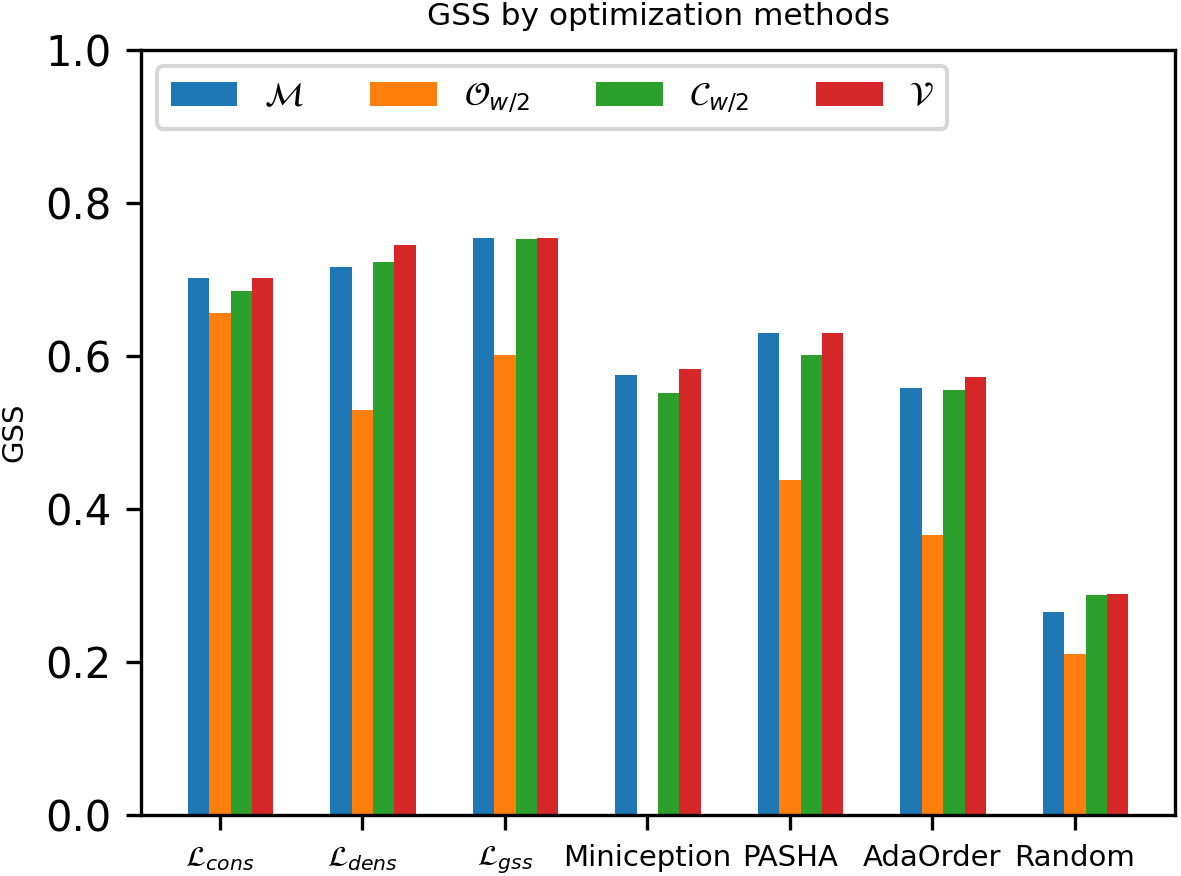
Comparing GSS of different masked minimizers using various optimization methods with
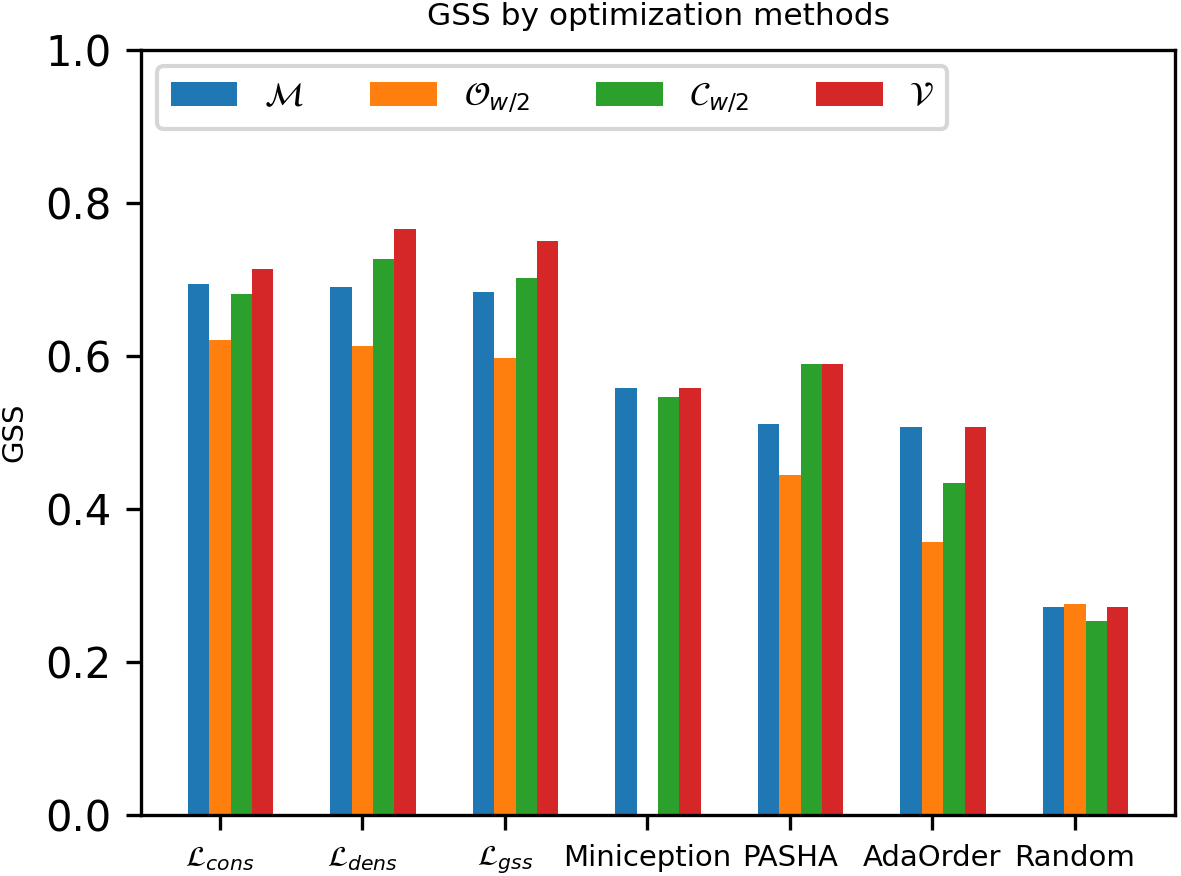
Comparing GSS of different masked minimizers using various optimization methods with
Among different masks of the same optimization method, we observe that the optimized mask
Interestingly, when combined with the
Among the best performing masks found by our optimization routine (Section 8.1.3), we observe that there is no fixed mask that consistently performs the best across all experiments. In addition, the maximum pruning depth observed is 3 (e.g., the algorithm terminates after 3 iterations of the outer loop because no possible GSS improvement can be found), which implies that dense masks are generally better for our benchmark sequences.
In contrast, the best performing masks reported by Dutta et al. (2022) are significantly sparser, such as
We visualize the distribution of GSS across different masks. Figure 5 (left) shows the scatter plot of all
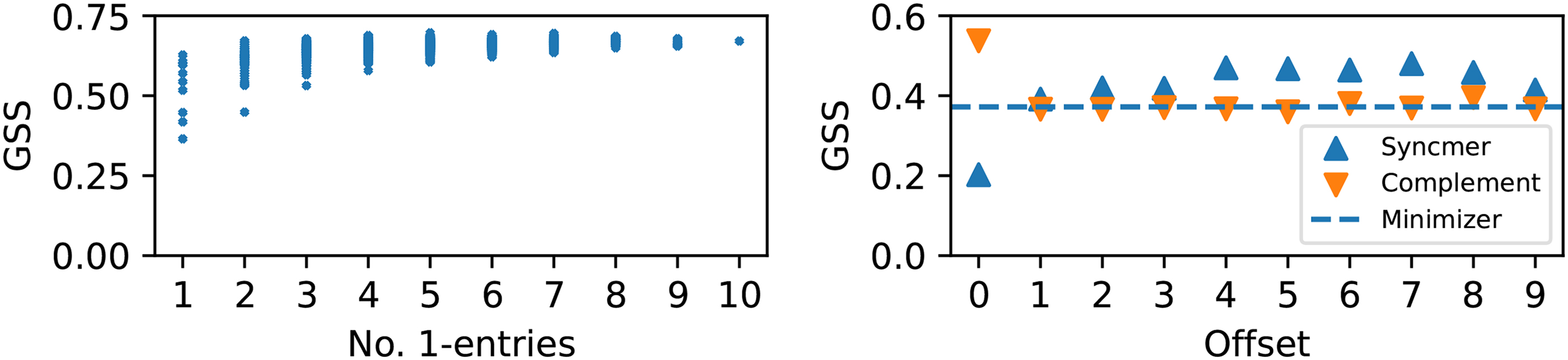
Left: GSS versus number of 1-entries of all masked minimizers trained on the bacterial genome
One advantage of open-syncmers with
In particular, because of the left-most tie breaking rule, every scheme whose mask contains the offset 0 (e.g., the minimizer mask, the open-syncmer mask with
CONCLUSION
We study the masked minimizer sketching scheme that applies the parameterized syncmer sampling rules (Dutta et al., 2022) to the window sampling mechanism of minimizers. We develop a bi-level optimization framework to design masked minimizers for a specific reference sequence. To account for the conflicting sketching metrics (e.g., density, conservation, and coverage), we propose a new sketching metric called GSS.
We show that our algorithm finds combinations of masks and k-mer orderings that induce masked minimizer schemes with better GSS than other sketch construction methods. We additionally introduce a special category of complement masks that combine desirable properties of minimizers and syncmers. We demonstrate the robustness of these masks in both the standard setting and sketching a homopolymer-rich sequence that is known to be a pitfall for the minimizer method.
This research opens up new directions for systematic construction of sequence sketches that improve genomic analysis. A current shortcoming of our method is the heuristic search for the mask variable. This challenging combinatorial problem will be an interesting avenue for future work.
Footnotes
ACKNOWLEDGMENTS
The authors thank all Kingsford group members for providing insightful discussion and proofreading the manuscript. An earlier version of this work was deposited in the bioRxiv preprint server.
AUTHORs' CONTRIBUTIONS
M.H. came up with the concept and methodology, implemented the software, conducted an empirical study, and wrote the original draft. G.M. and C.K. provided a discussion to formalize the methodology, helped design the empirical study, and assisted in writing the manuscript.
AUTHOR DISCLOSURE STATEMENT
C.K. is a co-founder of Ocean Genomics, Inc. G.M. is the VP of Software Engineering at Ocean Genomics, Inc.
FUNDING INFORMATION
This work was supported in part by the U.S. National Institutes of Health [R01HG012470], the U.S. National Science Foundation [DBI-1937540, III-2232121] and by the generosity of Eric and Wendy Schmidt by recommendation of the Schmidt Futures program.
