Abstract
Abstract
Phylogenetic stochastic mapping is a method for reconstructing the history of trait changes on a phylogenetic tree relating species/organism carrying the trait. State-of-the-art methods assume that the trait evolves according to a continuous-time Markov chain (CTMC) and works well for small state spaces. The computations slow down considerably for larger state spaces (e.g., space of codons), because current methodology relies on exponentiating CTMC infinitesimal rate matrices—an operation whose computational complexity grows as the size of the CTMC state space cubed. In this work, we introduce a new approach, based on a CTMC technique called uniformization, which does not use matrix exponentiation for phylogenetic stochastic mapping. Our method is based on a new Markov chain Monte Carlo (MCMC) algorithm that targets the distribution of trait histories conditional on the trait data observed at the tips of the tree. The computational complexity of our MCMC method grows as the size of the CTMC state space squared. Moreover, in contrast to competing matrix exponentiation methods, if the rate matrix is sparse, we can leverage this sparsity and increase the computational efficiency of our algorithm further. Using simulated data, we illustrate advantages of our MCMC algorithm and investigate how large the state space needs to be for our method to outperform matrix exponentiation approaches. We show that even on the moderately large state space of codons our MCMC method can be significantly faster than currently used matrix exponentiation methods.
1. Introduction
P
Stochastic mapping, initially developed by Nielsen (2002) and subsequently refined by others (Lartillot, 2006; Hobolth, 2008), assumes that discrete traits of interest evolve according to a continuous-time Markov chain (CTMC). Random sampling of evolutionary histories, conditional on the observed data, is accomplished by an algorithm akin to the forward filtering-backward sampling algorithm for hidden Markov models (HMMs) (Scott, 2002). However, since stochastic mapping operates in continuous time, all current stochastic mapping algorithms require computing CTMC transition probabilities via matrix exponentiation—a time consuming and potentially numerically unstable operation, when the CTMC state space grows large. de Koning et al. (2010) recognized the computational burden of the existing techniques and developed a faster, but approximate, stochastic mapping method based on time discretization. We propose an alternative, exact stochastic mapping algorithm that relies on recent developments in the continuous-time HMM literature.
Rao and Teh (2011) used a CTMC technique called uniformization to develop a method for sampling hidden trajectories in continuous time HMMs. The use of uniformization in this context is not new, but all previous methods produced independent samples of hidden trajectories with the help of matrix exponentiation—an operation with algorithmic complexity
As in the original method of Rao and Teh (2011), our new stochastic mapping method must pay a price for bypassing the matrix exponentiation step. The cost of the improved algorithmic complexity is the replacement of Monte Carlo in the state-of-the-art stochastic mapping with MCMC. Since Monte Carlo, if practical, is generally preferable to MCMC, it is not immediately clear that our new algorithm should be an improvement on the original method in all situations. We perform an extensive simulation study, comparing performance of our new MCMC method with a matrix exponentiation method for different sizes of the state space. We conclude that, after accounting for dependence of trait history samples, our new MCMC algorithm can outperform existing approaches even on only moderately large state spaces (s ∼ 30). We demonstrate additional computational efficiency of our algorithm when taking advantage of sparsity of the CTMC rate matrix. Since we suspect that our new method can speed up computations during studies of protein evolution, we examine in detail a standard GY94 codon substitution model (s = 61) (Goldman and Yang, 1994). We show that our new method can reduce computing times of state-of-the-art stochastic mapping by at least a factor of ten when working with this model. The last finding is important, because state-of-the-art statistical methods based on codon models often grind to a halt when applied to large datasets (Valle et al., 2014).
2. Methods
2.1. CTMC model of evolution
We start with a trait of interest, X(t), and a rooted phylogenetic tree with n tips and 2n − 2 branches. We assume that the phylogeny and its branch lengths,
A substitution history for a phylogenetic tree is the complete list of transition events (CTMC jumps), including the time of each event (location on the tree) and the type of the transition event (e.g., 2 → 1 transition). This state history can be encoded in a set of vectors, two vectors for each branch. Suppose branch i has ni transitions so the full state history for branch i can be described by a vector of state labels,
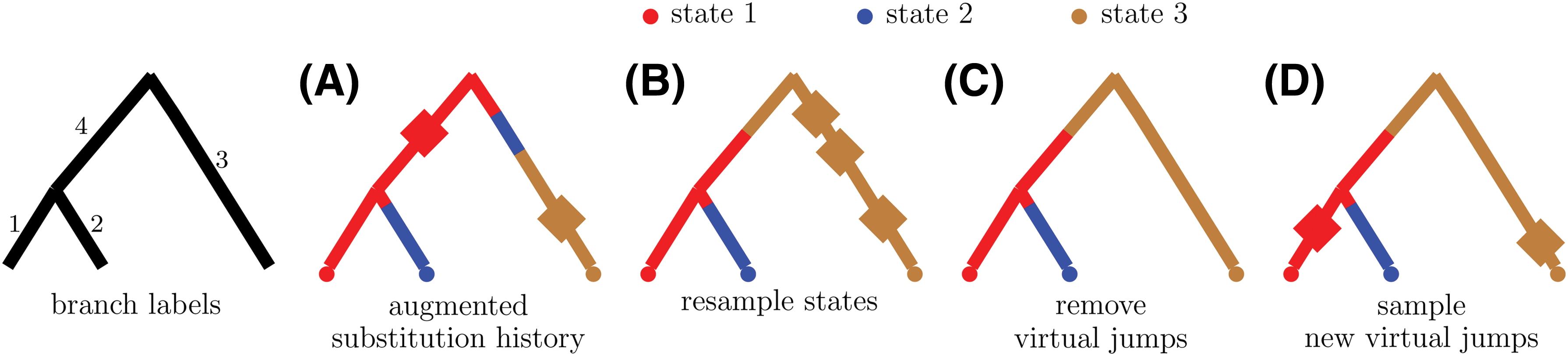
An example of applying the two Markov kernels of our MCMC sampler to an augmented substitution history. The diamonds represent virtual transitions.
The goal of stochastic mapping is to be able to compute properties of the distribution of the substitution history of a phylogenetic tree conditional on the observed states at the tips of the tree,
2.2. Nielsen's Monte Carlo sampler
Nielsen (2002) proposed the basic framework that state-of-the-art phylogenetic stochastic mapping currently uses. His approach samples directly from the conditional distribution,
2.3. CTMC uniformization
An alternative way to describe the CTMC model of evolution on a phylogenetic tree uses a homogenous Poisson process coupled with a discrete time Markov chain (DTMC) that is independent from the Poisson process. The intensity of the homogenous Poisson process, Ω, must be greater than the largest rate of leaving a state, max
k
|qkk|. The generative process that produces a substitution history on a phylogenetic tree first samples the total number of DTMC transitions over the tree, N, drawn from a Poisson distribution with mean equal to
Again, the uniformized generative process samples a state at the root of the tree and works down the tree sampling the state of each branch segment sequentially. Conditional on the previous/ancestral segment being in state k, we sample the current segment's state from a multinomial distribution with probabilities
An augmented substitution history of a phylogenetic tree encodes all the information in
2.4. New MCMC sampler
Equipped with notation describing the CTMC model of evolution and a companion uniformization process, we now turn our attention to making inference about a phylogenetic tree state history conditional on observed data. We investigate the situation where the tree topology is fixed, branch lengths are fixed, and the rate matrix parameters are all known and fixed. The goal is to construct an ergodic Markov chain on the state space of augmented substitution histories with the stationary distribution
Our MCMC sampler uses two Markov kernels to create a Markov chain whose stationary distribution is
2.4.1. Sampling states from
\documentclass{aastex}\usepackage{amsbsy}\usepackage{amsfonts}\usepackage{amssymb}\usepackage{bm}\usepackage{mathrsfs}\usepackage{pifont}\usepackage{stmaryrd}\usepackage{textcomp}\usepackage{portland, xspace}\usepackage{amsmath, amsxtra}\pagestyle{empty}\DeclareMathSizes{10}{9}{7}{6}\begin{document}
$$\textbf{\textit{p}} ( { \cal V} \mid { \cal W} , \textbf{\textit{y}} )$$
\end{document}
Our strategy for sampling states is to make a draw from the full conditional of internal node states and then to sample the states along each branch conditional on the branch's parent and child nodes. It is useful to remember that when conditioning on the number of virtual and real jumps, and locations of these jumps on the tree, our data-generating process becomes a DTMC with transition probability matrix
We start by using Felsenstein's algorithm to compute a partial likelihood matrix
where
After working our way up the tree we have the matrix of partial likelihoods,
Once we calculate the probability of the root being in each possible state we sample the state of the root from the multinomial distribution with probabilities we just computed. Next, we sample all nonroot, nontip nodes. Without loss of generality, let us consider node c connected to its parent node, node p, by branch i. Suppose node p's previously sampled state is h and the number of jumps on branch i is mi. The vector-containing observed tip states at the eventual descendants of node c is
Starting with the root we can now work our way down the tree sampling the states of each node from the multinomial distributions with probabilities we just described. Equation (1) may suggest that sampling internal nodes has algorithmic complexity
After sampling the states along each branch we have completed one cycle through the first Markov kernel by sampling from
2.4.2. Sampling virtual jumps from
\documentclass{aastex}\usepackage{amsbsy}\usepackage{amsfonts}\usepackage{amssymb}\usepackage{bm}\usepackage{mathrsfs}\usepackage{pifont}\usepackage{stmaryrd}\usepackage{textcomp}\usepackage{portland, xspace}\usepackage{amsmath, amsxtra}\pagestyle{empty}\DeclareMathSizes{10}{9}{7}{6}\begin{document}
$${ p} ( { \cal U} \mid { \cal S} , { \cal T} , \textbf{\textit{y}} )$$
\end{document}
After sampling the states on each branch,
Suppose there are m jumps along a branch including real transitions and virtual transitions. Let vd be the state of the chain after the dth transition and let
The density as written above has four parts, the probability of starting in state v0 = s0, the probability of m transition points, the density of the locations of m unordered points conditional on there being m points, and the probability of each transition in a discrete time Markov chain with transition matrix
The density of the augmented substitution history of one branch, p(
where
The full conditional density (3) is a density of an inhomogenous Poisson process with intensity r(t) = Ω + qX(t). This intensity is piecewise constant so we can add self-transition locations/times to a branch segment in state sa by drawing a realization of a homogenous Poisson process with rate
3. Assessing Computational Efficiency
3.1. Algorithm complexity
State-of-the-art stochastic mapping approaches rely on exponentiating CTMC rate matrices, requiring
3.2. Effective sample size
When comparing timing results of our MCMC approach and a matrix exponentiation approach, we need to account for the fact that our MCMC algorithm produces correlated substitution histories. One standard way to compare computational efficiency of MCMC algorithms is by reporting CPU time divided by effective sample size (ESS), where ESS is a measure of the autocorrelation in a stationary time series (Holmes and Held, 2006; Girolami and Calderhead, 2011). More formally, the ESS of a stationary time series of size N with stationary distribution ν is an integer Neff such that Neff independent realizations from ν have the same sample variance as the sample variance of the time series. The ESS of a stationary time series of size N is generally less than N and is equal to N if the time series consists of independent draws from ν.
In MCMC literature, ESSs are usually calculated for model parameters, latent variables, and the log-likelihood. Since we are fixing model parameters in this article, we monitor ESSs for our latent variables—augmented substitution history summaries—and log
3.3. Matrix exponentiation
In all our simulations we compare timing results of our new MCMC approach with another CTMC uniformization approach that relies on matrix exponentiation (Lartillot, 2006). For the matrix exponentiation approach we recalculate the partial likelihood matrix at each iteration, which involves re-exponentiating the rate matrix. We do so in order to learn how our MCMC method will compare to the matrix exponentiation method in situations where the parameters of the rate matrix are updated during a MCMC that targets the joint posterior of substitution histories and CTMC parameters (Lartillot, 2006; Rodrigue et al., 2008). Since matrix exponentiation is a potentially unstable operation (Moler and Van Loan, 1978), we do not repeat it at each iteration in our simulations. Instead, we precompute an eigen decomposition of the CTMC rate matrix once, cache this decomposition, and then use it to exponentiate
In one of our simulation studies, when we consider the effect of sparsity in the rate matrix, we depart from this matrix exponentiation regime. Instead of exponentiating the rate matrix at each iteration we exponentiate the rate matrix
3.4. Implementation
We have implemented our new MCMC approach in an R package phylomap, available online (Irvahn and Minin, 2014). The package also contains our implementation of the matrix exponentiation–based uniformization method of Lartillot (2006). We reused as much code as possible between these two stochastic mapping methods in order to minimize the impact of implementation on our time comparison results. We coded all computationally intensive parts in C++ with the help of the Rcpp package (Eddelbuettel and François, 2011). We used the RcppArmadillo package to perform sparse matrix calculations (Eddelbuettel and Sanderson, 2014).
4. Numerical Experiments
4.1. General setup
We started all of our simulations by creating a random tree with 50 or 100 tips using the diversitree R package (FitzJohn, 2012). For each simulation that required the construction of a rate matrix, we set the transition rates between all state pairs to be identical. We then scaled the rate matrix for each tree so that the number of expected CTMC transitions per tree was either 2 or 6. These two values were intended to mimic slow and fast rates of evolution. Six expected transitions in molecular evolution settings is usually considered unreasonably high but six transitions (or more) is reasonable in other settings like phylogeography. For example, investigations of Lemey et al. (2009) into the geographical spread of human influenza H5N1 found on the order of 40 CTMC transitions on their phylogenies. To obtain each set of trait data we simulated one full state history after creating a tree and a rate matrix. We used this full state history as the starting augmented substitution history for our MCMC algorithm.
To ensure our implementation of the matrix exponentiation approach properly sampled from
4.2. MCMC convergence
Although we have outlined a strategy for taking MCMC mixing into account via the normalization by ESS, we have not addressed possible problems with convergence of our MCMC. Examination of MCMC chains with an initial distribution different from the stationary distribution showed very rapid convergence to stationarity, as illustrated in Figure C-1 of the Supplementary Materials. Such rapid convergence is not surprising in light of the fact that we jointly update a large number of components in our MCMC state space without resorting to Metropolis-Hastings updates.
4.3. Effect of state space size
Our MCMC method scales more efficiently with the size of the state space than matrix exponentiation methods, so we were first interested in comparing running times of the two approaches as the size of the CTMC state space increased. In Figure 2, we show the amount of time it took the matrix exponentiation method to obtain 10,000 samples for different state space sizes, and we show the amount of time it took our MCMC method to obtain an ESS of 10,000 for different state space sizes. The size of the state space varied between 4 states and 60 states. The tuning parameter, Ω, was set to 0.2, ranging between 15 and 103 times larger than the largest rate of leaving a state.
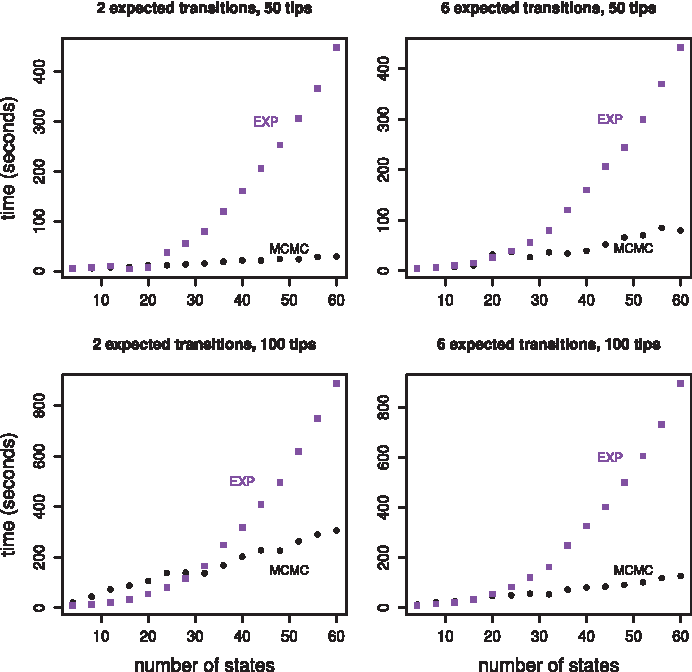
State space effect. All four plots show the amount of time required to obtain 10,000 effective samples as a function of the size of the state space for two methods, matrix exponentiation in purple squares and our MCMC sampler in black circles. The two plots in the top row show results for a randomly generated tree with 50 tips. The two plots in the bottom row show results for a randomly generated tree with 100 tips. The two plots in the left column show results for a rate matrix that was scaled to produce two expected transitions while the two plots in the right column show results for a rate matrix that was scaled to produce six expected transitions.
Figure 2 contains timing results for four different scenarios. We considered two different rates of evolution corresponding to two expected transitions per tree and six expected transitions per tree, and we considered two different tree tip counts, 50 and 100. The MCMC approach started to run faster than the matrix exponentiation approach when the size of the state space entered the 25 to 35 state range. At 60 states the MCMC approach was clearly faster in all four scenarios. For the senario involving 100 tips, 2 expected transitions, and 60 states, the MCMC method was almost three times faster than the matrix exponentiation approach. For the scenario involving 50 tips, 2 expected transtions, and 60 states, the MCMC method was about 15 times faster than the matrix exponentiation approach.
Our MCMC approach scales well beyond state spaces of size 60 though matrix exponentiation does not. Timing results for our MCMC approach at larger state space sizes can be found in Figure D-1 of the Supplementary Materials. Matrix exponentiation-based stochastic mapping was prohibitively slow on state spaces reported in Figure D-1.
4.4. Effect of the dominating Poisson process rate
Our tuning parameter, the dominating Poisson process rate Ω, balances speed against mixing for our MCMC approach. The larger Ω is the slower the MCMC runs and the better it mixes. The optimal value for Ω depends on the CTMC state space and on the entries of the CTMC rate matrix. In our experience, it is not difficult to find a reasonable value for Ω for a fixed tree and a fixed rate matrix by trying different Ω values. We show the results of this exploration in Figure 3 for two different values of the state space size, 4 and 60, and for two different trees, with 50 and 100 tips.
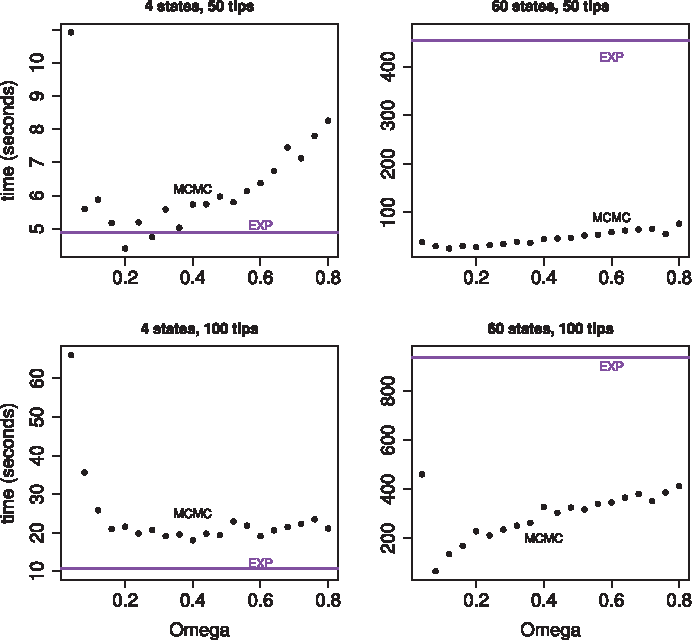
Time to obtain 10,000 effective samples as a function of the dominating Poisson process rate, Ω. All four plots show results of our MCMC sampler in black. Timing results for the matrix exponentiation method are represented by a purple horizontal line because the matrix exponentiation result does not vary as a function of Ω. The two plots in the top row show results for a randomly generated tree with 50 tips. The two plots in the bottom row show results for a randomly generated tree with 100 tips. The rate matrix for the plots in the left column had four states. The rate matrix for the plots in the right column had 60 states.
The top left plot in Figure 3 shows the balance between speed and mixing most clearly. The optimal value for Ω appears to be around 0.2 for 4 states and 50 tips. Our MCMC approach is clearly faster than the matrix exponentiation approach for a wide range of Ω values when the size of the state space is 60. When the size of the state space is 4 the matrix exponentiation approach can be faster, which is not surprising given the small size of the state space. Matrix exponentiation is about two times faster than our MCMC approach for the 100 tip tree with four states. Our MCMC approach can yield comparable speeds to the matrix exponentiation approach for the 50 tip tree with four states.
4.5. Effect of sparsity
Unlike matrix exponentiation methods, our new MCMC sampler is able to take advantage of sparsity in the CTMC rate matrix. There are three steps in our algorithm that can take advantage of sparsity: computing the partial likelihood matrix, sampling internal node states, and resampling branch states. In all three situations we need to multiply
Speed increases due to sparsity depend on the size of the state space and the degree of sparsity in the probability transition matrix,
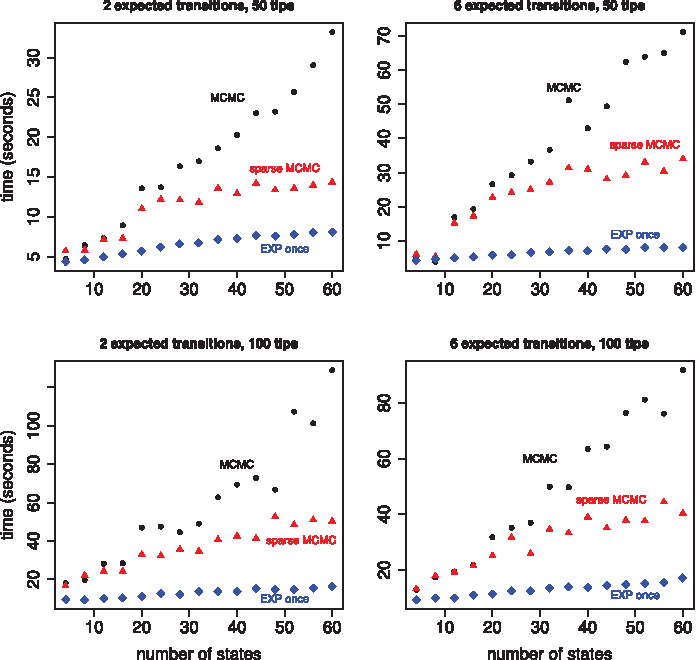
Time to obtain 10,000 effective samples as a function of the size of the state space. All four plots show results for three different implementations, our MCMC sampler in black, a sparse version of our MCMC sampler in red, and a matrix exponentiation approach that only exponentiates the rate matrix once per branch in blue. The rate matrix is tridiagonal and scaled to produce two expected transitions per tree (in the left column) or six expected transitions per tree (in the right column). The two plots in the top row show results for a randomly generated tree with 50 tips. The two plots in the bottom row show results for a randomly generated tree with 100 tips. The dominating Poisson process rate, Ω, is 0.2.
For a state space of size 60, the sparse implementation is about two times faster than the nonsparse implementation. Exponentiating the rate matrix once was always faster than the sparse implementation, sometimes by a factor of four. We used uniformization to sample substitution histories for individual branches within the matrix exponentiation algorithm. This portion of the algorithm can take advantage of sparsity, but there was not a large overall difference in run times between the sparse and nonsparse implementations.
4.6. Models of protein evolution
We now turn to the investigation of efficiency of our new phylogenetic stochastic mapping in the context of modeling protein evolution. Evolution of protein coding sequences can be modeled on the following state spaces: state space of 4 DNA bases/nucleotides, state space of 20 amino acids, and state space of 61 codons—nucleotide triplets—excluding the three stop codons. The codon state space is the most computationally demanding of the three, causing existing phylogenetic mapping approaches to slow down considerably. The increased complexity that comes from modeling protein evolution at the codon level enables investigations into selective pressures and makes efficient use of the phylogenetic information for phylogeny reconstruction (Ren et al., 2005).
In our numerical experiments, we use the Goldman-Yang-94 (GY94) model—a popular codon substitution model proposed by Goldman and Yang (1994), where the rate of substitution between codons depends on whether the substitution is synonymous (the codon codes for the same amino acid before and after the substitution) or nonsynonymous and whether the change is a transition (A ↔ G, C ↔ T) or a transversion. The rate matrix is parameterized by a synonymous/nonsynonymous rate ratio, ω, a transition/transversion ratio, κ, and a stationary distribution of the CTMC,
The diagonal rates are determined by the fact that the rows of
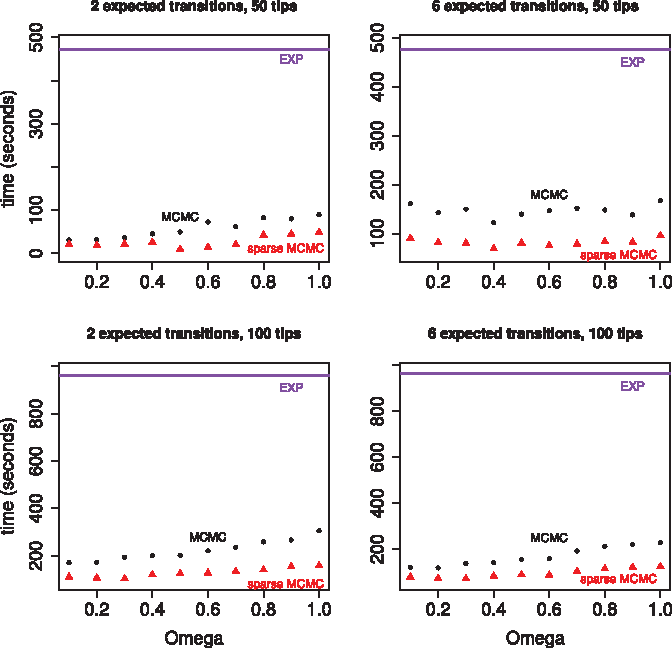
Time to obtain 10,000 effective samples as a function of the dominating Poisson process rate, Ω, for the GY94 codon rate matrix. All four plots show results for three different implementations: our MCMC sampler in black, a sparse version of our MCMC sampler in red, and a matrix exponentiation approach in purple. The GY94 rate matrix was scaled to produce 2 expected transitions per tree (in the left column) or 6 expected transitions per tree (in the right column). The two plots in the top row show results for a randomly generated tree with 50 tips. The two plots in the bottom row show results for a randomly generated tree with 100 tips.
Encouraged by the computational advantage of our method on the codon state space, we also compared our new algorithm and the matrix exponentiation method on the amino acid state space. We used an amino acid substitution model called JTT, proposed by Jones et al. (1992). The results can be found in Figure B-1 of the Supplementary Material. We found that our MCMC approach is competitive even on the amino acid state space, but does not clearly outperform the matrix exponentiation method. This finding is not surprising in light of the fact that the size of the amino acid state space is three times smaller than the size of the codon state space.
5. Discussion
We have extended the work of Rao and Teh (2011) on continuous time HMMs to phylogenetic stochastic mapping. Our new method avoids matrix exponentiation, an operation on which all current state-of-the-art methods rely. There are two advantages to avoiding matrix exponentiation: 1) matrix exponentiation is computationally expensive for large CTMC state spaces and 2) matrix exponentiation can be numerically unstable. In this article we concentrated on the former advantage, because it is easier to quantify. However, it should be noted that numerical stability of matrix exponentiation is an obstacle faced by all phylogenetic inference methods. Currently, the most popular approach is to employ a reversible CTMC model, whose infinitesimal generator is similar to a symmetric matrix and therefore can be robustly exponentiated via eigendecomposition (Schabauer et al., 2012). Researchers typically shy away from nonreversible CTMC models, to a large extent, because of instability of the matrix exponentiation of these models' infinitesimal generators (Lemey et al., 2009). In our new approach to phylogenetic stochastic mapping, we do not rely on properties of reversible CTMCs, making our method equally attractive for reversible and nonreversible models of evolution.
We believe our new method will be most useful when integrated into a larger MCMC targeting a joint distribution of phylogenetic tree topology, branch lengths, and substitution model parameters. Our optimism stems from the fact that stochastic mapping has already been successfully used in this manner in the context of complex models of protein evolution (Lartillot, 2006; Rodrigue et al., 2008). These authors alternate between using stochastic mapping to impute unobserved substitution histories and update model parameters conditional on these histories. We plan to incorporate our new MCMC algorithm into a conjugate Gibbs framework of Lartillot (2006) and Rodrigue et al. (2008). Since such an MCMC algorithm will operate on the state space of augmented substitution histories and model parameters, replacing Monte Carlo with MCMC in phylogenetic stochastic mapping may have very little impact on the overall MCMC mixing and convergence. A careful study of properties of this new MCMC will be needed to justify this claim.
The computational advances made in Lartillot (2006) and Rodrigue et al. (2008) are examples of considerable research activity aimed at speeding up statistical inference under complex models of protein evolution, prompted by the emergence of large amounts of sequence data (Lartillot et al., 2013; Valle et al., 2014). Challenges encountered in these applications also appear in statistical applications of many other models of evolution that operate on large state spaces: models of microsatellite evolution (Wu and Drummond, 2011), models of gene family size evolution (Spencer et al., 2006), phylogeography models (Lemey et al., 2009), and covarion models (Penny et al., 2001; Galtier, 2001). Our new phylogenetic stochastic mapping without matrix exponentiation should be a boon for researchers using these models and should enable new analyses that, until now, were too computationally intensive to be attempted.
Footnotes
Acknowledgments
We thank Jeff Thorne and Alex Griffing for helpful discussions and for feedback on an earlier version of this manuscript, and Jane Lange for pointing us to the work of Rao and Teh (2011). We thank Erick Matsen, Nicolas Lartillot, and Michael Landis for the comments and suggestions they made on the “phylobabble” discussion board. V.N.M. was supported in part by the National Science Foundation grant DMS-0856099 and by the National Institute of Health grant R01-AI107034. J.I. was supported in part by the UW NIGMS-sponsored Statistical Genetics Training grant NIGMS T32GM081062.
Author Disclosure Statement
No competing financial interests exist.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
