Abstract
Abstract
Many estimation problems in bioinformatics are formulated as point estimation problems in a high-dimensional discrete space. In general, it is difficult to design reliable estimators for this type of problem, because the number of possible solutions is immense, which leads to an extremely low probability for every solution—even for the one with the highest probability. Therefore, maximum score and maximum likelihood estimators do not work well in this situation although they are widely employed in a number of applications. Maximizing expected accuracy (MEA) estimation, in which accuracy measures of the target problem and the entire distribution of solutions are considered, is a more successful approach. In this review, we provide an extensive discussion of algorithms and software based on MEA. We describe how a number of algorithms used in previous studies can be classified from the viewpoint of MEA. We believe that this review will be useful not only for users wishing to utilize software to solve the estimation problems appearing in this article, but also for developers wishing to design algorithms on the basis of MEA.
1. Introduction
Problem 1 (Discrete-Space Point Estimation Problem [DSPEP])
Given data D and a discrete space Y correlated to D, find a point y in Y.
In this review, Y is called a predictive (solution) space, and it contains all the possible solutions (for the data D) of the target problem. For example, prediction of the secondary structure of an RNA sequence x is formulated as Problem 1 where
To solve this estimation problem, a score model S(y|D) (which gives a score of
Assumption 1
In Problem 1, a (posterior) probability distribution p(y|D) on a predictive space Y is given.
It is difficult to design reliable estimators for Problem 1. This is because there are an immense number of candidate solutions, and therefore, any point estimation, even if it is the prediction with the highest probability, is not reliable as its probability is extremely small. Hence, maximum likelihood (ML) and maximum score (minimum energy) estimators (both of which have been widely utilized) are not sufficient in those estimation problems. Moreover, as pointed out in Carvalho and Lawrence (2008), consistency, asymptotic normality, and asymptotic efficiency are not established for the ML estimator for Problem 1, although those properties have been established for the ML estimator on continuous spaces. Carvalho and Lawrence (2008) also pointed out that there is no reason for the ML estimation to be a representative solution in Y, because ML estimators do not consider the entire distribution of solutions.
When accuracy measures of a target problem are given (e.g., sensitivity, positive predictive value [PPV], Matthew's correlation coefficient [MCC], or F-score [Baldi et al., 2000]) (see Section A.1 in the Appendix), it is reasonable to design estimators that are suited to those accuracy measures. Maximizing expected accuracy (MEA) estimators, which are the main focus of this study, are able to consider both accuracy measures of the target problem and an entire distribution of solutions, and have been successfully applied to a number of estimation problems in bioinformatics (Do et al., 2006a; Sahraeian and Yoon, 2010; Lu et al., 2009; Nánási et al., 2010). In this article, we classify existing algorithms and software from the viewpoint of MEA, which will provide useful information not only for users but also for developers of such software.
This rest of this review is organized as follows. In Section 2, we explain the concepts of maximizing expected accuracy (MEA) estimation. In Section 3, we present a classification of existing algorithms from the viewpoint of MEA; therein, in Table 1, we summarize the classification. In Section 4, we discuss additional issues related to MEA estimations. In Section 5, we conclude, and in Section 6, we provide an Appendix.
This table is sorted by “Target problem.”
L and B mean Y ⊂ Ln and Y ⊂{0, 1} n , respectively.
Gain function. See Section 2.3 for definitions.
The use of approximated MEA estimators in Section 2.4.2.
The use of representative MEA estimators in Section 2.4.1.
Methods for computing estimation: DP (dynamic programming), IP (integer programming), SS (stochastic sampling), GS (Gibbs sampling).
Transmembrane topology predictions, signal peptide predictions, protein secondary structure prediction, etc.
An extension of
2. Concepts of Maximizing Expected Accuracy (MEA) Estimation
2.1. Maximizing expected gain (MEG) estimator
In Problem 1 with Assumption 1, the following estimator is called a Maximum expected gain (MEG) estimator (Hamada et al., 2011a).
where G(θ, y) on Y × Y is called a gain function, which gives higher values (gains) when θ and y are similar.
This MEG estimator is closely related to statistical decision theory, in which an estimator that minimizes expected loss is often considered (Carvalho and Lawrence, 2008). In order to facilitate the understanding of the relationship with MEA, in this review we use a gain function that should be maximized instead of minimizing a loss.
When the gain function G is designed according to the accuracy measures of the target problem (e.g., MCC, F-score, PPV and Sensitivity), the MEG estimator is called a maximum expected accuracy (MEA) estimator. (This does not mean the gain function is exactly equal to the accuracy measure.) On the other hand, when G(θ, y) is equal to the delta function, δ(θ, y), that is 1 only when θ is exactly equal to y, the estimator is called a maximum likelihood (ML) estimator. Note that it is quite unreasonable to employ the delta function as the accuracy measure, because the condition described by the delta function is too strict. ML estimators are, therefore, unsuitable as accuracy measures in many bioinformatics problems, and the gain function should be designed more carefully.
In the following two subsections, we introduce several commonly used predictive spaces and gain functions, which are used in the classification in Section 3 (and Table 1 therein).
2.2. Commonly used predictive (solution) spaces, Y
2.2.1. Y is a subset of Ln for |L| < ∞
Typically, L is a set of labels and the data D is a biological sequence with length n (e.g., DNA, RNA, or protein sequence) as in the following examples.
Example 1 (The space of protein secondary structures:
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$${ \cal P} ( x )$$\end{document}
)
For a protein sequence x and L = {α-helix, β-strand,loop} (a set of labels for components of protein secondary structures), a protein secondary structure y (of x) can be represented as
Example 2 (The space of gene structures:
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$${ \cal G} ( x )$$\end{document}
)
For a genome sequence x and L = {exon, intron, intergenic} (a set of labels for components of gene structures), a gene structure y can be represented as
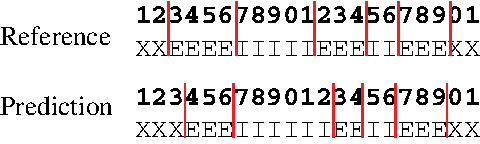
Example of gene prediction. The top and bottom figures are a reference gene structure θ and a predicted gene structure y, respectively. The labels X, E, and I indicate intergenic regions, exons, and introns, respectively. The vertical lines in red show boundaries (exon-intron and intergenic region-exon boundaries). We compute G(label) (θ, y) = 19 and
In general, Y is not equal to Ln but is a subset of Ln, which means that the labels of each dimension (position) in a prediction are mutually correlated and cannot be estimated independently.
2.2.2. Y is a subset of {0 1}n
Although this is a special case of the predictive space described in Subsection 2.2.1 (where L = {0, 1}), we consider it separately for convenience. In this case, 0 and 1 in a binary vector
Example 3 (The space of secondary structures of an RNA sequence:
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$${ \cal S} ( x )$$\end{document}
)
For an RNA sequence x, a secondary structure of x is represented as a upper triangular binary-valued matrix, y = {yij}1≤i≤j≤|x|, where yij = 1 means xi and xj (the i-th and j-th bases of x) form a base pair and yij = 0 means xi and xj do not form a base pair.
Example 4 (The space of alignments of two sequences:
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$${ \cal A} ( x , x^ \prime )$$\end{document}
For two biological sequences x and x′, a pairwise alignment y between x and x′ is represented as a binary-valued matrix
Note that the above predictive spaces are a subset of binary space, which means that every element in the predictive space has complicated constraints.
2.3. Commonly used gain functions
2.3.1. A gain function for Y ⊂ Ln: label gain function
For θ,
Here, I(condition) is the indicator function that returns 1 only when condition is true. When θ is a correct (reference) sequence and y is a prediction, Eq. (2) is equal to the number of correctly predicted labels. The MEG estimator of this gain function, therefore, maximizes the expected number of correctly predicted labels.
Example 5 (G(label) for gene prediction)
In gene prediction from a genomic sequence, when θ is a reference sequence and y is a prediction, G(label) (θ, y) is the number of correctly predicted labels. For example, in Figure 1, G(label) (θ, y) = 19.
2.3.2. A gain function for Y ⊂ Ln: boundary gain function
For θ,
where B is the list of all pairs of labels corresponding to a boundary (e.g., an exon-intron boundary for gene prediction). When θ is a correct prediction and y is a prediction, Eq. (3) is equal to a weighted sum of the number of correctly predicted boundaries and non-boundaries. The MEG estimator of this gain function is, therefore, suitable for accurate prediction of boundary of annotation (boundary accuracy).
Example 6 (
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$$\bf G_ \gamma^{{ \rm \bf ( boundary ) }}$$\end{document}
for gene prediction)
In gene prediction, when θ is a reference genomic sequence and y is a prediction,
The γ in Eq. (3) is a parameter that adjusts between the sensitivity and PPV of a prediction. Using larger γ leads to more boundaries (that is, more genes) in the prediction.
2.3.3. A gain function for Y ⊂ {0, 1}n: γ-centroid gain function
For θ,
where γ ≥ 0 is a weight parameter. When y is a prediction and θ is a reference sequence, this gain function is equal to a weighted sum of the number of TP and TN. (This gain function was originally proposed in the context of RNA secondary structure prediction, in Hamada et al. [2009a].)
Example 7 (
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$$G_ \gamma^{{ \rm ( centroid ) }}$$\end{document}
for RNA secondary structure)
For two secondary structures y and θ in

Example of RNA secondary structure prediction. The top and bottom structures are a reference θ and prediction y, respectively. (
Example 8 (
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$$G_ \gamma^{{ \rm ( centroid ) }}$$\end{document}
for pairwise alignment)
For two secondary structures y and θ in
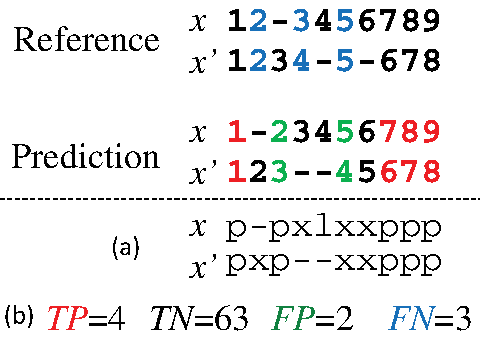
Example of pairwise alignment. The top and bottom alignments are a reference θ and prediction y, respectively. (The numbers indicate positions in the sequences and “-” indicates a gap.) (
An MEG estimator with this gain function is often called a γ-centroid estimator. The parameter γ in the γ-centroid estimator can be naturally introduced based on the criterion that more true predictions and fewer false predictions are required (Hamada et al., 2011a). The parameter is used for adjusting between the sensitivity and PPV of a prediction. It is easily seen that the MEG estimator of
2.3.4. A gain function for Y ⊂ {0, 1}n: MCC/F-score
For θ,
where Acc is either MCC or F-score (Baldi et al., 2000), both of which are accuracy measures providing a balance between sensitivity and PPV. If G(θ, y) = MCC(θ, y) or F-score(θ, y), where θ is a reference and y is a prediction, the MEG estimator of the gain function maximizes the expected accuracy (Acc).
Example 9 (G(Acc) for RNA secondary structure prediction)
For θ,
Unlike the γ-centroid estimators, the MEG estimator of this gain function does not contain any parameter. However, it is generally difficult to compute the estimator. Instead, Hamada et al. (2010) have proposed an approximate method to maximize expected MCC/F-score. In Hamada et al. (2010), the authors focused on RNA secondary structure prediction, but the method is applicable to other problems.
2.3.5. A gain function for Y ⊂ {0, 1}n × {0, 1}m
Suppose that each binary vector has two indices, that is, Y ⊂ {0, 1}
n
× {0, 1}
m
(like
is introduced. The second and third terms in the right-hand side are equal to 1 when θij = yij = 0 for all j and θij = yij = 0 for all i, respectively. If the products (Π i and Π j ) are replaced by sums (Σ i and Σ j ), the gain function is equal to (twice) the γ-centroid gain function, Eq. (4).
Interestingly, this gain function was independently proposed in Do et al. (2006a) (in the context of RNA secondary structure prediction) and in Schwartz et al. (2005) (in the context of pairwise alignment).
Example 10 (
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$$\bf G_ \gamma^{ ( 2{ \rm \bf dim} ) }$$\end{document}
for RNA secondary structure)
When Y is the space of secondary structures of a given RNA sequence x (i.e.,
Example 11 (
\documentclass{aastex}
\usepackage{amsbsy}
\usepackage{amsfonts}
\usepackage{amssymb}
\usepackage{bm}
\usepackage{mathrsfs}
\usepackage{pifont}
\usepackage{stmaryrd}
\usepackage{textcomp}
\usepackage{portland, xspace}
\usepackage{amsmath, amsxtra}
\pagestyle{empty}
\DeclareMathSizes {10} {9} {7} {6}
\begin{document}
$$G_ \gamma^{ ( 2{ \rm dim} ) }$$\end{document}
for pairwise alignment)
When Y is the space of possible pairwise alignments between two sequences x and x′ (i.e.,
2.4. Two variants of MEG/MEA estimators
The following two variants of an MEG/MEA estimator were proposed in Hamada et al. (2011a) (in the context of a restricted class of MEA estimators, that is, the γ-centroid estimators).
2.4.1. Representative/common MEG/MEA estimator
In some cases, the data D consists of several data-points dn, for example,
Example 12 (Common RNA secondary structure prediction)
Given a set of RNA sequences
Example 13 (Sequence feature prediction in a multiple alignment)
Given a set of biological sequences
For those problems, the following estimator (called a representative MEG estimator) can be introduced. It gives a consensus or common prediction for probability distributions of every data point:
where
Example 14
For Example 12, the estimators used in McCaskill-MEA and PETfold can be considered representative estimators of the
We remark that the following example can be also considered as a similar problem by taking dn = di,k = {x(i), x(k)} for
Example 15 (Pairwise alignment between two multiple alignments)
Given two multiple alignments A1 and A2, predict a pairwise alignment between A1 and A2.
We will describe further applications of representative estimators in Section 3. See also the column “Rep” in Table 1 below.
2.4.2. Approximated MEG/MEA estimator with additional information
In Problem 1, by employing additional information appropriately, it is possible to improve accuracy.
Example 16 (RNA secondary structure prediction with homologous sequences)
Given a (target) RNA sequence x and its homologous sequence h, predict a secondary structure
Example 17 (Pairwise alignment with homologous sequence information)
For two biological sequences x and x′ and their homologous sequence h, predict a pairwise alignment
Example 18 (RNA alignment with common secondary structure information)
For two RNA sequences x and x′, predict a pairwise alignment
Ideally, a (refined) probability distribution on the predictive space Y is given by marginalizing onto
By using these marginal probability distributions on a predictive space Y, the MEG estimators are introduced directly. However, the computational cost of computing this MEG estimator is generally huge, and several heuristic methods are, therefore, employed, including a factorization of the probability distribution p(y′∣D, A). (For example, a probability distribution of possible structural alignments between x and h is factorized into the distributions of secondary structures of x and x′, and the distribution of pairwise alignments.) The factorization generally leads to a number of inconsistencies in the distribution and those inconsistencies should be resolved when the gain function is designed.
We call this type of estimator an “approximated MEA estimator” (Hamada et al., 2011a).
Example 19
For Examples 16, 17, and 18, approximated MEA estimators are employed in CentroidHomfold (Hamada et al., 2009c), ProbCons (Do et al., 2005), and CentroidAlign (Hamada et al., 2009b), respectively.
We will also describe further applications of this type of estimator in Section 3. See also the column “Apr” in Table 1 below.
2.5. Commonly used approaches to compute MEG/MEA estimators
To obtain a final prediction of MEG/MEA (and related) estimators, we need to compute the “argmax” operation in Eq. (1). There are several commonly used approaches:
DP algorithms are widely used in bioinformatics, including alignment and RNA secondary structure prediction (Smith and Waterman, 1981). IP is also employed in bioinformatics problems (Sato et al., 2011; Kato et al., 2010). Stochastic sampling enables us to sample directly from the posterior distribution p(y∣D). This approach has been proposed for pairwise alignments (Webb-Robertson et al., 2008), RNA secondary structure predictions (Ding et al., 2005), and structural alignments of RNA sequences (Harmanci et al., 2009).
In methods described in the next section, one of the above techniques is employed to compute a final prediction; see the “Comp” column in Table 1 below.
3. Classification of Various Estimators in Bioinformatics From The Viewpoint of MEA
In this section, we classify various estimators appearing in bioinformatics from the viewpoint of MEA. The classification considers the type of predictive space, the gain function, and the optimization method. For a summary of the classification, see Table 1.
3.1. Feature predictions in biological sequence
3.1.1. Transmembrane topology prediction and signal peptide prediction
For the prediction of sequence features like transmembrane topology, signal peptides, coil-coil structures, and protein secondary structures (which are formulated as Problem 1; for example, see Example 1), the “Optimal accuracy decoding” method used in Kall et al. (2005) can be considered as the MEA estimator of the gain function G(label) (Eq. (2)). Also, in transmembrane topology prediction and signal peptide prediction, the authors showed that this estimator achieved superior performance to the ML estimator and a (heuristic) posterior decoding method (cf. Section 4.2) proposed by Fariselli et al. (2005).
Moreover, the authors proposed an improved method for the problem which incorporated homologous sequence information (given by sequences aligned to the target sequence). This method can be considered as a representative MEA estimator (Section 2.4.1; Example 13) of the gain function G(label). In their article, the authors showed that prediction accuracy was substantially improved by employing homologous sequence information.
3.1.2. Gene prediction
Gene prediction is formulated as Problem 1 with
3.1.3. HIV recombination detection
For the problem of detecting recombination in the genome of the human immunodeficiency virus (HIV) with jumping hidden Markov models (HMMs) (Schultz et al., 2006), Nánási et al. (2010) proposed using the highest expected reward decoding (HERD) for the HMMs. This is a kind of MEA estimator with a special gain function that is an extension of
3.2. Pairwise/multiple/local alignment of biological sequences
3.2.1. Pairwise alignment
For the problem of (pairwise) alignment of two sequences x and x′ (Problem 1 with
Miyazawa (1995) proposed an estimator for pairwise alignments, which constructs alignments by using all the aligned bases whose posterior probabilities are larger than 0.5. Interestingly, a set of aligned bases whose probability is larger than 0.5 always produces a consistent alignment (Miyazawa, 1995; Carvalho and Lawrence, 2008) (i.e., one contained in
Miyazawa's approach (Miyazawa, 1995) typically gives rise to an incomplete alignment that contains a number of unaligned residues (because all the paired residues whose posterior probability is less than 0.5 are unaligned). As an alternative, Holmes and Durbin (1998) proposed an estimator that maximizes the sum of posterior probabilities of aligned bases. This estimator is equivalent to the MEG estimator of
Recently, Frith et al. (2010) employed the MEG estimator of the gain function
On the other hand, the alignment method proposed in Schwartz et al. (2005) and Schwartz, (2007) is equivalent to the MEA estimator of the gain function
It should be emphasized that each of the above estimators can be efficiently computed by a Needleman-Wunsch-style DP algorithm in O(∣x∣∣x′∣) time. The recursive equation of the DP is written as
where Mi,k stores the optimal value of the alignment between two sub-sequences
For the alignment method proposed by Holmes and Durbin (1998), Xik is set to be pik, the marginal probability that xi and
3.2.2. Multiple alignment of DNA/protein sequences
In most multiple alignment algorithms, pairwise alignments (according to a guide tree) are first made in order to obtain a final multiple alignment of a set of sequences S. In this step, pairwise alignment between x and x′ in S can be estimated by using the homologous sequence information of the other sequences, H = S \{x, x′} (cf. Example 17). An approximated MEA estimator of the gain function
In the (progressive) alignment procedure, pairwise alignment between two multiple alignments (Example 15) is employed. A representative MEA estimator has been utilized in several multiple alignment algorithms, including ProbCons (Do et al., 2005). Note that the final multiple alignment of these algorithms is obtained by using a DP algorithm.
On the other hand, the estimator used in AMAP (Schwartz and Pachter, 2007) is equivalent to the MEA estimator of the gain function
3.2.3. Local alignment of DNA/protein sequences
Frith et al. (2010) employed the MEA estimator of the gain function
3.3. Sequence analyses of RNAs
This field is one of the most successful applications of MEA estimation. The importance of sequence analysis of RNAs has increased due to the recent discovery of (functional) non-coding RNAs (Carninci and Hayashizaki, 2007; Mattick, 2005).
3.3.1. RNA secondary structure prediction
RNA secondary structure prediction (i.e., Problem 1 with
There exist several state-of-the-art probabilistic models for secondary structures of a given RNA sequence: (a) the McCaskill model (McCaskill, 1990) with experimentally determined energy parameters (Mathews et al., 1999), (b) the McCaskill model with Boltzmann likelihood (BL) parameters (determined by a machine learning method) (Andronescu et al., 2010, 2007), (c) the CONTRAfold model (Do et al., 2006a) based on the conditional random field (CRF) model, and (d) the stochastic context free grammar (SCFG) model (Dowell and Eddy, 2004). Those models can be utilized as the probability distribution on the predictive space
The estimator used in Sfold (Ding et al., 2005) can be considered as the MEG estimator of the gain function
CONTRAfold (Do et al., 2006a) utilized the MEA estimator of the gain function
On the other hand, Hamada et al. (2009a) proved that the MEA estimator of the gain function
If we have the homologous sequences of the target RNA sequence (Example 16), the probability distribution of secondary structures of the target RNA sequence should be provided by the marginalized probability distribution of structural alignments between the target sequence and homologous sequences. An approximated MEA estimator with this probabilistic distribution has also been proposed (Hamada et al., 2009c). (The software implementing this approach is called CentroidHomfold.) In Hamada et al. (2009c, 2011c), the authors showed that the accuracy of secondary structure prediction was greatly improved by employing homologous sequence information.
The computation of most of the estimators described above is conducted by using a Nussinov-type DP algorithm (Nussinov et al., 1978) in O(∣x∣3) time:
where Mi,j stores the best score of the sub-sequence
Although no efficient method has been reported to maximize expected Acc, where Acc is equal to MCC or F-score (i.e., the MEG estimator with the gain function G(Acc) [Eq. (5)]), Hamada et al. (2010) have recently proposed an approximate method that uses a pseudo expected MCC or F-score that is a quite good approximation to the expected MCC or F-score, respectively.
3.3.2. Common secondary structure prediction of multiple alignment of RNAs
The problem is to predict a secondary structure whose length is equal to the length of an alignment. This is often called a common or consensus secondary structure (Example 12). The RNAalifold model (Bernhart et al., 2008; Hofacker et al., 2002) and the Pfold model (Knudsen and Hein, 1999, 2003) directly provide a probability distribution p(θ∣D) for the common secondary structures of a given alignment D. Those probabilistic models are then used in the following MEA estimators.
The estimator used in the latest version of Pfold (Knudsen and Hein, 2003) is the MEA estimator of the gain function
RNAalifold (Bernhart et al., 2008) employs the centroid estimator (the MEA estimator of the gain function
McCaskill-MEA (Kiryu et al., 2007b) is deemed to be a representative MEA estimator (Section 2.4.1) of the gain function
The estimator used in PETfold (Seemann et al., 2008) can be considered as a representative MEA estimator of the gain function
Recently, Hamada et al. (2011b, 2009a) also utilized a representative MEA estimator (Section 2.4.1) of the gain function
All the estimators described above can be computed by a DP algorithm similar to Eq. (9) (Hamada et al., 2011b).
3.3.3. Multiple alignment of RNAs
Because secondary structures are closely related to the functions of (functional) non-coding RNAs, the standard multiple alignment method (Section 3.2.2) is generally insufficient for aligning RNA sequences. Instead, structural alignment is appropriate where both consensus secondary structure and alignment are simultaneously estimated and optimized. However, it is known that the computational cost of structural alignment is high (Sankoff, 1985).
In Hamada et al. (2009b), the authors proposed a fast and accurate method for aligning multiple RNA sequences (CentroidAlign). Their estimator is equivalent to an approximate MEA estimator, which is an approximation of the MEA estimator of the gain function
3.3.4. Local alignment of RNAs
Tabei and Asai (2009) proposed a method (SCARNA-LM) for computing local alignment of RNAs. They utilized the MEA estimator with the gain function
3.3.5. RNA-RNA interaction prediction
RactIP (Kato et al., 2010) estimates RNA-RNA interactions, that is, joint secondary structures of two interacting RNA sequences. The method used in RactIP can be seen as an approximated MEA-based estimator with the gain function
Seemann et al. (2011) proposed an algorithm (PETcofold) to predict an RNA-RNA interaction between two multiple alignments of RNA sequences. The aim is to predict conserved interactions (and joint secondary structures) between the two multiple alignments, which is similar to the idea of predicting pairwise alignments and common secondary structure from a given multiple alignment of RNA sequences. Their algorithm can be seen as a representative MEA estimator with the gain function
3.4. Phylogenetic tree (topology) estimation
Phylogenetic tree (topology) estimation is a classic and important problem in sequence analysis (Durbin et al., 1998). A phylogenetic tree for a given operational taxonomic unit (S) is represented as a binary vector with 2n−1 − n − 1 dimensions, where n is the number of units in S, based on partitions of S formed by cutting every edge in the tree. The topological accuracy measure for estimated trees is often based on the partitions (e.g., Robinson-Foulds [RF] measure [Robinson and Foulds, 1981]; Section 2.4 in Zhang et al. [2011]). A sampling algorithm can be used to estimate the partitioning probabilities (Metropolis et al., 1953).
Felsenstein (1985) proposed the X%-consensus tree, and the 50% consensus tree is equivalent to the tree of the centroid estimation (i.e., the centroid tree). Moreover, it is easily seen that the X%-consensus tree is equivalent to the MEG estimator with the gain function
4. Discussion
4.1. Avoiding point estimations
As described in Section 1, it is difficult to design reliable point estimators for Problem 1. Although point estimation based on the viewpoint of MEA provides a promising approach to the problem, solutions still have extremely low probability. It is, therefore, desirable to avoid point estimation if possible. When a pipeline is developed by combining several estimation algorithms, point estimation should be avoided in the middle of the pipeline even if the final prediction is a point estimation. For example, when a phylogenetic tree is estimated from several unaligned sequences, one standard approach is to predict a multiple alignment of the sequences and then estimate a phylogenetic tree from the predicted multiple alignment. This approach would not be appropriate because point estimation of multiple alignments is uncertain (i.e., results have low probability). Hence, if possible, a phylogenetic tree should be estimated considering all the possible multiple alignments. Although, in general, the computational cost might be increased by considering all the possible alignments, an approach similar to that in Section 2.4.2 is useful for reducing computational cost. It should be noted that the credibility limit of a point estimation (Webb-Robertson et al., 2008; Newberg and Lawrence, 2009) is also useful, because it is considered as a global measure of the estimation.
Another possible approach for avoiding the unreliability of point estimation for Problem 1 is to predict several suboptimal solutions (Steffen et al., 2006; Wuchty et al., 1999), giving up point estimations. It would also be useful to cluster solutions in the predictive space and estimate a solution for every cluster (Ding et al., 2004). Note that we can employ MEA-based estimators (e.g., with the gain function
4.2. Posterior decoding methods (PDMs)
MEA/MEG estimators are considered as a special case of posterior decoding methods (PDM). In posterior decoding methods, several marginal probabilities are (heuristically) employed in order to obtain (decode) a final point estimation. Although it is often difficult to interpret PDMs from the viewpoint of MEG/MEA, we now list posterior decoding methods appearing in bioinformatics.
For sequence feature prediction (Section 3.1), Fariselli et al. (2005) proposed a posterior decoding method to predict the topology of all beta membranes proteins.
For pairwise/multiple alignment of biological sequences (Sections 3.2.1 and 3.2.2), ProDA (Phuong et al., 2006) produces local multiple alignment of protein sequences, in which a posterior decoding method with marginal probabilities for the unaligned (flanking) regions was employed. GRAPE (Lunter et al., 2008) utilizes a posterior decoding method similar to the MEA estimator of
For RNA secondary structure prediction (Section 3.3.1), ProbKnot (Bellaousov and Mathews, 2010) uses a kind of posterior decoding method to predict secondary structure with pseudo-knots. It seems difficult to consider their estimator from the viewpoint of MEA, although the authors call their method “maximum expected accuracy.”
For (structural) RNA alignments (Section 3.3.3), PARTS (Harmanci et al., 2008), RAF (Do et al., 2008), and Murlet (Kiryu et al., 2007b) employ posterior decoding methods based on the Sankoff algorithm (Sankoff, 1985). R-coffee (Wilm et al., 2008), PicXAA-R (Sahraeian and Yoon, 2011), and MAFFT (Katoh and Toh, 2008) use a posterior decoding method similar to CentroidAlign (Hamada et al., 2009b) (Section 3.3.3) and do not produce structural alignment.
For (Bayesian) co-estimation of phylogeny and sequence alignment, Lunter et al. (2005) utilized a posterior decoding method.
4.3. Training probabilistic models from the viewpoint of MEA (MEA training)
In this review, we assumed that a probability distribution p(y∣D) on a predictive space Y is obtained beforehand in Problem 1. It is, however, important to design the probability distribution p(y∣D) itself. Distributions given by a probabilistic model such as an HMM or CRF contain a number of parameters. It would, therefore, be useful to train the parameters in the probability distribution with respect to the target accuracy measures. This type of training is called “MEA training” in general, and there have been several studies of MEA training in the field of machine learning: (Suzuki et al., 2006; Gross et al., 2007b; Jansche, 2007). There are, however, few studies applying MEA training to problems in bioinformatics (Gross et al., 2007a), and further studies in that area would be enlightening.
5. Conclusion
In this review, we have briefly described the concepts of MEA estimators, which are an alternative approach to conventional maximum likelihood or maximum score estimators. We then classified existing algorithms used in bioinformatics from the viewpoint of MEA. We believe that this review will be useful not only for users of the software mentioned in this review but also for developers wishing to design algorithms on the basis of MEA.
6. Appendix A
A.1. Accuracy measures based on TP, TN, FP, and FN
There are several measures for evaluating a prediction in estimation problems for which we have a reference (correct) prediction (Problem 1). The sensitivity (SEN), positive predictive value (PPV), Matthew's correlation coefficient (MCC), and F-score for a prediction are defined as follows:
where TP, TN, FP, and FN are defined by
where
Footnotes
Acknowledgments
We are grateful to Dr. Martin C. Frith for commenting on the manuscript. This work was supported in part by a Grant-in-Aid for Scientific Research on Innovative Areas in Japan.
Disclosure Statement
No competing financial interests exist.
