Abstract
Background:
Biobanking of prostate cancer tissue is crucial for advancing biomarker-guided precision medicine. However, there is no standardized optimal protocol for biobanking prostatectomy specimens. This study aims to compare the representativeness and sustainability of two biobanking protocols—“Punch” and “Slice”—currently used in Norway.
Methods:
Fresh frozen tissue from 40 radical prostatectomy specimens was biobanked using both the Punch and Slice protocols. Following macroscopic evaluation, a 2 mm thick transverse slice of the prostate (Slice protocol) was collected and stored in an ultra-freezer for future drill biopsy subsampling, guided by histopathological assessment of adjacent formalin-fixed, paraffin-embedded tissue sections. After the slice was collected, five cylindrical tissue samples were punched from the cut surfaces (Punch protocol). Statistical analyses were conducted to compare the sampling precision and time consumption of both protocols.
Results:
Cancerous tissue was successfully sampled in 87.5% of cases using the Punch protocol and 75% of cases using the Slice protocol. Both methods yielded comparable results in terms of the number of cancerous cores and the ability to sample tissue representing the highest Gleason grade. The mean biobanking time of tissue slices was 4.9 minutes compared to 15.1 minutes for the ready-to-use tissue punches. Both methods have previously been shown to provide high-quality RNA extracts.
Conclusion:
Both biobanking protocols are effective for sampling prostate cancer tissue, with no significant difference in precision or quality. The choice between protocols should consider factors such as resource availability, tissue quantity, and specific research needs. The Punch protocol is less resource-intensive overall, while the Slice protocol collects vastly more tissue, has a shorter period of ischemia, and provides detailed mapping of biobanked components, allowing for further subsampling at multiple time points.
Introduction
The overall increase in prostate cancer cases, coupled with the fact that many patients still succumb to the disease, underscores the need for better patient stratification and biomarker-guided precision medicine in prostate cancer care. Access to well-defined biobanks with extensive metadata is crucial in helping researchers gain deeper biological insights into the behavior of prostate cancer.
Despite more than 30 years of prostate biobanking, no gold standard exists for processing and collecting biological material from radical prostatectomy specimens. Biobanking methods vary widely, with some protocols collecting systematic or targeted punch biopsies from the cut prostate,1–5 while others preserve entire transverse slices for later extraction of desired tissue elements.6–9 Choosing a method requires balancing tissue quality and quantity, as well as simplicity and duration of the biobanking, without disrupting routine pathological evaluation.
Established in 2010, Biobank Norway is a large-scale national research infrastructure for population-based and disease-specific clinical biobanks. Recognizing the imperative for multicenter biobanking of heterogeneous diseases, Biobank Norway launched a pilot to establish a federated prostate tissue biobank in 2018.
This study aimed to compare two main prostate cancer biobanking protocols in Norway: the collection of cross-sectional prostatectomy slices and punch biopsies. We evaluated the representativeness of the collected material and addressed the sustainability of each method for prospective biobanking of fresh tissue from our extensive prostate cancer cohorts.
Materials and Methods
Patient selection
Specimens from 40 patients who underwent robotic-assisted laparoscopic radical prostatectomy (RALP) at Oslo University Hospital (OUS) between February and July 2018 were prospectively collected given that written consent to the Research Biobank of Cancer at OUS (REC 23207) had been signed.
Preoperative data collection on tumor localization
Information on potential RALP patients was received before the morning of surgery. The suspected tumor area was identified based on diagnostic core needle biopsy pathology reports and preoperative MRI data. A sketch marking the suspicious area(s) on a stylized prostate was drawn (Fig. 1, top) and later used to determine the location of the cross-sectional slice for biobanking.
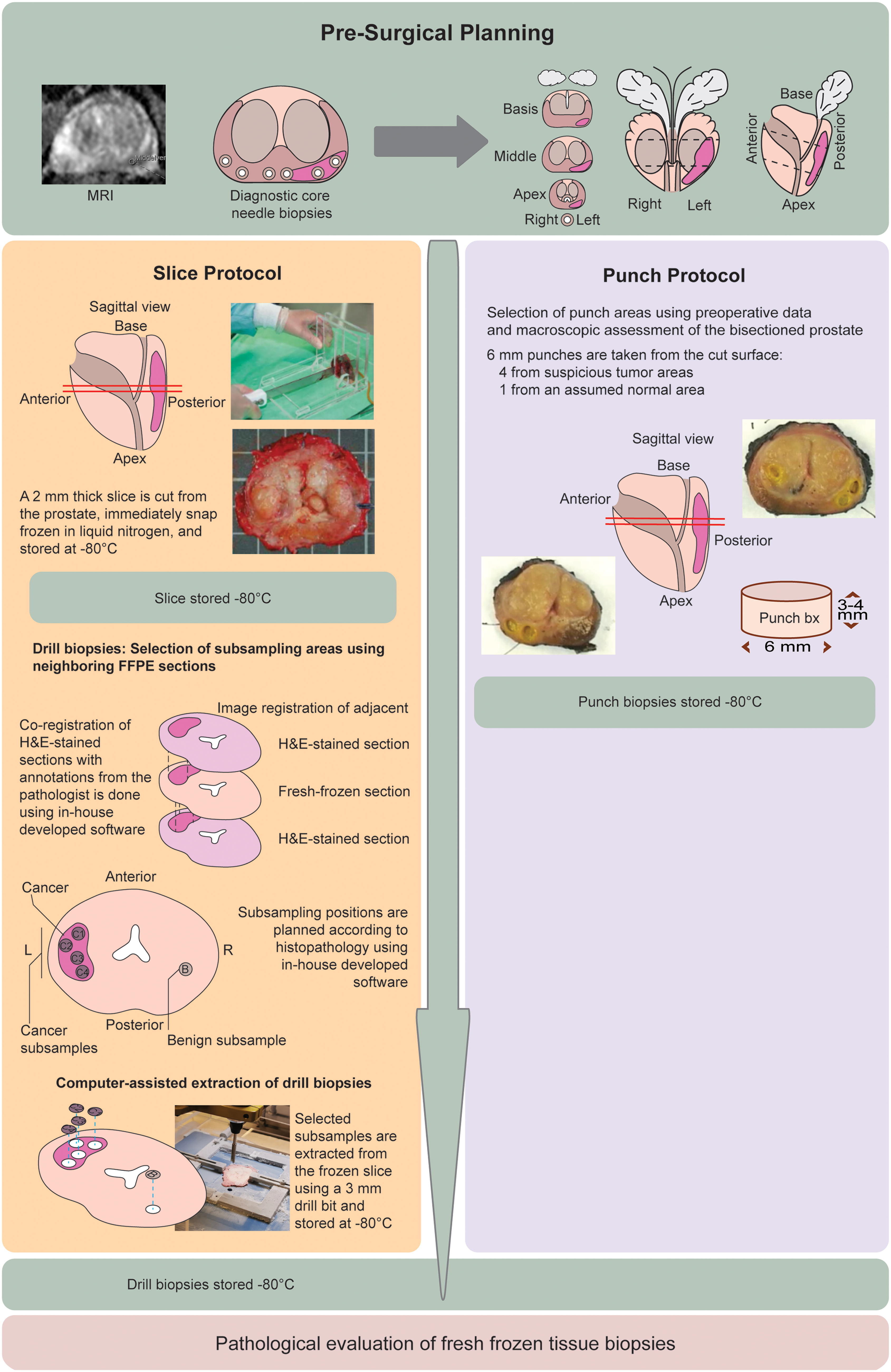
Workflow for prostate biobanking and research-ready sample preparation. The figure outlines the biobanking process for prostatectomy specimens and the subsequent steps required to prepare samples for research use. The workflow begins with the pre-surgical planning phase (
Collection of biobank material; Slice protocol and Punch protocol
Upon receiving the prostate at the Department of Pathology, it was oriented, with the dorsal surface marked black, the right side green, and the left side blue using tissue marking dye (Electron Microscopy Sciences, 63020-BK/GR/BL). Measurements were taken for anterior-posterior diameter, largest transverse dimension, urethral length, and right/left vesicles/ducts. Biobanking was then conducted using both methods, with the Slice protocol (Fig. 1, left panel) performed before the Punch protocol (Fig. 1, right panel).
The Slice protocol
A 2 mm cross-sectional prostatectomy slice was collected as described by Bertilsson et al., 6 with the following modification: the slice location was determined by prostate palpation and the preoperative information described above. The prostate was positioned in a plastic rack, right side up and anterior surface facing the biobanker, and adjusted to produce a slice including the tumor-suspicious area.
The Punch protocol
After removing the transverse slice, the macroscopic appearance of the remaining cut surfaces guided the extraction of four punch cylinders (6 mm diameter) from cancerous tissue and one from normal tissue, using a new needle for each sample. The tissue samples were transferred to numbered and 2D-coded cryotubes. Scalpels and tweezers were washed with absolute ethanol between extractions. The tubes were collected in an ID-marked bag, snap-frozen (−80°C) in a mixture of isopentane and dry ice, and then transferred to an ultra-freezer for storage. The punched holes were highlighted in yellow, and the cut surfaces with their orientation and numbering were photographed and attached to the pathology journal (Fig. 1, right panel).
The Slice protocol—post-biobanking assessment and subsampling
Drill biopsies (3 mm diameter) were collected from the frozen 2 mm thick cross-sectional prostatectomy slices based on histopathological evaluation of adjacent tissue (Fig. 1, left panel). Gleason scoring is a cornerstone of histological assessment in prostate cancer, based on pathologist evaluation of the architectural patterns in the malignant tissue.
10
Currently, three malignant Gleason patterns (GP) are widely recognized: GP3 (least aggressive), GP4, and GP5 (most aggressive). The presence and relative proportions of these patterns (also called grades) are used to determine the Gleason score (GS), which is calculated by combining the predominant pattern with the highest-grade pattern remaining (biopsies) or the second most prevalent pattern (prostatectomies); e.g., GP4 + GP5 = GS9.
11
This score plays a crucial role in guiding patient prognosis and treatment decisions. After annotating areas of different Gleason patterns in adjacent hematoxylin and eosin (H&E) slides, subsampling was performed. Four biopsies targeted cancerous areas, while one targeted benign tissue, as for the Punch protocol (see subsection:
Histopathological assessment of tissue sections adjacent to the biobanked slice
For each of the 40 prostatectomy specimens, formalin-fixed paraffin-embedded (FFPE) transverse tissue sections adjacent to the biobanked slice, both basally and apically, were stained with hematoxylin and eosin (H&E) and evaluated microscopically to predict areas of cancer in the biobanked slice located between them. These H&E sections were annotated to create a tissue map, guiding the later extraction of drill biopsies. Annotation was done using a microscope (Olympus BX50) and marker pens of different colors (Edding® 140S).
In 32 cases, the resulting H&E slides depicted the transverse prostatic surface on both sides of the biobanked slice, with each surface visualized by 1–3 large slides (76 × 50 mm). In the remaining 8 cases, collection of drill biopsies was based on H&E slides from only one side due to the biobanked slice’s proximity to the prostatic apex or base, totaling 100 slides for the 40 cases. Each of the 100 slides was manually annotated using different pen colors and line types to outline seven tissue categories: Gleason pattern 3, Gleason pattern 4, Gleason pattern 5, high-grade prostatic intraepithelial neoplasia (HGPIN), stroma, atrophy, and nodular hyperplasia.
The histopathological evaluation was conducted by two pathologists, including an experienced uropathology specialist (E.G.). In cases of disagreement, a third uropathology specialist was consulted (O.G.A). We defined a tumor focus as malignant tissue surrounded by a 2–3 mm rim of benign stromal or glandular elements, using this to distinguish closely related tumor foci on the same slide. The only size criterion was a minimum focus diameter of 5 mm.
Selection of subsampling areas for drill biopsies
H&E-stained sections with overlaid histopathology annotations were scanned at 600 dpi using a standard document scanner. An in-house designed template was used for size calibration and automated import into an in-house developed software for image registration, core subsampling planning, and computer-guided subsampling. Histopathology annotations and section outlines were traced to create vectorized objects linked to the H&E image and its different annotation classes. These images and annotation objects were registered to a digital image of the fresh-frozen sample through rotation and translation. Overlapping annotation objects were used to map and plan the subsampling of drill biopsies (Fig. 2).
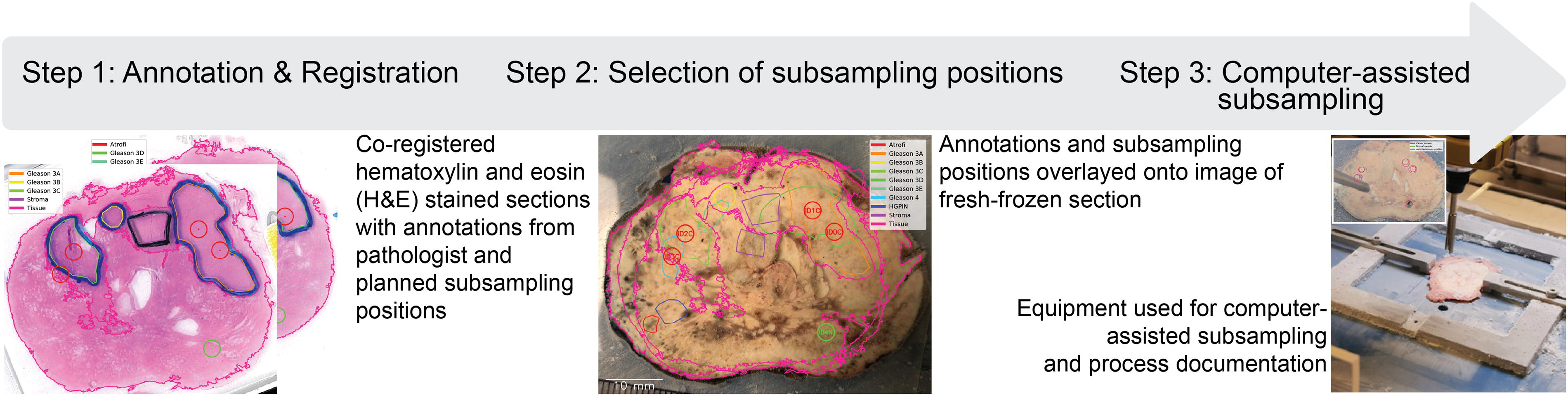
Steps involved in computer-assisted subsampling of the biobanked slice. The figure illustrates the process of subsampling specific tissue regions from the biobanked slice as part of the Slice protocol. H&E-stained tissue sections containing pathologist annotations are digitized and aligned with images of the frozen slice using specialized software to precisely identify subsampling positions. Cores are then extracted from selected regions using an in-house developed drilling station for accurate, reproducible sample collection.
Sampling was performed using an in-house designed drill station based on Bertilsson et al., 6 enhanced with a live camera image overlay showing the planned subsampling position and the outline of the fresh frozen tissue. The frozen tissue was secured by two clamps within a square frame connected to a commercially available coordinate table, positioning it on a liquid nitrogen-cooled sample stage to align with the tissue outline on the live camera for precise subsampling. After visually confirming optimal placement, subsampling was done with a 3 mm diameter hollow drill bit, 6 replaced after each case. Images were captured immediately before and after the procedure, and each subsample was retrieved with forceps and stored at −80°C in a cryovial.
Pathological assessment of frozen sections from the punch and drill biopsies
From each of the 400 cylindrical biopsies (5 per method from each of the 40 cases), one cryo-section was obtained (Slice protocol: 10 µm at −20°C using a Thermo Fisher Cryostar NX70 cryostat; Punch protocol: 5 µm at −15°C using a Leica CM 1950 cryostat), H&E stained, and digitized using an Olympus VS120-S5 slide scanner. Images were evaluated using the open-source image viewer QuPath 12 on a medical-grade EIZO RX850 monitor. The evaluation followed the same methodology as previously described and was conducted by the same three pathologists, who were blinded to the targeted tissue of each biopsy. We assessed the presence of cancerous tissue, its area percentage, and the quantity of different Gleason patterns, as well as significant areas of (HGPIN), stroma, atrophy, and nodular hyperplasia.
Biobanking duration data
We define “biobanking duration” as the time period from when a specimen is received at the pathology department until the sampled tissue is snap-frozen for long-term storage. Since both biobanking protocols were applied to all 40 specimens in this study, internal duration data were unsuitable for comparison. Instead, we obtained external data: for the Punch protocol, we used retrospective duration data from biobanking at OUS between 2010 and 2023 (
Statistical analyses
McNemar’s test was used to investigate differences between the two sampling protocols for categorical variables at the patient level (successful sampling of tumor core, normal core, diagnostic Gleason grade, and core with >75% tumor cells). The Wilcoxon signed-rank test was used for patient-level comparison of continuous variables (number of cancer cores and number of cores with >75% tumor cells per patient). The Mann–Whitney U test was used for comparisons at the core level (tumor content and Gleason grade). Biobanking duration was compared using the unpaired
Ethics
All patients provided consent for inclusion in the Research Biobank of Cancer at OUS (REC 23207). As the project falls outside the Health Research Act’s scope (Section 2), it did not require approval from the Regional Committee for Medical and Health Research Ethics (REC 605138).
Results
Histomorphological assessment of slice-adjacent tissue sections
Cancerous tissue was detected in H&E-stained sections from all 40 specimens. In 28 cases, the highest Gleason pattern identified matched the diagnostic prostatectomy report. Malignant areas were identified both apically and basally to the biobanked slice in 29 specimens, exclusively apically in 5, and exclusively basally in 6. On average, 4.6 cancer foci per case were observed, evenly distributed between apical and basal slides.
Histomorphological assessment of frozen punched and drilled cores
A case-level overview comparing the two protocols is provided in Table 1. Cancer was identified in at least one core for 75% of cases with the Slice protocol and 87.5% of cases with the Punch protocol (Table 1;
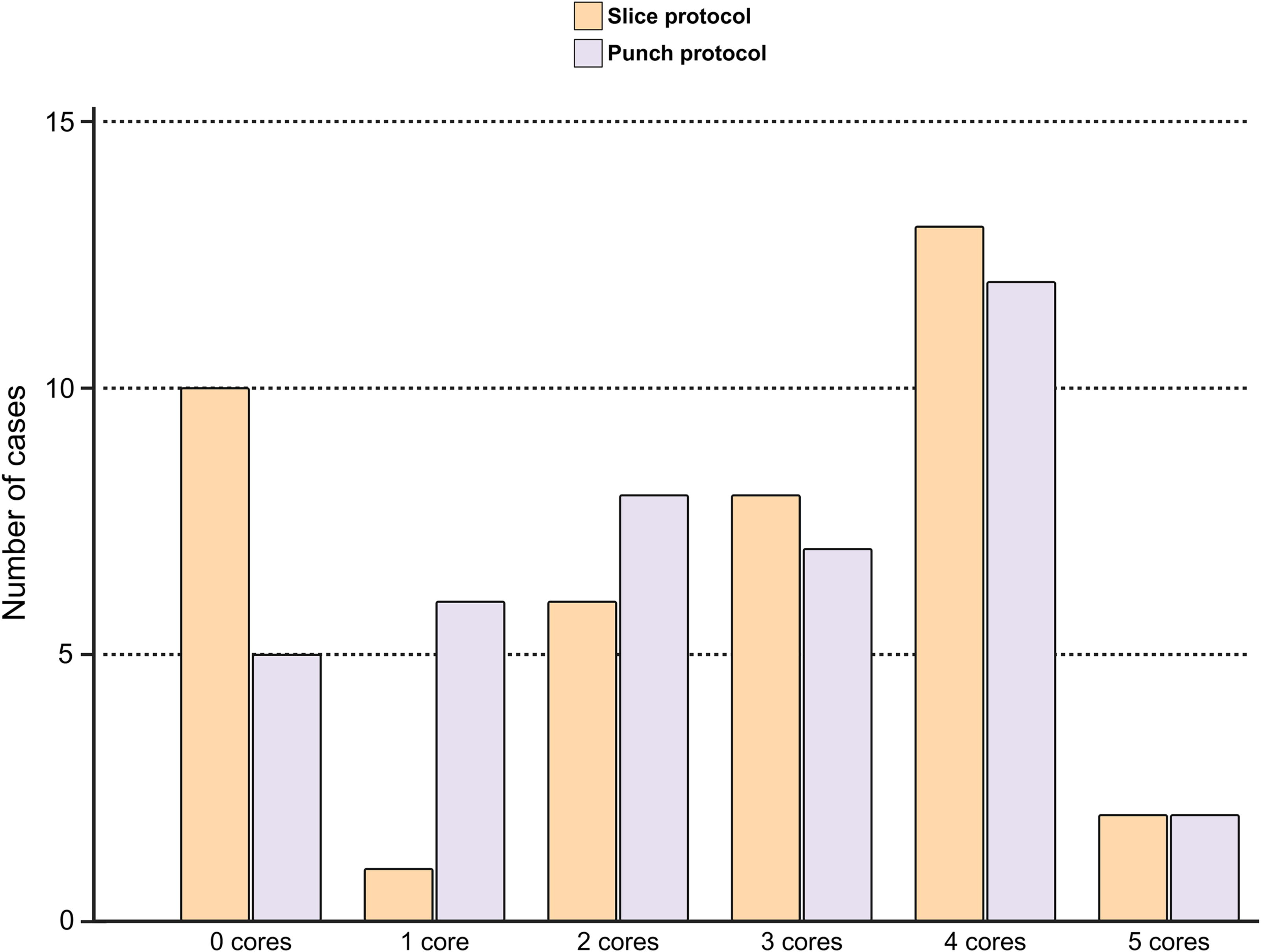
Number of cancerous cores sampled per case. The number of cancerous cores sampled from each of the 40 prostatectomy cases using the Slice (
Comparison of Core Biopsy Contents for the Two Protocols (Case-Level,
No significant difference between protocols (
In many biomarker discovery studies, there is a notable interest in using material with high purity of malignant tissue obtained from the region that represents the highest Gleason grade. We found that over half of the biobanked cores contained more than 50% tumor, regardless of the method (Fig. 4). At the case level, at least one core with more than 75% tumor content was present in 23 cases for the Slice protocol and 20 cases for the Punch protocol, with no significant differences between the two (Table 1,
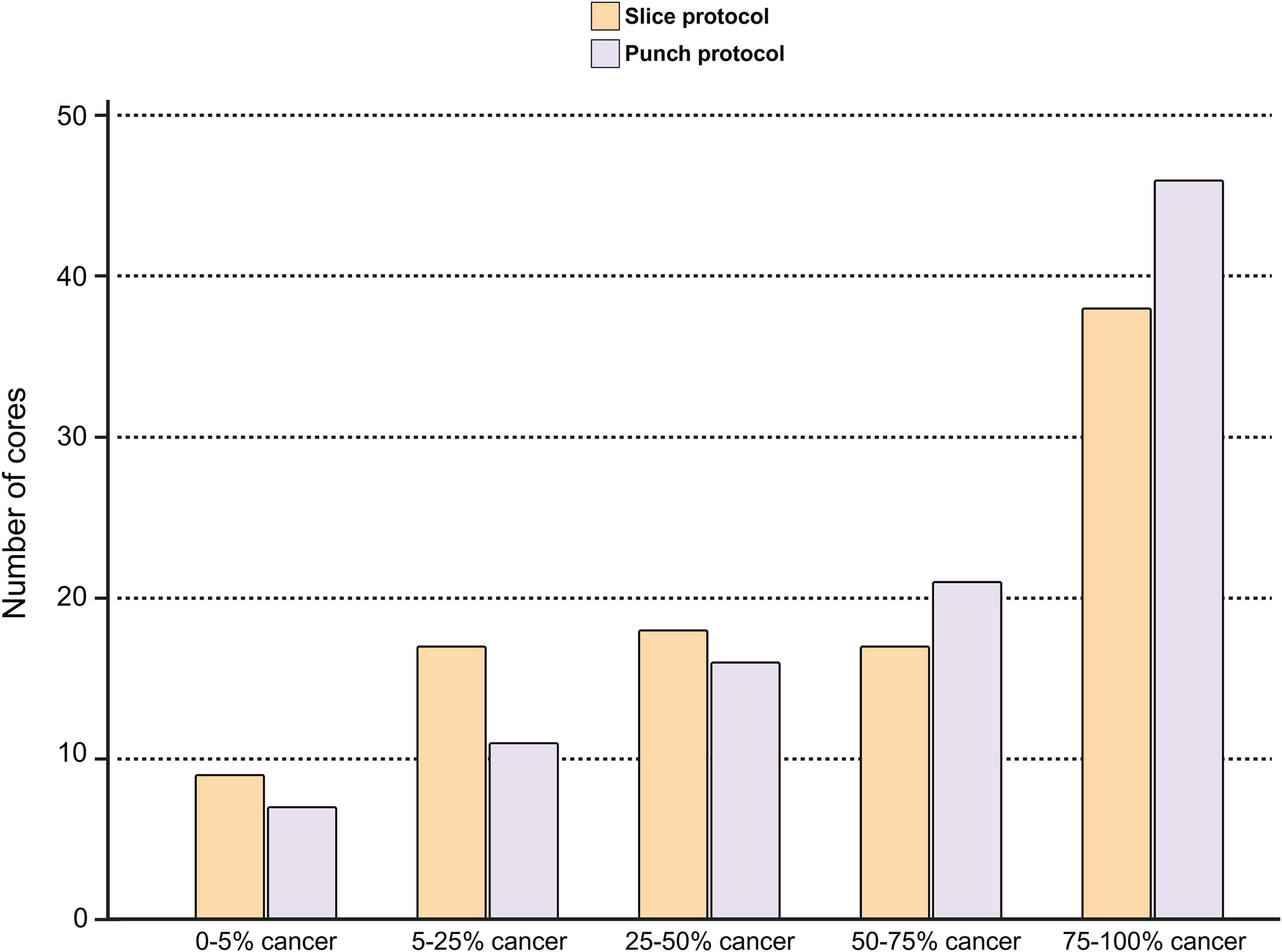
Proportion of cancerous tissue per core. The proportion of cancerous tissue sampled per cancer-positive core is important, as it largely determines the potential utility of biobanked material for future studies. Displayed here is the proportion of cancerous tissue in cancer-positive cores from both sampling methods, showing that most cores are cancer-rich regardless of protocol used.
Putative applications of biobanked fresh-frozen tissue
The Slice and Punch protocols have been successfully used for a variety of bulk omics analyses, including RNA, protein, and metabolite studies,13–16 as well as for spatial transcriptomics, proteomics, and metabolomics.13–16 Both protocols yield high-quality RNA extracts, with Slice samples averaging a RNA Integrity Number (RIN) of 9.16, 6 and over 99% of Punch samples having RIN values above 9.0.14–16
Biobanking duration
The Slice and Punch protocols had a mean biobanking time of 4.9 and 15.1 minutes, respectively. The biobanking time was significantly different for the two methods (
Discussion
This study demonstrates that the Slice and Punch protocols biobank malignant and benign prostate tissue with equally high precision, and thus the choice between protocols should be guided by other factors such as available resources and the specific requirements of downstream analyses. Given their distinct methodologies, each protocol has its own strengths and weaknesses. The Slice protocol collects significantly more tissue and provides a detailed microscopic mapping of the biobanked material, from which subsamples can later be extracted at any time and over multiple instances without thawing the tissue slice. 17 The Punch protocol, on the other hand, is overall less complicated and cheaper, does not rely on specialized equipment or software, and requires very little resources after the initial biobanking procedure.
Macroscopic evaluation of the cut prostatic surface has been described as challenging due to the multifocal and heterogeneous nature of prostate cancer.8,18 To enhance the chance of targeting the index tumor, we used preoperative information to determine the initial level of prostate bisectioning, which is standard for the Punch protocol. In contrast, the Slice protocol typically does not rely on macroscopic evaluation and involves placing the initial transverse cut at the mid-prostate without considering information regarding tumor localization. Thus, we are comparing the Punch protocol with a modified version of the original Slice protocol. While the Punch protocol requires collecting and interpreting preoperative information, the Slice protocol biobanking can be performed without any prior knowledge of tumor localization. Our study design does not provide an optimal basis for comparing the two protocols in terms of sampling heterogeneous tumor tissue, which could potentially be evaluated using immunohistochemistry (IHC) or genomic markers. In the Slice protocol, sampling was partially directed toward the most aggressive areas of the digital tissue map, even when within the same larger tumor focus. This approach limits our ability to accurately assess the protocols for heterogeneity-focused endpoints. However, previous research has explored this aspect using both biobanking protocols independently.19–22
The Slice protocol requires specialized equipment, including a prostate holder with adjustable vertical walls to stabilize the organ during transverse slice harvesting and a double-bladed knife for cutting the organ. The resulting slice is placed in a pre-cooled cryoclamp to prevent vapor insulation, ensuring a flat, frozen specimen suitable for subsampling. Subsamples are extracted from the frozen slice using an in-house developed drilling device, without thawing. 6 Our optimized study protocol involves using H&E-stained transverse large slides obtained from processing of the specimen, immediately adjacent to the biobanked slice both apically and basally, to determine areas for subsequent drill biopsy collection. This requires an additional cryosection, increasing the workload for both the lab and the pathologist. Alternatively, the protocol can be adapted to use only one H&E-stained adjacent section, combined with increased sampling near suspected target areas in the frozen slice to counteract the increased uncertainty of tumor foci localization. In addition, alignment of the frozen slice and annotated H&E sections requires custom software that can correct for changes in tissue morphology between FPPE and fresh-frozen prostate tissue—a process taking approximately 10–15 minutes per case. Implementing this method in a pathology laboratory is estimated to cost between 2.500 and 3.000 EUR. In contrast, the Punch protocol does not require any specialized instruments, extra sections, or computer software, and the pre-biobanking preparations (MRI and biopsy review) take approximately 5 minutes per case.
In terms of time efficiency, we found a statistically significant difference in biobanking duration between the two methods, with the Slice protocol on average being 10 minutes faster to perform than the Punch protocol. The biological processes triggered by ischemia following organ extraction could potentially give rise to bias in metabolomics studies using biobanked material, making it desirable to minimize the ischemia period. However, the Slice protocol relies on subsequent pathologist annotations to map tissue components within the biobanked slice, as well as computer alignment and mechanized drilling of subsamples, all of which extend the time required before the material can be made available for research.
Research projects examining field effects or tumor microenvironment-cancer cell interactions are likely to benefit from using the Slice protocol. This method allows for more precise subsampling of cores within the tissue slice, in contrast to the Punch protocol, where cores are extracted during the biobanking procedure. Furthermore, the Slice protocol allows post-biobanking study design, offering flexibility to adapt study plans or explore additional aspects in follow-up studies by enabling multiple rounds of resampling from the frozen slices. On the contrary, if the protocol needs to be adapted for biobanking of living cells or tissue explants, the Punch protocol is likely better suited. With the Punch protocol, living cells can be prepared for culturing from the tissue core prior to freezing, while the Slice protocol would in essence require integration of the Punch protocol to obtain such material. The differences between the two protocols across various aspects are illustrated in Figure 5.
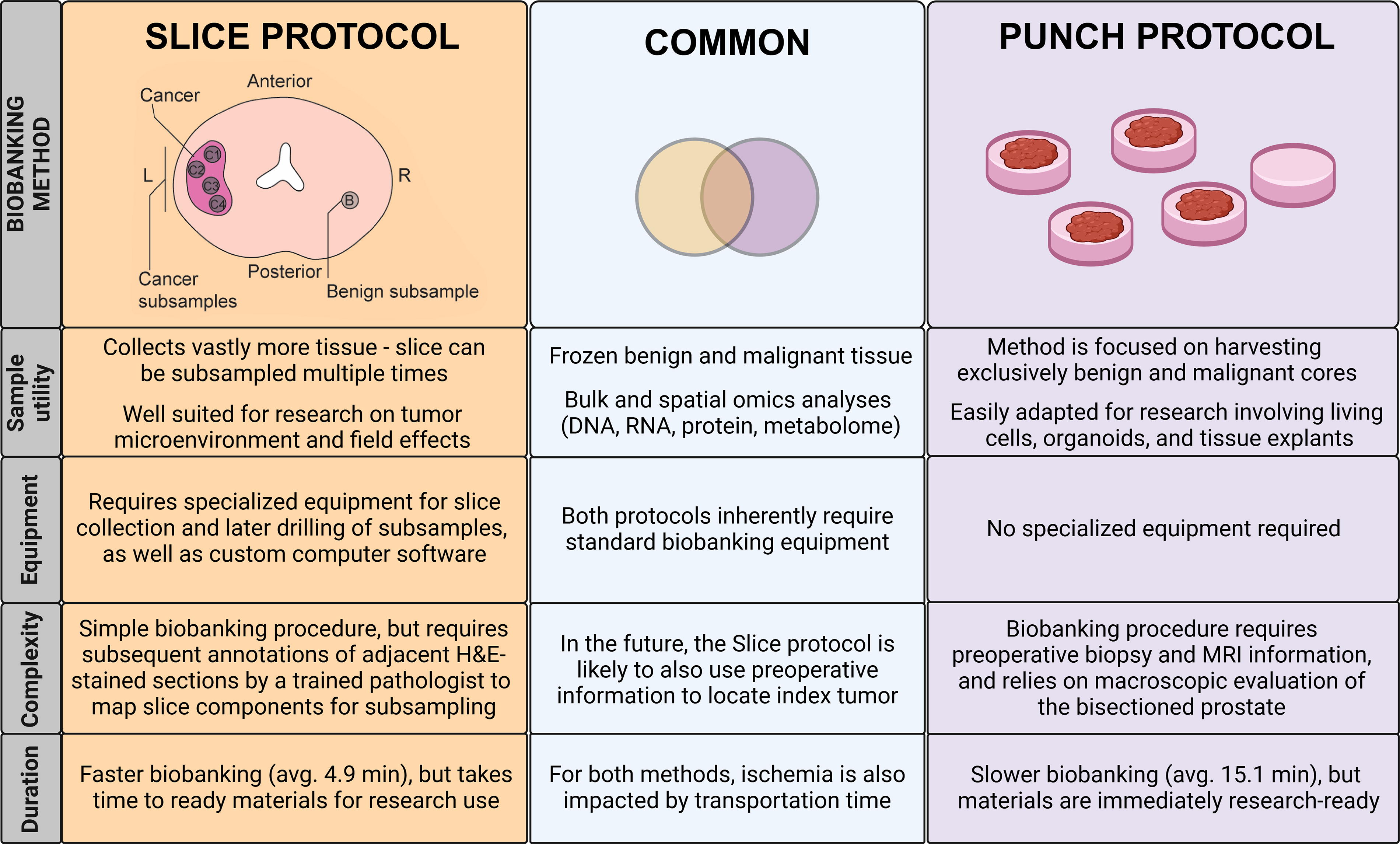
Comparison of Slice and Punch biobanking protocols. The figure highlights key differences in sample utility, equipment needs, procedural complexity, and biobanking duration between the Slice and Punch protocols, as well as illustrating their shared features.
The chosen study design has some drawbacks warranting further discussion. Since each surgical specimen had five core biopsies (Punch protocol) extracted immediately after removal of the central 2 mm slice, the adjacent H&E-stained large slides did not represent a complete morphological equivalent to the biobanked slice. These areas sampled by the Punch protocol would not be available for histomorphological evaluation and, in effect, could not be subsampled by the Slice protocol. This might explain the slightly lower precision in tumor sampling observed with the Slice protocol, though the difference was not statistically significant. In addition, the size of the core biopsies collected by the two protocols differs, with cores from the Slice protocol and Punch protocol having a diameter of 3 mm and 6 mm, respectively. As we wanted to compare the two protocols in their original form, we did not make any adjustments to the diameters of extracted cores. This difference could, however, impact the probability of sampling cancer, as well as the proportion of malignant tissue present in the cores.
The discrepancies between the highest Gleason grades observed in our biobanked material and the highest Gleason grades in the pathology reports could, partly, be attributed to freeze artifacts that distort glandular morphology and obscure the nuclear characteristics of the prostatic epithelium, making precise histopathological assessment difficult.23,24 Embedding tissue in OCT-gel before sectioning has previously been shown to improve tissue morphology and reduce freeze artifacts; 8 however, this would interfere with downstream molecular analyses and is therefore not recommended.25,26
Regarding future perspectives, the Slice protocol is currently being enhanced to replace core drilling with an automated tissue knife capable of extracting custom shapes from annotated tissue directly after digital slide analysis. Of note, the widespread adoption of digital pathology has enabled pathologists to analyze images and annotate specific tissue regions with greater speed and accuracy than before. This technological shift not only enhances efficiency but also facilitates easier sharing and collaboration on annotated images across different sites. There is also ongoing work to integrate preoperative MRI data in selecting the level of slice collection. In contrast, the Punch protocol is a well-established, cost-effective method having incorporated MRI-guided fresh tissue core sampling for over a decade.
Conclusions
Our results show that both biobanking protocols effectively sample tumor and benign tissue from radical prostatectomy specimens, with no statistically significant differences in sampling precision. The Slice protocol harvests vastly more tissue, making it better suited for studying tumor-stroma interactions and field effects, as well as having a shorter ischemia period favoring this method for metabolomics studies. The Punch protocol is overall simpler, less resource-demanding, and can easily be adapted to facilitate work on living cells, tissue explants, and organoids. The choice between methods should primarily consider factors beyond sampling precision.
Footnotes
Acknowledgments
The authors extend their gratitude to Biobank1 27 and the Prostate Biobank at Oslo University Hospital 28 for providing invaluable resources and support. They also thank the patients who generously contributed to this research. Special acknowledgment goes to consultant pathologists Liljana Vlatkovic and Kathrine Lie for their role in establishing the Punch protocol at Oslo University Hospital. Finally, they express their gratitude to the MR Core Facility at NTNU for conducting the drill biopsy sampling, the Cellular & Molecular Imaging Core (CMIC) Facility at NTNU for their work in cryo-sectioning, H&E staining, and digitizing drill biopsies, and Don Trinh at OUH for cryo-sectioning and H&E-staining punch biopsies.
Data Availability
The data from this study is not publicly accessible in compliance with Norwegian privacy laws. Physical histology slides are securely archived at the Department of Pathology, Stavanger University Hospital, Norway. Digital slides and patient data are stored on secure servers managed by the participating institutions.
Authors’ Contributions
A.B.: Data curation, formal analysis, investigation, visualization, writing—original draft, writing—review and editing. G.F.G.: Formal analysis, writing—review and editing. A.N.: Data curation, investigation, methodology, resources, writing—review and editing. S.K.: Investigation, software, visualization, writing—review and editing. E.M., E.G., and O.G.A.: Data curation, investigation, writing—review and editing. H.S.: Conceptualization, funding acquisition, writing—review and editing. J.H.: Conceptualization, methodology, writing—review and editing. E.A.M.J.: Conceptualization, supervision, writing—original draft, writing—review and editing. M.-B.T.: Conceptualization, resources, writing—review and editing. K.A.T.: Conceptualization, funding acquisition, project administration, supervision, visualization, writing—original draft, writing—review and editing.
Author Disclosure Statement
The authors have no conflicting interests to declare.
Funding Information
This work was funded by Biobank Norway 2 (245464/F50) and Biobank Norway CS1 Quality Biobanking, with additional personnel and infrastructure support from Oslo University Hospital, Stavanger University Hospital, and St. Olavs Hospital. Additional funding for some authors was provided by the European Research Council (758306), the Norwegian Cancer Society (208263–2019), and the Central Norway Regional Health Authority (26094).
