Abstract
As scientific investigations increasingly adopt Open Science practices, reuse of data becomes paramount. However, despite decades of progress in internet search tools, finding relevant astrobiology datasets for an envisioned investigation remains challenging due to the precise and atypical needs of the astrobiology researcher. In response, we have developed the Astrobiology Resource Metadata Standard (ARMS), a metadata standard designed to uniformly describe astrobiology “resources,” that is, virtually any product of astrobiology research. Those resources include datasets, physical samples, software (modeling codes and scripts), publications, websites, images, videos, presentations, and so on. ARMS has been formulated to describe astrobiology resources generated by individual scientists or smaller scientific teams, rather than larger mission teams who may be required to use more complex archival metadata schemes. In the following, we discuss the participatory development process, give an overview of the metadata standard, describe its current use in practice, and close with a discussion of additional possible uses and extensions.
Introduction
In response to a White House directive for open and equitable research (White House, 2023), the National Aeronautics and Space Administration (NASA), along with other federal agencies, has declared 2023 to be the Year of Open Science (NASA TOPS, 2023). Open Science entails the sharing of scientific data, software, and other research products (Fecher and Friesike, 2014). This is important because reuse and repurposing of legacy data is increasingly commonplace, with over half of the investigations funded by NASA's Science Mission Directorate (SMD) based on archival data (Big Data Task Force, 2017), for example. However, discovery of relevant astrobiology datasets remains difficult in part due to the lack of precise dataset descriptors tailored toward the needs of researchers (Aydinoglu et al., 2014). Hence, we have developed the Astrobiology Resource Metadata Standard (ARMS) (Keller et al., 2019), an evolving comprehensive standard for the description, access, and discovery of information related to all areas relevant to astrobiology. ARMS goes beyond just datasets and can describe any product of astrobiology research, including physical samples, software, publications, and so on. From an astrobiology viewpoint, the advantage of using ARMS over a generic metadata standard comes from the inclusion of metadata specific to astrobiology, which allows the researcher to describe their information more precisely and in much greater detail.
ARMS is not a static metadata standard, but one that will evolve based on community feedback, emerging trends in scientific focus, and changing data management needs. Developed within the context of the Astrobiology Habitable Environments Database (AHED) (AHED Team, 2023a), ARMS is nonetheless a standalone metadata standard independent of AHED, fully available to be used in novel contexts. The ARMS standard is described online (AHED Team, 2023b) and available as an Extensible Markup Language Schema (World Wide Web Consortium, 2004). By making ARMS publicly available, we support NASA's vision of an Open Science ecosystem that will lead to a transformation in science, increasing accessibility to knowledge and accelerating scientific discoveries (Strategic Data Management Working Group, 2019; NASA Science Mission Directorate, 2022).
Background
General guidance exists on what characteristics a metadata standard should have, as well as its usage. The FAIR principles recommend optimizing specific qualities of data (Wilkinson et al., 2016), such that they should be Findable, Accessible, Interoperable and Reusable, hence the acronym. Many of these principles relate to design of metadata. TRUST (Lin et al., 2020) is another set of principles that are more geared toward properties of repositories, advocating for Transparency, Responsibility, User Focus, Sustainability, and Technology. NASA's SMD requires that all newly SMD-funded research shall make its publications, data, and software publicly available and that its data should follow the FAIR principles (NASA Science Mission Directorate, 2022). Greenberg et al. (2009) argue that a scientific metadata architecture should be easy to use, interoperable with other standards, and suitable for machine processing. However, a survey of 16 metadata standards found that these aims are rarely met (Qin and Li, 2013). Although there are many semantic commonalities among the standards, differences in terminology create barriers to interoperability and tools reuse. Complexity in some standards also creates an undue burden on the creators of the metadata.
Several standards for scientific metadata have informed and influenced our development of ARMS. Darwin Core (Wieczorek et al., 2012) was developed for biodiversity informatics, influenced by the seminal Dublin core metadata standard (Weibel et al., 1998). In turn, the Dryad repository developed their own metadata standard (Greenberg et al., 2009), based in part on the Darwin Core, with a focus on evolutionary biology. The Investigation/Study/Assay Metadata Framework (Johnson et al., 2021) is concentrated on life science and structured around the three concepts in its name. Similarly, the Core Scientific Metadata Model (Matthews et al., 2010) was developed to describe data from large-scale facilities, with its core organizing concept being a scientific study. NASA's Planetary Data System (Planetary Data System, 2022a) has its own metadata standard for encoding the data (Planetary Data System, 2022b), with its obvious focus on planetary missions. Finally, the Site-Based Data Curation Project developed a metadata standard to describe scientifically significant data from Yellowstone's hot springs research (Palmer et al., 2017), a valuable site for a variety of astrobiological investigations.
As a multidisciplinary field, describing astrobiology metadata is challenging due to the varied data sources, measurements, and formats in use (Detweiler et al., 2019). We found no existing metadata standard had both the scope and detail needed—either the standard failed to cover the breadth of astrobiology or was so high-level as to be of limited use. Moreover, we wanted to make ARMS available quickly, keeping the standard lightweight initially. This would ease early adoption, as metadata would initially be created largely by hand, with tools and automation allowing us to add more semantics in later years. Thus, we made the pragmatic choice of creating ARMS as an independent standard. The downside of initially avoiding some complexity is that future integration with existing standards and improving semantics becomes more difficult.
Development Methodology
We used a participatory design approach (Spinuzzi, 2005), where users of the desired product (in this case, ARMS) actively participate in the design process. The development team consisted both of computer scientists well-versed in metadata and standards as well as astrobiologists who would be the eventual end-users. In addition, we regularly met with scientists outside our team to vet our modeling choices and to ensure ARMS represented the larger field. We also consulted the literature, particularly the standards mentioned in Section 2. To develop the astrobiology-specific elements, specifically a set of keywords, we reviewed astrobiology journal keywords from the prior 10 years as well as keywords independently developed by Taşkın and Aydinoglu (2015) and Miller et al. (2014). We validated the ARMS keywords by searching for their occurrences in the abstracts of the 2019 Astrobiology Science Conference (Meadows, 2019). Over 98% of these abstracts contained at least one of our keywords, indicating good coverage; conversely, about one-third of the keywords were not found in any abstract (Lafuente et al., 2019). The extensive coverage from our initial set of keywords could mask a lack of specificity in certain areas, where a general keyword is found in the text, but a more appropriate, specific keyword is missing from ARMS. To discover new potential keywords, we identified groups of similar publications and then identified distinguishing keywords of each, mimicking an artificial intelligence technique known as topic modeling (Blei, 2012). We used abstracts from the 2019 Astrobiology Science Conference as the corpus for our study. We used the partitioning around medoids (PAM) algorithm (Schubert and Rousseeuw, 2019) to organize the abstracts into disjoint clusters, with the number of clusters chosen empirically to yield clusters that were neither too broad nor too narrow. From each cluster, we automatically identified terms that were rare in other clusters but prevalent within the cluster as potential keywords to add to ARMS. From this, we identified 36 keywords that we added into ARMS, including new areas such as education and outreach.
Description
To be ARMS compliant, the astrobiology resource (e.g., dataset, software, physical sample) must be accompanied by a standard set of metadata that describes the resource and characterizes its content. These metadata fall into two categories: resource identification metadata (describing provenance, points of contact, versioning information, funding/support, and associated geospatial collection information) and content metadata (describing the actual content of the resource and its relation to the broader astrobiological context), as shown in Fig. 1. Some metadata elements are free text, while others are restricted or complex types.
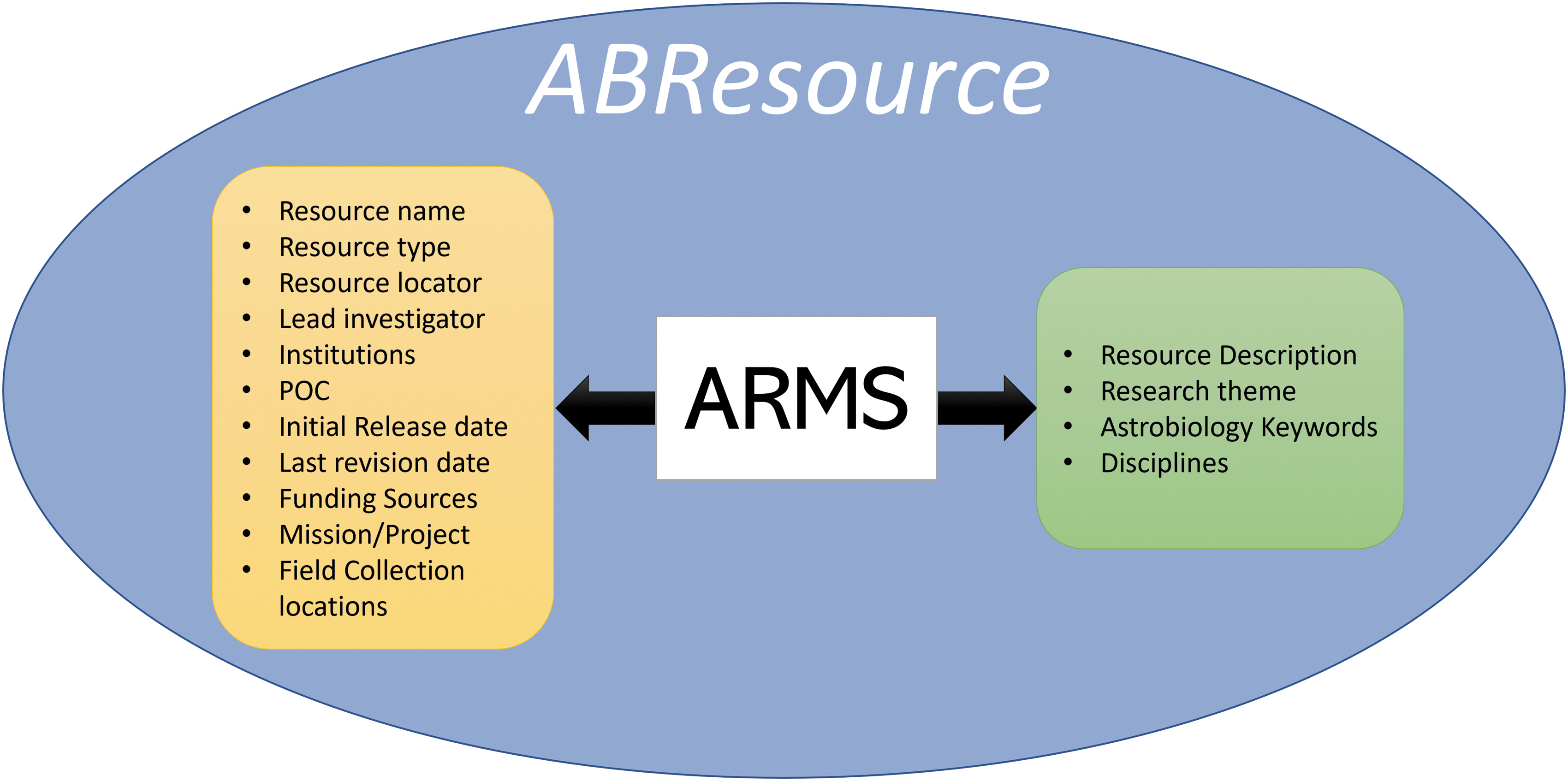
Top-level elements of an ARMS AstroBiology Resource (ABResource), with identification metadata on the left and content metadata on the right.
Of particular interest are the metadata elements that are especially useful as search parameters, namely
Research Theme, the broad research area(s) most relevant to the resource, focused on those identified in the 2015 NASA Astrobiology Strategy Document (Hays et al., 2017);
Project or Mission during which the resource was collected;
Discipline(s) most relevant to the creation or use of the resource; and
Keyword(s) that best characterizes the astrobiology resource described by the ARMS metadata.
The keywords are organized into a four-tier hierarchy, under a top-level structure of 11 categories (AHED Team, 2023b). These top-level categories (in italics) contain keywords that describe the resource in several ways: further detailing the scientific discipline (astronomical, biological, chemical, or geological) or physical process; characterizing the collection (environmental [in situ], exploration [of unusual environments], planetary [beyond Earth]); the analysis methods or computational methods employed; and the institutional support for acquiring the resource. An example of some of these metadata is given in Fig. 2.

Subset of ARMS metadata for (
ARMS was initially developed to standardize the metadata contained in AHED (AHED Team, 2023a), a long-term repository and productivity platform for the storage, discovery, and analysis of data relevant to the field of astrobiology (Bristow et al., 2021). The consistent structure established by ARMS greatly facilitates precise querying and rapid interpretation of search results. The AHED Web Portal (AHED Team, 2023a) hosts an online dataset creation tool, letting users rapidly and intuitively archive ARMS-labeled files or links to other online resources (Fig. 3). AHED also provides an interactive, multifaceted search interface for AHED datasets based on ARMS metadata.
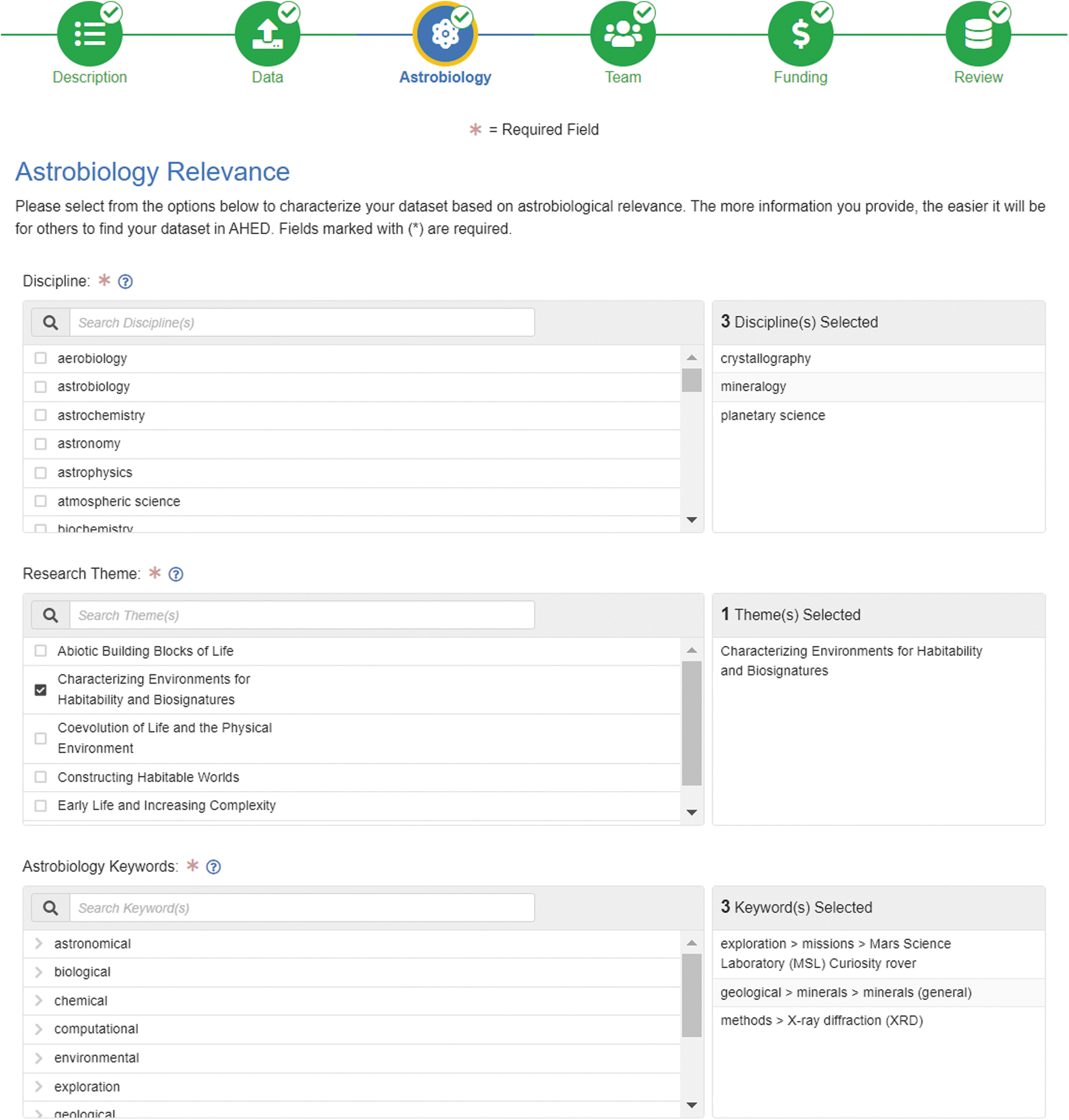
Excerpt of AHED online tool to create ARMS metadata.
Beyond AHED, ARMS can be used to inform the astrobiology portion of a comprehensive science modeling effort, or even be directly incorporated into such. For instance, we have participated in the NASA Science Mission Directorate's Data Catalog effort, which seeks to build a cross-science search capability (Bugbee et al., 2022) and provided ARMS to inform their modeling efforts. In a similar vein, portions of ARMS may be useful for indexing astrobiology text documents, particularly the extensive keyword hierarchy. Examples of this include indexing astrobiology research papers, conference submissions, or proposals for funding. Finally, ARMS could be used to support cross-system communication and representation of astrobiology resources. In fact, the AHED system uses the Open Data Repository (Lafuente et al., 2018), a general data repository, as its backend.
The transformation to Open Science cannot happen by fiat; it must receive the necessary institutional and technological support. Toward this end, we have created ARMS to describe products of astrobiology research. Standardizing the metadata aids the discovery and interpretation of astrobiological datasets, supporting the aims of Open Science. Our approach can also serve as a blueprint for similar endeavors in other science disciplines. As discussed above, ARMS is not the only metadata standard with relevance to astrobiology. Efforts should be made to harmonize ARMS with other relevant standards, such as Darwin Core and PDS-4, potentially translating ARMS to or from these standards. As the discipline and focus of astrobiology evolves, so must ARMS; in particular the keywords, funding sources, missions/projects, and science disciplines will need to be updated over time. The methods outlined in Section 3 to identify keywords can be used to find new keywords in the future, but these too should evolve. Ultimately, this approach could be expanded to not only update the existing structure but to facilitate generating initial keyword structures, so that our approach can be applied to new science domains. Finally, more tools should be developed to assist the labeling of astrobiology datasets with ARMS metadata, leveraging recent advances in natural language processing as appropriate. Within AHED, we have strived to streamline this process, but nonetheless it is a multistep process that can take tens of minutes. Choosing appropriate keywords can be particularly daunting. However, much of this information can be easily gleaned from available sources, for instance associated publications that can be provided as part of the submission process. The challenge is to extract this information automatically from such documents.
Footnotes
Acknowledgments
The authors would like to thank Caleb Scharf and Daniel Berrios for their reviews and comments.
Author Disclosure Statement
No competing financial interests exist.
Funding
This work was funded by NASA SMD Planetary Science Division's Science Enabling Research Activity (SERA) program.
Abbreviations Used
Associate Editor: Michael C. Storrie-Lombardi
