Abstract
Growing evidence of the potential habitability of Ocean Worlds across our solar system is motivating the advancement of technologies capable of detecting life as we know it—sharing a common ancestry or physicochemical origin with life on Earth—or don't know it, representing a distinct emergence of life different than our one known example. Here, we propose the Electronic Life-detection Instrument for Enceladus/Europa (ELIE), a solid-state single-molecule instrument payload that aims to search for life based on the detection of amino acids and informational polymers (IPs) at the parts per billion to trillion level. As a first proof-of-principle in a laboratory environment, we demonstrate the single-molecule detection of the amino acid L-proline at a 10 μM concentration in a compact system. Based on ELIE's solid-state quantum electronic tunneling sensing mechanism, we further propose the quantum property of the HOMO–LUMO gap (energy difference between a molecule's highest energy-occupied molecular orbital and lowest energy-unoccupied molecular orbital) as a novel metric to assess amino acid complexity. Finally, we assess the potential of ELIE to discriminate between abiotically and biotically derived α-amino acid abundance distributions to reduce the false positive risk for life detection. Nanogap technology can also be applied to the detection of nucleobases and short sequences of IPs such as, but not limited to, RNA and DNA. Future missions may utilize ELIE to target preserved biosignatures on the surface of Mars, extant life in its deep subsurface, or life or its biosignatures in a plume, surface, or subsurface of ice moons such as Enceladus or Europa.
One-Sentence Summary: A solid-state nanogap can determine the abundance distribution of amino acids, detect nucleic acids, and shows potential for detecting life as we know it and life as we don't know it.
Introduction
Widespread synthesis of complex organics (amino acids, nucleobases, sugars) occurred early in the history of the solar nebula due to radiation processing of ices (Nuevo et al., 2009, 2012; Ciesla and Sandford, 2012; Meinert et al., 2016), which is likely to occur throughout the universe. These building blocks of “life as we know it” are believed to have been delivered to all potentially habitable zones in the solar system by comet and meteorite impacts (Engel and Macko, 1997; Martins et al., 2008; Schmitt-Kopplin et al., 2010; Callahan et al., 2011; Cooper et al., 2011). Life may have arisen under similar physicochemical environments, such as within alkaline vent systems (Martin and Russell, 2007; Martin et al., 2008) or impact-driven hydrothermal systems (Osinski et al., 2013) on early Earth, Mars, or within subsurface oceans on icy worlds (Vance et al., 2007; Russell et al., 2014; Barge and White, 2017). Because these building blocks form in interstellar space (Nuevo et al., 2009, 2012; Ciesla and Sandford, 2012), in reducing planetary atmospheres (Hörst et al., 2012), and through aqueous chemistry (Patel et al., 2015; Steel et al., 2017; Ménez et al., 2018), we hypothesize that amino acids are a common component of life and that nucleic acids or related informational polymers (IPs) might be a common solution for information storage and heredity.
Amino acids can be stable over geologic time (Engel and Macko, 1997; Bada et al., 1998; Glavin et al., 1999; Pizzarello and Cronin, 2000), and their structural complexity and abundance distribution in extraterrestrial material, as well as under terrestrial abiotic production, represent a null hypothesis for amino acid distributions in the search for ancient or extant life (Davila and McKay, 2014; Reh et al., 2016). Quantum chemical calculations and biochemical experiments have further shown that the chemical reactivity of proteinogenic amino acids exhibits a notable pattern related to the posited temporal emergence in the genetic code (Granold et al., 2018): amino acids added to the genetic code later in evolutionary history have greater reactivity than those found in the Murchison meteorite or recovered from the Miller-Urey experiment (Higgs and Pudritz, 2009; Trifonov, 2009). Here, we show a correlation between the HOMO–LUMO gap (energy difference between a molecule's highest occupied molecular orbital, HOMO, and lowest unoccupied molecular orbital, LUMO) of the proteinogenic amino acids and their tunneling conductance through a nanogap system. We argue that the quantum property of the HOMO–LUMO gap can be utilized as a new measure of amino acid complexity and possibly as an indicator of life that utilizes regulated electron transfer mechanisms.
Therefore, the detection of amino acids and their abundance distribution is a potential biosignature that could allow for the differentiation between widespread abiotic chemistry and potentially ancient or extant life. Alternatively, charged backbones, like the polyphosphates of DNA and RNA, may be universal for aqueous life (Benner, 2004, 2017): they separate a polymer's physical properties from its associated information content, facilitating replication and enabling evolution. Quantifying the presence of linear charged polymers with encoded information content could provide evidence of life, even without knowing the structure of the informational units (such as nucleobases) or their sequence. However, recent work demonstrating the abiotic synthesis of RNA polymers 100–300 nucleotides in length on mineral glass surfaces demonstrates the potential for linear charged polymers in a prebiotic setting (Jerome et al., 2022).
Life elsewhere may also use similar building blocks because of shared ancestry through meteoritic exchange, a scenario most plausible for Earth and Mars (Gladman and Burns, 1996; Gladman et al., 1996; Weiss et al., 2000; Carr, 2022; Fritz et al., 2005; Shuster and Weiss, 2005). Instruments targeting RNA or DNA, such as the Search for Extra-Terrestrial Genomes (SETG) (Ruvkun et al., 2002; Isenbarger et al., 2008a, 2008b; Lui et al., 2011; Carr et al., 2016) are in development but, at present, rely on biological reagents that must be stabilized, and require a complex series of steps to prepare nucleic acids for analysis. Solid-state nanopores have demonstrated the detection and characterization of nucleic acids and have been proposed to potentially detect other polymers (Rezzonico, 2014; Xia et al., 2022). However, such nanopores require high (MHz) sample rates and ultra-low noise, and even so, solid-state nanopore detection of individual bases has not yet been realized (Shekar et al., 2016).
Here we propose the Electronic Life-detection Instrument for Enceladus/Europa (ELIE), a solid-state single-molecule detector. ELIE relies on quantum electronic tunneling nanogap sensors (Fig. 1), which can detect and discriminate among single α-amino acids (Ohshiro et al., 2014), and detect RNA and DNA, including oligomers and individual nucleotides and, with caveats, short oligonucleotide sequences (Ohshiro et al., 2012, 2014, 2018). We aim to primarily target amino acids and IPs, although we are developing ELIE with the potential to target other types of molecules (Fig. 2), including abiotic or prebiotic polymers. This versatility of detecting a wide array of molecules and their abundance patterns would allow ELIE to identify forward contamination but also ancestrally related life—and possibly mirror-image life or alternative polymers. Here we focus on amino acid detection because amino acids exhibit relatively high stability under harsh conditions in comparison to IPs, and are potential targets for characterizing abiotic and prebiotic environments, and classifying ancient or extant life.
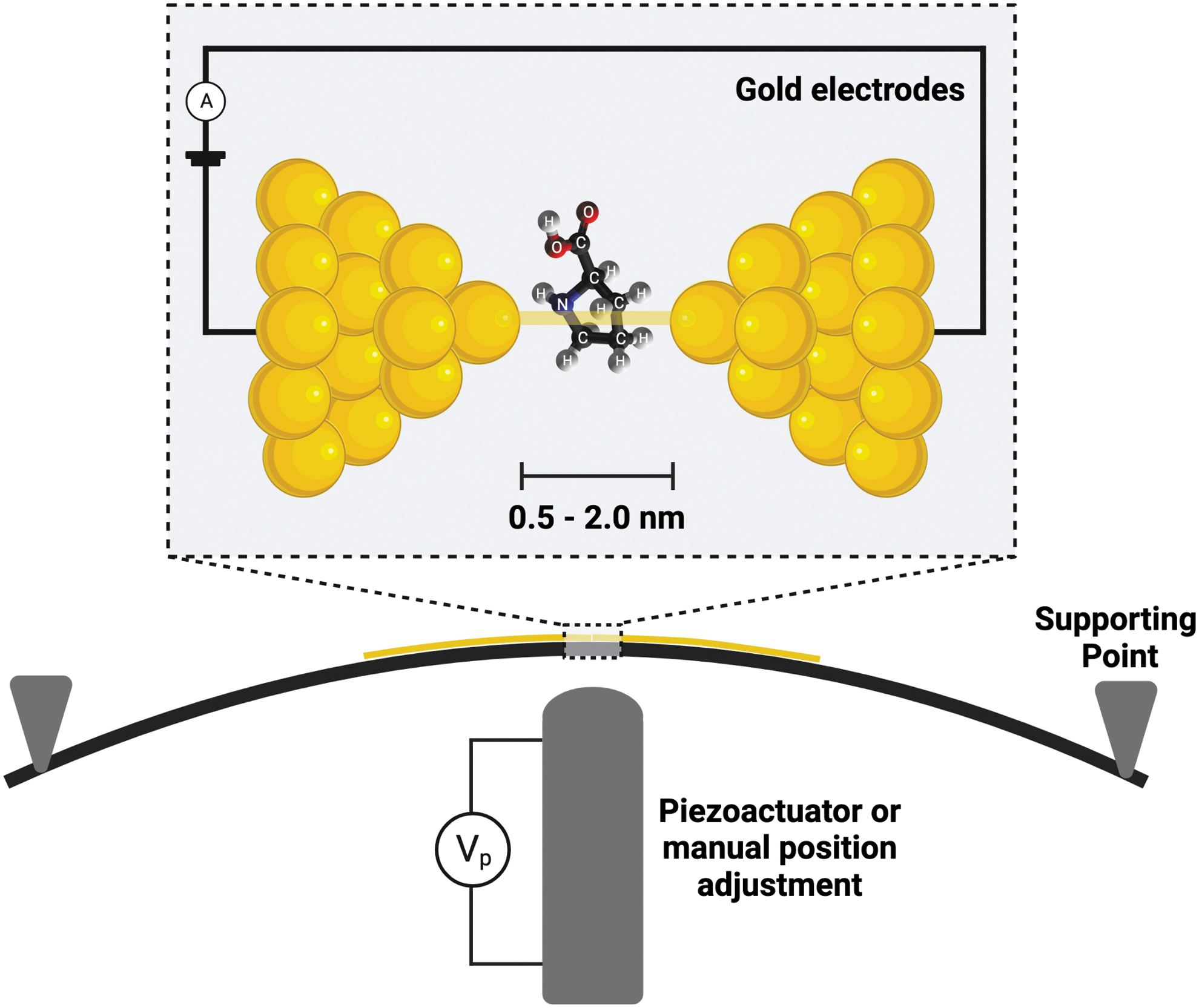
Single-molecule detection with a quantum electronic tunneling nanogap sensor, using the Mechanically Controlled Break Junction method. Three-point bending mechanically breaks a gold nanowire, forming two gold nanoelectrodes. Additional adjustments of the actuator or manual micrometer change the gap size. Target molecules are detected by variations in electrical current between the electrodes due to the dependency of electron tunneling on the electronic structure of any intervening molecule (Di Ventra and Taniguchi, 2016). Created with
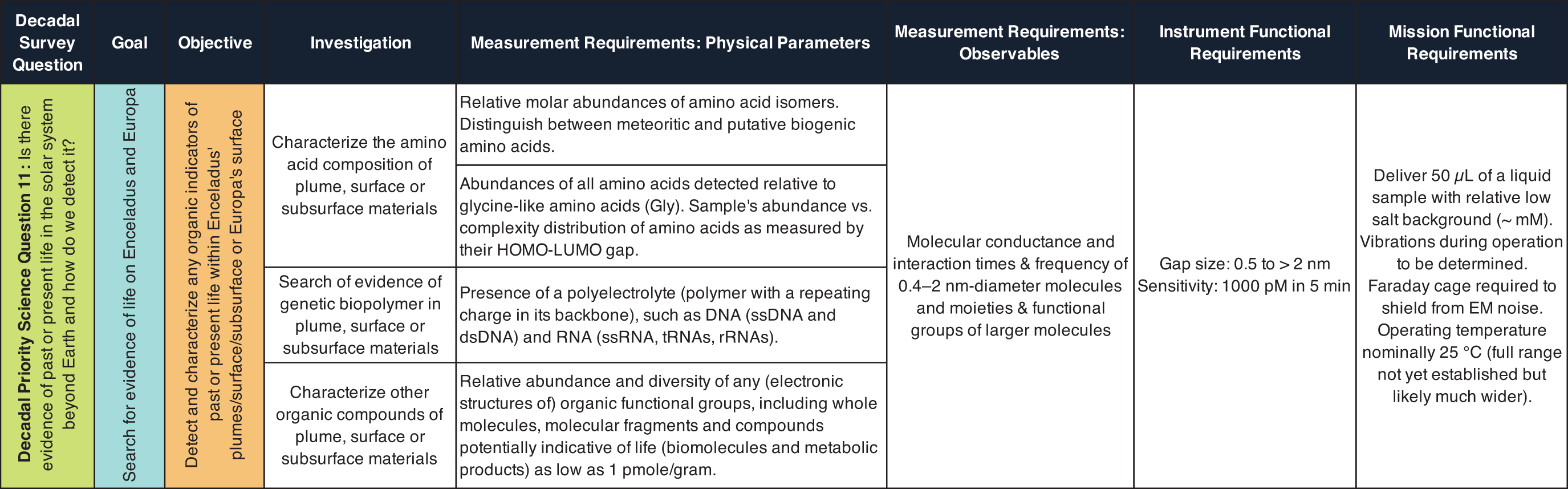

Inferred knowledge of Enceladus and its habitability can provide a rough requirement for analytical sensitivity. The discovery of Enceladus' plume of icy grains and gas by the Cassini mission (Porco et al., 2006; Waite et al., 2009) and the tentative evidence for transient water vapor plumes at several locations on Europa found in Hubble Space Telescope observations (Roth et al., 2014; Sparks et al., 2016) support the idea of material transport from the interior of these moons and, in concert with other studies, the presence of global subsurface oceans (Kivelson et al., 2000; Zimmer et al., 2000; Thomas et al., 2016). Previous work based on inferred amino acid degradation rates suggests that detection of amino acids in these predicted subsurface oceans above a concentration of 1 nM would indicate active production by geochemical or biotic pathways (Truong et al., 2019). Additionally, Steel et al. (2017) predict that methanogen-based microbial life and abiotic hydrothermal processes could produce a maximum of 90 μM and 104 μM concentrations of amino acids, respectively, in the predicted subsurface oceans of these moons.
In this study, we validate the single-molecule detection of L-proline at 10 μM with our first low–technology readiness level (TRL) ELIE prototype and use this result to extrapolate an order-of-magnitude sensitivity of 1 nM. Furthermore, we provide evidence that the quantum property of the HOMO–LUMO gap describes the physical mechanisms underlying the conductance patterns of different proteinogenic amino acids obtained from the solid-state single-molecule detector. Using amino acid abundance distributions reported in the literature, we then show the potential for ELIE to discriminate between abiotically and biotically derived α-amino acids. Finally, this work provides preliminary evidence for the feasibility of integrating ELIE into future in situ life-detection missions.
ELIE instrument hardware
The starting point for ELIE was a benchtop nanogap system developed by the Taniguchi laboratory at Osaka University as reported in Tsutsui et al. (2008a), Ohshiro et al. (2012), Ohshiro et al. (2014), and Ohshiro et al. (2018) (Fig. 3a). The benchtop system consists of a picoammeter, a National Instruments computer, a custom Faraday cage enclosing stepping and piezo motors, and a jig structure holding a nanogap chip, a piezo controller, and battery banks. One of the goals of the subsequently developed ELIE prototype is to demonstrate a significant mass and volume reduction relative to this benchtop system. Compared to the benchtop system, the ELIE system consists of a low-noise amplifier for voltage supply, a laptop for data collection, and a Faraday cage holding an amplifier head stage, a nanogap chip, and a manual micrometer (Fig. 3b, 3c).

ELIE instrument prototype. (
The nanogap chips (Fig. 3d) are fabricated as previously reported by Tsutsui et al. (2008a). First, a silicon substrate is spin-coated with a thin polyimide layer. Gold nanojunctions (100 × 100 nm) are then patterned on top using a standard electron-beam lithography and lift-off technique. Subsequently, the polyimide beneath the junction is carved away by isotropic reactive ion etching, using O2/CF4 gas, to generate a free-standing gold nanowire bridge; this bridge is later formed into a nanometer-scale gap prior to measurement using the Mechanically Controlled Break Junction (MCBJ) method. An optional addition is a cover made of polydimethylsiloxane (PDMS), attached to the silicon substrate, and treated with ozone plasma for bonding. This cover contains a microchannel that allows for the containment of a sample solution and its retention through reduced evaporation under room-temperature conditions. The Faraday cage is connected to a low-noise and high-bandwidth amplifier (Chimera Instruments, VC100, 8 pA RMS at 100 kHz, with sampling up to 4 MHz) that supplies and controls the bias voltage (clamp ±1V) and current measurements (± 20 nA) of the nanogap.
The chip is mounted in a three-point bending mechanism in a jig within the Faraday cage, rinsed with 10% ethanol (as a wetting agent), and bent mechanically with a manual micrometer to form an atomically sharp gap. A 10 kΩ resistor is connected in series to protect the nanogap junction from over-current breakdown due to ohmic heating while in the connected state. In order to form the nanogap, the chip is bent by the micrometer to an open state while monitoring the current. Once the gap is formed and in the open (unconnected) state, the resistor is removed from the circuit so that the current resulting from the applied voltage difference will reflect the gap conductance. As described by Tsutsui et al. (2008b), the resulting tunneling current (I) can be defined as an exponential function of the gap distance (d) as I ∼ exp (βd) with the decay constant 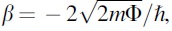 , where m,
, where m, 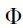 , and
, and 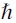 are the electron mass, the work function of a gold atom (Au), and reduced Planck's constant, respectively. The gap is typically characterized in a dry state or in the presence of a buffer prior to making measurements with a sample. The distance is adjusted through chip bending, which produces changes in the gap distance that are a fraction of the vertical motion of the adjustment mechanism. The distance is monitored by the (VC100) measured current, which represents the sum of offset error (set to zero when bias is set to 0 mV), ionic current, and tunneling current. In practice, the baseline is zeroed during analysis so that differences in current reflect the enhancement of tunneling current when a molecule of interest occupies the gap.
are the electron mass, the work function of a gold atom (Au), and reduced Planck's constant, respectively. The gap is typically characterized in a dry state or in the presence of a buffer prior to making measurements with a sample. The distance is adjusted through chip bending, which produces changes in the gap distance that are a fraction of the vertical motion of the adjustment mechanism. The distance is monitored by the (VC100) measured current, which represents the sum of offset error (set to zero when bias is set to 0 mV), ionic current, and tunneling current. In practice, the baseline is zeroed during analysis so that differences in current reflect the enhancement of tunneling current when a molecule of interest occupies the gap.
ELIE instrument L-proline measurement
L-proline was obtained from Sigma (81709), and a 10 μM L-proline solution was prepared by dilution in nuclease-free water with a background of 1 mM phosphate buffer, pH 7.4 at 25°C (Sigma P3619). L-proline was selected based on prior use as a nanogap standard and the ability to detect L-proline at a range of gap sizes (e.g., in Ohshiro et al., 2014). The solution was introduced through the microchannel, and the current was recorded for 5 min. Current measurements were conducted with an applied voltage of 100 mV and a sampling rate of 4 MHz.
ELIE instrument tunneling current data analysis
Data were low pass filtered to 25 kHz and downsampled to 100 kS/s to reduce the amount of electrical noise and the amount of data storage, respectively. Baseline adjustment was defined by estimating a kernel density over windows of 0.1 s with a 0.001 s sliding step. Events were detected using OpenNanopore v1.2, a tool that employs adaptive thresholding (by adjusting to low-frequency variations in the baseline) and a cumulative sum (CUSUM) fit to detect events (Raillon et al., 2012). Low-level noise signals within the set of events were subsequently identified using a 6σ threshold below the inferred mean of the events' tunneling currents and were excluded from further analysis. Features extracted from each of the events include the tunneling current (mean current level within a single event) and dwell time (duration) (Fig. 4a).
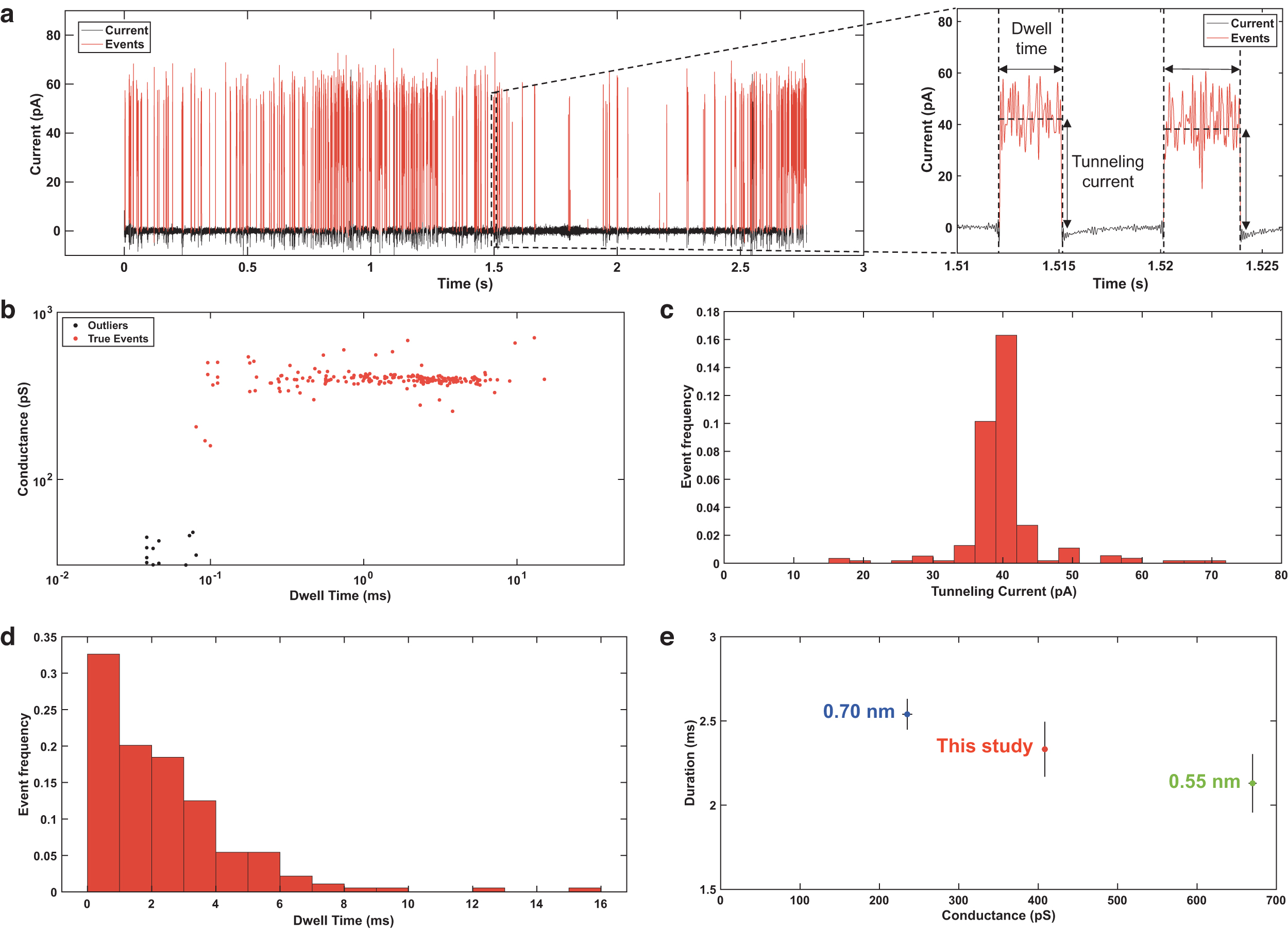
Single-molecule identification of L-proline. (
To gain insight into determinants of tunneling current and dwell time, linear regression models were fitted by plotting the values of various physical and quantum properties of proteinogenic amino acids (molecular weight, molecular volume, and HOMO–LUMO gap) against the conductance and dwell time values of amino acids measured by Ohshiro et al. (2014) (using the 0.55 nm and 0.70 nm nanogap electrodes; refer to Supplementary Figure S1). The HOMO–LUMO gap values of the proteinogenic amino acids were reported by Granold et al. (2018) and determined by semiempirical calculations using the Molecular Orbital PACkage (MOPAC) 2003 AM1 Hamiltonian method in the MOPAC interface of Chem3D 9.0.
Predicting differences in distribution patterns between meteoritic and biologically produced amino acids
We generated a literature-based database of reported abundances of meteoritic α-amino acids and amino acids extracted from diverse environmental samples (Martins et al., 2007, 2015; Hou et al., 2009; Glavin et al., 2010, 2020, 2021; Fuchida et al., 2014; Noell et al., 2018; Aerts et al., 2020; Aponte et al., 2020; Elsila et al., 2021). The analysis only considered meteoritic α-amino acids observed in multiple meteorite samples and reported by multiple authors. The studied meteorite samples were classified based on meteorite groups, following mineralogy and elemental and isotopic composition (Supplementary Table S1). Environmental samples considered in the analysis cover some of the closest terrestrial analogs to the surface of Mars and predicted subsurface environments of Enceladus and Europa (Supplementary Table S2). The analysis only included the reported abundances of the α-amino acids that were present across all samples. Estimated event rates that would be expected for each amino acid by source (environmental or meteoritic) were determined by using an approximate standard event rate observed from the ELIE data in Section 2.4 of 50 events/s in 10 μM aqueous solutions and assuming a linear relationship between concentration and event rate. Finally, single-molecule conductance values of α-amino acids reported by Ohshiro et al. (2014) (using the 0.70 nm nanogap electrodes) were plotted against the predicted event rates. For the α-amino acids not reported by Ohshiro et al. (2014), the expected nanogap conductance signals were estimated using the HOMO–LUMO gap regression model obtained in Section 2.5.
Results
Proof of principle: Single-molecule detection of L-proline
Single-molecule detection was demonstrated with the ELIE system using 10 μM L-proline aqueous solutions and manual adjustment of the gap size between the electrodes. Maintaining a gap in the unconnected state within a measurable gap size range was limited to short intervals on the scale of seconds because gap adjustment was manual. The current–time profile obtained from this experiment is shown in Fig. 4a. After low-pass filtering, downsampling, and further processing through the OpenNanopore v1.2 software tool, 203 events were detected across the 2.7 s signal. By subsequently applying a 6σ threshold below the inferred mean of the event tunneling currents, 12 events were identified as outliers and removed, leaving 191 events to be analyzed (Fig. 4b). As observed by the current and time histograms in Fig. 4c and 4d, detected events had average current signals of 40.21 pA ± 6.12 pA and durations of 2.30 ms ± 2.21 ms. Extrapolation of this experimental result suggests a sensitivity of 1 nM or <100 ppt in 5 min, corresponding to 1 pmol/g. Lastly, the determined conductance was compared to measurements by Ohshiro et al. (2014) using the 0.55 nm and 0.70 nm nanogap electrodes to test for L-proline; Fig. 4e suggests that the gap size produced by the manual micrometer was intermediate between the sizes tested by Ohshiro et al. (2014). These results confirm the relationship between the gap distance and tunneling current previously outlined by Tsutsui et al. (2008b), where a larger gap distance generates lower tunneling currents and, therefore, conductance.
Linear regression models
Supplementary Table S3 summarizes the statistics obtained from the linear regression models, fitted by plotting conductance and dwell time values of amino acids against the specific amino acid's molecular weight, molecular volume, and HOMO–LUMO gap. The HOMO–LUMO gap showed the best linear relationship to the conductance and dwell time values obtained by the 0.70 nm (Fig. 5a) and the 0.55 nm nanogap (Fig. 5b) electrodes of all tested covariates. However, only the linear relationship observed between the HOMO–LUMO gap and the conductance values of amino acids obtained from the 0.70 nm nanogap electrodes is interpreted as statistically significant. Lower HOMO–LUMO gap values of the proteinogenic amino acids analyzed with the 0.70 nm nanogap electrodes resulted in higher conductance values. Furthermore, no statistically significant relationship was observed between the dwell time and the other physical properties tested. Supplementary Figs. S2 and S3 showcase the remaining linear regression plots obtained from Section 2.5.
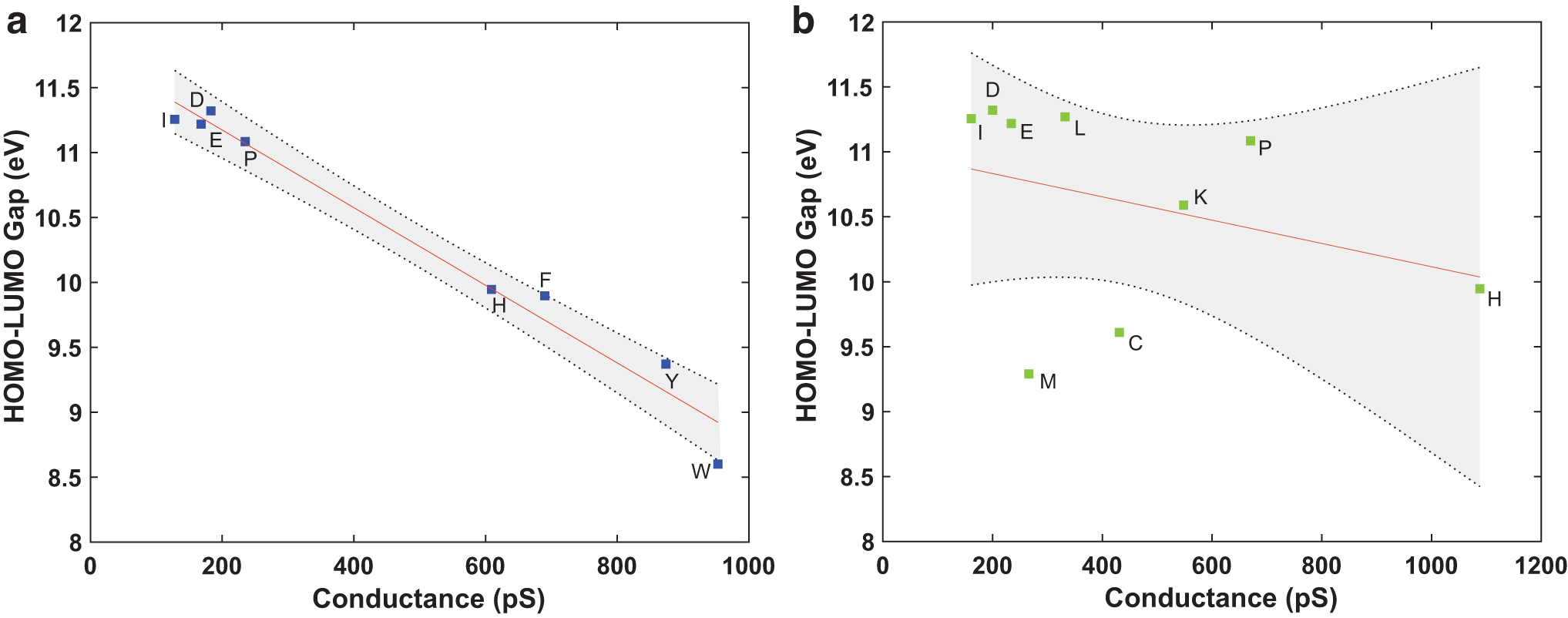
HOMO–LUMO gap of amino acids as a defining property for conductance levels using higher nanogap sizes. (
Event rates are estimated from the abundance of each amino acid. The conductance-event rate plots of amino acids by sample and category predict event rates that differ by orders of magnitude between meteorite samples and marine and land environments with high biological productivity (Fig. 6; Supplementary Fig. S4a, S4b). An overlap is observed in the event rates between the carbonaceous chondrite samples and those from environmental samples with low biological productivity and harsh conditions like Antarctica and the Atacama Desert. However, as depicted in Supplementary Table S4, some of the abundances of ɑ-amino acids within the carbonaceous chondrites are the result of terrestrial contamination. Furthermore, an overlap of estimated event rates is observed between all the higher HOMO–LUMO gap amino acids in the genetic code (Fig. 6), which also coincide with those α-amino acids found in meteorites, following the meteoritic reported amino acids gathered in the database. In summary, while there is overlap in the total event rates estimated, high conductance (low HOMO–LUMO gap) events are diagnostic for life (Fig. 6).
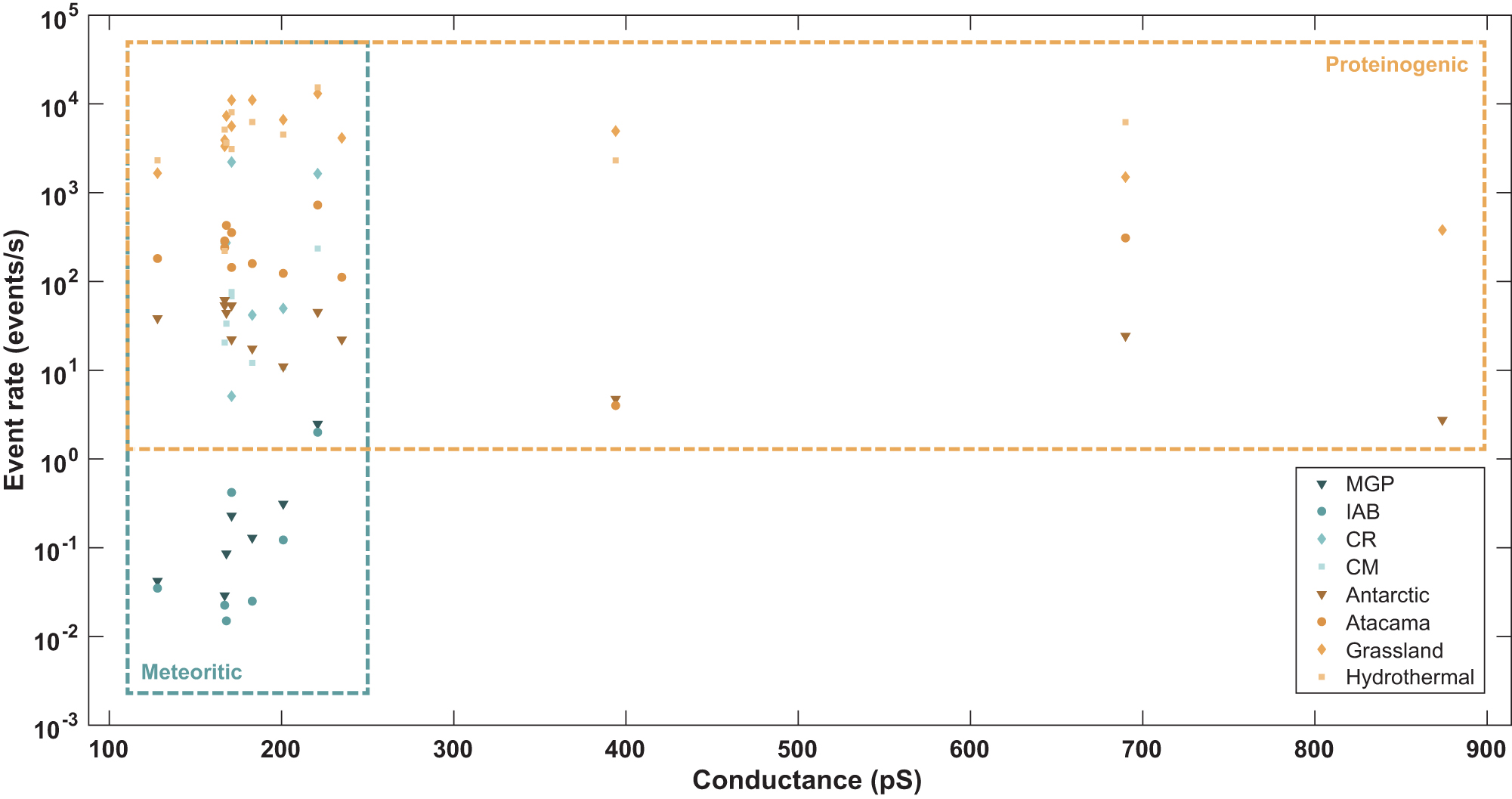
Potential discrimination between biotically and abiotically derived ɑ-amino acids. Distribution of predicted and experimentally validated conductance values of the 13 amino acids considered within the database as a function of the estimated event rates expected to be recorded from ELIE, on each of the meteorite (blue shades) and environmental (orange shades) samples. Meteorite group key: MGP, Main Group Pallasites; IAB, Iron meteorites group AB; CR, Renazzo-type chondrites; CM, Mighei-type chondrites.
Nanogap technology for single-molecule detection and potential for space compatibility
Improving levels of detection toward single-molecule sensitivity, in the picomol/g range, remains an ongoing goal for in situ life-detection technologies. This is a significant issue because signatures of life or abiotically produced organics in other planetary bodies across our solar system may be present at exceedingly low abundances, or only accessible in low abundances (Eigenbrode et al., 2021). Here, we demonstrate that nanogap systems show great potential in this critical regard by performing as ultra-sensitive agnostic detectors without sacrificing the capacity to discriminate among molecules or chemical moieties (such as between different amino acids or DNA bases). While further work is required to rigorously evaluate the analytical sensitivity and limit of detection, the extrapolated sensitivity meets thresholds proposed for organics in the Europa Lander notional Organic Compositional Analyzer baseline payload (Hand et al., 2017) and amino acids in the Enceladus Orbilander notional separation mass spectrometer and microcapillary electrophoresis with laser-induced fluorescence (MacKenzie et al., 2021). In comparison to other state-of-the-art instruments, such as gas chromatography–mass spectrometry (GC-MS) or capillary electrophoresis coupled to laser-induced fluorescence (CE-LIF), nanogap systems do not require derivatization agents for detection and separation of the targeted biomarkers. Many derivatization agents have shown preferential reactions with water that produce increased background signals and low derivatization efficiencies of the targeted molecules (Leshin et al., 2013; Goesmann et al., 2017) and/or form by-products that interfere with the targeted molecules at low concentrations (Stalport et al., 2012; Creamer et al., 2017). Furthermore, the nanogap adjustment mechanism provides a general detection approach of nanometer-scale molecules (e.g., individual amino acids) or moieties (e.g., nucleobases) that are part of longer biomarkers, such as peptides, nucleic acids, or other polymers.
HOMO–LUMO gap as an indicator of amino acid biogenicity
Linear regression models, obtained by using data of single-molecule conductance measurements from nanogap sensing experiments of single amino acids (Ohshiro et al., 2014), demonstrated a relationship between the level of conductance and the HOMO–LUMO gap values of 8 proteinogenic amino acids at higher nanogap sizes. Higher conductance values are obtained for amino acids with lower HOMO–LUMO gap values; a quantum property that defines the energy difference between a molecule's highest energy-occupied molecular orbital (HOMO) and lowest energy-unoccupied molecular orbital (LUMO) in the ground state. These two molecular orbitals underpin frontier molecular orbital theory (Fukui et al., 1954; Hoffmann, 1971; Fukui, 1982), a practical model used to describe the reactivity and regioselectivity of, and thus the stability of, chemical systems. The theory shows that the reaction rate is determined by the energy gap between the HOMO of the nucleophile and the LUMO of the electrophile; therefore, smaller energy gaps would result in greater attraction between these two molecular orbitals and a faster reaction toward a transition state. Based on empirical measurements, we can postulate an analogy to the nanogap, in which a molecule in the gap acts as both an “electrophile” and a “nucleophile,” in that it is both accepting and donating electrons, making the HOMO-LUMO gap relevant.
This relationship between the HOMO–LUMO gap of molecules and their chemical reactivity may reflect the importance of electron transfer as an evolutionary adaptation of life as we know it. Electron transfer pathways are essential for metabolism, energy storage, biosynthesis, metal ligation, and counteracting changes in the environment, such as coping with oxidative stress by cells that initially evolved to thrive in anoxic environments (Cavalier-Smith, 2006; Weiss et al., 2016; Moosmann, 2021). Comprehensive analyses considering multifactor theories, complexity frameworks, protein compositions, concentrations in extraterrestrial and early Earth–simulated environments, and thermodynamic properties all suggest that most proteinogenic amino acids with low HOMO–LUMO gaps (e.g., histidine, phenylalanine, tyrosine, and tryptophan) should have been incorporated late into the genetic code (Fig. 7 and Supplementary Fig. S5a–S5d) (Trifonov, 2000; Higgs and Pudritz, 2009; Meringer et al., 2013; Marshall et al., 2017). This may have enabled increasingly reactive amino acids to act as organic semiconductors, mediated by the incorporation of aromatic systems and less electronegative heteroatoms into their structures (Fig. 8) (Bender et al., 2008; Gray and Winkler, 2015). The catalytic propensity of amino acids (Bartlett et al., 2002) has been previously described as a potential approach in the search for extraterrestrial life (Georgiou, 2018). In contrast (e.g., to Fig. S5a–S5d), proteinogenic amino acids with low HOMO–LUMO gaps show both high and low catalytic propensity (Fig. S5e). This suggests that catalytic propensity, as a feature of life, could represent a complementary measure to the HOMO–LUMO gap.
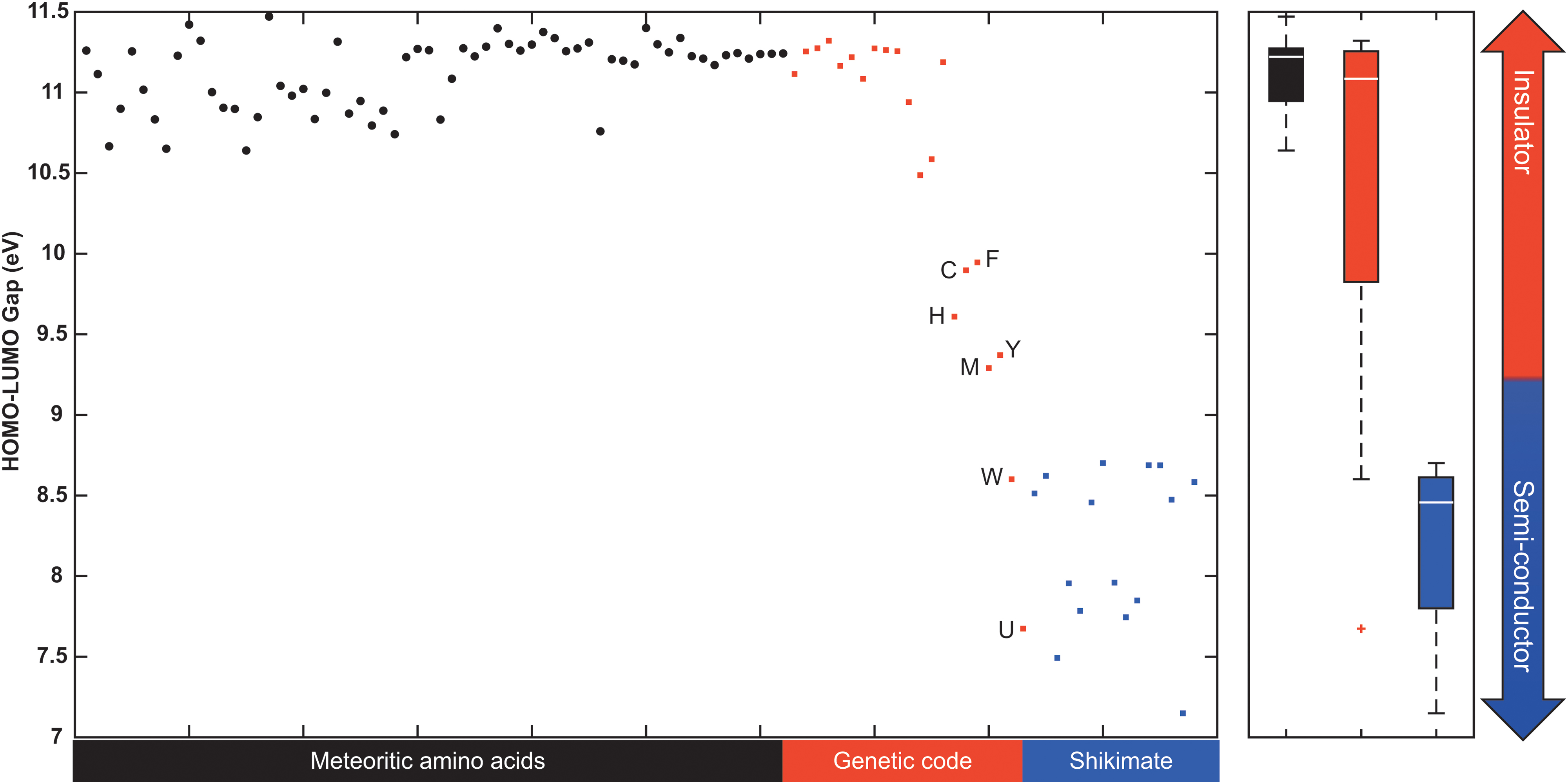
Amino acid HOMO–LUMO gaps. Distributions of the HOMO–LUMO gaps of 62 Murchison meteorite amino acids, 21 genetically encoded amino acids, and various metabolic descendants of the shikimate pathway (a biosynthetic pathway that leads to the formation of F, Y, and W). The quantum-chemical properties were determined by Granold et al. (2018) using semiempirical calculations with the MOPAC 2003 AM1 software package. The meteoritic amino acids are plotted based on the number of carbon atoms within their structure. The 21 proteinogenic amino acids (including selenocysteine) are plotted in the order of their evolutionary appearance according to Trifonov (2000), derived from 40 criteria considering multifactor theories, experimental data simulating primordial conditions, and amino acid complexity, among others. The metabolic descendants of the shikimate pathway are plotted in alphabetical order. Late additions to the genetic code are labeled with their respective one-letter code (F: phenylalanine; H: histidine; C: cysteine; Y: tyrosine, M: methionine; W: tryptophan, U: selenocysteine). Refer to Supplementary Dataset S3 for individual names and HOMO–LUMO gaps of each amino acid and metabolic descendant of the shikimate pathway.
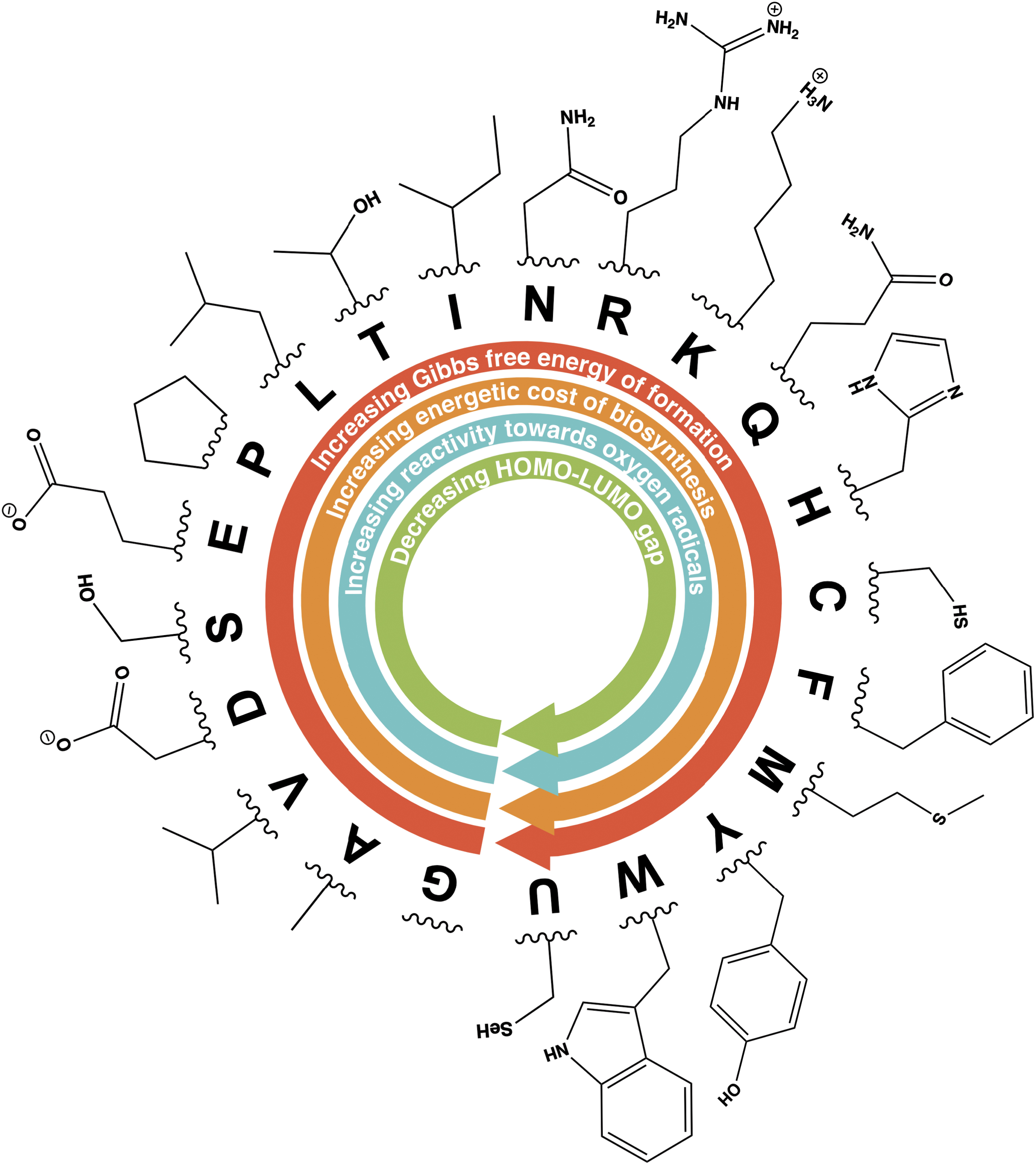
Predicted order of appearance of proteinogenic amino acids within the genetic code by Trifonov (2000), along with their functional groups. Arrows show increasing and decreasing patterns of some amino acids quantum, redox, and thermodynamic properties as discussed and reported by Akashi and Gojobori (2002), Amend and Shock (1998), and Granold et al. (2018). Arrows indicate trends with quantitative support (Supplementary Fig. S10). One-letter amino acid designations are provided in Supplementary Dataset S3.
From a quantum physical context, the relationship between the level of conductance and the HOMO–LUMO gap in solid-state single-molecule detectors is attributed to the energy barrier controlling the electron transport within molecular junctions. The band structures of the gold nanojunctions within the nanogap chip are composed of crystal orbitals that conduct electrons at the Fermi level, a continuous energy level at room temperature (Taniguchi, 2017). Once these orbitals couple with another molecule passing through the nanogap, electronic interactions occur, shifting electron transmission within the nanogap and, in turn, generating changes in the conductance band that are mainly dependent on the energy barrier: the energy difference between the Fermi level and the HOMO or LUMO. Molecules with lower HOMO–LUMO gaps tend to have their HOMO (or LUMO) orbital overlapping to a greater extent with the Fermi level of the nanoelectrodes, thus permitting greater electron transmission (Taniguchi, 2019). Therefore, this quantum property of amino acids explains the trends observed in the resulting conductance of amino acids sensed by the nanogap system. Here we have found that the HOMO–LUMO gap is an accurate predictor of amino acid conductance, and thus amino acid complexity, in nanogap systems at nanogap sizes that are significantly higher than the targeted molecules (Fig. 5). In contrast, this relationship with conductance is not true for the mass and spatial size (molecular volume) of amino acids (Supplementary Fig. S2), which seem to be the determining mechanisms responsible for electrical sensing via ionic current blockade in biological and solid-state nanopores (Wei et al., 2020).
In contrast, no statistically significant relationship was observed between the dwell time from the nanogap sensing experiments of single amino acids by Ohshiro et al. (2014) and the proteinogenic amino acid's physical and quantum properties considered in this study (Supplementary Dataset S1 and Fig. S3). However, as seen in Supplementary Fig. S6, there is an evident pattern in the dwell time at lower nanogap sizes, in which dwell time decreases in relation to the functional group of the amino acids tested. This can be explained by the electronic coupling (bond) strength between the molecule and the electrodes, the other major property influencing electron transport through the molecular junction, which is primarily determined by the chemical nature of the anchoring groups to the nanoelectrodes (Huang et al., 2015). It has been previously reported that thiol and amine groups form strong-stable and weak-covalent bonds on gold surfaces, respectively (Leff et al., 1996; Xue et al., 2014). Furthermore, imidazole groups are known to coordinate with gold nanoparticles and contribute significantly to interactions between metallic surfaces and biological filaments (Zhou et al., 2010). These observations suggest that the bond strength between the gold atoms and amino acids like histidine, cysteine, and lysine allow for longer periods of single-molecular junctions at low nanogap sizes. Also, the observed trend at lower nanogap sizes (thiol > amine > carboxyl) agrees with previous works studying the single-molecule conductance of alkenes and diphenylacetylene-type molecules with different anchoring groups (Chen et al., 2006; Hong et al., 2012). It has also been argued that the breakage of the gold contacts when the nanogap is being formed can lead to different rearrangements of gold atoms within each electrode, which can ultimately impact the coupling strength with a passing molecule (Arroyo et al., 2011). Further studies are needed to assess other factors (e.g., electrical potential, molecular volume) and their possible combinatory influence on the dwell time at longer nanogap sizes. These results also highlight the need for automated gap control in future ELIE systems to maintain the gap at optimal distances for the molecular junction to be formed with the targeted molecules.
Detection of low HOMO–LUMO gap small molecules may be indicative of life, but such molecules are not unique to living systems. Some amino acids in the genetic code with lower HOMO–LUMO gaps (<10 eV; 5σ below the mean HOMO–LUMO gap of Murchison meteorite amino acids, as reported by Granold et al. [2018]) have been reported to be produced abiotically in terrestrial environments and endogenously in extraterrestrial materials. Ménez et al. (2018) demonstrated the abiotic formation of tryptophan, an aromatic α-amino acid with a low HOMO–LUMO gap, during hydrothermal alteration of mantle rock in the Atlantis Massif. Moreover, Friedel-Crafts-type reactions are predicted to be responsible for the formation of this amino acid during the hydrothermal alteration of oceanic peridotites; a different pathway from the Strecker-cyanohydrin synthesis, which has been predominantly supported for the formation of endogenous α-amino acids in CM and CR chondrites (Ehrenfreund and Charnley, 2000; Martins et al., 2007; Glavin et al., 2020).
A capability of the ELIE instrument in development is to assess the abundance distributions of α-amino acids within the sample and thereby assess the abiotic or biotic origin of these targeted molecules. Because the abiotic production of amino acids is thermodynamically driven (Higgs and Pudritz, 2009), lower molecular weight amino acids, with lower values of Gibbs free energy of formation (Fig. 8), are often found in high abundances within abiotic samples (Supplementary Fig. S7). In contrast, the biotic production of amino acids is driven by functionality, yielding higher abundances of high molecular weight species with low HOMO–LUMO gaps (e.g., phenylalanine and tyrosine) relative to lower molecular weight amino acids. Furthermore, analysis of the HOMO–LUMO gaps of the 62 meteoritic amino acids analyzed by Granold et al. (2018), of which only 16 are known to occur in biology, as a function of an increasing number of carbon atoms per molecule, reveals a convergence to a high HOMO–LUMO gap (Supplementary Fig. S8). This is in opposition to the amino acids within the genetic code, where the HOMO–LUMO gap varies significantly yet trends to lower HOMO–LUMO gap values with more carbon atoms (Supplementary Fig. S9). This further suggests that amino acids with lower HOMO–LUMO gaps were probably incorporated into the genetic code as the result of biological pressures. Therefore, if the mechanisms of formation mentioned above are ruled out, and the abundances of low HOMO–LUMO gap amino acids are high, then such detections (many events at high conductance) would be consistent with a potential biological source of amino acids or other small molecules.
Advancing ELIE
Future work includes evaluating ELIE's potential for the detection of biomolecular mixtures, nonstandard nucleic acid bases, and other polymers to expand the structural diversity scope of detection and reduce the risk of false negatives. This will allow the development of a biomolecule detection/characterization algorithm with the potential for molecule classification using supervised and unsupervised methods (Taniguchi et al., 2019). Furthermore, we seek to develop ELIE into a breadboard system with long-term expectations of being applicable to life-detection missions in Ocean Worlds, satisfying the mass, power, and volume constraints needed for such missions. To reach these next steps, we are improving some of the instrument's current limitations by (i) integrating an automated piezoactuator for gap control, (ii) optimizing the Mechanically Controlled Break Junction process, (iii) characterizing gap quality through monitoring of quantized gap conductance, (iv) automating sample handling, and (v) reducing the size of the low-noise amplifier. These improvements will allow us to better assess the performance of ELIE in relevant environments, including the influence of salt, vacuum exposure, vibration tolerance, and the effects of space radiation.
Conclusions
The discovery of a second genesis of life on Europa or Enceladus would immediately suggest that life is common in the universe, addressing fundamental questions about life's origins and distribution across the universe, which would change humanity's perspective. The present efforts are a proof of concept of an instrument capable of detecting single amino acids, at a sensitivity that is not currently feasible for most of the emerging technologies for in situ amino acid detection proposed for planetary missions to Ocean Worlds. The detection of the amino acid L-proline was validated in the μM range, and extrapolation suggests an ultimate sensitivity in the nM (pmol/g) range. Furthermore, the quantum property of the HOMO–LUMO gap is shown to best describe the conductance levels of proteinogenic amino acids and, thus, proposed as a novel approach to measure amino acid complexity. Finally, by predicting event rates based on amino acid distributions in terrestrial and extraterrestrial samples, we suggest a potential approach to distinguish between biotically and abiotically derived amino acid distributions. Ongoing efforts aim to extend ELIE's capabilities for the detection of other biomolecules and to develop the instrument into a breadboard system for potential opportunities for infusion into Ocean Worlds missions.
Footnotes
Acknowledgments
We thank Bernd Moosmann for providing HOMO–LUMO gap estimates for the Murchison meteorite amino acids, the genetically encoded amino acids, and the various metabolic descendants of the shikimate pathway and the data of the peroxyl radical scavenging assays. We are grateful to the reviewers for their helpful and detailed comments that strengthened the manuscript. This paper was previously submitted to the BioXriv preprint server at
Author Contributions
C.E.C. and J.L.R.C. conceived and wrote the paper; C.E.C. collected the data; C.E.C. and S.L. built the hardware; M.T., T.O., and Y.L. provided the nanogap sensors; C.E.C and D.D. designed the L-proline experiment; all authors edited and approved the manuscript.
Author Disclosure Statement
The authors declare no competing interests.
Funding
This work was supported by NASA awards 80NSSC19K1028 and 80NSSC22K0188 to C.E.C.
Data Availability
Raw data and scripts are available at
Supplementary Material
Supplementary Dataset S1
Supplementary Dataset S2
Supplementary Dataset S3
Supplementary Dataset S4
Supplementary Dataset S5
Supplementary Dataset S6
Supplementary Dataset S7
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
Supplementary Table S5
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Figure S6
Supplementary Figure S7
Supplementary Figure S8
Supplementary Figure S9
Supplementary Figure S10
Abbreviations Used
Associate Editor: Christopher McKay
