Abstract
Many drug concentration-effect relationships are described by nonlinear sigmoid models. The 4-parameter Hill model, which belongs to this class, is commonly used. An experimental design is essential to accurately estimate the parameters of the model. In this report we investigate properties of D-optimal designs. D-optimal designs minimize the volume of the confidence region for the parameter estimates or, equivalently, minimize the determinant of the variance-covariance matrix of the estimated parameters. It is assumed that the variance of the random error is proportional to some power of the response. To generate D-optimal designs one needs to assume the values of the parameters. Even when these preliminary guesses about the parameter values are appreciably different from the true values of the parameters, the D-optimal designs produce satisfactory results. This property of D-optimal designs is called robustness. It can be quantified by using D-efficiency. A five-point design consisting of four D-optimal points and an extra fifth point is introduced with the goals to increase robustness and to better characterize the middle part of the Hill curve. Four-point D-optimal designs are then compared to five-point designs and to log-spread designs, both theoretically and practically with laboratory experiments.
D-optimal designs proved themselves to be practical and useful when the true underlying model is known, when good prior knowledge of parameters is available, and when experimental units are dear. The goal of this report is to give the practitioner a better understanding for D-optimal designs as a useful tool for the routine planning of laboratory experiments.
Many drug concentration-effect relationships are described by nonlinear sigmoid models. The 4-parameter Hill model, which belongs to this class, has been commonly used to characterize the concentration dependence of many biochemical, physiological and pharmacological responses (Holford and Sheiner, 1981). In our experience with several hundred concentration-effect laboratory experiments, the Hill model has been found to fit data exceedingly well. Parameter estimation with the Hill model is a special case of the general problem of parameter estimation in nonlinear regression models (Box and Lucas, 1959; Bates and Watts, 1988; Seber and Wild, 1989). Both the method of estimation (Sheiner and Beal, 1985; Giltinan and Ruppert, 1989; Amisaki and Eguchi, 1999) and the experimental design (Merle and Mentre, 1997) are essential in order to obtain reliable estimates of the pharmacokinetic parameters. While the two components of a sound estimation procedure are intimately related, we focus on the design part in this report. More specifically, we investigate properties of D-optimal designs (Atkinson and Donev, 1992). D-optimality is a popular criterion since it is geometrically intuitive when a model is linear. In this case the confidence region for the model's parameters is ellipsoidal. A D-optimal design minimizes the content of this confidence region and so minimizes the volume of the ellipsoid. A-optimality and E-optimality are two other criteria directly related to the shape of the ellipsoid. The two criteria are algebraically expressed in terms of the lengths of the axes of the confidence ellipsoid. G-optimality is yet another criterion. While it is not directly linked to the confidence ellipsoid, G-optimality is intimately related to D-optimality. The celebrated General Equivalence Theorem establishes this relationship and thus offers a useful tool to test for D-optimality. There exist also other criteria of optimality along with the ones mentioned above. An advantage of D-optimality is that the optimal designs do not depend upon the scale of the variables. Linear transformations do not change D-optimal designs, which is not in general true for A- and E-optimal designs. In nonlinear regression, a linear approximation to the nonlinear model is used. Unlike in a linear case, D-optimal designs for nonlinear models generally depend on the assumed parameter values. However, even when these preliminary guesses about the parameter values are appreciably different from the true values of the parameters the D-optimal designs produce satisfactory results. This property of D-optimal designs is called robustness and can be quantified by D-efficiency. We show that D-optimal designs, pure or modified, are acceptably robust with respect to the deviation of the prediction of the parameter vector from the true one. Our work extends research on optimal designs for the 3-parameter Hill model by Endrenyl and his group (Bezeau and Endrenyi, 1986; Endrenyi et al., 1987). The goal of this paper is to give the practitioner a better understanding for D-optimal designs as a useful tool for the routine planning of laboratory experiments.
THEORETICAL SECTION
We shall assume that the relationship between observed responses

where ɛ

Graph of the 4-parameter Hill model. The following parameter values have been assumed:
We assume that in Eq. 1 there are no systematic errors, which means that the expected values of the observations are the true responses,

Here the parameter λ is a nonnegative real number, is the proportionality parameter. Constant variance is implied when λ = 0, whereas λ = 1 corresponds to constant coefficient of variation. The variance of the Poisson distribution behaves as (2) with λ = 0.5. The power model (2) is commonly used for heteroscedastic regression modeling in pharmacokinetics. Our laboratory experience with several hundred concentration-effect experiments confirmed the appropriateness of (2) for modeling data variation (Levasseur, et al., 1995; Levasseur et al., 1998).
Once (2) has been assumed to be an appropriate random model for data variation, then

The weighting matrix appears in the equations below to account for heteroscedasticity. Clearly, in a homoscedastic situation, this matrix becomes the identity matrix.
The volume of the ellipsoidal confidence region is proportional to the determinant of the variance-covariance matrix of the estimated parameters. The information matrix

Here ξ = (
Define
For many D-optimal designs, the design contains

where
The D-efficiency of any design ξ is computed according to the following formula:

The reciprocal of D-efficiency can be thought of as the factor by which a given design ξ should be replicated in order to achieve a precision of parameter estimates equal to that of D-optimal design. For example, 2 replicates of a design with D-efficiency of 0.5 are needed to achieve the same precision as that of the D-optimal design ξ*.
When a maximization procedure is used to satisfy D-optimality criterion (Eq. 4), one needs to be aware that a local, rather than a global, maximum could be reached. One way to ascertain that a found candidate design is indeed D-optimal is to apply the General Equivalence Theorem. In order to state it, we introduce G-optimality. The objective of a G-optimal design is to minimize the standardized variance of the predicted response (the weighting factor being accounted for): ξ* is called G-optimal if
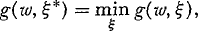
where

Here
A sufficient condition for ξ* to satisfy (Eq. (7)) is

The General Equivalence Theorem states that conditions (4), (7), and (9) are equivalent.
The condition (9) means that the function
Thus the General Equivalence Theorem is of great practical importance because it can be invoked to confirm D-optimality.
METHODS
Our work was aimed at generating D-optimal designs for the model described by (1–2) and then investigating properties of such designs. The effort was focused on finding practical constrained designs rather than unconstrained ones. Therefore, our design region is a closed interval 0 ≤
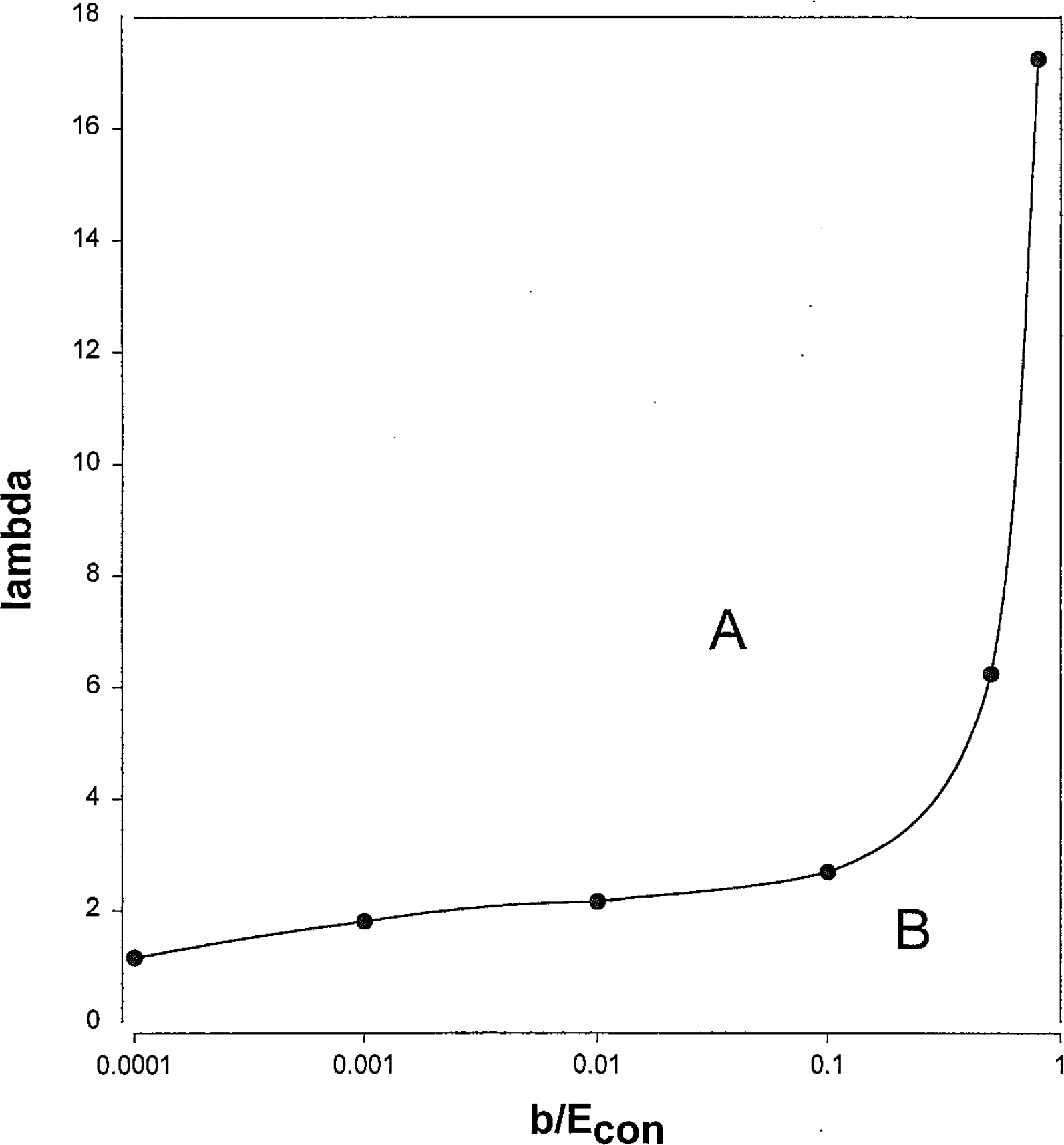
Two regions: A, conforming to the D-optimal designs not having
Figure 3 illustrates what happens to the D-optimal points when changes are made to one parameter at a time. The graph in the upper left corner of the figure conforms to

Influence of changing parameters on 4 D-optimal design points. The values are changed one at a time starting with a graph in the upper left corner of the figure that conforms to
Along with the 4-point D-optimal design, we explored two five-point designs which we call design A and design B, respectively. We intended to create a more robust design by adding a fifth additional point. Design A consists of the four D-optimal points plus an additional one computed as a geometric mean of the two middle D-optimal points. Design B merely replicates one of the D-optimal points (dose2 in Table 1). When all of the assumed parameter values are the true ones, the D-efficiencies for the 3 designs are: 4-point D-optimal, 1.00; 5-point design B, 0.951, 5-point design A, varies from about 0.93 to 0.94, depending upon the set of true parameters.
D-optimal designs used in laboratory experiments a

Assumed parameter values are:
Dose2 and dose3 are two middle points in the 4-point D-optimal design. Dose5 is the fifth point used in design A together with the 4 D-optimal points.
Dmax = 1500 has been used in this case because a concentration as high as Dmax = 111000 (1000 times the
D-optimal designs were used in real laboratory cell growth inhibition studies with each of seven anticancer drugs combined with the concentration of 78 μM of folic acid (FA) in the medium. Concentration-effect experiments were conducted in a 96-well plate growth inhibition assay (Levasseur et al., 1995). Briefly, exponentially growing HCT-8 (human ileocecal adenocarcinoma) cells were plated in wells with RPMI 1640 medium supplemented with 10% dialysed horse serum on day 0, treated on day 1 and incubated at 37°C in a 5% humidified atmosphere. Treatments were randomized on each plate. Cell growth was measured on day 5 with the SRB protein dye assay.
Methotrexate (MTX) and Trimetrexate (TMTX) are inhibitors of the enzyme, dihydrofolate reductase. Tomudex (ZD1694) is an inhibitor of thymidylate synthase, and was a gift from Zeneca Pharmaceuticals (Macclesfield, England). AG2032 and AG2034 (inhibitors of glycinamide ribonucleotide formyltransferase), AG2009 (inhibitor of aminoimidazolecarbox-amide ribonucleotide formyltransferase) and AG337 (inhibitor of thymidylate synthase) were gifts from Agouron Pharmaceuticals, Inc. (San Diego, CA).
Assumed true parameter values for D-optimal design calculations were taken from previously-completed concentration-effect experiments. For each previous experiment, A. was estimated by fitting (2), after logarithmically transforming both sides of the equation, with unweighted linear regression, to the variances and means of each set of replicates at each design point. Also, for each experiment the
RESULTS
Table 2 gives the D-efficiencies of the 4-point D-optimal design, five-point design A, and a log-spread design, respectively, assuming that: (a) our prior estimates of the parameter values that we guessed to be the true ones were in fact the true values; or (b) the analysis of data from the log-spread design resulted in the true parameter values (although the actual design points are the same as in (a)). The 4-point D-optimal design included 1 control point at
D-efficiencies of a set of designs for anticancer drug concentration-effect curves
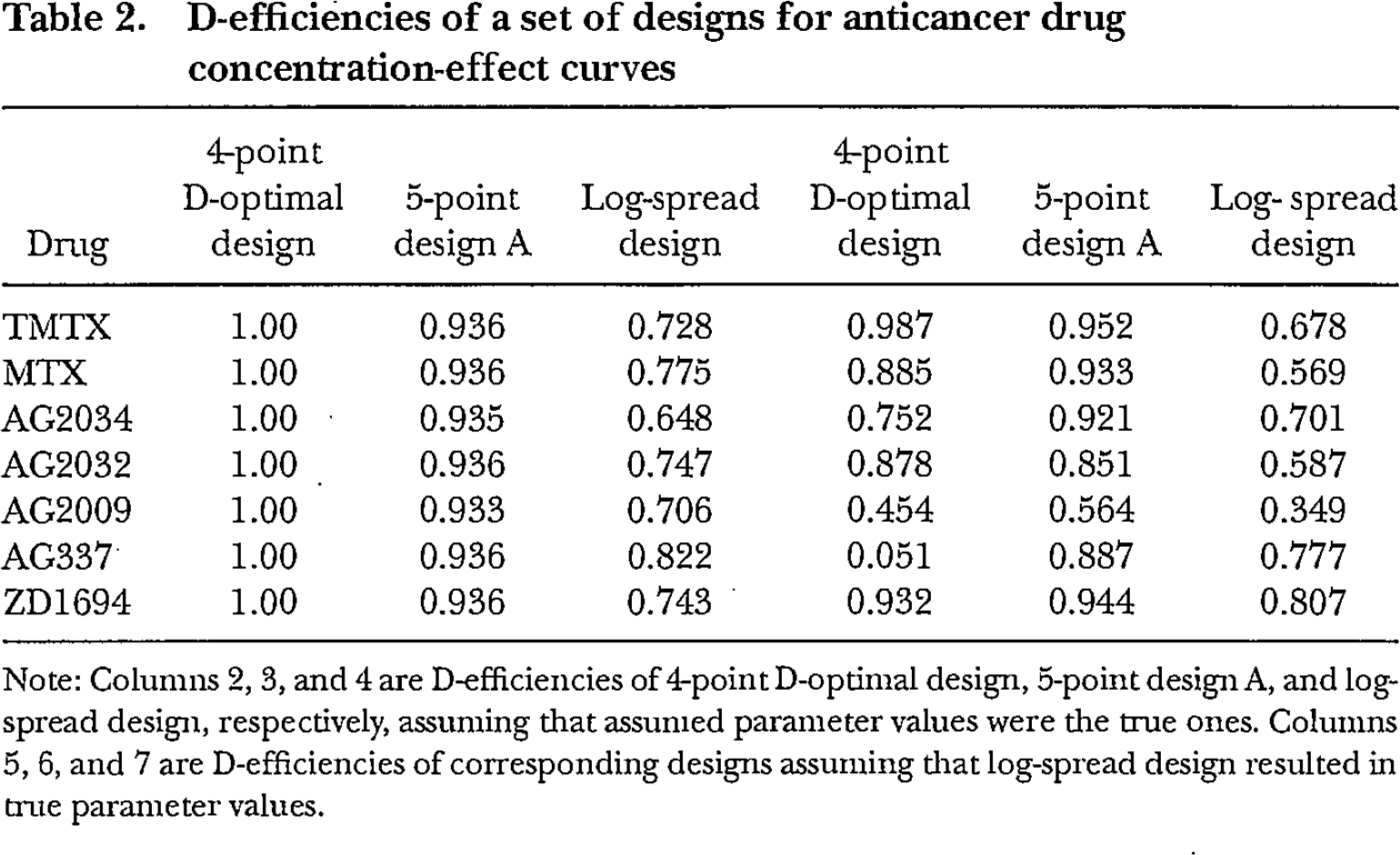
Note: Columns 2, 3, and 4 are D-efficiencies of 4-point D-optimal design, 5-point design A, and log-spread design, respectively, assuming that assumed parameter values were the true ones. Columns 5, 6, and 7 are D-efficiencies of corresponding designs assuming that log-spread design resulted in true parameter values.
For the real laboratory experiments, the 4-point D-optimal design included 10 replicates per design point, for a total of 40 data points per experiment; the 5-point design A included 10 replicates per design point for a total of 50 data points per experiment; and the 12-point log-spread design included 5 replicates per design point for a total of 60 data points per experiment. The estimates of the parameters needed to generate the D-optimal designs for these experiments were pooled from the results of extensive past studies conducted by our group on the same drug and cell line. The data for each of the 7 drugs for 3 designs (21 total experiments) were analyzed by fitting model (1) to data with iteratively-reweighted nonlinear regression as described above. The fitted concentration-effect curves for three compounds, MTX, TMTX, and AG2009, with 78 μM folic acid in the medium, for the 3 designs, are displayed in Figure 4. MTX is a representative example of a drug with a relatively steep concentration-effect curve, AG2009 has a very shallow curve, and TMTX has a curve with intermediate steepness. Note that for the log-spread design, for MTX only 1 design point is located on the falling part of the curve, for TMTX there are 3 points, and for AG2009 there are 5–7 points. Overall, the parameters were well estimated; the standard errors were around 10% or less of the parameter estimates. Exceptions include AG337 for the 4-point D-optimal design. For these 2 cases, the design points missed the falling portion of the concentration-effect curve. Overall, the corresponding parameter estimates are very similar among the 3 designs (Table 3) and the previous experiments.

The fitted concentration-effect curves for three compounds, methotrexate, trimetrexate, and AG2009. Curves are displayed, along with the mean observed responses and standard deviations at the design points. Solid points represent observed data. Open triangles represent the predicted design points. Open squares represent control responses (always the first D-optimal design point).
Parameters estimated from laboratory experiments

Replication of design points is important in creating practical designs. It is known from theory (Bates and Watts, 1988; Seber and Wild, 1989) that the standard error of an individual parameter estimate decreases proportionally to
DISCUSSION
In this paper we examined different designs for the estimation of parameters in the Hill model. We assume that the true underlying model is known and that it is the Hill model. We also assume that the variance component is given by the power model (2). Our laboratory experience supports these assumptions. One should be aware that these assumptions may not hold for a different type of data. For instance, the binomial variance model σ2i =
Both
