Abstract
There is substantial evidence that inflammation within the central nervous system contributes to stroke risk and recovery. Inflammatory conditions increase stroke risk, and the inflammatory response is of major importance in recovery and healing processes after stroke. We investigated the role of inflammatory genes IL1B, IL6, MPO, and TNF in stroke susceptibility and recovery in a population sample of 672 patients and 530 controls, adjusting for demographic, clinical and lifestyle risk factors, and stroke severity parameters. We also considered the likely complexity of inflammatory mechanisms in stroke, by assessing the combined effects of multiple genes. Two interleukin 6 (IL6) and one myeloperoxidase (MPO) single-nucleotide polymorphisms were significantly associated with stroke risk (0.022correctedP < 0.042), highlighting gene variants of low to moderate effect in stroke risk. An epistatic interaction between the IL6 and MPO genes was also identified in association with stroke susceptibility (P = 0.031 after 1,000 permutations). In a subset of 546 patients, one IL6 haplotype was associated with stroke outcome at 3 months (correctedP = 0.024), an intriguing finding warranting further validation. Our findings support the association of the IL6 gene and present novel evidence for the involvement of MPO in stroke susceptibility, suggesting a modulation of stroke risk by main gene effects, clinical and lifestyle factors, and gene–gene interactions.
Introduction
The brain was once regarded as an ‘immune privileged’ organ, neither susceptible to inflammation nor affected by systemic inflammatory responses. This view has, however, completely changed, and the brain is nowadays known to exhibit key features of inflammation, such as synthesis of cytokines and glial activation, and to intervene in the regulation of systemic inflammation and in acute phase response after brain injury (reviewed in Lucas et al (2006)). There is also substantial evidence that inflammation within the central nervous system has a role in many brain disorders including stroke, a major cause of death and significant disability in western countries.
Stroke pathophysiology is likely regulated by a combination of environmental/lifestyle and unclear genetic risk factors. Increasingly, research studies are suggesting that inflammation significantly contributes to stroke risk, progression, and outcome (Rodríguez-Yáñez and Castillo, 2008; Wang et al, 2007). For instance, known clinical risk factors for stroke, like atherosclerosis, diabetes, obesity, hypertension, and peripheral infection, are associated with an elevated systemic inflammatory profile (Bastard et al, 2006; Hansson and Libby, 2006; Moutsopoulos and Madianos, 2006). Atherosclerosis, in particular, is an inflammatory disease and a major contributor to stroke, either through thromboembolism, which results from the rupture of atherosclerotic plaques, or indirectly through cardioembolism (Hansson and Libby, 2006).
Inflammation is equally of major importance in the acute phase of stroke and in the recovery process. It is known that the inflammatory response that follows ischemic or hemorrhagic stroke contributes to exacerbate the initial injury, but that neuroprotective and regenerative molecules are secreted at different stages after a stroke event (Correale and Villa, 2004; Lakhan et al, 2009; Lucas et al, 2006). Clearly, inflammation in central nervous system injury in general, and in stroke in particular, cannot be classified straightforwardly as harmful. Although there are many inflammatory mediators with detrimental effects, some can be beneficial and others may have dual roles, suggesting a complex orchestration in the acute and recovery phases after stroke (Lucas et al, 2006).
An inflammatory process is thus implicated in pathological conditions that increase stroke risk, in the injury mechanisms upon stroke and in the recovery pathways that mediate stroke outcome. It is likely that variants of genes encoding inflammatory molecules will influence not only individual stroke risk, but also the extension of the injury and the recovery process, and a number of studies have assessed this hypothesis. For instance, the tumor necrosis factor (TNF) gene has been associated with subarachnoid hemorrhage (Yamada et al, 2006), and polymorphisms in the interleukin 1 beta (IL1B) and interleukin 6 (IL6) genes have been associated with ischemic stroke, and with ischemic stroke and intracerebral hemorrhage, respectively (Bis et al, 2008; Yamada et al, 2006). However, conflicting results have been obtained for the IL6 gene (Tso et al, 2007). Other lines of evidence show that inflammatory molecules influence the extension of injury and the recovery process. Interleukin 1 beta and TNF-alpha are known to be released by neurons and endothelial cells in response to ischemia, initiating an inflammatory response and inducing IL-6 and IL-8, with deleterious consequences (Rodríguez-Yáñez and Castillo, 2008). The myeloperoxidase (MPO) gene is another intriguing candidate, as the encoded enzyme catalyzes the formation of MPO-derived reactive species that may contribute to atherosclerosis progression and destabilization of atherosclerotic plaques (reviewed in Schindhelm et al (2009)). Reinforcing the hypothesis of a role in stroke, MPO polymorphisms have been associated with the size of the brain infarct and functional outcome (Hoy et al, 2003).
In this study, we tested the genetic association of major inflammatory players IL1B (2q14), IL6 (7p21), TNF (6p21.3), and MPO (17q23.1) with stroke susceptibility and stroke outcome at 3 months. The apparent complexity of the inflammatory mechanisms in stroke and the multiplicity of players involved suggest a concerted process, in which implicated molecules interact tightly to regulate each other. Still, nonadditive interactions or epistasis are generally overlooked in genetic studies. Epistasis is a plausible explanation for the lack of replication across different populations in candidate gene studies or in genome-wide association studies, where it is particularly difficult to assess because of the large dimension of the data (Lanktree et al, 2010; Moore, 2003). We, therefore, examined both independent gene effects and the occurrence of gene–gene interactions among the tested inflammatory genes in stroke risk and stroke recovery.
Materials and methods
Study Population
The study population included 672 first-ever stroke patients, recruited through Neurology and Internal Medicine Departments of several hospitals in Portugal. Stroke definition and the protocol for clinical assessment of patients were previously described (Krug et al, 2010; Manso et al, 2010). Five hundred thirty healthy controls with no clinical history of stroke were also enrolled. As stroke is a late-onset disease, we included older healthy individuals to reduce the probability of misclassification as controls. Information on clinical and lifestyle risk factors, matching the data available for patients, was obtained by direct interview of control subjects. A subset of 546 patients was included in the outcome analysis. These patients were classified into two groups based on the modified Rankin Scale (mRS) at 3 months: patients with mRS ≤ 1 were scored as ‘good recovery’ and with mRS > 1 were scored as ‘poor recovery’, as previously described (Manso et al, 2010).
The study was approved by the Ethics Committee of the Portuguese Dr Ricardo Jorge National Institute of Health and other hospitals involved, and participants gave their informed consent.
Single-Nucleotide Polymorphism Genotyping
To tag the genetic variation in the IL1B, IL6, MPO, and TNF gene regions, single-nucleotide polymorphisms (SNPs), located within and up to 5 kb upstream and downstream of those genes, were selected using the H-clust method (Rinaldo et al, 2005) (HapMap Release 21/phase II July 2006). Three SNPs in IL1B, six in IL6, two in MPO, and three in TNF were genotyped using Sequenom iPLEX assays with allele detection by mass spectroscopy, using Sequenom MassARRAY technology (Sequenom, San Diego, CA, USA) and following the manufacturer's protocol. Primer sequences were designed using Sequenom's MassARRAY Assay Design 3.0 software (Sequenom, San Diego, CA, USA). Quality control analyses were performed based on the genotyping of eight HapMap individuals, duplicated samples within and across genotyping plates, Mendelian segregation in three pedigrees and no-template samples. For each SNP, call rate <90% and deviation from Hardy–Weinberg equilibrium (P < 0.05) were checked.
Statistical Analysis
To identify potential confounders, univariate analyses were performed comparing demographic and clinical and lifestyle risk factors between patients and controls, using the Pearson's χ2 test and the Mann–Whitney test for discrete and continuous variables, respectively. Variables with a P < 0.25 in univariate analysis (Table 1) or of particular clinical relevance were included in a logistic regression model using forward selection (Hosmer and Lemeshow, 2000), and were maintained in the model if they were associated with stroke susceptibility at a P ≤ 0.05 level. The selected covariates were not correlated (−0.5 < interaction i < 0.5). Logistic regression analyses 17 were then used to determine the effect of each genetic variable on stroke susceptibility after adjustment for the significant covariates. Odds ratio (OR) and 95% confidence intervals (95% CIs) were computed for the log-additive model. A similar procedure was followed for the analysis of stroke outcome. Demographic and clinical data reflecting the severity of stroke were compared between patients with poor (mRS > 1) and good (mRS ≤ 1) outcome at 3 months to identify potential confounders (Table 2). Logistic regression analyses were then used to determine the effect of each genetic variable on patient's outcome after adjustment for the significant covariates. OR and 95% CI were also computed for the log-additive model.
Demographic and clinical characteristics of the population sample
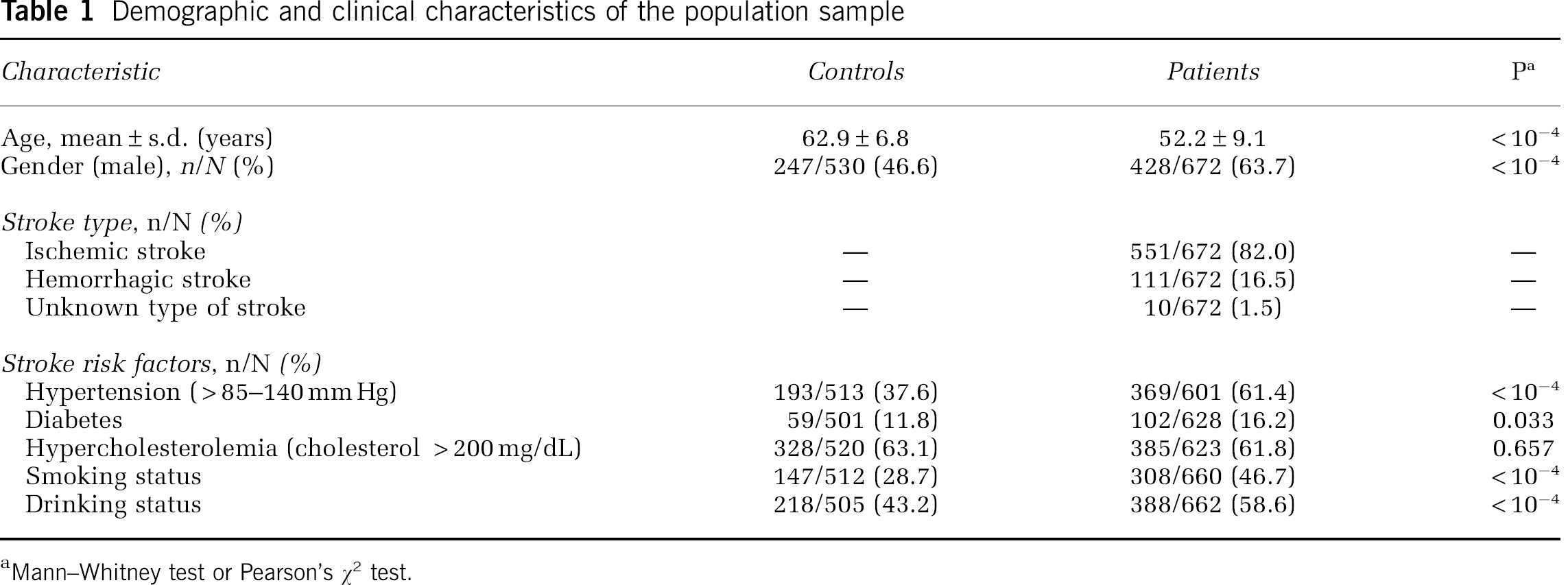
aMann–Whitney test or Pearson's χ2 test.
Demographic and clinical characteristics of stroke patients analyzed for outcome at 3 months
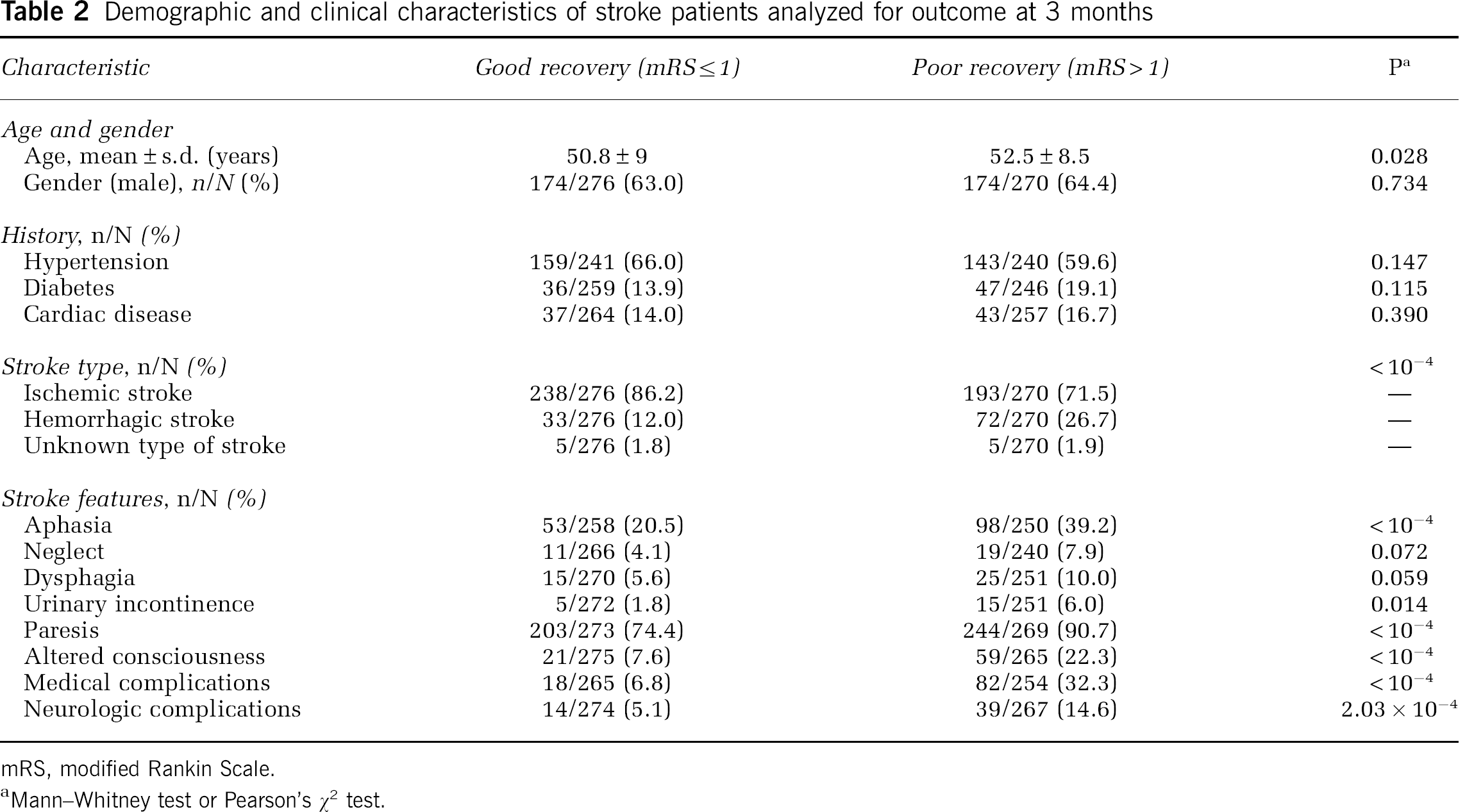
mRS, modified Rankin Scale.
Mann–Whitney test or Pearson's χ2 test.
Univariate and logistic regression analyses were performed using MASS and SNPassoc packages of the R software (R: A language and Environment for Statistical Computing, 2004) (v2.6.0). Haplotype blocks in the four genes were determined using the default method (Gabriel et al, 2002) of the Haploview software (Barrett et al, 2005) (v4.0) and haplotype-based association analyses were performed. This algorithm may select different haplotype blocks for the entire sample and the outcome study subset, as occurred specifically for the IL6 gene, resulting in different haplotypes being tested for association with stroke susceptibility and outcome. Bonferroni correction for multiple testing was used to correct significant associations in individual SNP analysis, as well as haplotype-based association analysis.
Testing for genetic interactions in association with stroke susceptibility and outcome was performed using the multifactor-dimensionality reduction (MDR) method (Ritchie et al, 2001) (v2.0, beta 7.2), a nonparametric and genetic model-free approach. Briefly, by pooling multilocus genotypes into high- and low-risk groups, the MDR reduces the dimensionality of the data from N dimensions to one dimension. The new multilocus genotype attribute is then tested for its ability to classify and predict disease status, or good/poor outcome at 3 months. False-positive results due to multiple testing are reduced through combination of the cross-validation strategy and permutation testing (Moore, 2003; Ritchie et al, 2001). As the MDR method does not accept missing data, we imputed missing genotypes for each SNP using the PLINK software (http://pngu.mgh.harvard.edu/purcell/plink/) (Purcell et al, 2007) and CEU HapMap genotype data were used as reference. Best models for each group of genes were chosen from among the best 2, 3, and 4-factor models, based on the testing balanced accuracy (TBA) and the cross-validation consistency (CVC) of 10 cross-validation intervals. TBA measures how often individuals are correctly classified in relation to disease or outcome status, and CVC measures the number of times the MDR found the same set of loci across the cross-validation subsets. A model with a TBA > 0.6 is almost always statistically significant, while a TBA > 0.55 is considered interesting; TBA = 0.5 is random (http://compgen.blogspot.com/2006/12/mdr-101-part-4-results.html). The statistical significance of the best models was calculated after 1,000 permutations using the MDR Permutation Tool (v1.0, beta 2). Odds ratio-based MDR (OR-MDR) (v1.3-1) was used to determine the OR and 95% CI for each genotype combination as a quantitative measure of disease risk (Chung et al, 2007). To determine the gain in information about disease or outcome status by combining two variables together over that provided by the independent analysis of these variables, the MDR uses entropy measures (Jakulin and Bratko, 2003). Evidence for a synergistic interaction occurs when the combination of two or more SNPs gives a positive information gain. If the information gain is negative, then there is evidence for redundancy or correlation between SNPs; and if the information gain is 0, then the SNPs have independent effects. Entropy-based interaction dendrograms are used for interpreting epistasis models (Moore et al, 2006).
Results
The demographic and clinical characteristics of our population sample are presented in Table 1. Univariate analysis showed that four stroke risk factors—hypertension, diabetes, smoking status, and alcohol consumption—were, as expected, significantly more frequent in patients than in control individuals. Gender and age were also significantly different between these two groups. During sample collection, the incidence of stroke was higher in males than in females, as expected in this age range; thus, the male/female ratio was higher in the patients' group. As stroke is a late-onset disease, we selected controls with a higher mean age than patients to reduce the probability of mis-classification as ‘stroke free.’ Multivariate analyses were performed after univariate analyses and nongenetic confounders were identified. Adjusting for these covariates was performed in the final logistic regression model, which also included genetic markers.
All SNPs tested were in Hardy–Weinberg equilibrium in controls and met quality control criteria, and were thus further analyzed. Two contiguous SNPs in the IL6 gene were associated with stroke susceptibility under a log-additive model (rs2069837: P = 0.005, OR (95% CI) = 0.66 (0.50 to 0.89); rs2069861: P = 0.007, OR (95% CI) = 1.74 (1.15 to 2.63)) (Figure 1, Table 3, Supplementary Table 1), after adjusting for covariates significant in the multivariate analysis model—gender, hypertension, diabetes, and smoking status. These associations with stroke susceptibility remained significant after Bonferroni correction for multiple testing (rs2069837: correctedP = 0.032; rs2069861: correctedP = 0.042). A three-marker haplotype containing the two SNPs individually associated with stroke susceptibility and a third SNP (rs10242595) contiguous to rs2069861 conferred an increased risk of stroke (A(rs2069837)–T(rs2069861)–G(rs10242595), P = 0.014) (Supplementary Table 2). This association did not withstand a Bonferroni correction, although this method may be overconservative because these SNPs are not fully independent. Overall, these results highlight a region in the IL6 gene as a likely susceptibility locus, with contiguous tagging SNPs associated with stroke susceptibility.
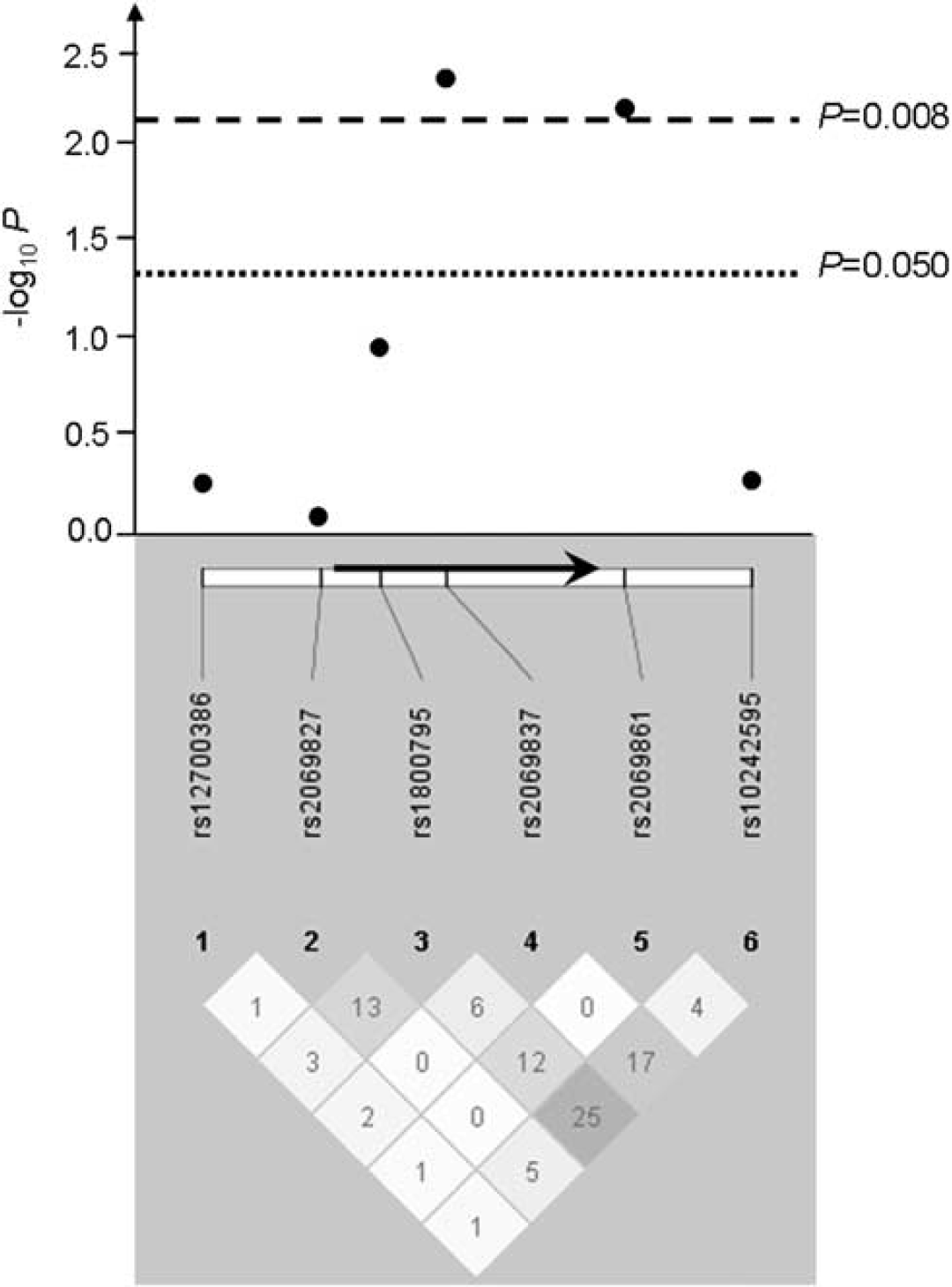
IL6 association results (−log10 P) with stroke susceptibility and pairwise r2 among genotyped SNPs in our population sample. The positions of the six SNPs relative to the IL6 gene (represented by an arrow) are indicated. The magnitude of linkage disequilibrium (r2) is represented by the white–black gradient shading and the values within each diamond. Association results above the line–log10 P = 1.3 are considered significant (P < 0.050); and those above–log10 P = 2.1 survive Bonferroni correction (P < 0.008). Linkage disequilibrium blocks were generated using the Gabriel et al (2002) method.
Genotype and allele frequency distribution, and association with stroke susceptibility for the IL6 and MPO SNPs
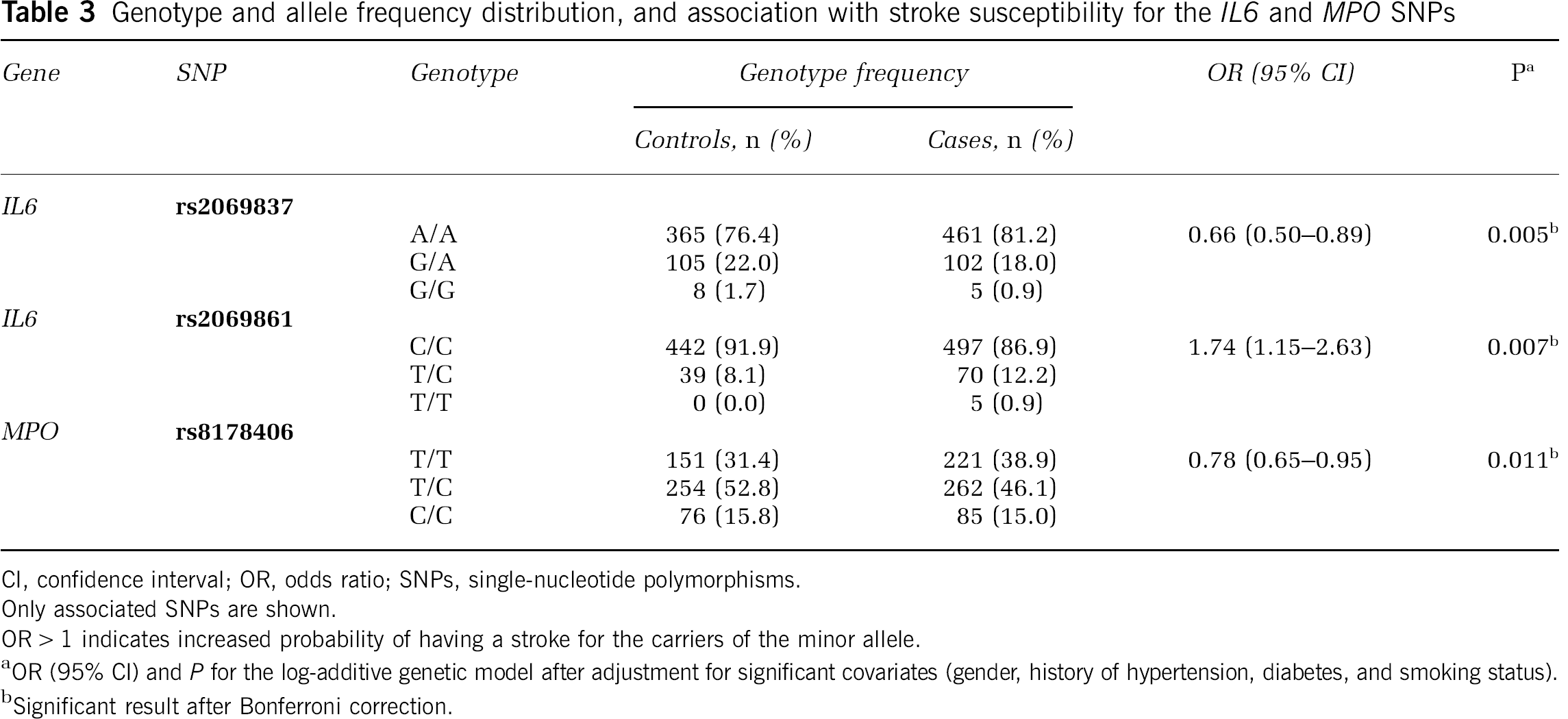
CI, confidence interval; OR, odds ratio; SNPs, single-nucleotide polymorphisms.
Only associated SNPs are shown.
OR > 1 indicates increased probability of having a stroke for the carriers of the minor allele.
OR (95% CI) and P for the log-additive genetic model after adjustment for significant covariates (gender, history of hypertension, diabetes, and smoking status).
Significant result after Bonferroni correction.
One SNP in the MPO gene was significantly associated with stroke susceptibility (rs8178406: P = 0.011, OR (95% CI) = 0.78 (0.65 to 0.95)), and this association survived Bonferroni correction (correctedP = 0.022) (Table 3). Interestingly, restricting the analysis to ischemic patients showed a more significant association (correctedP = 0.006). We did not attempt the independent analysis of hemorrhagic patients, as these were too few for adequate statistical power. None of the tested SNPs in IL1B and TNF was associated with stroke susceptibility in this sample (Supplementary Table 1).
Because gene interactions may have an important impact on complex phenotypes, including human disease susceptibility, we investigated the existence of genetic interactions that could contribute to stroke risk, using the MDR method. The most significant model for interaction was a two-marker combination between rs10242595 in the IL6 gene and rs8178406 in the MPO gene (Table 4). This model shows a moderately increased TBA of 0.556, thus correctly classifying 55.6% of the individuals tested (P = 0.031, based on 1,000-fold permutations), but a high CVC of 9/10, i.e., the model was selected 9 times out of 10 cross-validation subsets. The global OR for this model was 1.69 (95% CI = 1.31 to 2.19). Two genotype combinations of these SNPs contributed to stroke: AA(rs10242595)-CC(rs8178406) and GA(rs10242595)-TT(rs8178406) (OR (95% CI) = 2.80 (1.17 to 7.53) and 1.53 (1.22 to 1.99), respectively) (Table 5). The IL6 rs10242595 SNP is part of the three-marker haplotype associated with stroke susceptibility (Supplementary Table 2), while the MPO rs8178406 SNP was independently associated with stroke (Table 3). Interpretation of the genetic effects identified by the MDR is possible using the interaction dendrogram shown in Figure 2. Interaction between rs10242595 and rs8178406 shows a positive information gain, indicating a nonlinear, synergistic relationship between the IL6 (rs10242595) and the MPO (rs8178406) genes (i.e., epistasis). Our finding thus suggests that an interaction between two genetic variants in the IL6 and MPO genes contributes to stroke susceptibility, warranting confirmation in independent populations. Additional models were not significant, although there are trends possibly suggesting that interactions between IL6 (rs10242595) and TNF (rs909253), as well as MPO (rs2071590) and TNF (rs8178406) (permuted P = 0.054 and P = 0.060, respectively) may contribute to stroke susceptibility (Table 4).
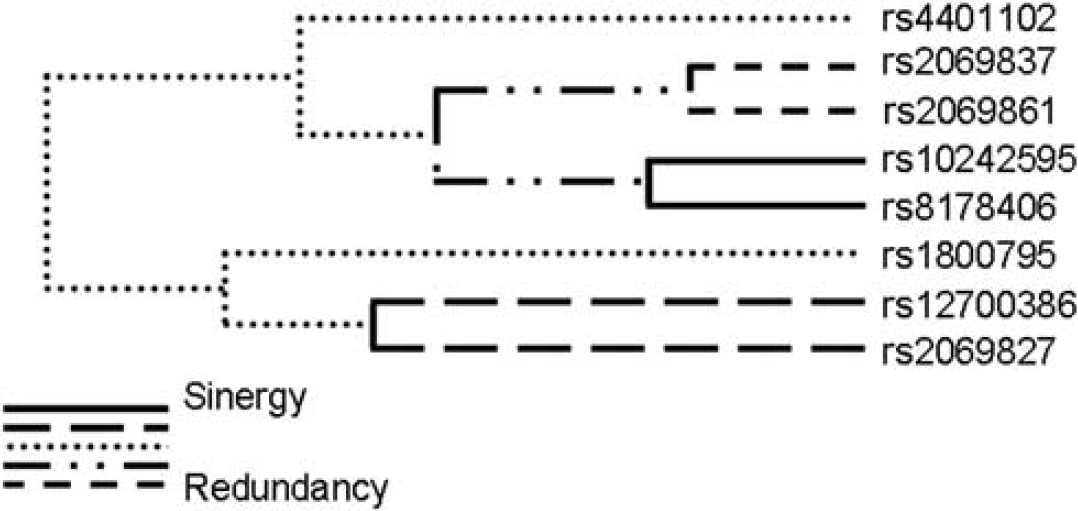
Interaction dendrogram for the IL6 and MPO polymorphisms in stroke susceptibility. The length of the dendrogram branch that connects two polymorphisms indicates the strength of interaction (the shorter the branch, the stronger is the interaction).
Gene × gene interaction models obtained using the multifactor-dimensionality reduction (MDR) method in stroke susceptibility
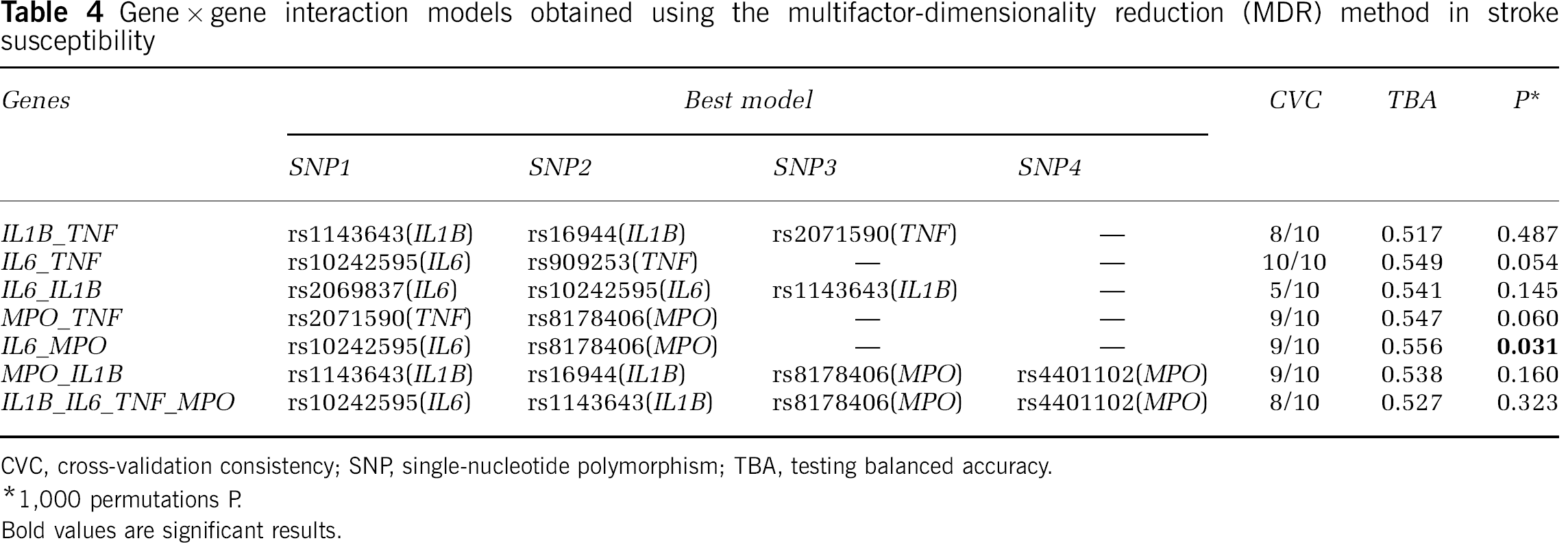
CVC, cross-validation consistency; SNP, single-nucleotide polymorphism; TBA, testing balanced accuracy.
1,000 permutations P.
Bold values are significant results.
Odds ratio (OR) of each genotype combination of IL6 rs10242595 and MPO rs8178406 obtained using the OR-based MDR
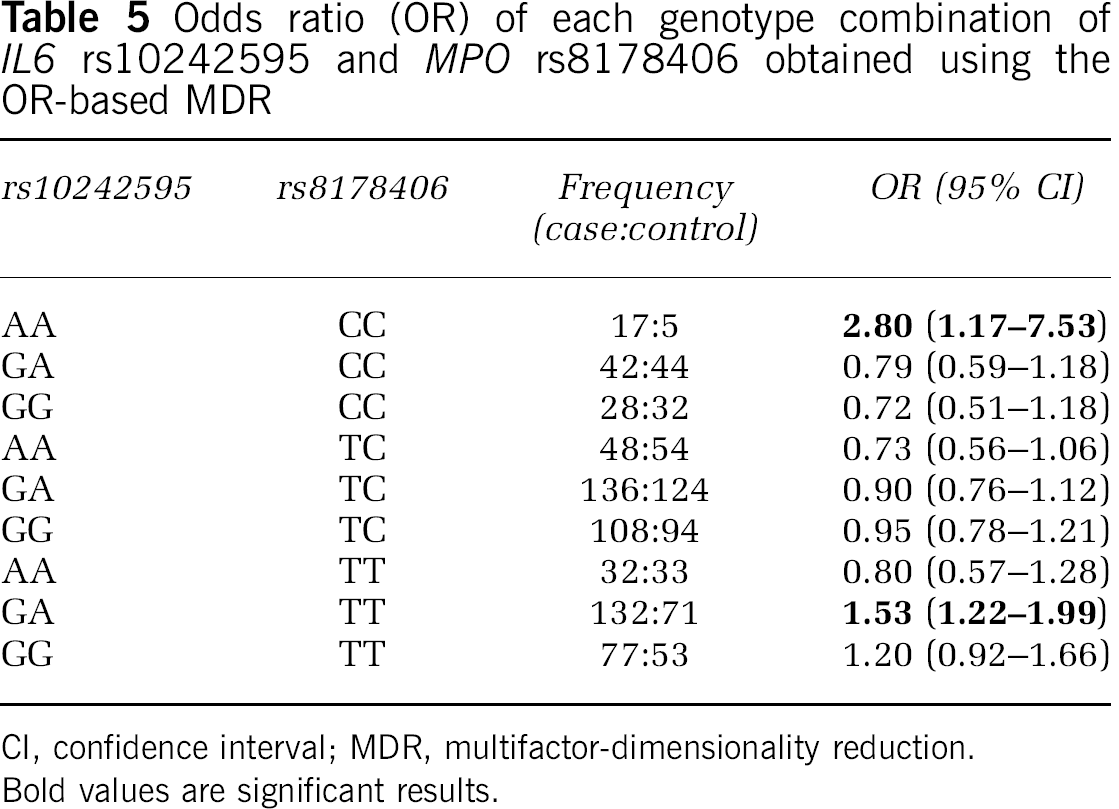
CI, confidence interval; MDR, multifactor-dimensionality reduction.
Bold values are significant results.
The impact of IL1B, IL6, MPO, and TNF genetic variants in patient's outcome at 3 months was investigated in the subset of 546 patients for whom clinical information during hospitalization at 3 months was available. The demographic and clinical characteristics of our population sample are presented in Table 2. Occurrence of aphasia, urinary incontinence, paresis, altered consciousness and medical and neurologic complications during hospitalization, which reflect stroke severity, was identified in univariate analysis as significant predictors of poor outcome. One SNP in the IL6 gene was associated with stroke outcome at 3 months (rs1800795: P = 0.011, OR (95% CI) = 1.52 (1.10 to 2.12)) (Supplementary Table 3), after adjusting for covariates significant in the multivariate model—type of stroke, history of hypertension, and occurrence of aphasia, paresis, altered consciousness, and complications during hospitalization, but did not remain significant after Bonferroni correction (correctedP = 0.066). One two-marker haplotype containing this SNP was associated with an increased probability of good recovery at 3 months (G(rs1800795)-A(rs2069837), P = 0.008), and this haplotypic association survived Bonferroni correction (correctedP = 0.048) (Supplementary Table 4). We found no evidence for an association of IL1B, MPO, or TNF with stroke outcome at 3 months (Supplementary Tables 3 and 4). No significant interaction model was found for stroke outcome at 3 months (Supplementary Table 5).
Discussion
The objective of the present study was to investigate the role of selected inflammatory genes in stroke susceptibility and recovery. We found evidence for a main effect of the IL6 and MPO genes in stroke risk, with specific polymorphisms significantly associated with stroke susceptibility, after adjustment for confounding demographic, clinical or lifestyle risk factors. We also report an epistatic gene interaction effect between IL6 and MPO in stroke susceptibility. Our genetic findings thus support previous evidence from other research areas for a role of inflammatory molecules in stroke.
Association analysis of the IL6 gene showed that two SNPs survived Bonferroni correction, highlighting a region in the IL6 gene that is likely to harbor risk variants of moderate to low effect size. The associated SNPs are contiguous to the IL6 SNP (rs1800795) that has been widely tested in multiple population sets, but are not in linkage disequilibrium with this functional polymorphism in our sample. We did not replicate the association with this SNP, suggesting that the present results are signaling a different causative variant in the IL6 gene, but still reinforce a role of the IL6 gene in stroke susceptibility. Accordingly, several previous studies have failed to confirm the association of rs1800795 with stroke, while others showed heterogeneity regarding the associated allele or genotype (Tso et al, 2007). These conflicting results may be due to allelic or genetic heterogeneity and/or limitations in study designs, or reflect true differences in stroke etiology between populations. Our study also provided novel evidence for the association of the MPO gene with stroke. Of the two SNPs tested, covering genetic variability in this region, one was associated with stroke risk. This effect seemed to be largely driven by the ischemic stroke subset where the strength of association was improved, perhaps reflecting somewhat distinct pathological mechanisms for the hemorrhagic and ischemic subtypes. Validation in independent populations is now warranted.
The identification of a synergistic interaction between IL6 and MPO contributing to stroke susceptibility highlights the importance of testing for epistasis and illustrates the complexity of the inflammatory processes in stroke. It indicates that susceptibility may be modulated not only by a variety of genetic factors but also by nonlinear gene–gene interactions, as had been previously shown by others (Flex et al, 2004; Liu et al, 2009; Palmer et al, 2010). Liu et al (2009), in particular, investigated the existence of gene–gene interactions between five candidate genes and stroke and found that individuals with a combination of polymorphisms in three of these genes had an increased risk of thrombotic stroke. Two additional studies also report that the risk of stroke increases with the number of high risk genotypes in proinflammatory gene polymorphisms carried by an individual, suggesting that such polymorphisms act synergistically (Flex et al, 2004; Palmer et al, 2010). Finally, our finding is in agreement with a previous in vitro functional study, showing that enzymatically inactive MPO induced IL-6 secretion in a dose and time-dependent manner by endothelial cells (Lefkowitz et al, 2000).
The genetic factors influencing outcome after a stroke event are far less studied than genetic risk factors. The importance of inflammation after stroke onset and the correlation between inflammatory marker levels and infarct volume or patient's outcome (Smith et al, 2004; Sotgiu et al, 2006), led us to investigate the role of several inflammatory genes in stroke outcome at 3 months. We found an IL6 two-marker haplotype associated with patient's outcome at 3 months. These results are intriguing, as IL-6 is one of the cytokines induced after stroke, having a fundamental role in the inflammatory injury that follows a stroke event, but equally known to have neuroprotective effects in later stages after stroke (Herrmann et al, 2003). Interleukin-6 levels have been correlated with stroke severity, 12 months mortality, clinical outcome, and brain infarct volume (Smith et al, 2004). However, our results require validation in independent, larger population samples.
Interleukin-6 is a pleiotropic cytokine, with both pro and antiinflammatory functions and a low level of expression in the brain under normal physiological conditions (Luheshi and Rothwell, 1996). However, increased levels of IL-6 have been detected after a stroke event (Clark et al, 1999). Increased expression of IL-6 has also been found in atherosclerotic plaques (Schieffer et al, 2000), suggesting that the identified IL6 association with stroke susceptibility may be mediated by atherosclerosis progression. Likewise, abundant MPO-positive cells are present in sites of atherosclerotic plaque rupture, and this molecule may contribute to stroke through destabilization of the atherosclerotic plaques (Sugiyama et al, 2001). It would be very interesting to correlate IL6 and MPO genetic variants with the carotid intimal-media wall thickness, which is a marker for atherosclerosis (Mattace Raso et al, 1999). However, this data are only available for a small percentage of our patients, precluding this analysis for now.
The complex interplay between genetic background, clinical and lifestyle factors and the environment may ultimately regulate the onset, acute phase, and outcome of stroke. In the present study, we present supporting evidence for a role of the IL6 and MPO inflammatory genes in stroke susceptibility, and show that stroke risk is modulated by main gene effects together with clinical and lifestyle factors as well as by gene–gene interactions. Our findings are compatible and strengthen previous genetic and biological observations, highlighting the need of further functional studies, particularly in view of the possible utility of IL-6 as a diagnostic and prognostic biomarker for stroke.
Footnotes
Acknowledgements
The authors are grateful to all study participants and their families. The authors thank Dr Marinho Falcão, Dr Carlos Dias, and the whole team at Instituto Nacional de Saäde Dr Ricardo Jorge, and all the clinicians that recruited study subjects from the following hospitals: H.S. João, H. do Espírito Santo, Centro Hospitalar do Funchal, H. Conde S. Bento, H.S. José, H.S. Marcos, H. Garcia de Orta, H. de Faro, Centro Hospitalar de Coimbra, Centro Hospitalar de Vila Nova de Gaia, H. Infante D. Pedro, Serviços de Assistência Médico-Social do Sindicato dos Bancários do Sul e Ilhas, H. Sto. António dos Capuchos, H. Sto. António, H. Distrital de Mirandela, H. Sta. Maria, H. de Egas Moniz, H. Prof. Doutor Fernando Fonseca and H.S. Pedro de Vila Real. The authors also thank the technical assistance provided by the Genotyping Unit at Instituto Gulbenkian de Ciência.
The authors declared no conflict of interest.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
